2 Indicator data: calculations and visualizations
The basis for the Nature Index (NI) calculations are the indicator data. These are observations, model estimates, or expert judgements that quantify the state (typically abundance) of the different indicator species, species groups, or habitat types.
The “raw” indicator data, including associated reference values, are stored in the NI database (https://naturindeks.nina.no/). As part of the presentation of NI2025, we want to present the following for each individual indicator:
- Raw indicator values, by area, as time-series plots
- Raw indicator values, by area, as pair of value and uncertainty maps
- Scaled indicator values, by area, as time-series plots
- Scaled indicator values, by area, as pairs of value and uncertainty maps
- Area-averaged indicator values (referred to as “indicator index” in the documentation from NI2020), as part of the time-series plots for scaled indicator values
The area-averaged indicator values are also required to be fed back into the NI database as a separate table that can be used for corresponding visualizations on the webpage.
2.1 General setup
There are a few variables that are used to control execution of parts of the code in this chapter. We start off by defining these here:
## Indicator focal years
ind_focalYears <- c(1990, 1995, 2000, 2005, 2010, 2014, 2019, 2024)
## Toggle for using/ignoring reference value uncertainty in visualizations
#uncertainRefValue <- TRUE
uncertainRefValue <- FALSE
## Toggle for visualizing scaled values truncated before or after summarising
truncAfterSummary <- TRUE
#truncAfterSummary <- FALSEAdditionally, we will define and set a seed to ensure reproducibility of the simulations:
2.2 Connect to the Nature Index Database
To access and download indicator data, we need to retrieve a token from the NI database. For that to work, we need access to our NI database access credentials; we retrieve those from environmental variables (if you have not saved them as environmental variables, you can find the code for doing that in chapter 0; alternatively, you can also manually set them here):
After retrieving login credentials, we can open a connection / request a token:
2.3 List indicators and retrieve data
If the person running this script has full access to the database (admin rights), they should be able to access the complete list of indicators by directly pulling it from the database using NIcalc::getIndicators():
indicatorList <- NIcalc::getIndicators() %>%
dplyr::arrange(name)
saveRDS(indicatorList, file = "data/NI2025_IndicatorList.rds")Alternatively, you can read in the saved version of the indicator list that comes with this repository:
The list is contains a total of 203 indicators, and is ordered alphabetically:
## id name
## 1 3 Alge på bjørk
## 2 250 Alke
## 3 5 Alm
## 4 7 Atlantisk høgmyr areal
## 5 50 Aure
## 6 9 Bananslørsopp
## 7 10 Begerfingersopp
## 8 11 Begroing elver eutrofierings indeks
## 9 233 Begroing elver forsurings indeks
## 10 12 Berggylte
## 11 399 Bjørkefink
## 12 23 Blåbær
## 13 16 Blåkveite
## 14 18 Blåskjell
## 15 390 Blåstrupe
## 16 20 Blåtopp i fuktig kystlynghei
## 17 343 Bløtbunn artsmangfold fauna hav
## 18 21 Bløtbunn artsmangfold fauna kyst
## 19 22 Bløtbunn eutrofiindeks
## 20 398 Bokfink
## 21 282 Boltit
## 22 27 Brosme
## 23 29 Brunbjørn
## 24 338 Brunmyrak
## 25 31 Bunndyr elver forventningsamfunn
## 26 228 Bunndyr-eutrofieringsindeks (ASPT)
## 27 231 Bunndyr-forsuringsindeks (Raddum 1)
## 28 351 Dagsommerfugler i skog
## 29 416 Dagsommerfugler i åpent lavland
## 30 340 Dikesoldogg
## 31 410 Dompap
## 32 314 Duetrost
## 33 36 Dyreplankton artssammensetning
## 34 38 Edelkreps
## 35 41 Eldre lauvsuksesjon (MiS)
## 36 42 Elg
## 37 43 Elvemusling
## 38 211 Engmarihand
## 39 150 Engvokssopper
## 40 301 Enkeltbekkasin
## 41 49 Fiolgubbe
## 42 268 Fiskemåke ferskvann
## 43 242 Fiskemåke kyst
## 44 284 Fjellerke
## 45 52 Fjellrev
## 46 53 Fjellrype
## 47 54 Fjellvalmue
## 48 286 Fjellvåk
## 49 392 Flaggspett
## 50 28 Flekkhvitkjuke
## 51 270 Fossekall
## 52 110 Fossenever
## 53 407 Fuglekonge
## 54 60 Gamle trær (MiS)
## 55 434 Gammel skog
## 56 62 Gaupe
## 57 408 Gjerdesmett
## 58 303 Gluttsnipe
## 59 317 Granmeis
## 60 318 Gransanger
## 61 66 Greplyng
## 62 414 Grå fluesnapper
## 63 245 Gråmåke
## 64 68 Grønlandssel
## 65 69 Grønn fåresopp
## 66 72 Grønngylte
## 67 394 Grønnspett
## 68 304 Grønnstilk
## 69 305 Gulerle
## 70 335 Gulsanger
## 71 412 Hagesanger
## 72 74 Hardbunn vegetasjon algeindeks
## 73 75 Hardbunn vegetasjon nedre voksegrense
## 74 287 Havelle
## 75 234 Havhest
## 76 235 Havsule
## 77 257 Havørn
## 78 288 Heilo
## 79 289 Heipiplerke
## 80 77 Hjort
## 81 359 Humler i skog
## 82 417 Humler i åpent lavland
## 83 337 Hvitmyrak
## 84 84 Hyse
## 85 319 Hønsehauk
## 86 85 Issoleie
## 87 86 Isterviersumpskog
## 88 400 Jernspurv
## 89 88 Jerv
## 90 89 Jordstjerner
## 91 90 Jordtunger
## 92 438 Karplanter i semi-naturlig eng
## 93 92 Klappmyss
## 94 95 Kolmule
## 95 322 Kongeørn
## 96 98 Kopperrød slørsopp
## 97 385 Krikkand
## 98 247 Krykkje
## 99 100 Kusymre
## 100 101 Kveite
## 101 386 Kvinand
## 102 94 Køllesopper
## 103 104 Laks - kyst og elver
## 104 360 Laks i havet
## 105 105 Lange
## 106 106 Lappkjuke
## 107 290 Lappspurv
## 108 439 Lavhei i villreinkommuner
## 109 108 Liggende død ved (MiS) – arealandel
## 110 419 Liggende død ved – mengde
## 111 109 Lirype
## 112 111 Lodde
## 113 251 Lomvi
## 114 253 Lunde
## 115 324 Løvsanger
## 116 115 Makrell
## 117 248 Makrellterne
## 118 326 Munk
## 119 307 Myrsnipe
## 120 125 Myrtelg
## 121 325 Måltrost
## 122 396 Nøtteskrike
## 123 130 Olavsstake
## 124 384 Orrfugl
## 125 134 Oter ferskvannsbestand
## 126 135 Oter kystbestand
## 127 142 Palsmyr areal
## 128 145 Planteplankton (Chl a)
## 129 146 Planteplankton innsjøer
## 130 252 Polarlomvi
## 131 149 Praktrødspore
## 132 151 Prestekrage
## 133 152 Purpurlyng
## 134 389 Ravn
## 135 156 Reker hav
## 136 409 Ringdue
## 137 391 Ringtrost
## 138 433 Rogn-Osp-Selje
## 139 153 Rådyr
## 140 249 Rødnebbterne
## 141 308 Rødstilk
## 142 406 Rødstjert
## 143 402 Rødstrupe
## 144 395 Rødvingetrost
## 145 165 Sei
## 146 167 Sennegras
## 147 298 Siland
## 148 369 Sild - havgående bestander
## 149 440 Sildemåke
## 150 310 Sivspurv
## 151 339 Smalsoldogg
## 152 370 Smågnagere - fjellbestander
## 153 332 Smågnagere - skogbestander
## 154 275 Smålom
## 155 311 Småspove
## 156 291 Snøspurv
## 157 180 Solblom
## 158 292 Steinskvett
## 159 333 Stokkand
## 160 186 Storfugl
## 161 241 Storjo
## 162 277 Storlom
## 163 187 Storpiggslekten (sopp)
## 164 236 Storskarv ssp carbo
## 165 237 Storskarv ssp sinensis
## 166 278 Strandsnipe
## 167 183 Stående død ved (MiS) – arealandel
## 168 418 Stående død ved – mengde
## 169 296 Svartand
## 170 246 Svartbak
## 171 328 Svarthvit fluesnapper
## 172 222 Svartkutling
## 173 403 Svartmeis
## 174 194 Svartnende kantarell
## 175 195 Svartsonekjuke
## 176 393 Svartspett
## 177 397 Svarttrost
## 178 196 Sveltstarr
## 179 312 Svømmesnipe
## 180 436 Tamrein
## 181 254 Teist
## 182 202 Tilstand kystlynghei
## 183 201 Tilstand semi-naturlig eng og strandeng
## 184 415 Tjeld
## 185 203 Tobis
## 186 279 Toppand
## 187 330 Toppmeis
## 188 239 Toppskarv
## 189 378 Torsk - havsbestander
## 190 313 Trane
## 191 405 Trekryper
## 192 401 Trepiplerke
## 193 207 Trær med hengelav (MiS)
## 194 209 Ulv
## 195 380 Ungsild (1-2 år)
## 196 208 Vanlig uer
## 197 213 Vannplanter innsjø
## 198 215 Vier alpint belte
## 199 216 Villrein
## 200 1 Ål
## 201 293 Ærfugl
## 202 141 ØyepålNow that we have all indicators listed we can download the entire set of indicator data. This takes a while to run, and we therefore save the data import as a local file after it finished downloading so that we do not have to re-download the entire dataset repeatedly.
indicatorImport <- list()
for(i in 1:nrow(indicatorList)){
indicatorImport[[i]] <- NIcalc::importDatasetApi(
username = NIdb_username,
password = NIdb_password,
indic = indicatorList$name[i]
)
}
names(indicatorImport) <- indicatorList$id
saveRDS(indicatorImport, "data/indicatorImport.rds")The next step in the NI workflow is “assembling” the data. This step makes sure that the parameters characterizing uncertainty distributions for indicator values are present for all data points (see documentation of NIcalc::assembleNiObject() for more details). This also requires some time to run, and like the previous step, we will save the output so we can work directly from that afterwards.
indicatorData <- list()
for(i in 1:nrow(indicatorList)){
indicatorData[[i]] <- NIcalc::assembleNiObject(
inputData = indicatorImport[[i]],
predefNIunits = c(allArea = TRUE, parts = TRUE, counties = FALSE),
indexType = "thematic"
)
}
names(indicatorData) <- indicatorList$id
saveRDS(indicatorData, "data/indicatorData_assembled.rds")2.4 Simulate indicator value distributions
For visualizing indicator values (both original and scaled), we need associated measures of uncertainty. While all indicator value have such information associated in the database, the format thereof varies, i.e. some indicators have lower and upper quartiles in addition to means reported, while others are specified via custom distributions.
We require a standardized way for presenting values including uncertainty across all indicators. In previous iterations of NI, it seems that results were displayed as median plus 95% confidence interval (= 2.5% and 97.5% quantiles, according to NINA Report 1990, “Nature Index System Documentation”). While working with alternative approaches to visualizing indicator data, we also experimented with using coefficient of variation (= relative standard deviation = sd/mean) for visualizing uncertainty in maps (see here https://ninanor.github.io/NIviz/maps-1.html), and displays of the entire indicator value distributions (e.g. 5.1 here: https://ninanor.github.io/NIviz/other-figures.html).
