workflow tutorial
example.Rmd
library(seatrackRgls)
knitr::opts_chunk$set(
collapse = TRUE,
comment = "#>"
)Prepare basic metadata for calibration. Basic metadata must have
logger_id, logger_model, species,
colony, date_deployed,
date_retrieved columns. It is expected to be one row per
logger/retrieval year combination.
print(example_metadata)
#> logger_id logger_model date_deployed date_retrieved colony
#> 1 C411 mk4083 2015-06-11 2017-06-11 Sklinna
#> species
#> 1 Black-legged kittiwakePrepare colony information. Colony information must have
colony, col_lat, col_lon
columns.
print(example_colony_info)
#> colony col_lat col_lon
#> 1 Sklinna 65.202 10.995Set your import directory, where your light data is placed. Light
data is expected to be in the format
<logger_id>_<year_retrieved>_<logger_model>,
e.g. C411_2017_mk4083
import_dir <- "light_data"
print(list.files(import_dir))
#> [1] "C411_2017_mk4083.lig"Also set up an export directory, where all outputs will be saved.
export_dir <- "processed_light_data"With all this loaded, you can now carry out the first step which is to calibrate your data. To assist in this, there is an initial round of processing that generates helpful plots to choose calibration values
prepare_calibration(
import_directory = import_dir,
metadata = example_metadata,
all_colony_info = example_colony_info,
output_dir = export_dir
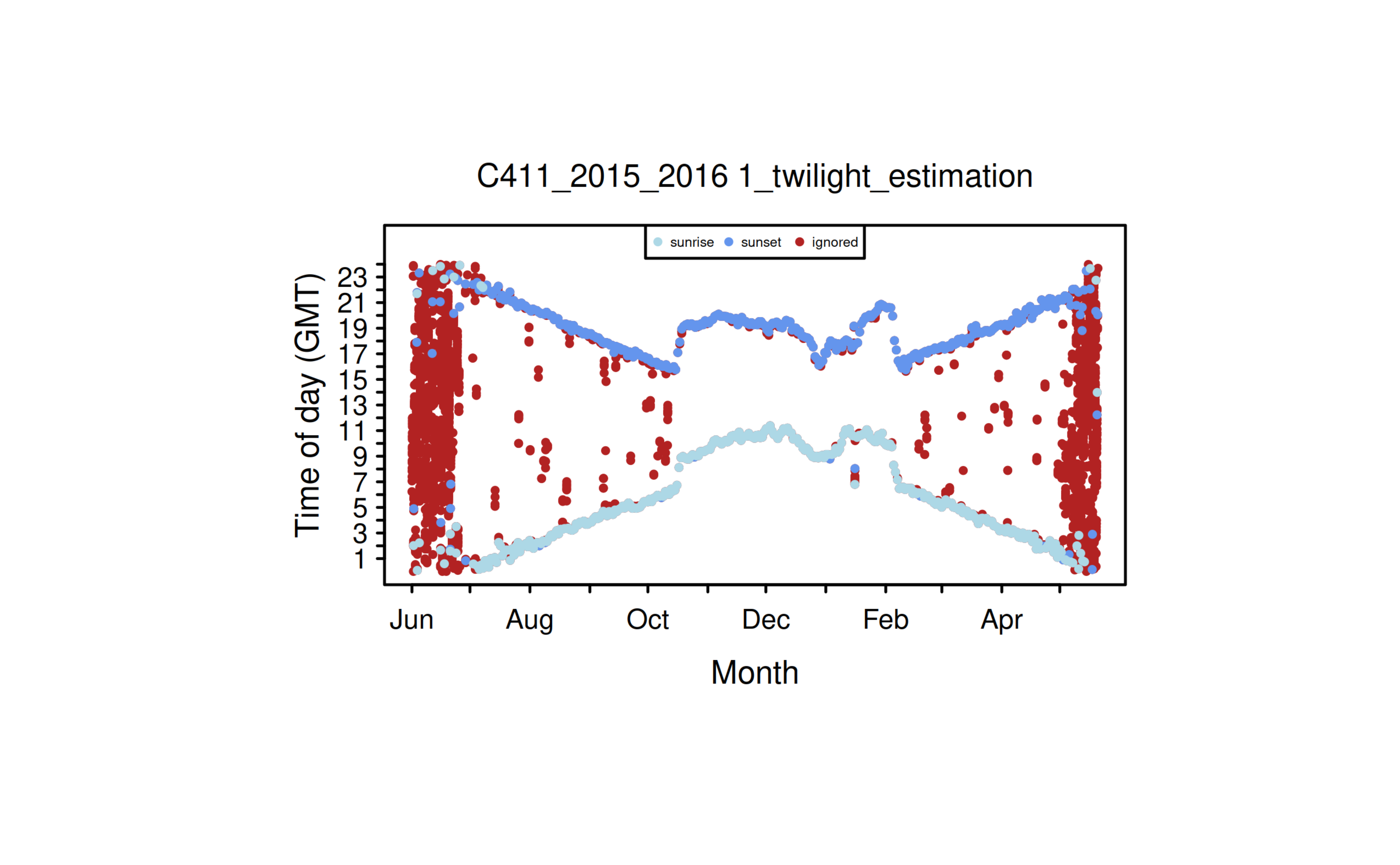
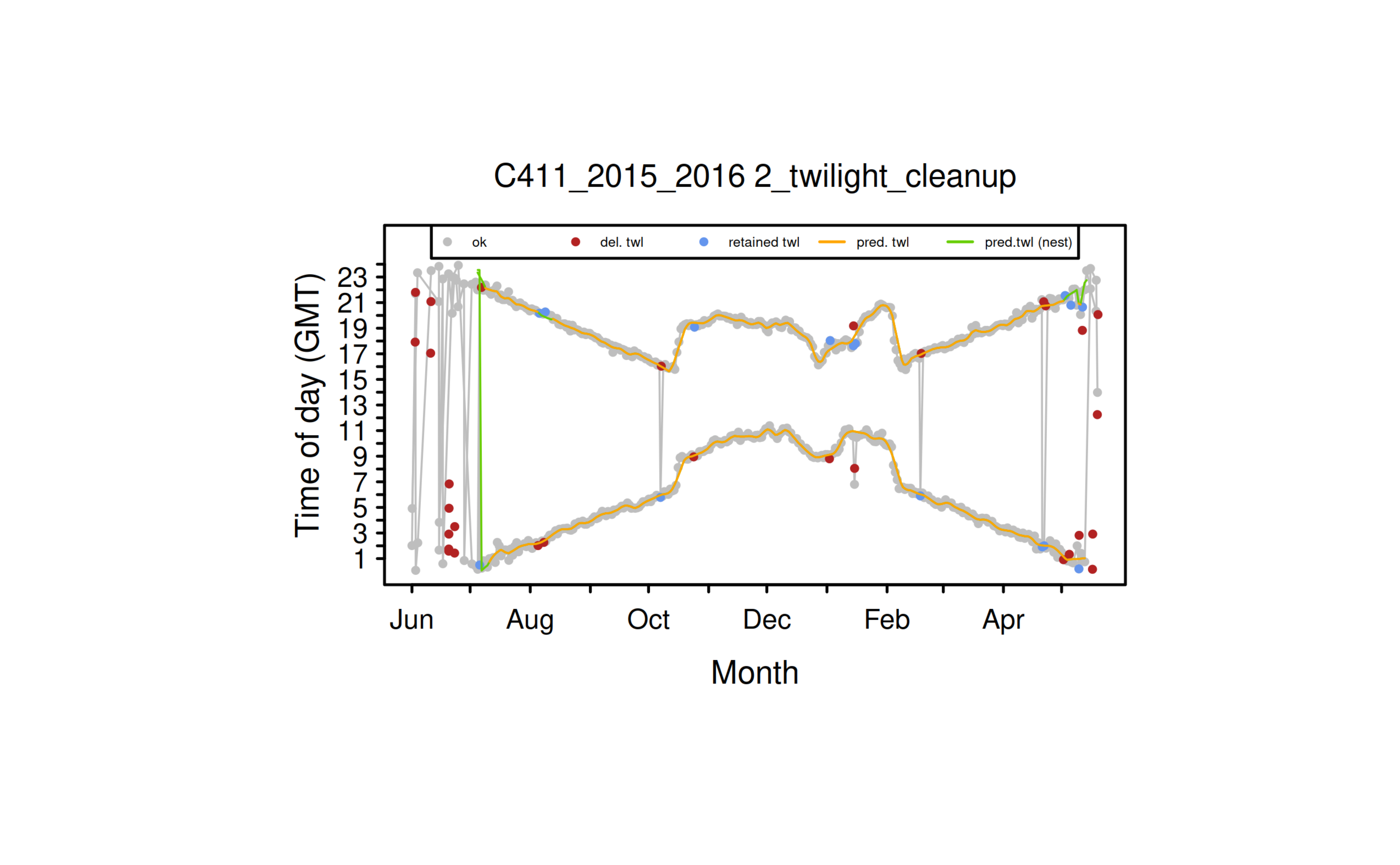
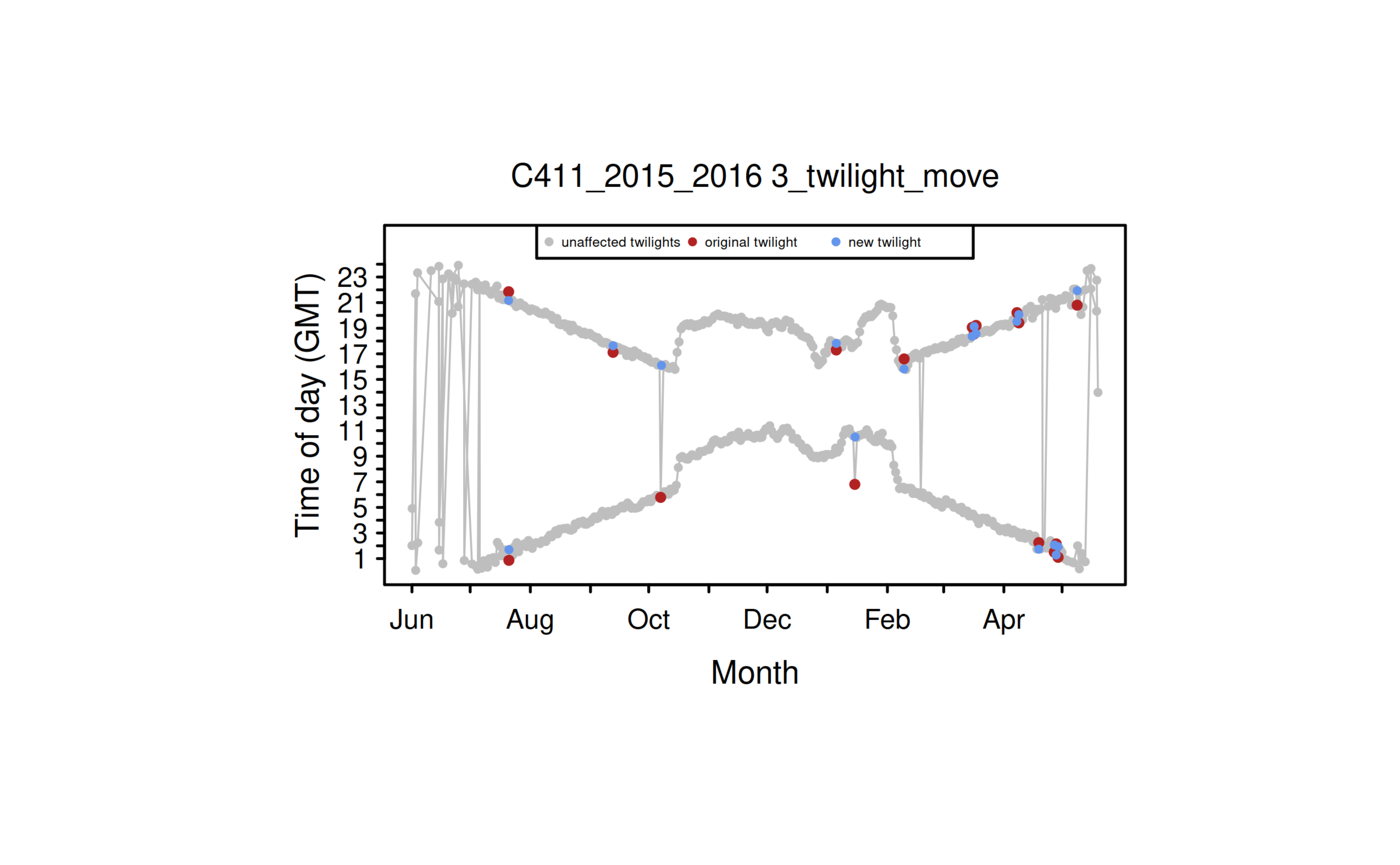
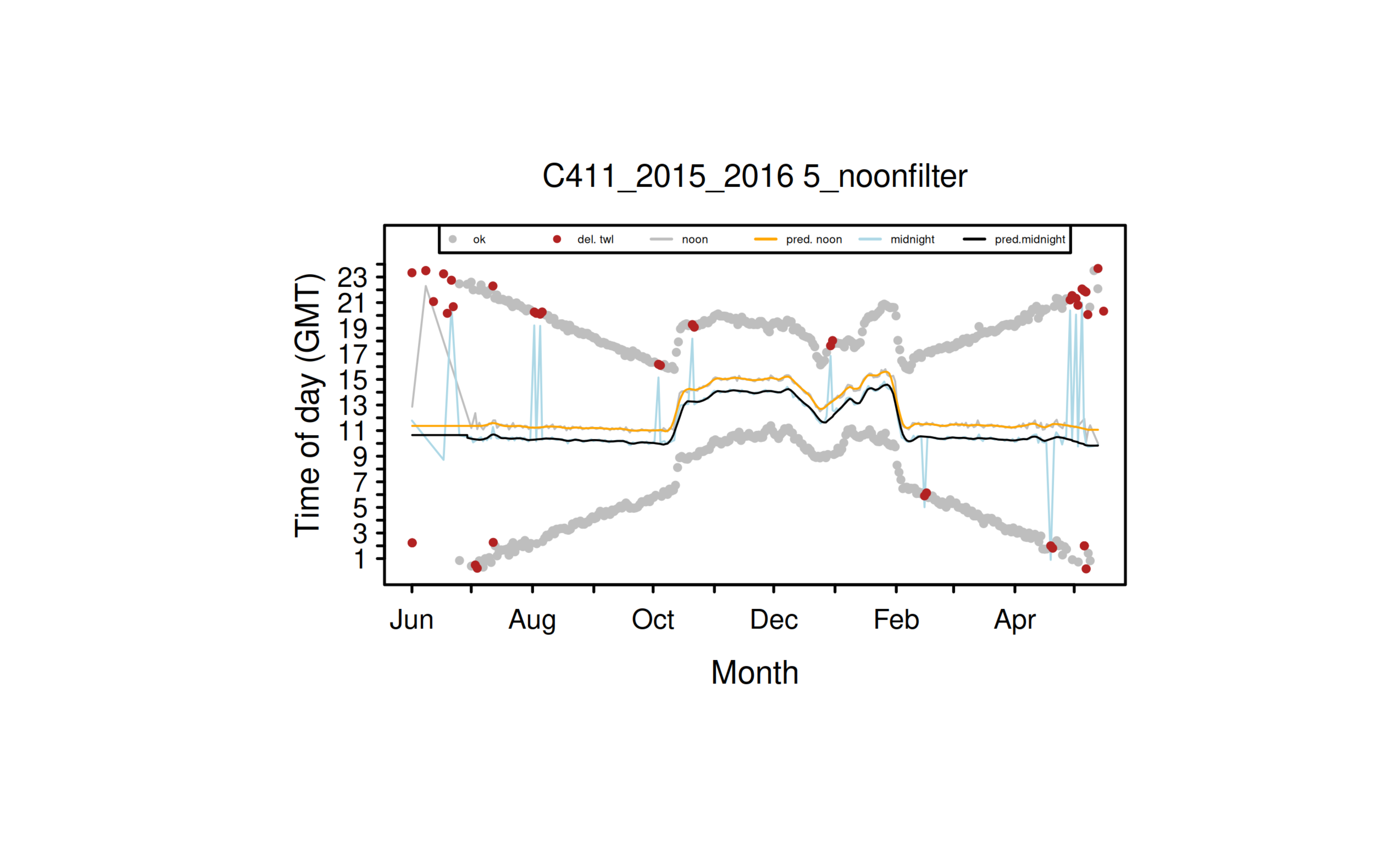
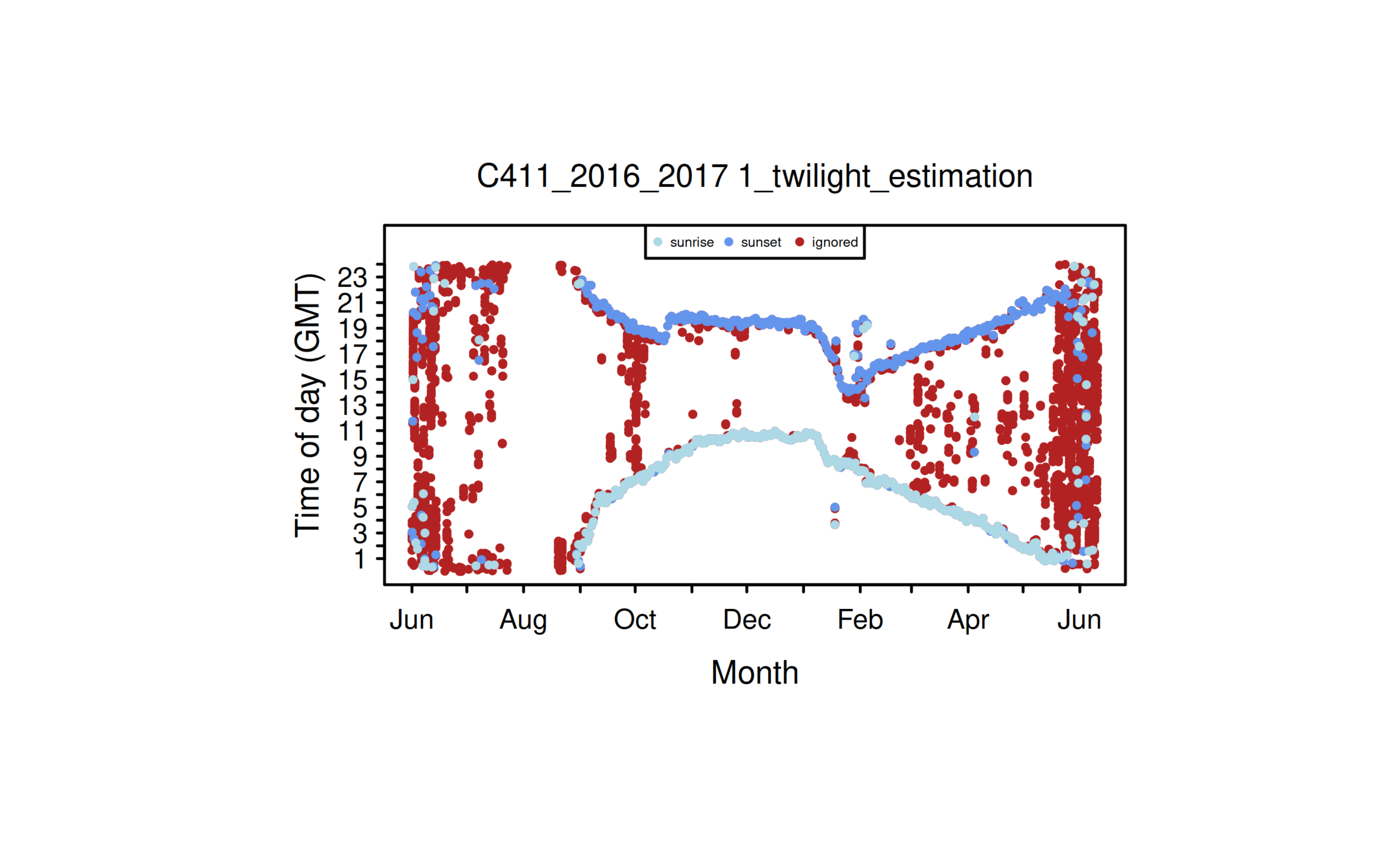
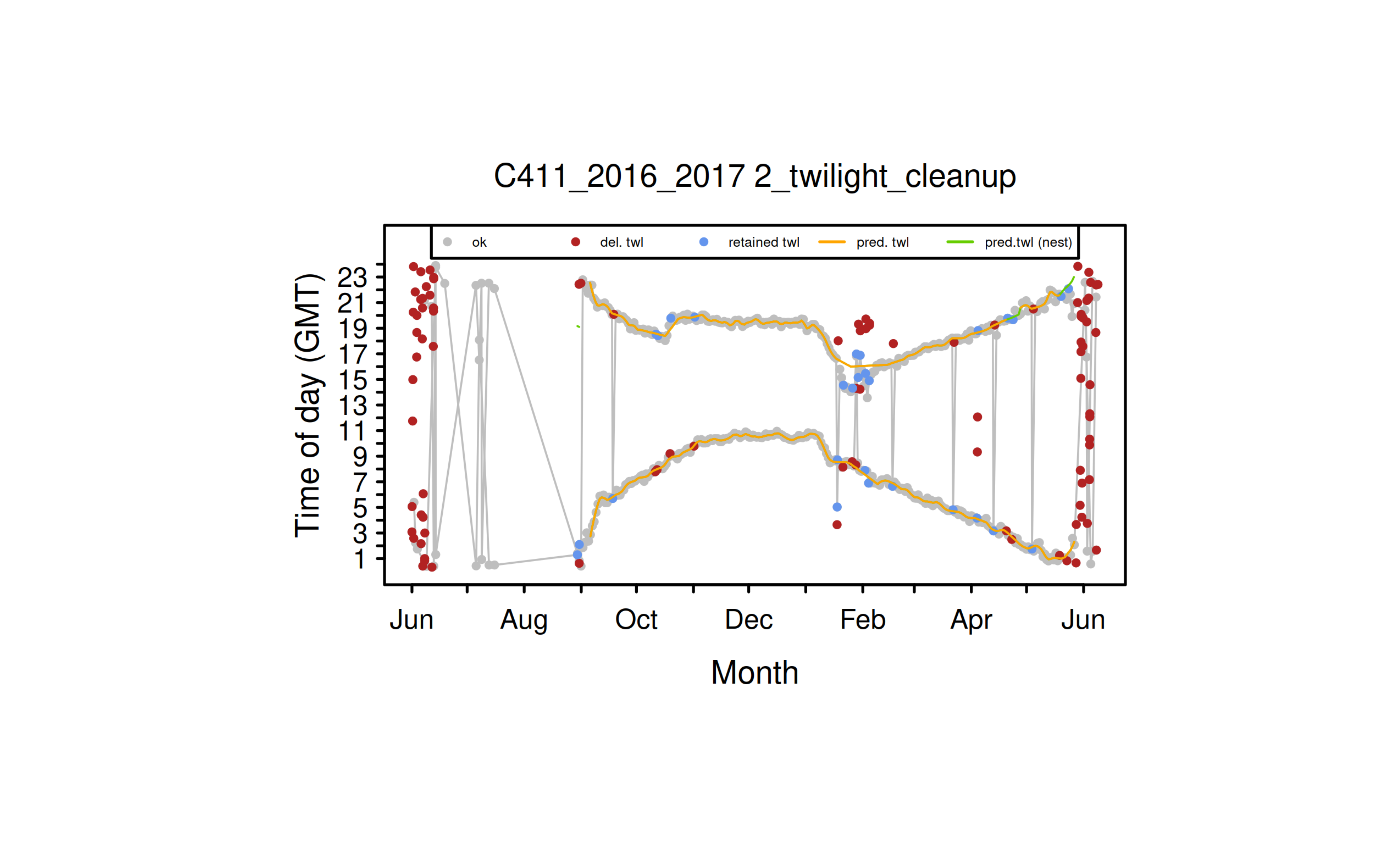
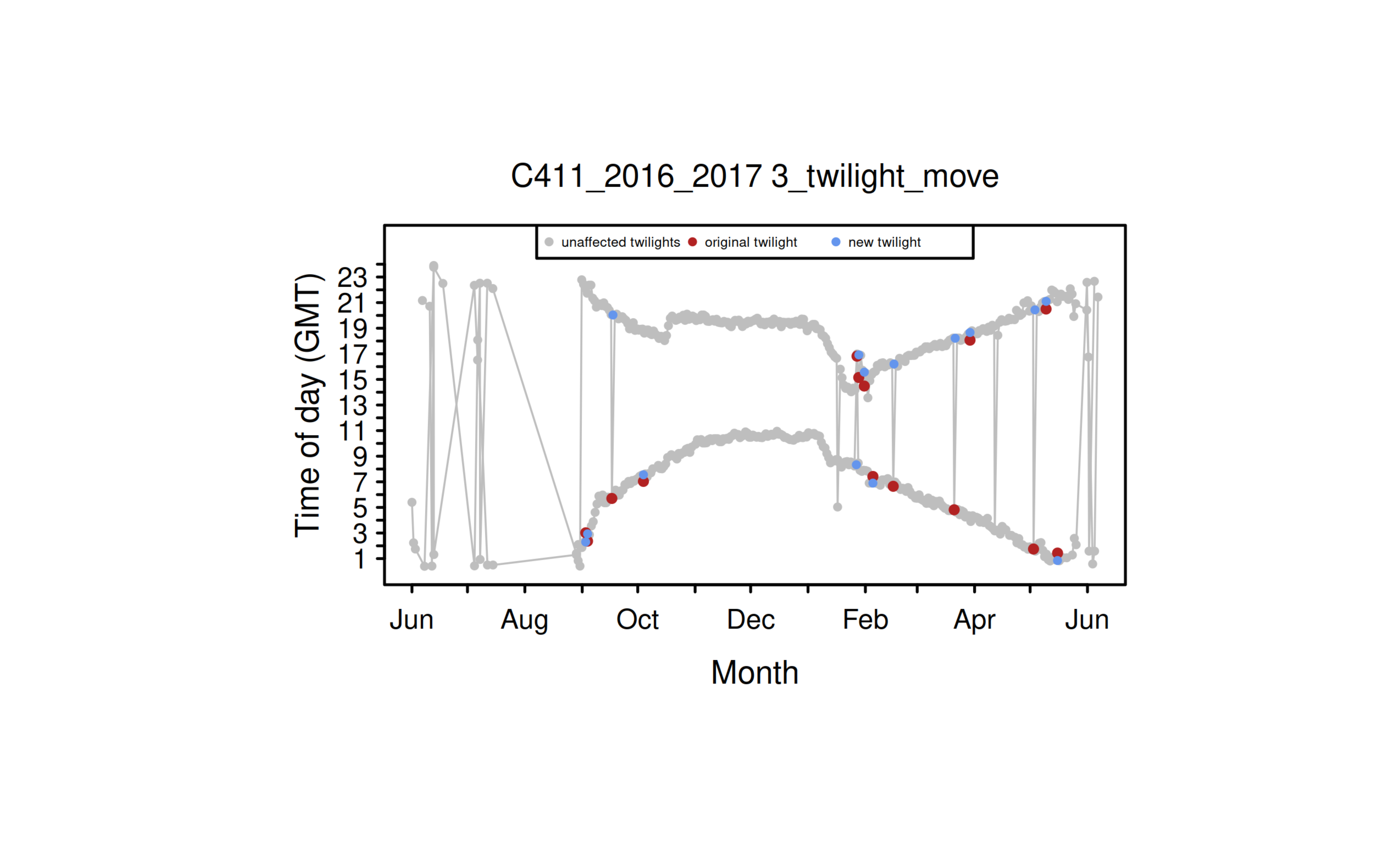
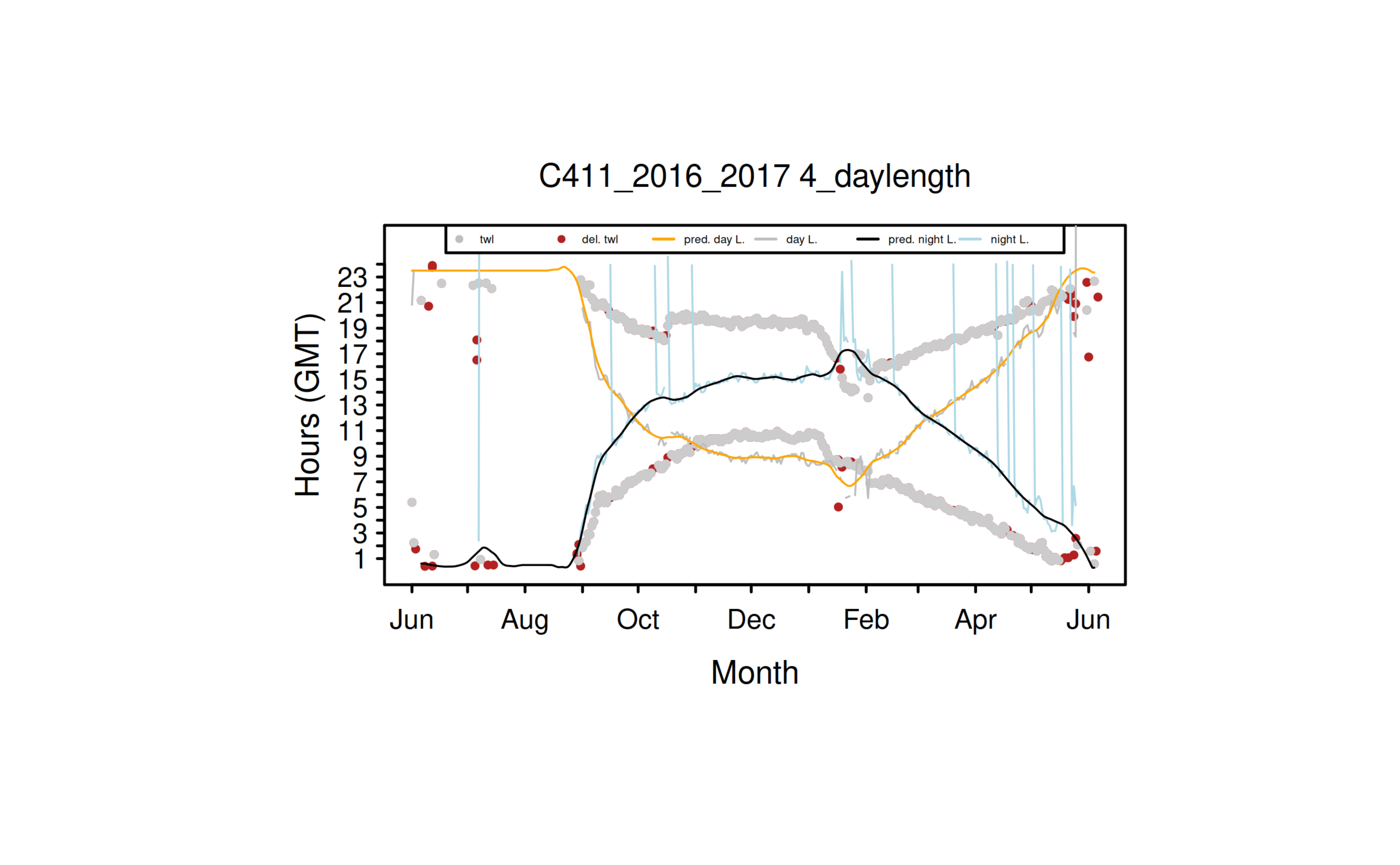
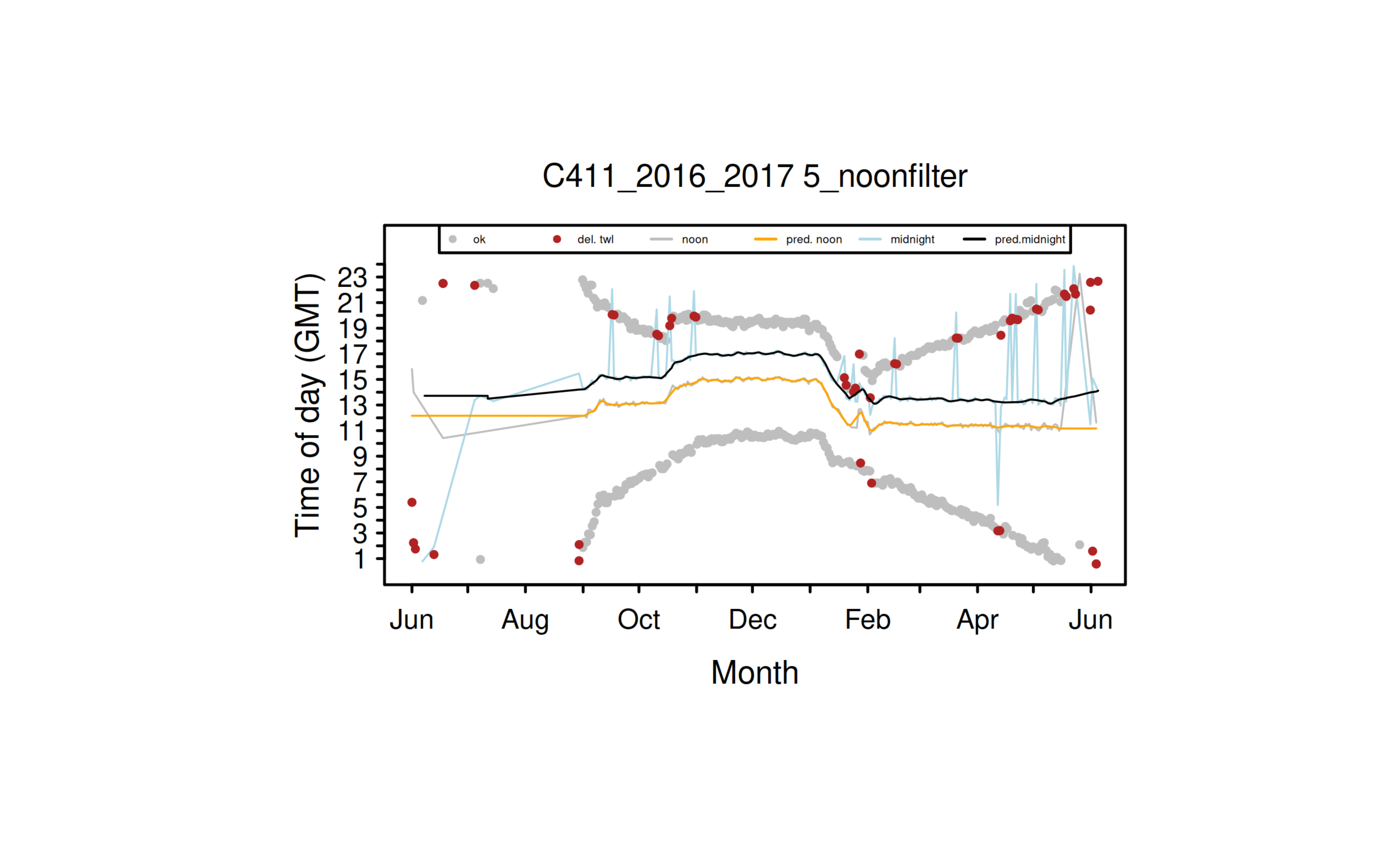
)You will find the calibration plots in the sun_calib
folder created on your output_dir.
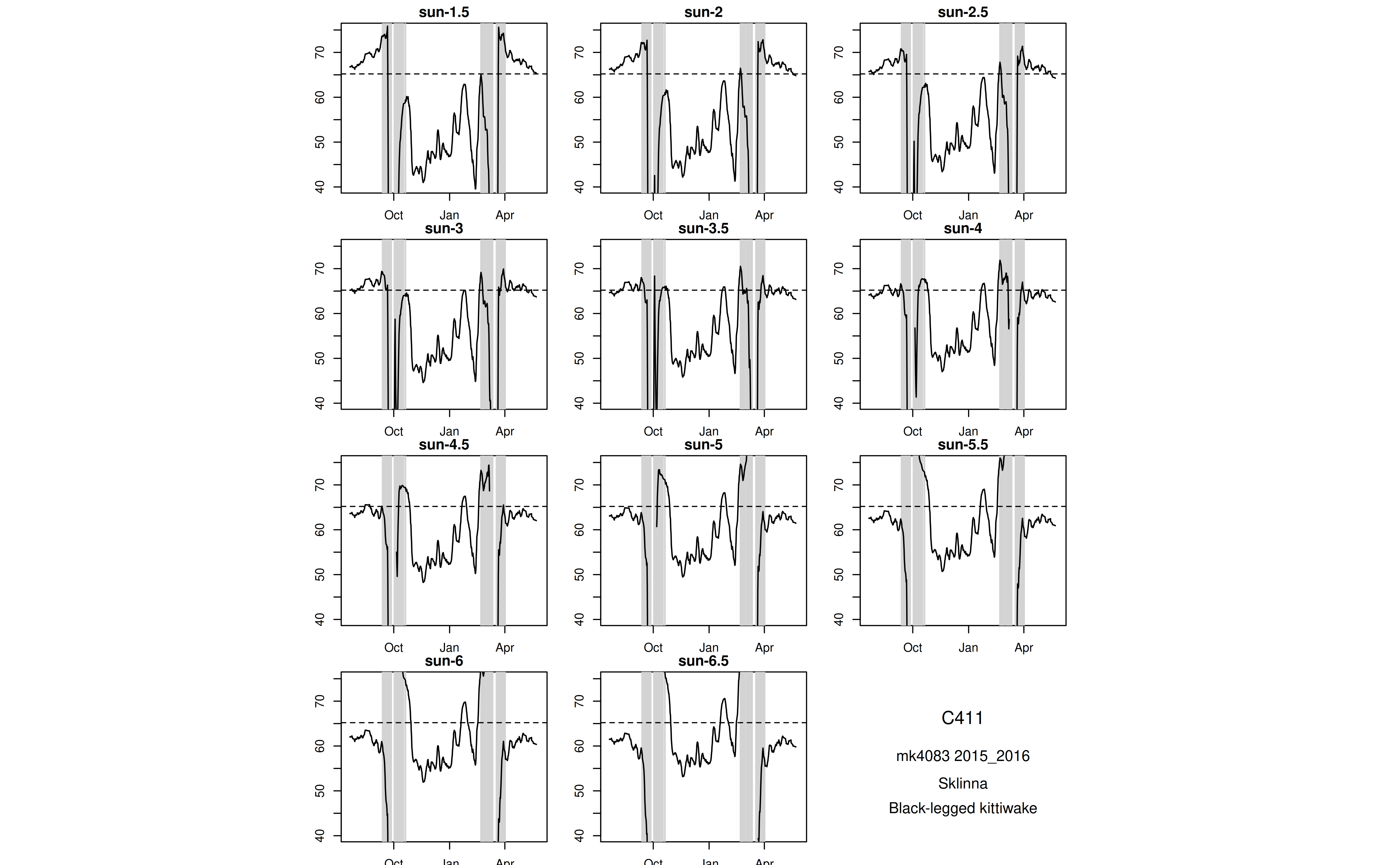
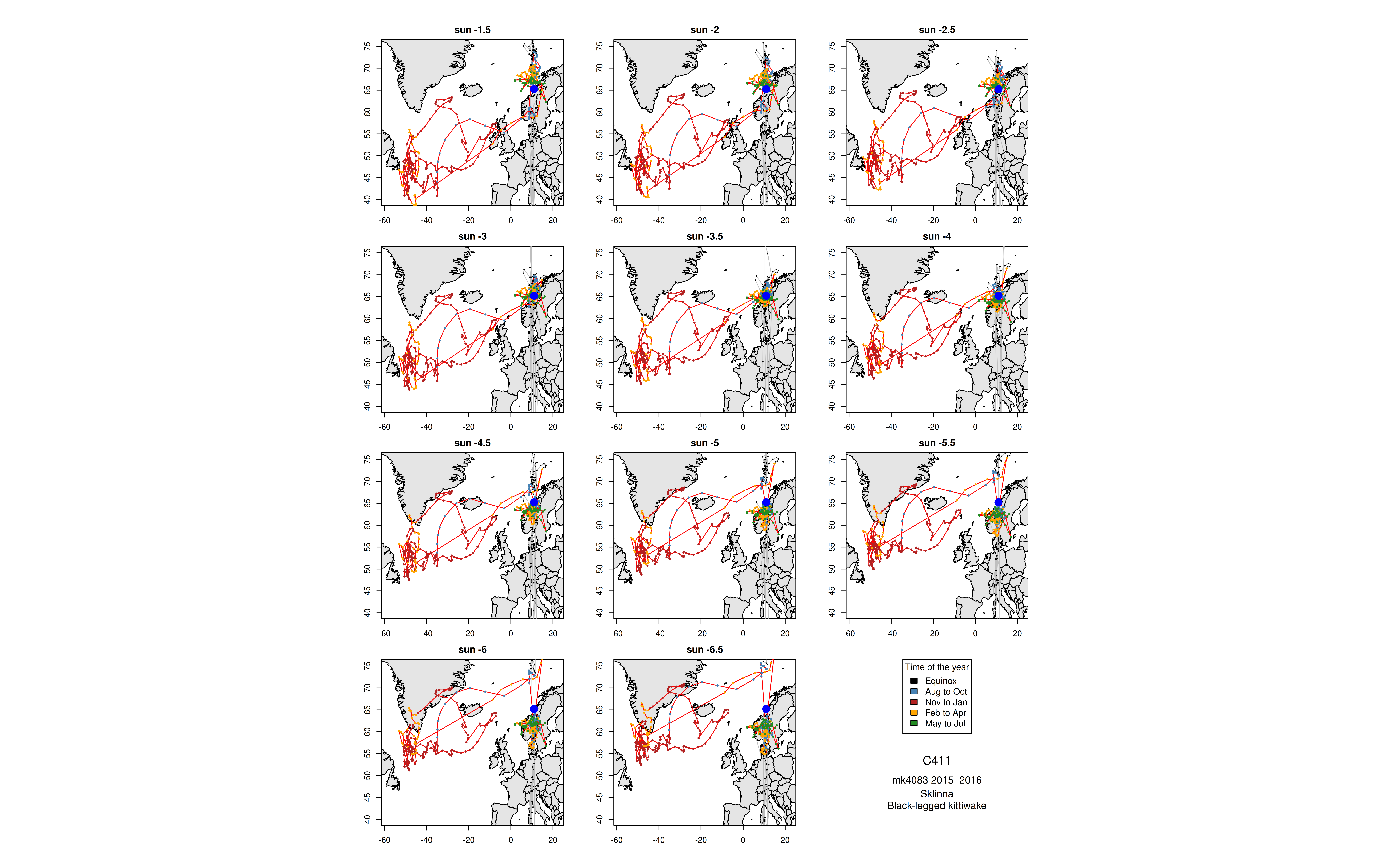
Stare at these plots. Use the force.
By default, this code will also have generated an excel file in the