Based on this, we have a need for a) different measures of uncertainty and b) samples of draws from the uncertainty distributions of all indicators. We can obtain all of this information from simulations. The function simulate_IndicatorData() can be used to run the corresponding simulations for all indicators and years with available data:
## Source functions
source("R/simulate_IndicatorData.R")
source("R/scale_IndicatorValues.R")
## Make folder for storing data as files
if(!dir.exists("data/simulated_IndicatorData")){
dir.create("data/simulated_IndicatorData")
}
## Set up list for storing all results
simData <- list()
## For all indicators, simulate data from distributions and save as files and in list
for(i in 1:nrow(indicatorList)){
message("")
message(crayon::bold(crayon::green(paste0(indicatorList$name[i]))))
simData_i <- simulate_IndicatorData(indData = indicatorData[[i]])
simData[[i]] <- simData_i
saveRDS(simData_i, file = paste0("data/simulated_IndicatorData/simData_id_", indicatorList$id[i], ".rds"))
message("")
}
## Save total list containing all datasets
saveRDS(simData, file = "data/simulated_IndicatorData/simData_all.rds")2.5 Summarize indicator value distributions
In the next step, we want to calculated some summary metrics for our indicator data.
The helper function summarise_IndicatorDist() takes the raw and scaled indicator datasets and calculates the following metrics for each year- and area-specific indicator value:
- Mean
- Median
- Standard deviation
- Relative standard deviation
- 2.5%, 5%, 25%, 75%, 95%, and 97.5% quantiles
We first source the function, and then run it on each of the three datasets (raw, scaled with fixed reference, scaled with sampled reference) available for all indicators. The results we will save as local .rds and .csv files.
## Source function
source("R/summarise_IndicatorDist.R")
## Make folder for storing data as files
if(!dir.exists("data/IndicatorData_StatSummaries")){
dir.create("data/IndicatorData_StatSummaries")
}
savePath <- "data/IndicatorData_StatSummaries/"
## Re-set seed
set.seed(mySeed)
## Summarise indicator value distributions for all datasets and indicators
for(i in 1:nrow(indicatorList)){
message(crayon::bold(crayon::green(paste0(indicatorList$name[i]))))
# Raw (unscaled data)
summarise_IndicatorDist(indSim = simData[[i]]$raw,
ind_id = indicatorList$id[i],
dataset_name = "raw",
savePath = savePath)
# Scaled using fixed reference
summarise_IndicatorDist(indSim = simData[[i]]$scaled_fixedRef,
ind_id = indicatorList$id[i],
dataset_name = "scaled_fixedRef",
savePath = savePath)
# Scaled using sampled reference
summarise_IndicatorDist(indSim = simData[[i]]$scaled_sampledRef,
ind_id = indicatorList$id[i],
dataset_name = "scaled_sampledRef",
savePath = savePath)
message("")
}2.6 Calculating indicator indices
Traditionally, time-series of area-aggregated indicator values have been published for a selection of indicators on the NI webpage. These are referred to as “indicator indices” and are calculated using the same framework as the NI for different ecosystems (via NIcalc::calculateIndex()). We calculate these here using the wrapper function calculate_IndicatorIndex().
The source code underlying this function originates from scripts from NI2020 stored on NINA’s internal servers under “R:/Prosjekter/12892000_naturindeks_rammeverk_database_og_innsynslosning/Naturindeks 2020/Beregninger/Indikatorer/”.
Back then, indicator indices were calculated on a per-ecosystem basis, likely to circumvent the issue that no alternative had been coded into “NIcalc” yet. In calculate_IndicatorIndex() here, we have made adjustments that allow calculating indicator indices taking into account all relevant ecosystems; in practice, this means that area weights are determined by the extent of area within each spatial unit covered by any of the ecosystems relevant to the indicator (as opposed to just one selected one). It’s still possible to calculate indicator indices specific to single ecosystems too by setting weigh_byEcosystem = TRUE and specifying which ecosystem in the call to the function. However, for the purpose of visualizing indicator data, we will work with indicator indices calculated for all relevant ecosystems.
Below, we loop over all indicators to calculate indicator indices, and save the results into a list for further use in R and as files into a designated folder:
## Source function
source("R/calculate_IndicatorIndex.R")
## Create folder for saving results
if(!dir.exists("results_Indicators")){
dir.create("results_Indicators")
}
## Set up list for storing results
indIndex <- list()
## Calculate indicator indices and save results, per indicator
for(i in 1:nrow(indicatorList)){
indIndex_i <- calculate_IndicatorIndex(dataSet = indicatorData[[i]],
testRun = FALSE,
NAimputation = FALSE,
weigh_byEcosystem = FALSE,
include_subAreas = TRUE,
truncAtRef = TRUE,
enforce_weightSum1 = TRUE)
indIndex[[i]] <- indIndex_i
readr::write_excel_csv(indIndex_i$summary, file = paste0("results_Indicators/indIndex_Summary_id_", indicatorList$id[i], ".csv"))
saveRDS(indIndex_i, paste0("results_Indicators/indIndex_id_", indicatorList$id[i], ".rds"))
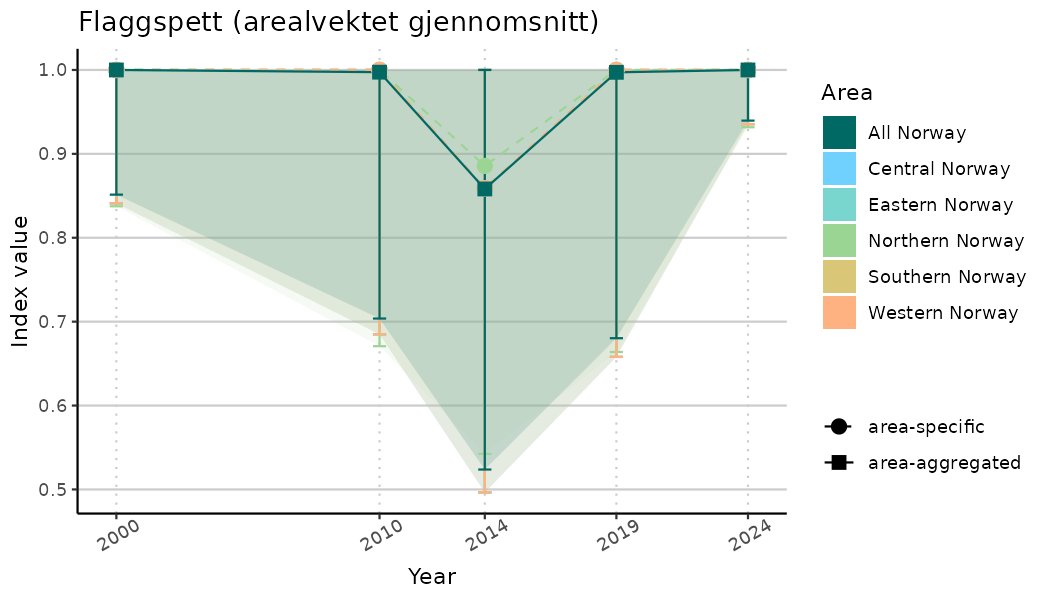
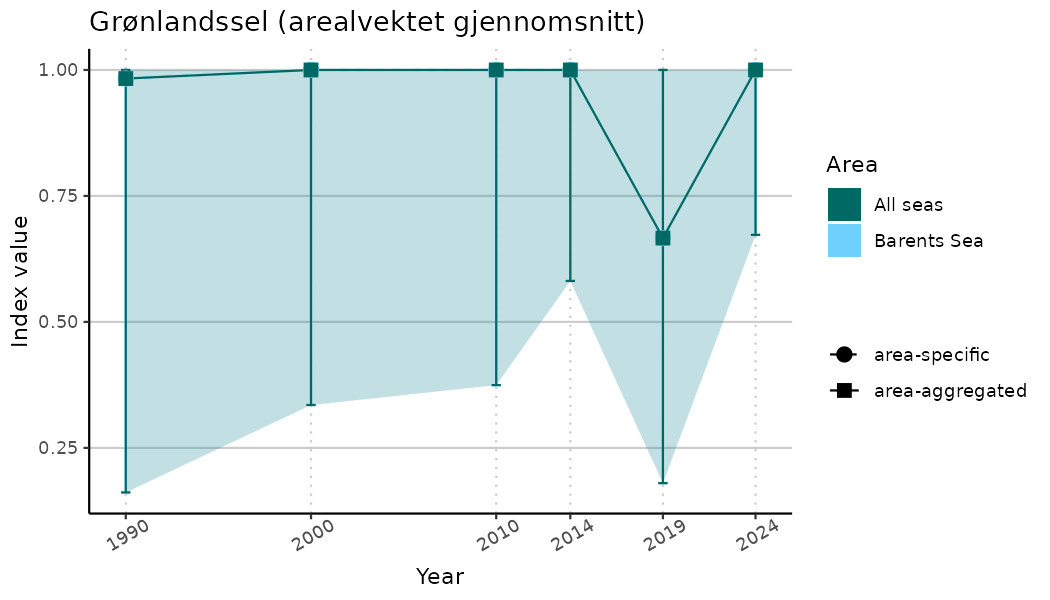
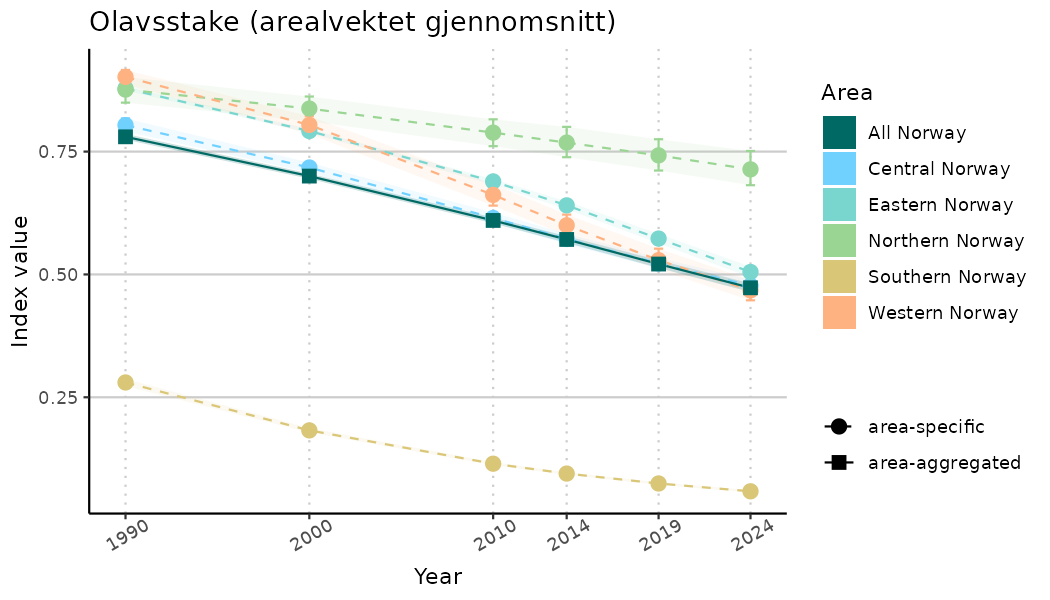
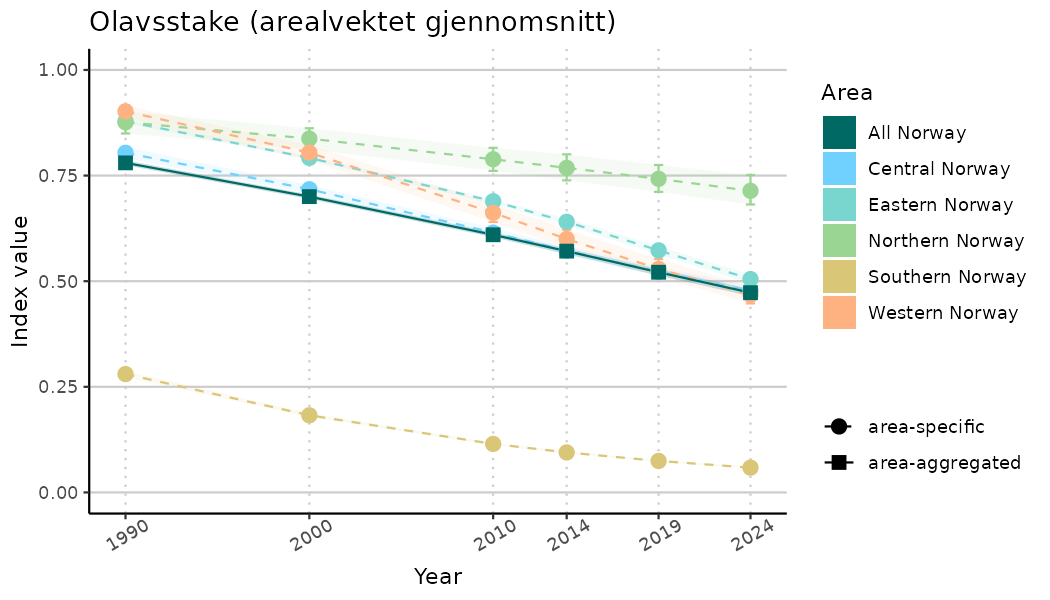
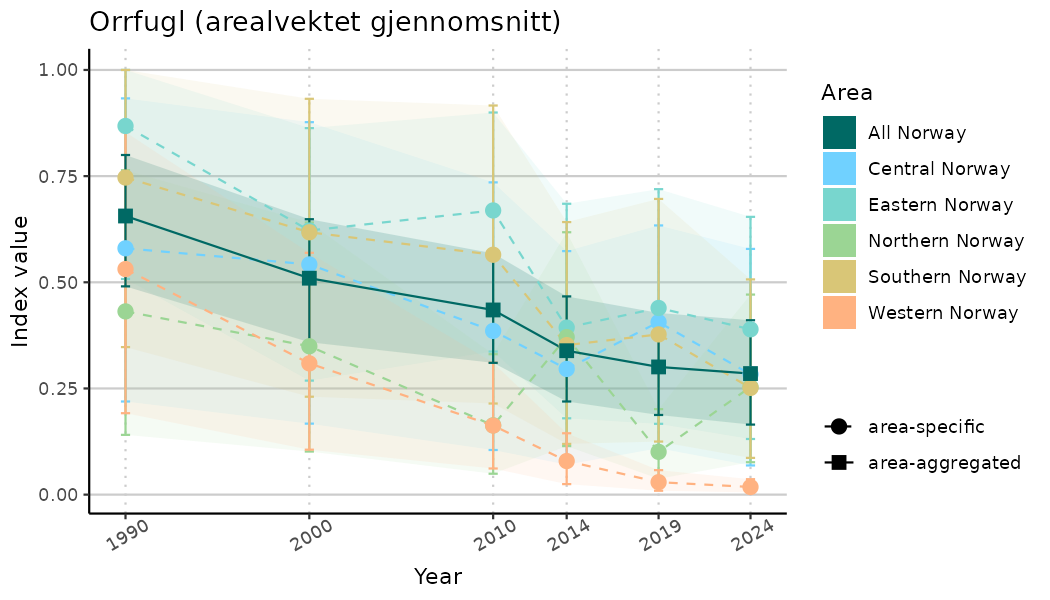
}Note that we are calculating indicator indices without imputations (NAimputation = FALSE). This is because imputation using the “mice” algorithm will not work very well for many indicators due to limited data and high collinearity. It works for indices combining many indicators because that translates to more (variable) data. Without imputations, indicator indices for selected years represent area-weighted means of values for the areas with available data in the specified year. Since different areas may have data available for different years, changes in calculated indicator indices over time have to be interpreted with caution as changes may indicate a) changes in average indicator values and/or b) changes in composition of areas with values. We assist this visually by not “connecting” the indicator index estimates across years below.
2.7 Visualizing raw and scaled indicator data
For the remainder of this chapter, we will work with one indicator at a time and read in the respective simulated datasets anew. This means we can remove the large objects containing data for all indicators simultaneously from the workspace:
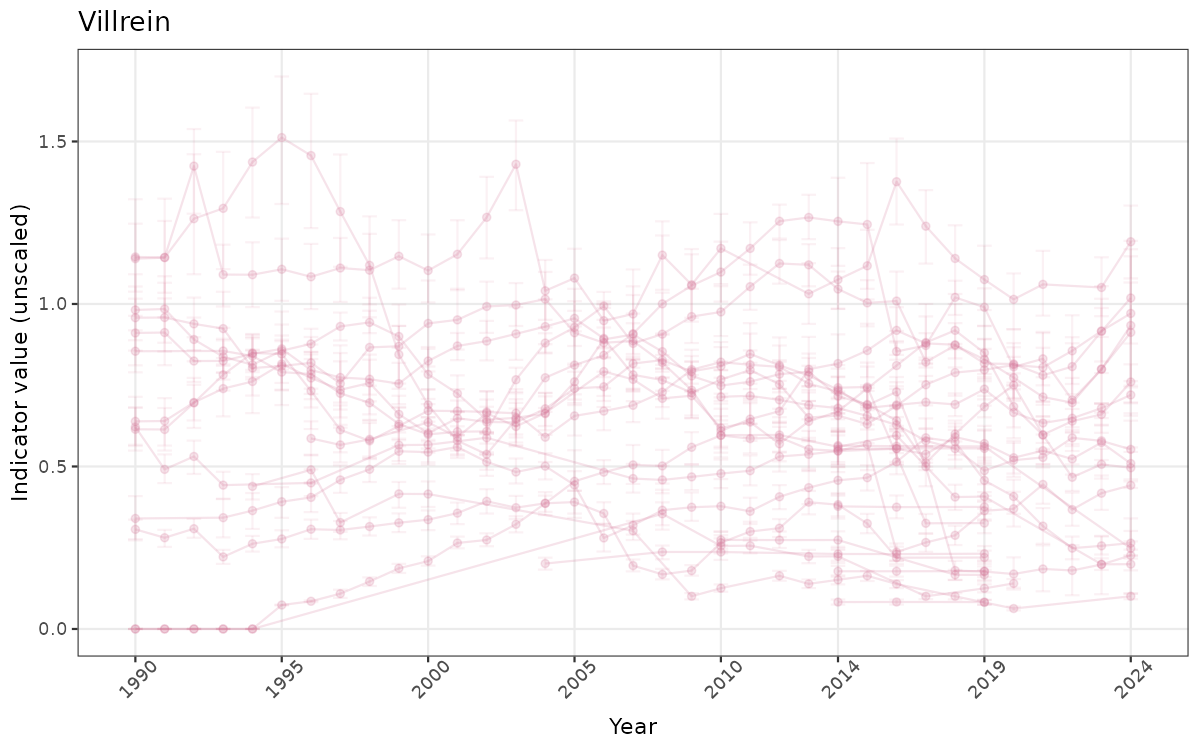
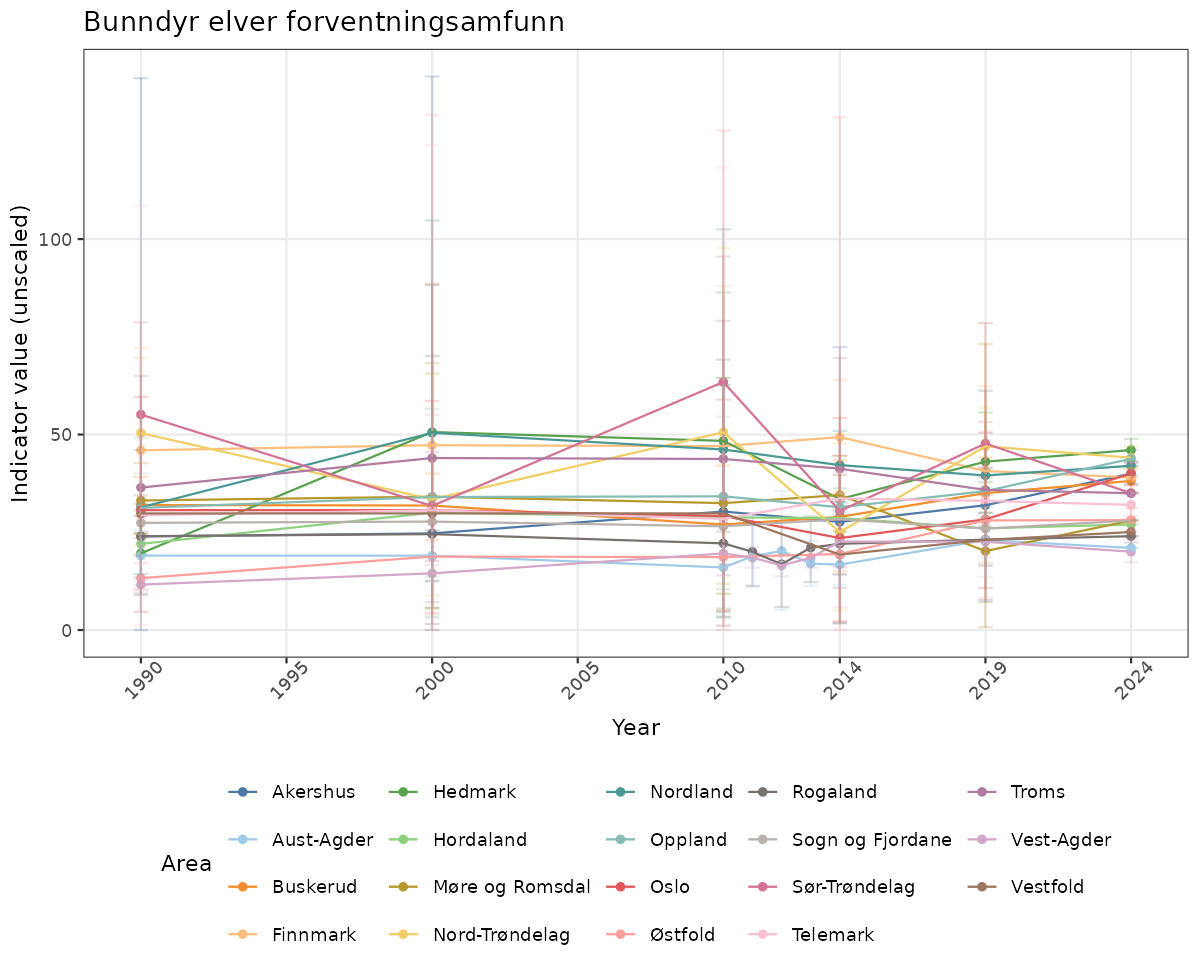
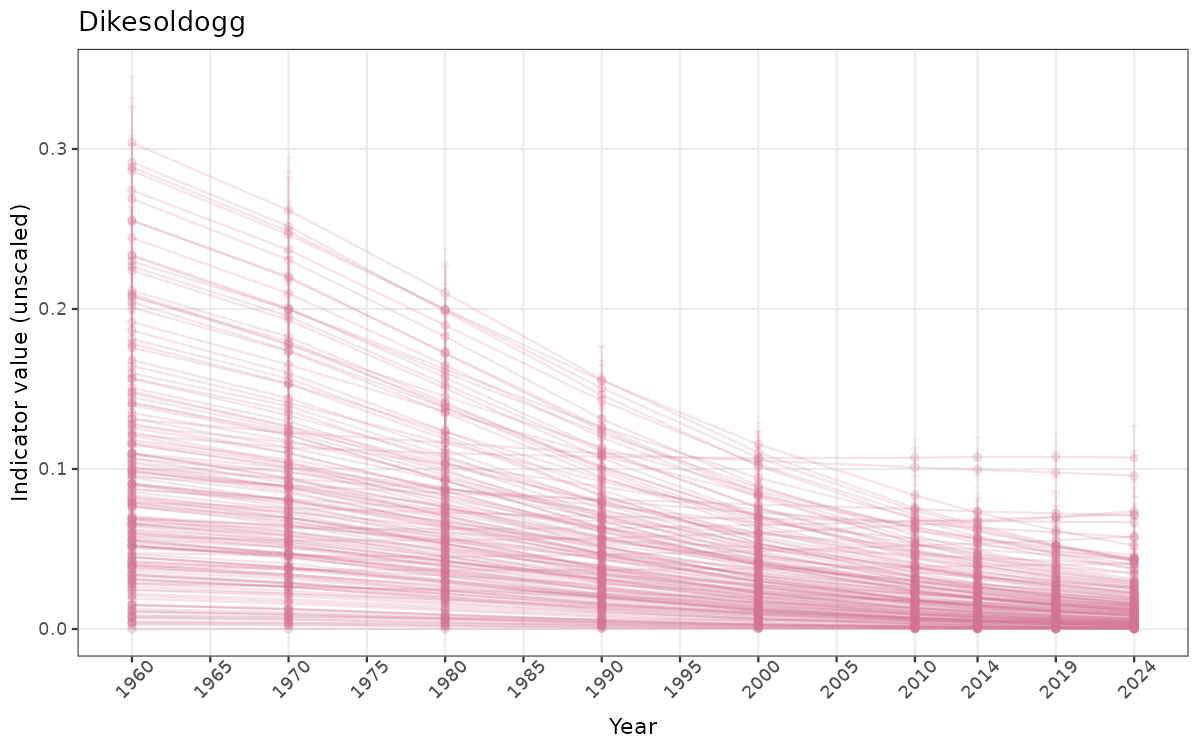