calibration folder. You can use this to enter your
calibration values.

You must fill in at least the sun_angle_start column. It is also reccomended to include your name in the analyzer column.
Once you have filled in your calibration template, you can use these values to process the light data and export positions.
We can pass a path to the calibration data file:
calibration_data_path <- file.path(export_dir, "calibration", "calibration.xlsx")At this stage, we might want to include some extra relevant information in the final data output.
print(example_extra_metadata)
#> logger_id date_retrieved logger_producer ring_number country_code
#> 1 C411 2017-06-11 Biotrack 6211704 NOSThe final positions are now exported to your
output_dir.
head(positions)
#> logger_id logger_id_year total_years_tracked logger_model start_datetime
#> 1 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> 2 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> 3 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> 4 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> 5 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> 6 C411 C411_2017 2015_2017 mk4083 2015-06-12
#> end_datetime year_tracked species
#> 1 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> 2 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> 3 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> 4 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> 5 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> 6 2016-05-31 23:59:59 2015_2016 Black-legged kittiwake
#> date_time sun_angle eqfilter lon_raw lat_raw lon_smooth1
#> 1 2015-06-22 11:29:56 -0.7500000 TRUE 8.001184 63.18181 8.001184
#> 2 2015-06-26 23:52:50.877155 -0.7487047 TRUE 2.513619 54.92256 2.513619
#> 3 2015-06-28 11:28:04 -0.7482729 TRUE 8.785596 61.70479 5.649608
#> 4 2015-07-05 00:34:04.6875 -0.7478411 TRUE -7.403305 59.32453 -7.403305
#> 5 2015-07-06 00:35:06.073771 -0.7474093 TRUE -7.615658 62.10852 -7.615658
#> 6 2015-07-08 23:42:43 -0.7469775 TRUE 5.600252 65.04015 5.600252
#> lat_smooth1 lon lat tFirst
#> 1 63.18357 8.191518 63.22370 2015-06-22 01:18:53
#> 2 54.92467 4.011154 56.62996 2015-06-26 20:31:44.37931
#> 3 58.31558 5.649608 58.31558 2015-06-28 01:43:58.375
#> 4 59.32636 -7.403305 59.32636 2015-07-04 21:44:05
#> 5 62.11016 -7.615658 62.11016 2015-07-05 22:16:10
#> 6 65.04161 8.031431 64.57426 2015-07-08 22:10:13
#> tSecond type colony col_lat col_lon sun_angle_start
#> 1 2015-06-22 21:40:59 1 Sklinna 65.202 10.995 -0.75
#> 2 2015-06-27 03:13:57.375 2 Sklinna 65.202 10.995 -0.75
#> 3 2015-06-28 21:12:09.625 1 Sklinna 65.202 10.995 -0.75
#> 4 2015-07-05 03:24:04.375 2 Sklinna 65.202 10.995 -0.75
#> 5 2015-07-06 02:54:02.147541 2 Sklinna 65.202 10.995 -0.75
#> 6 2015-07-09 01:15:13 2 Sklinna 65.202 10.995 -0.75
#> sun_angle_end light_threshold noon_filter daylength_filter coast_to_land
#> 1 -0.5 50 TRUE TRUE 100
#> 2 -0.5 50 TRUE TRUE 100
#> 3 -0.5 50 TRUE TRUE 100
#> 4 -0.5 50 TRUE TRUE 100
#> 5 -0.5 50 TRUE TRUE 100
#> 6 -0.5 50 TRUE TRUE 100
#> coast_to_sea loess_filter_k months_breeding_start months_breeding_end
#> 1 Inf 6 4 8
#> 2 Inf 6 4 8
#> 3 Inf 6 4 8
#> 4 Inf 6 4 8
#> 5 Inf 6 4 8
#> 6 Inf 6 4 8
#> boundary.box_xmin boundary.box_xmax boundary.box_ymin boundary.box_ymax
#> 1 -95 100 30 88
#> 2 -95 100 30 88
#> 3 -95 100 30 88
#> 4 -95 100 30 88
#> 5 -95 100 30 88
#> 6 -95 100 30 88
#> analyzer date_retrieved logger_producer ring_number country_code
#> 1 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> 2 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> 3 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> 4 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> 5 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> 6 Kate Kittiwake 2017-06-11 Biotrack 6211704 NOS
#> raw_data_file
#> 1 C411_2017_mk4083.lig
#> 2 C411_2017_mk4083.lig
#> 3 C411_2017_mk4083.lig
#> 4 C411_2017_mk4083.lig
#> 5 C411_2017_mk4083.lig
#> 6 C411_2017_mk4083.ligNote our extra metadata appended to the end.