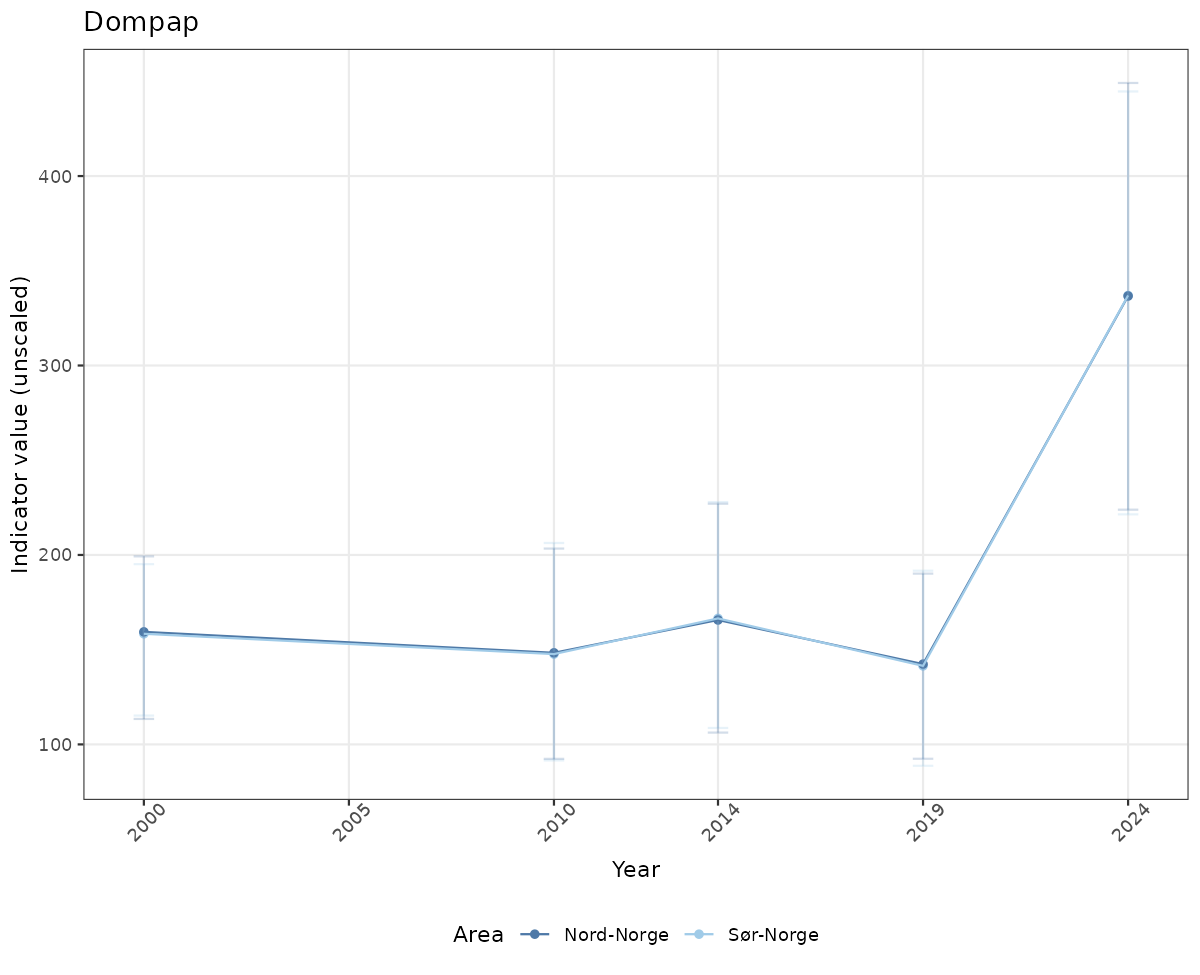
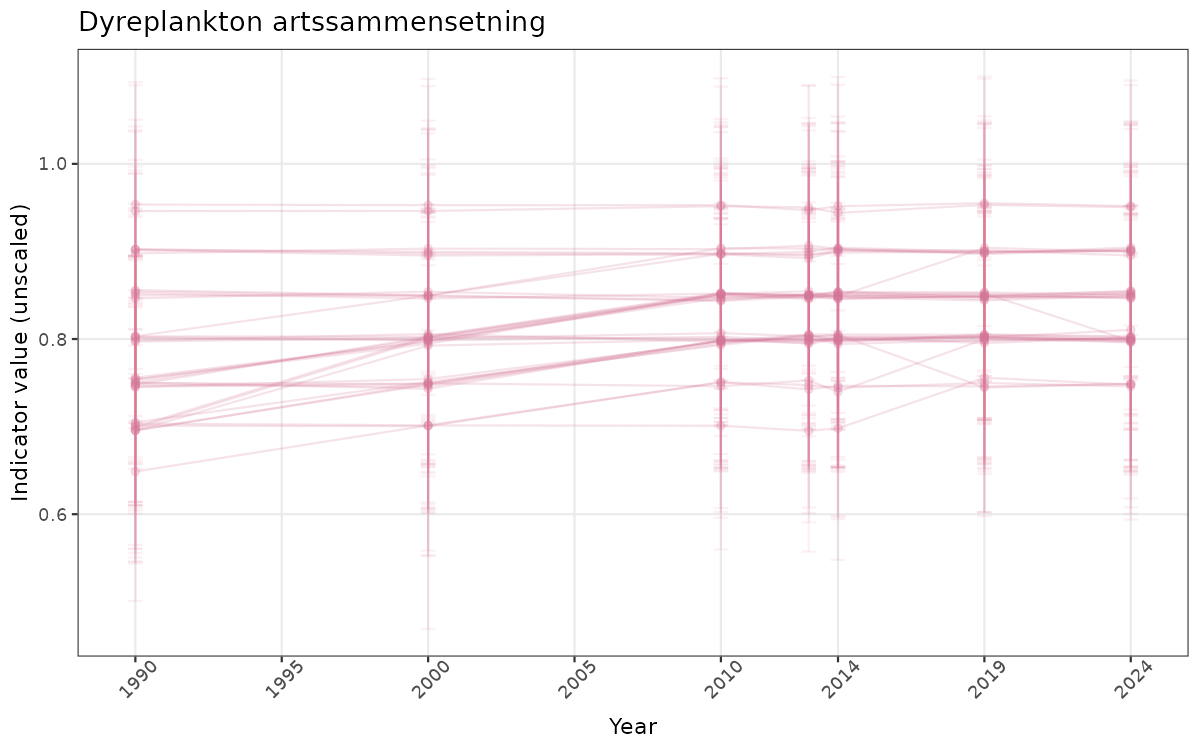
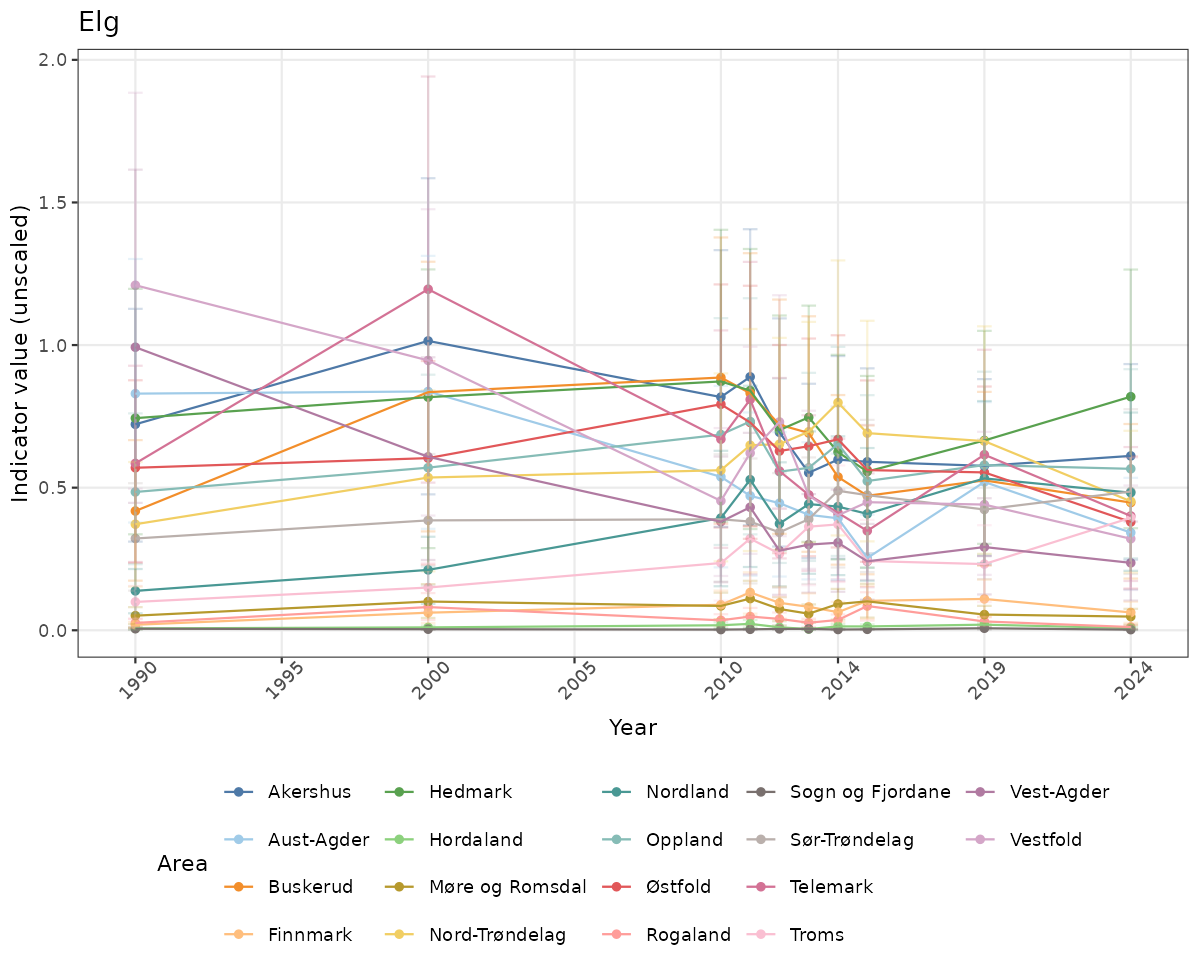
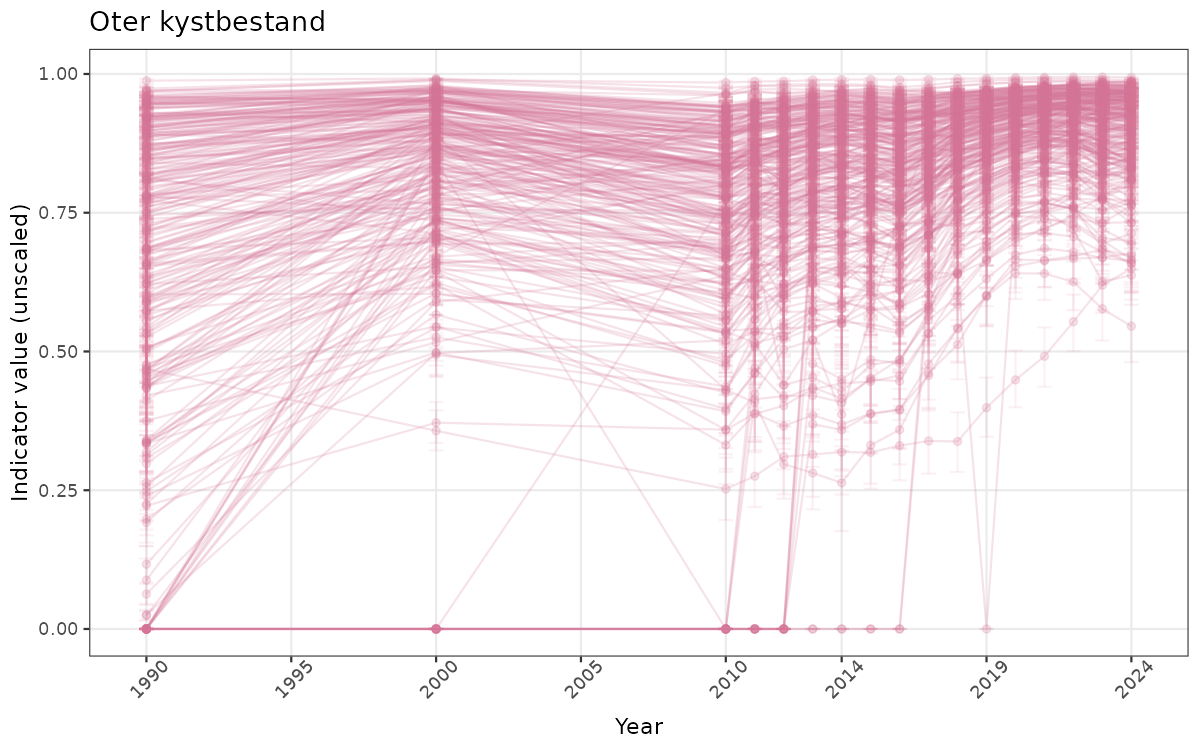
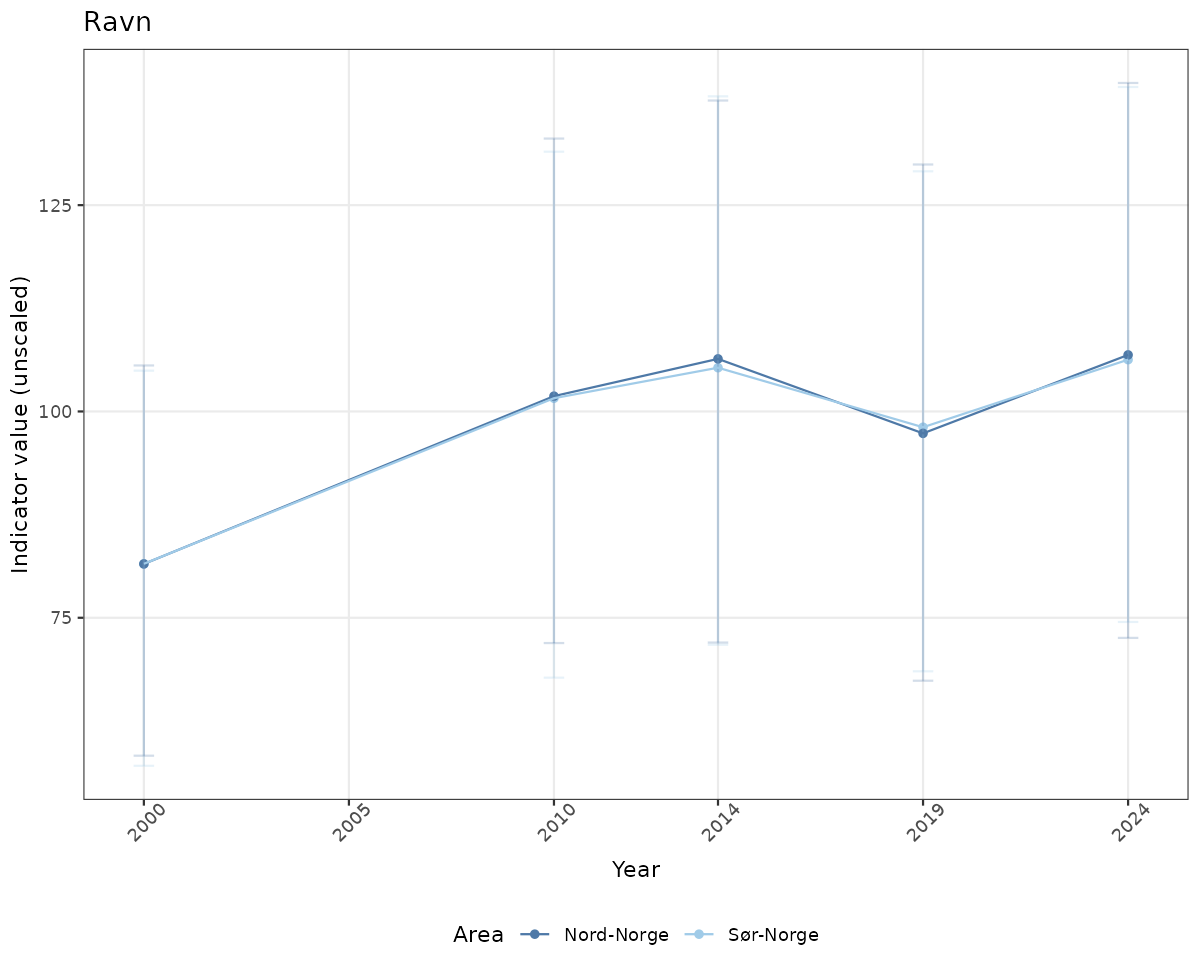
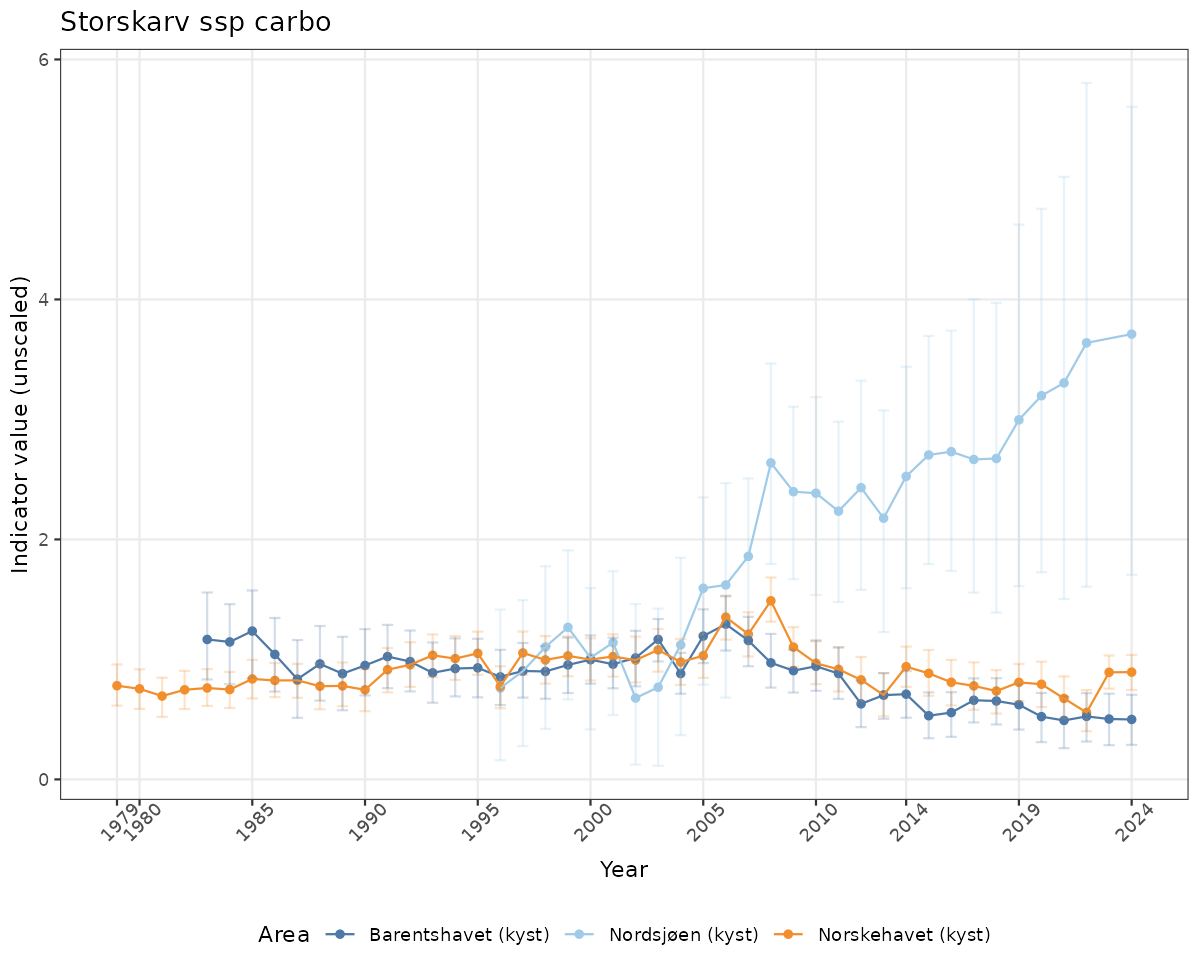
There are two types of visualizations of indicator data that we are aiming for:
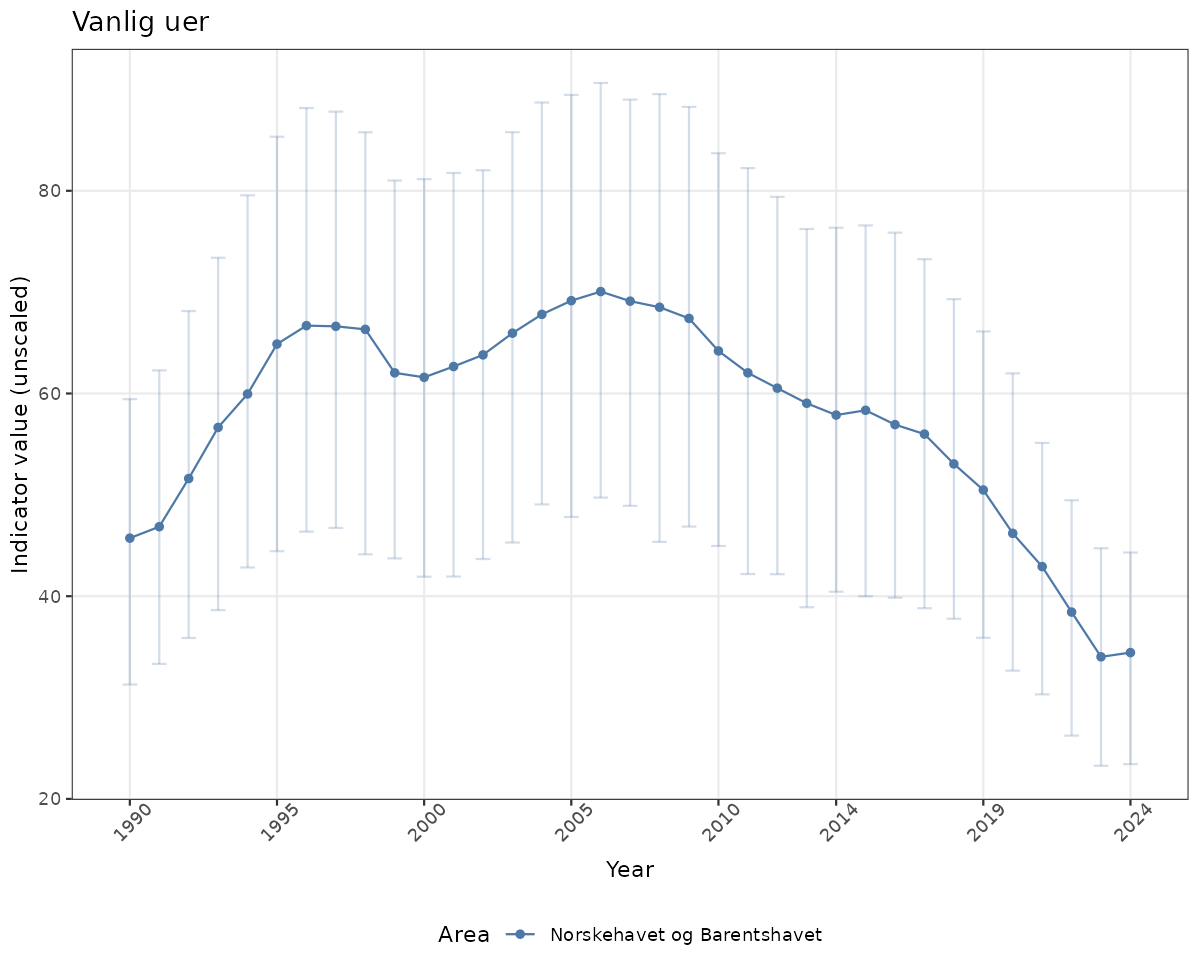