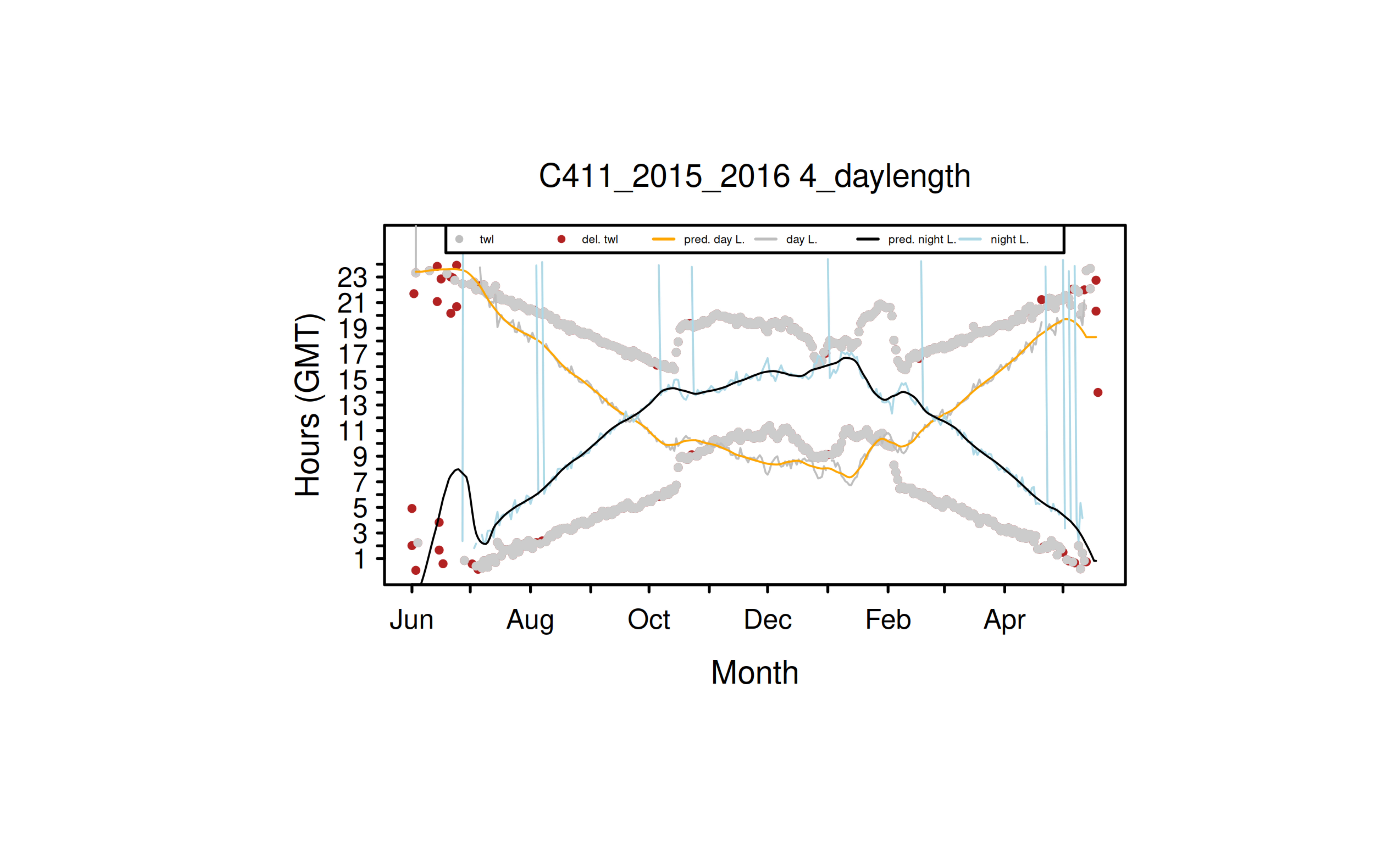
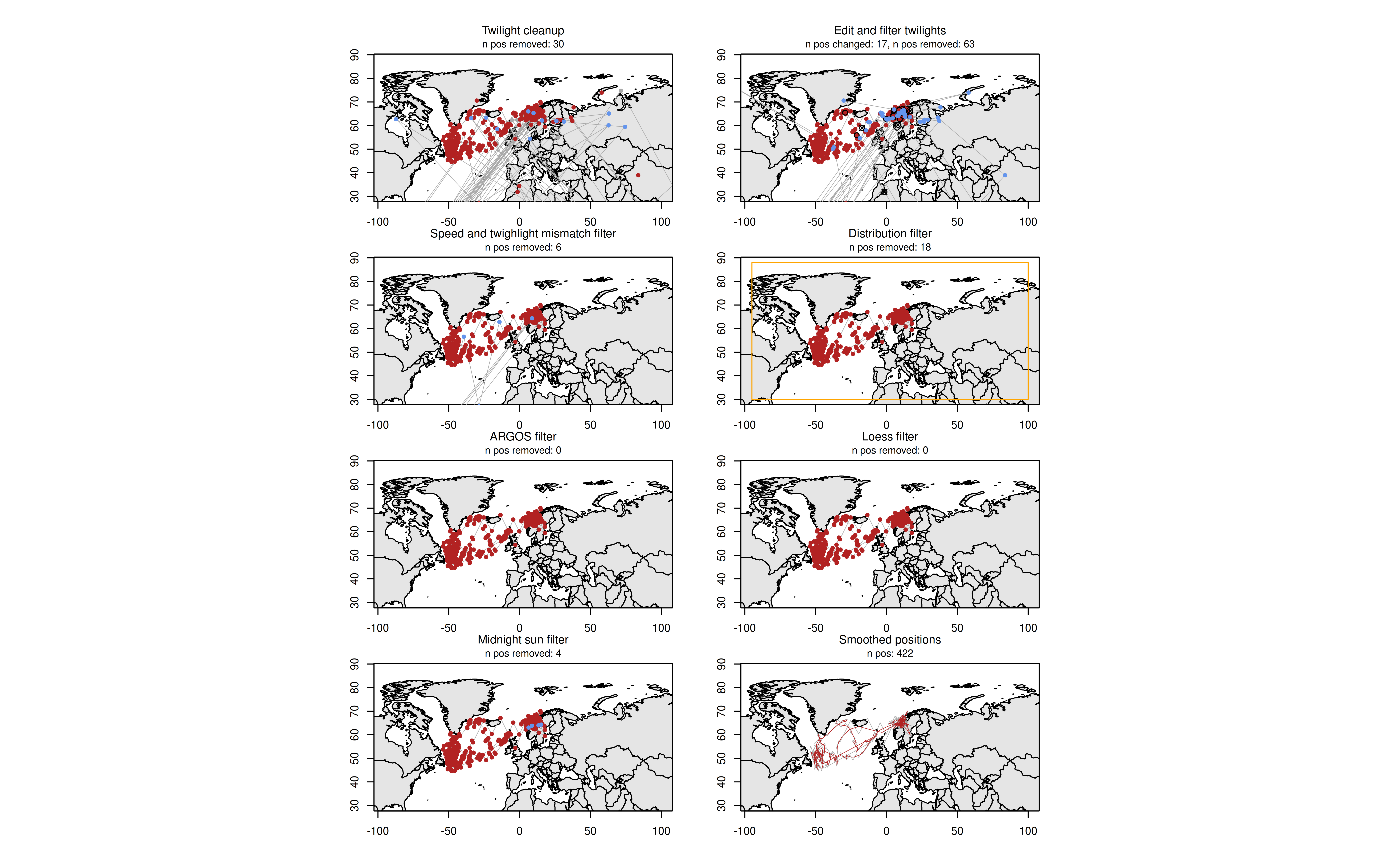
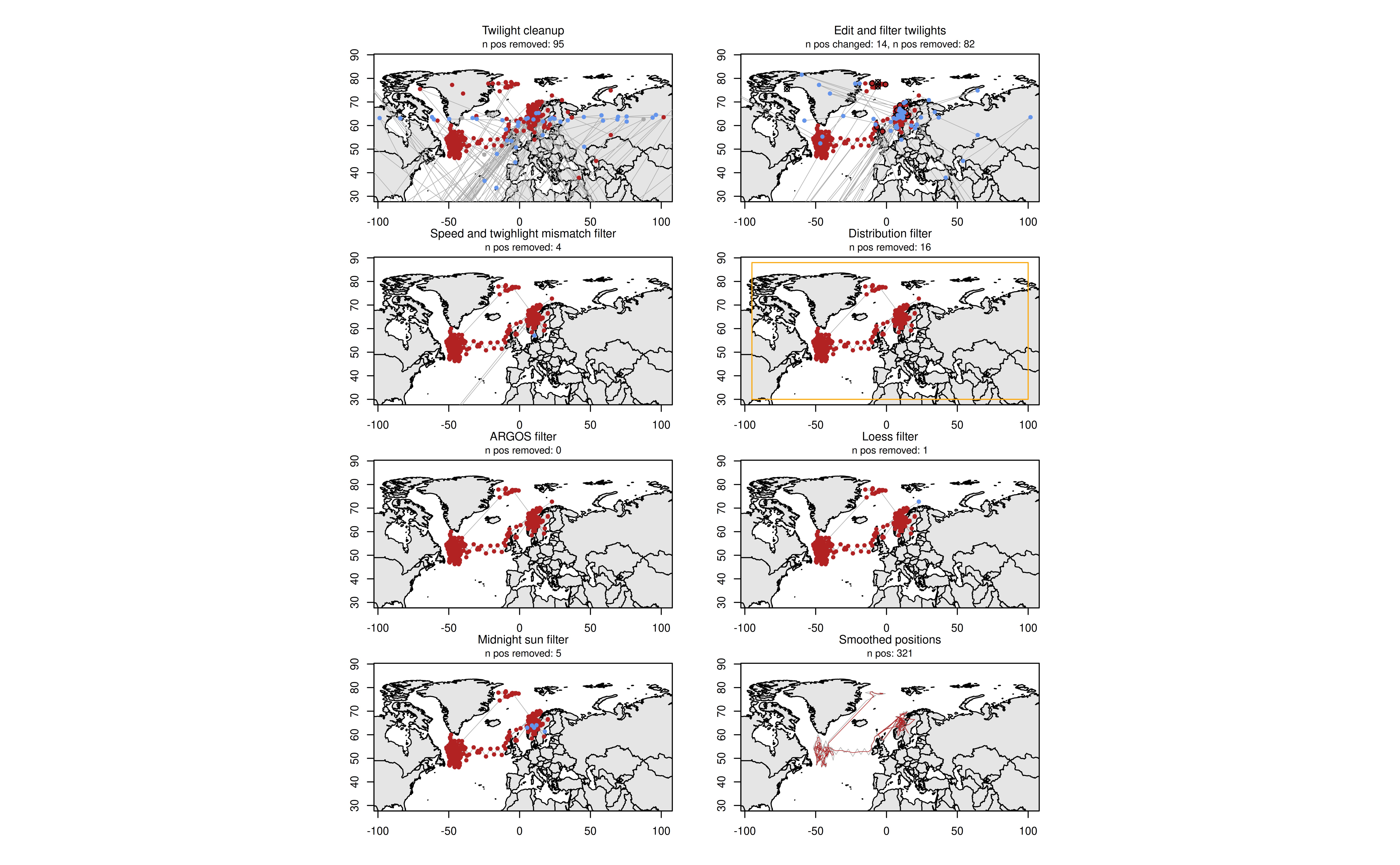