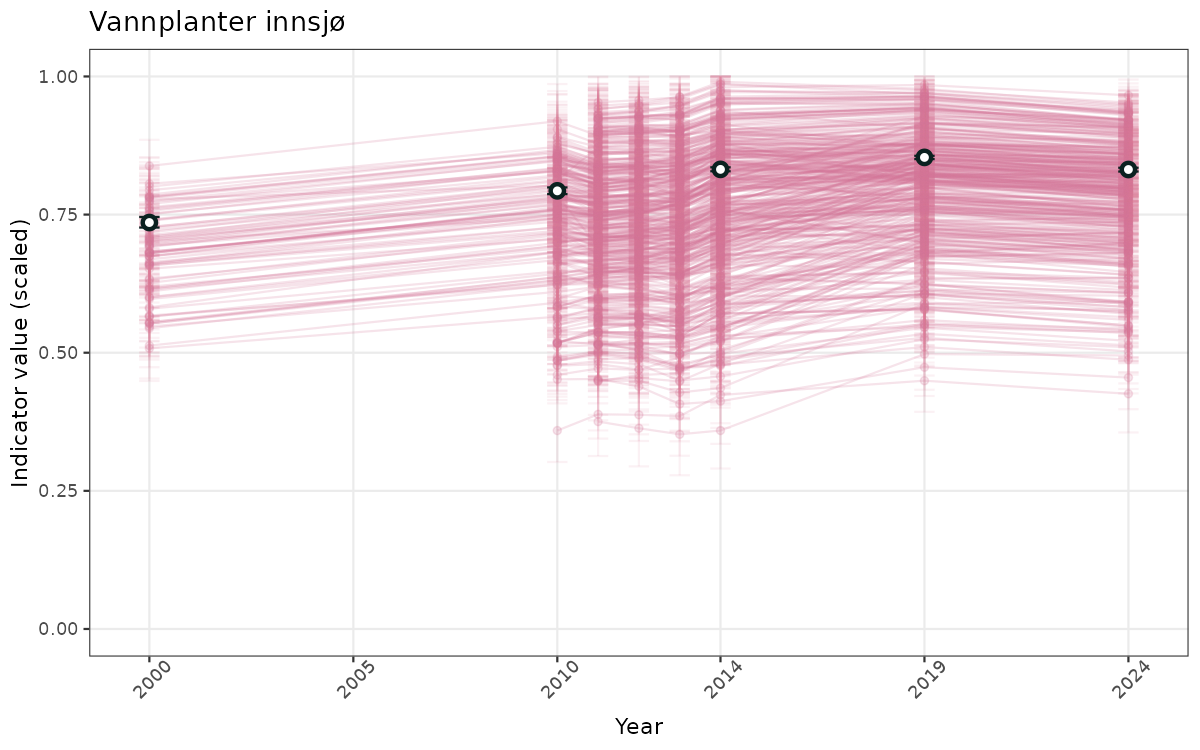
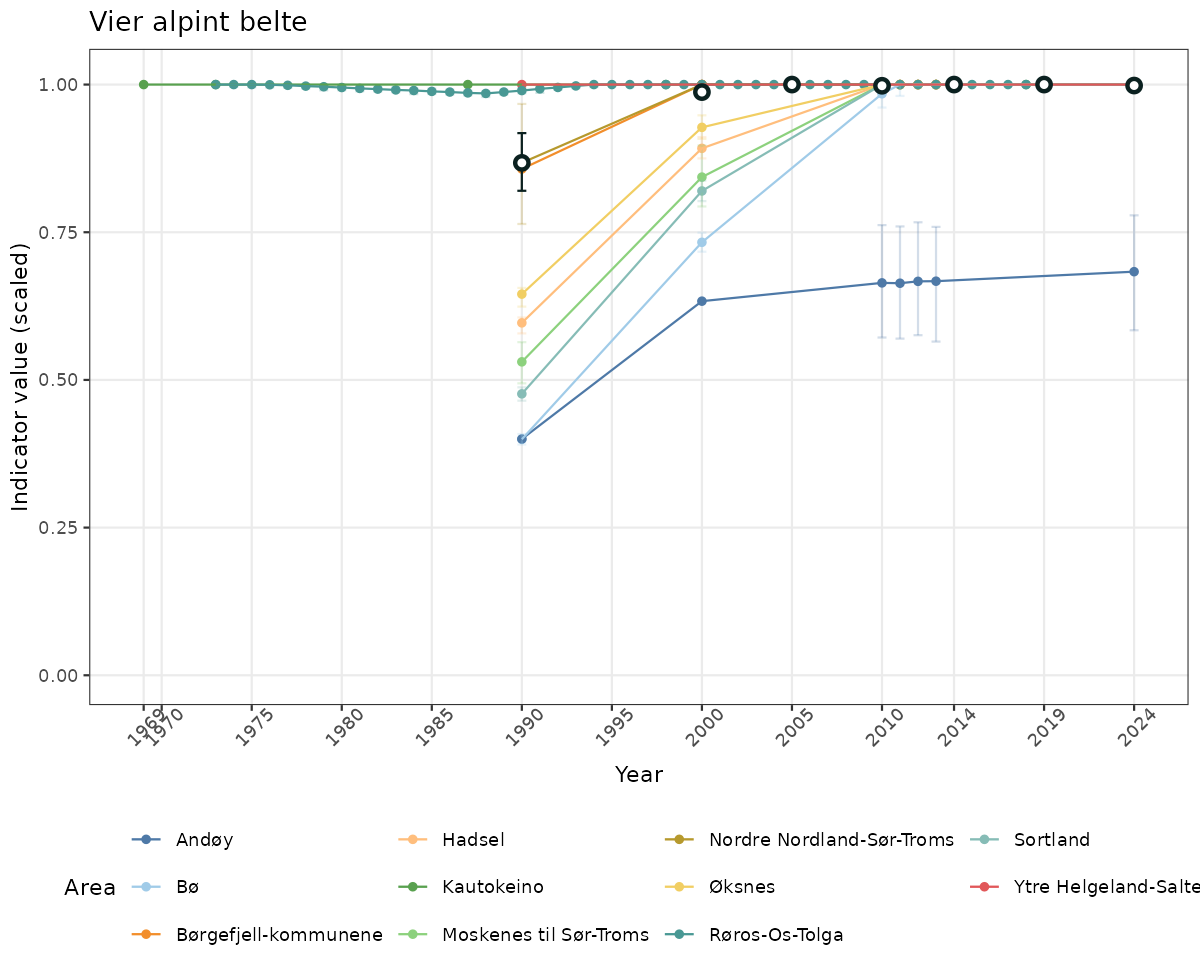
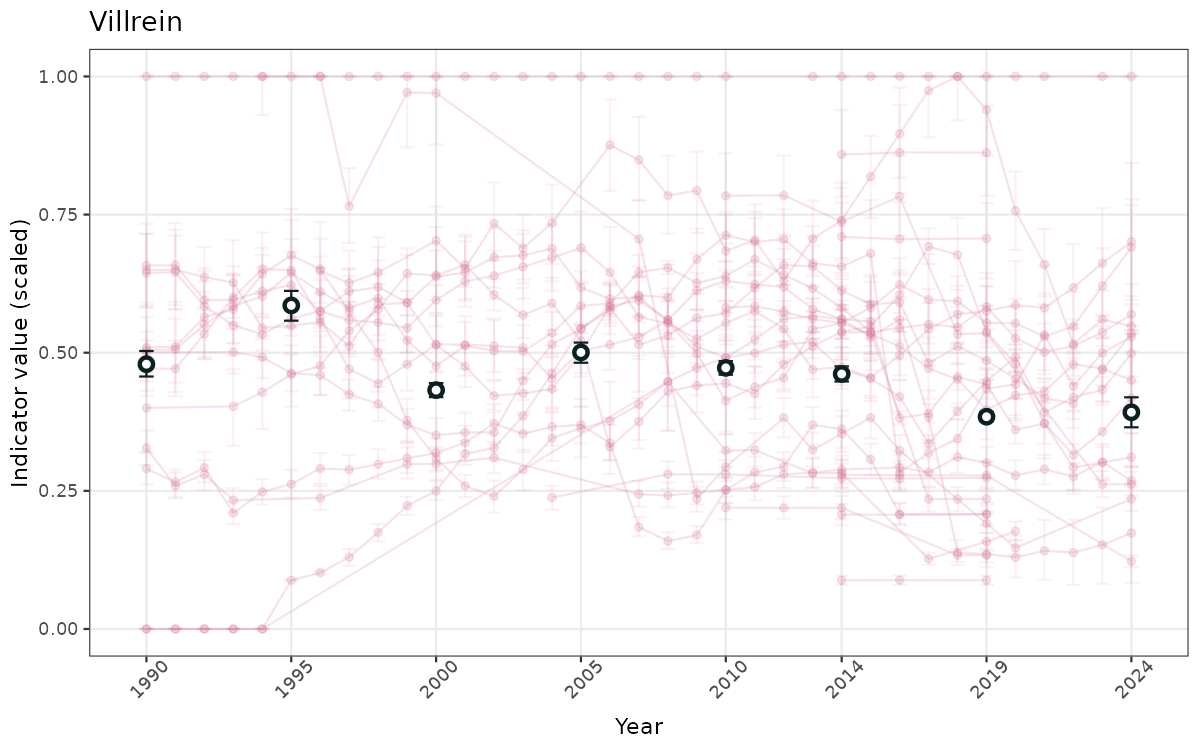
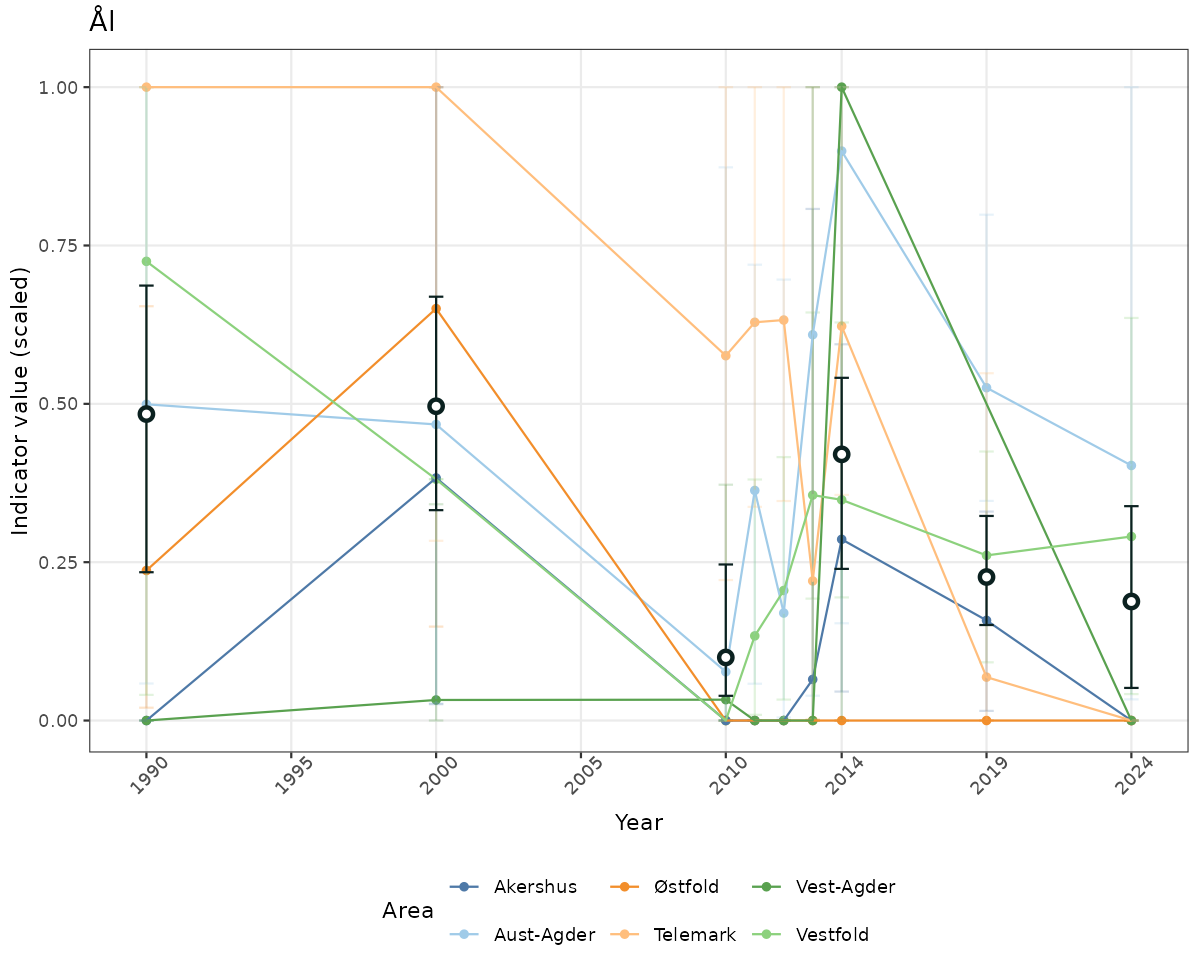
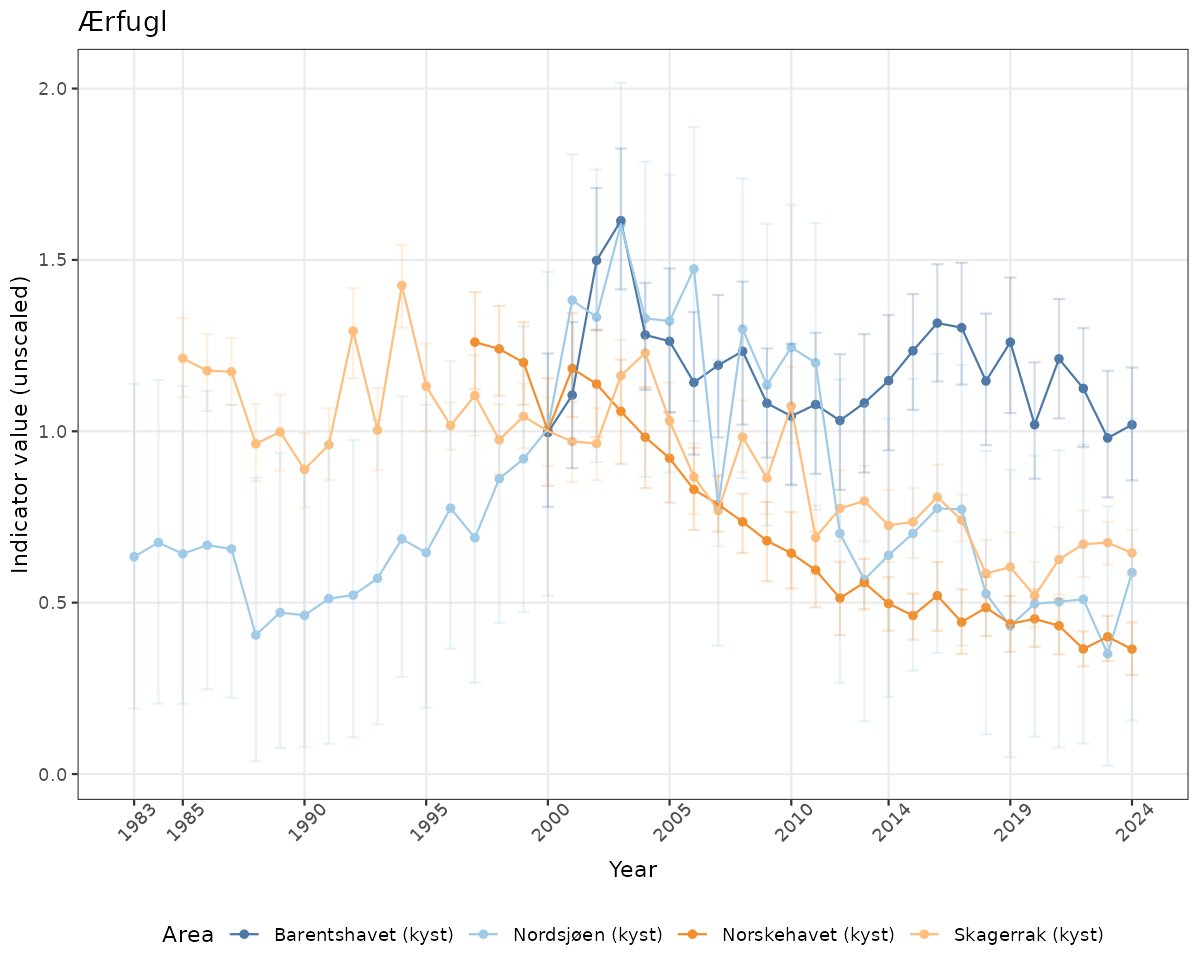