Maps are automatically exported.
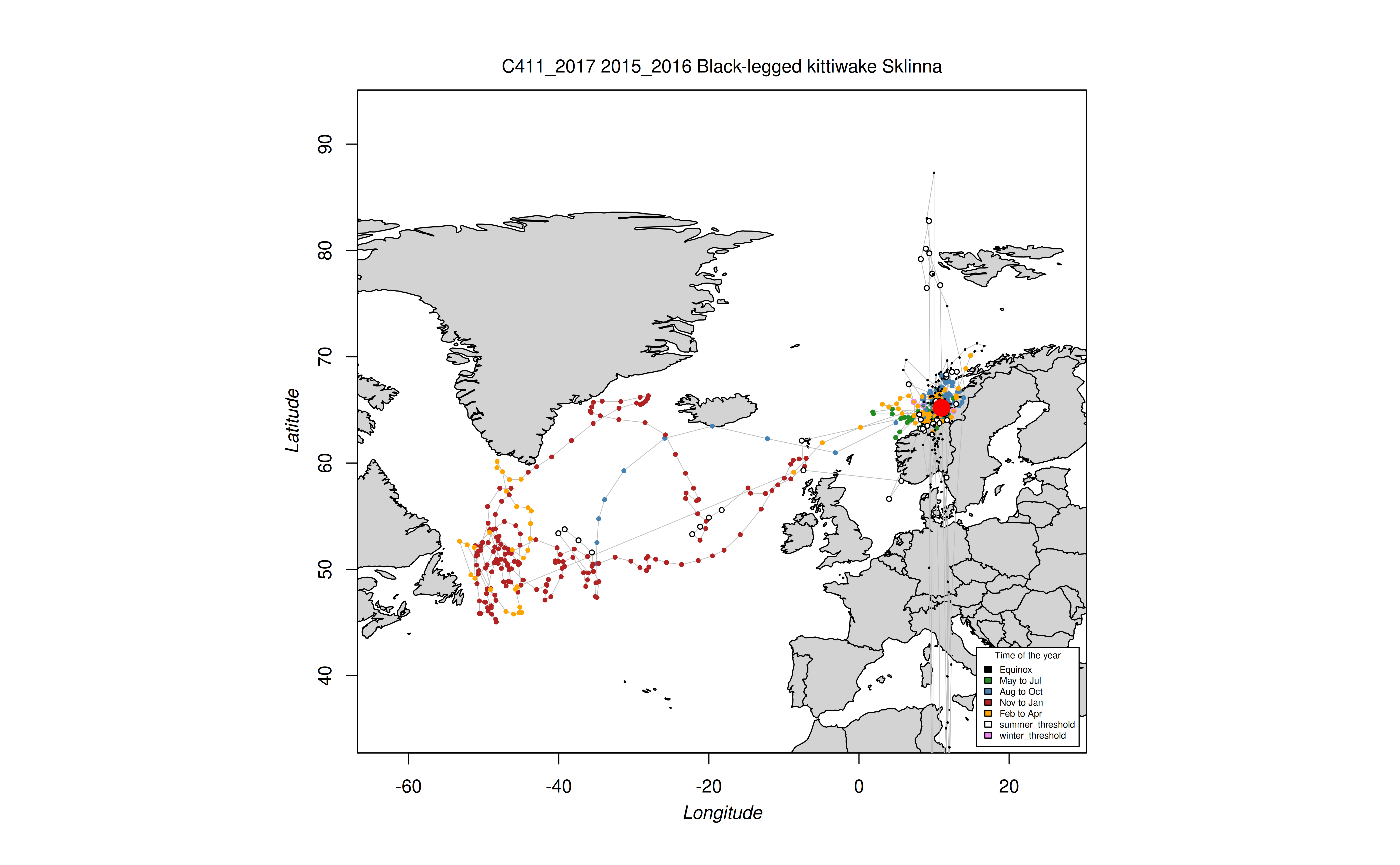
It is worth examining the filter plots too.