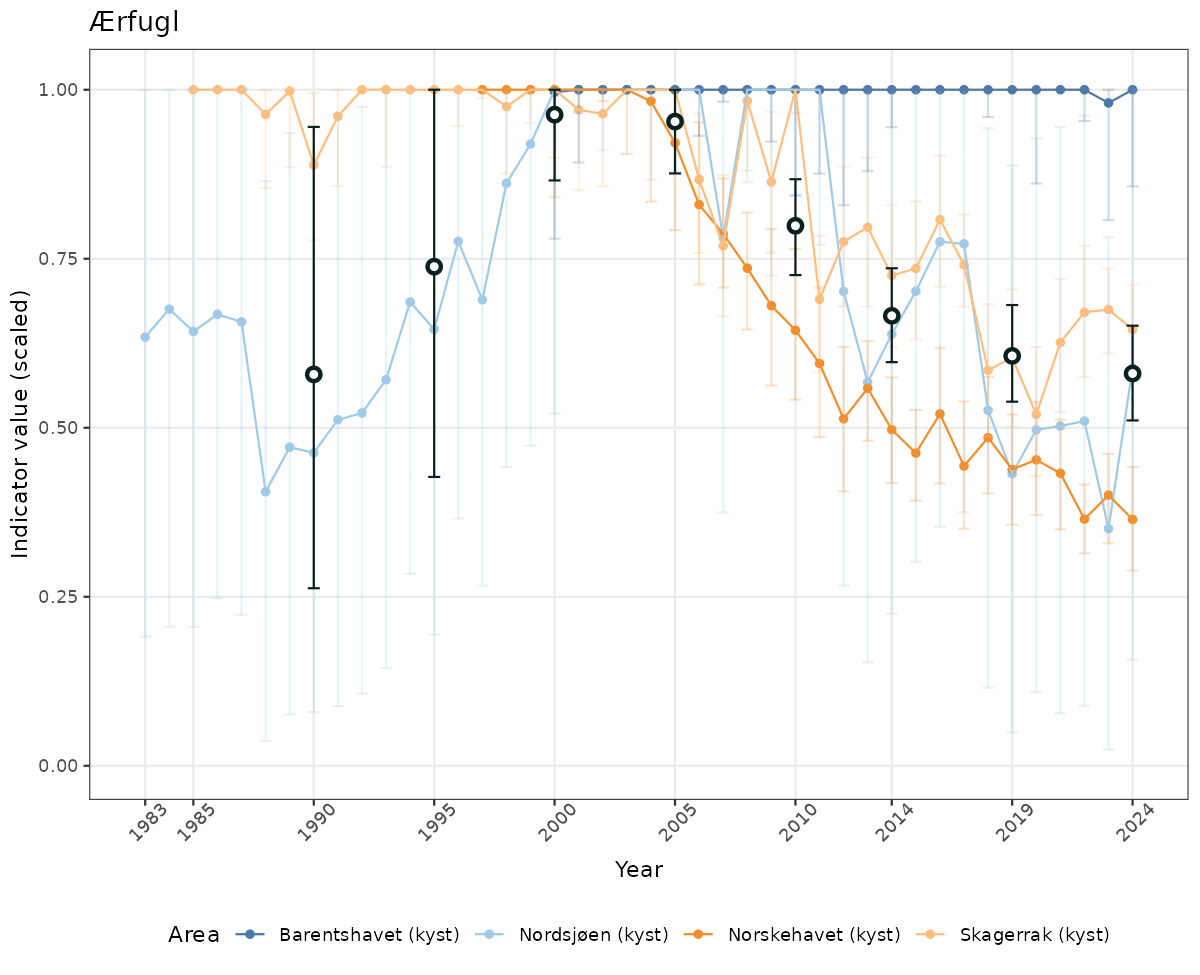
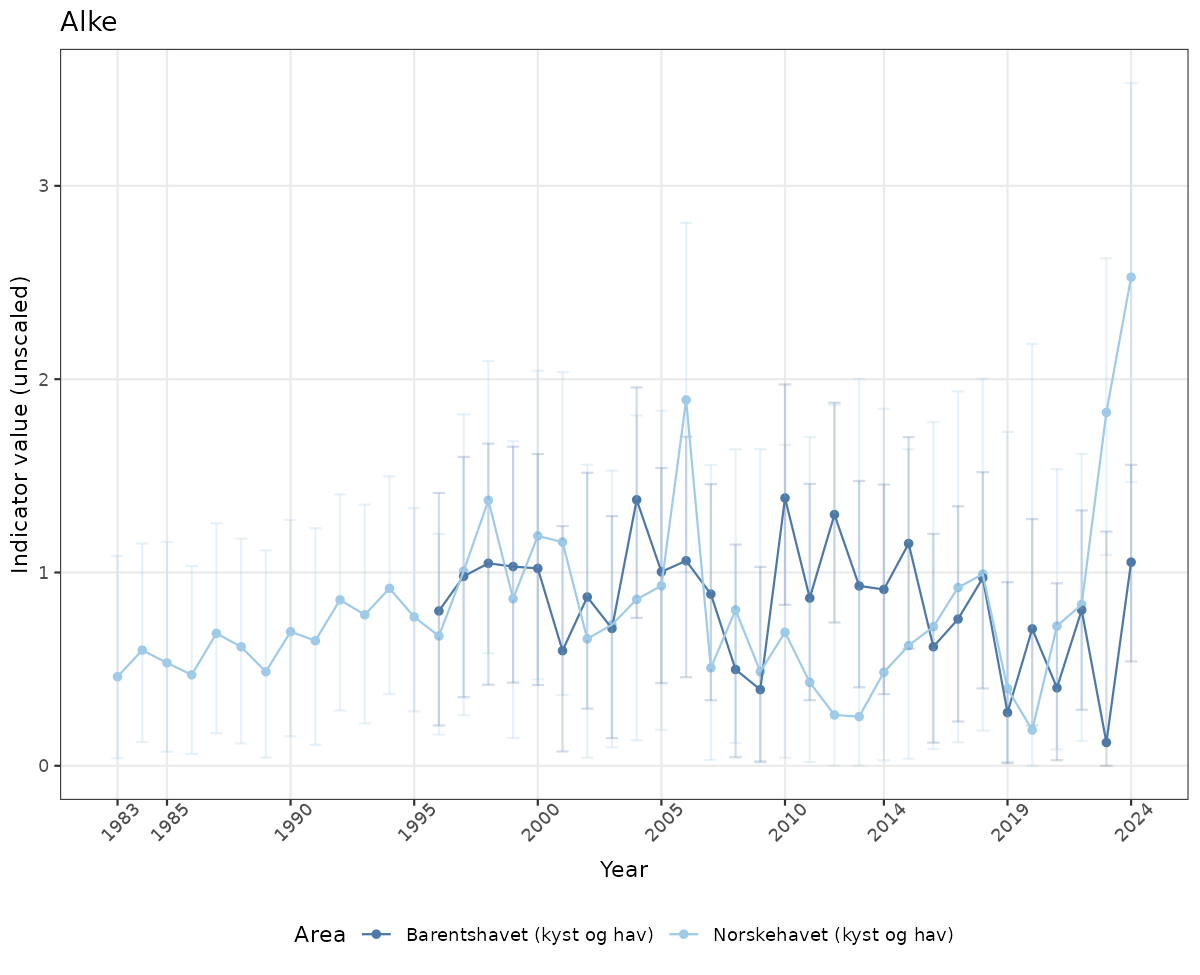
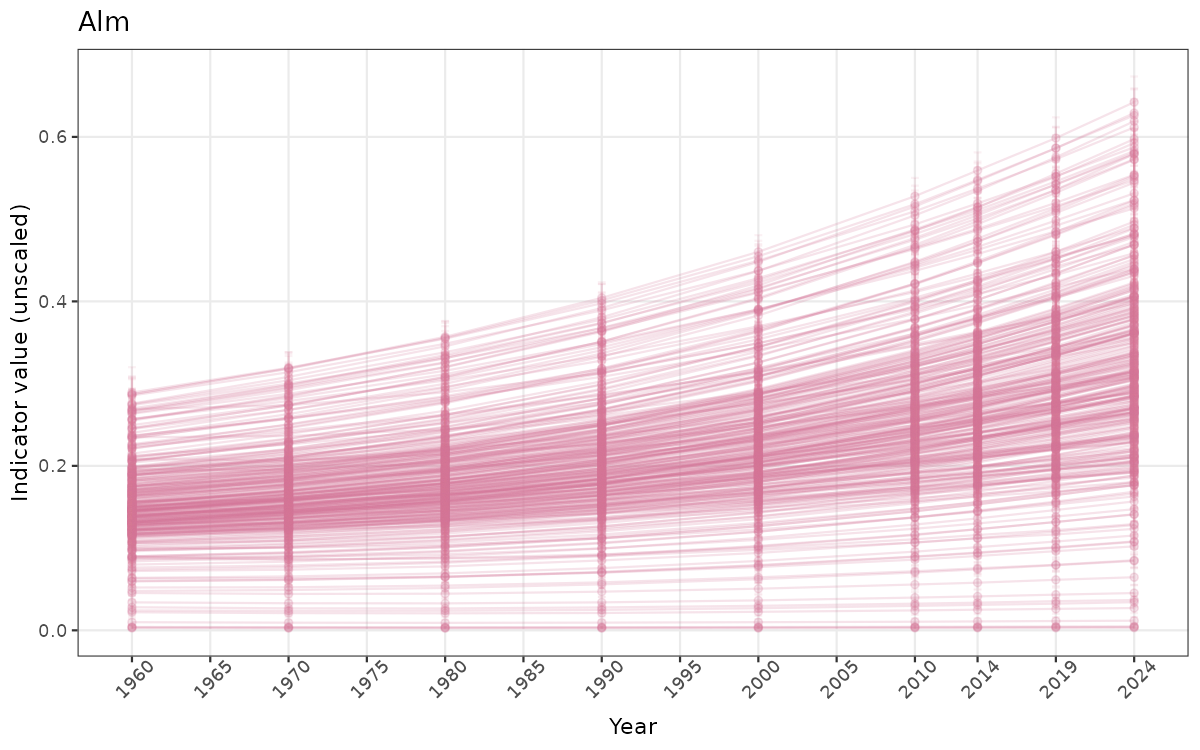
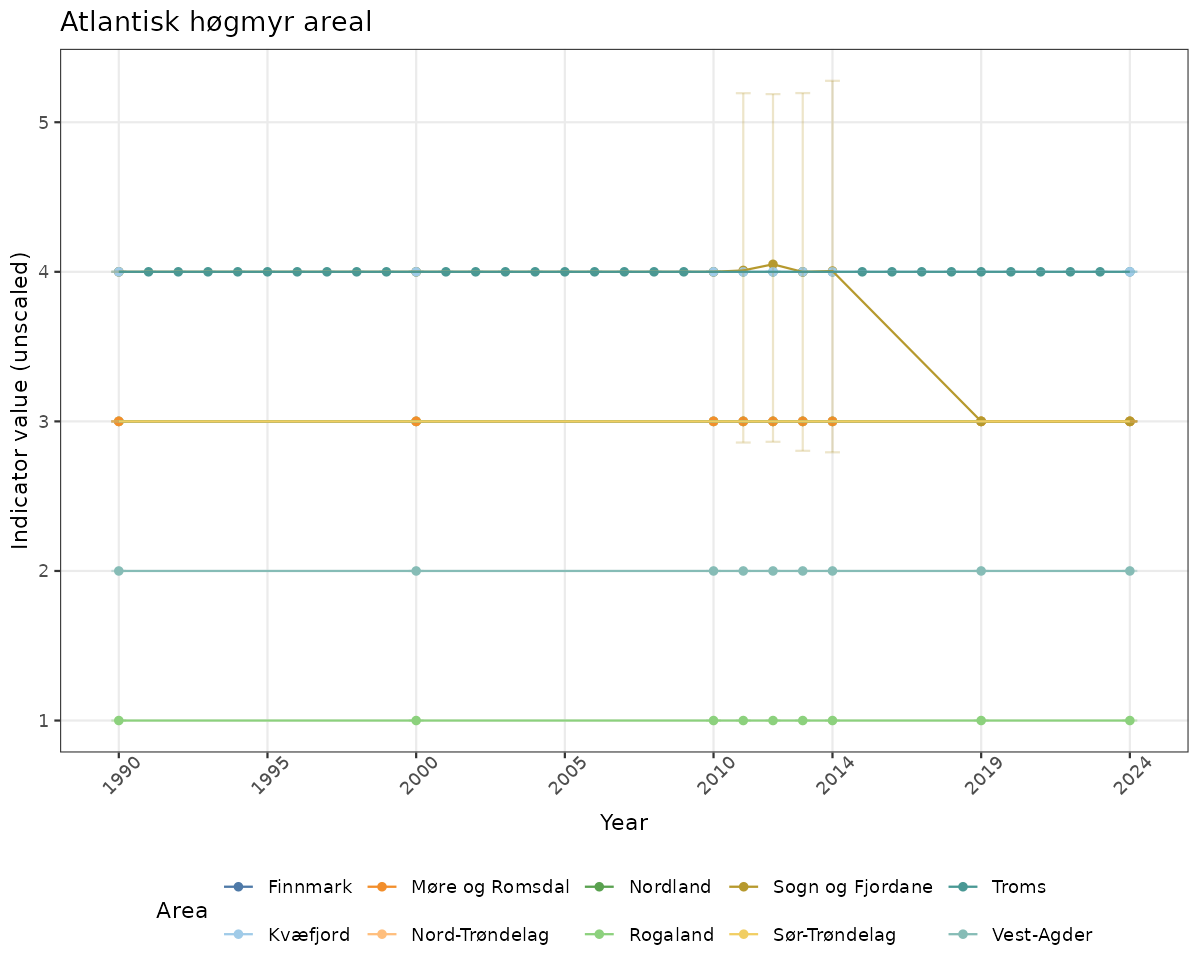
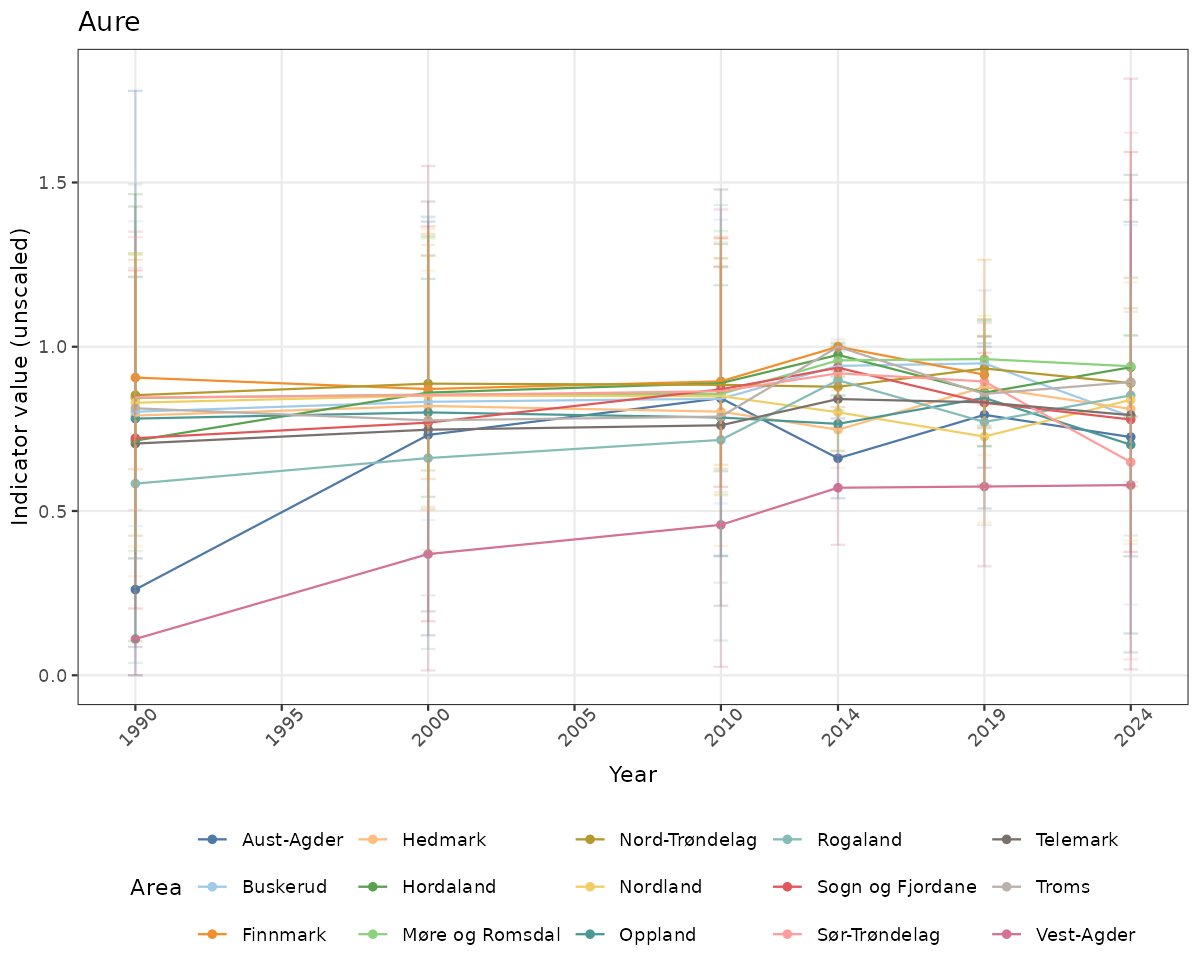
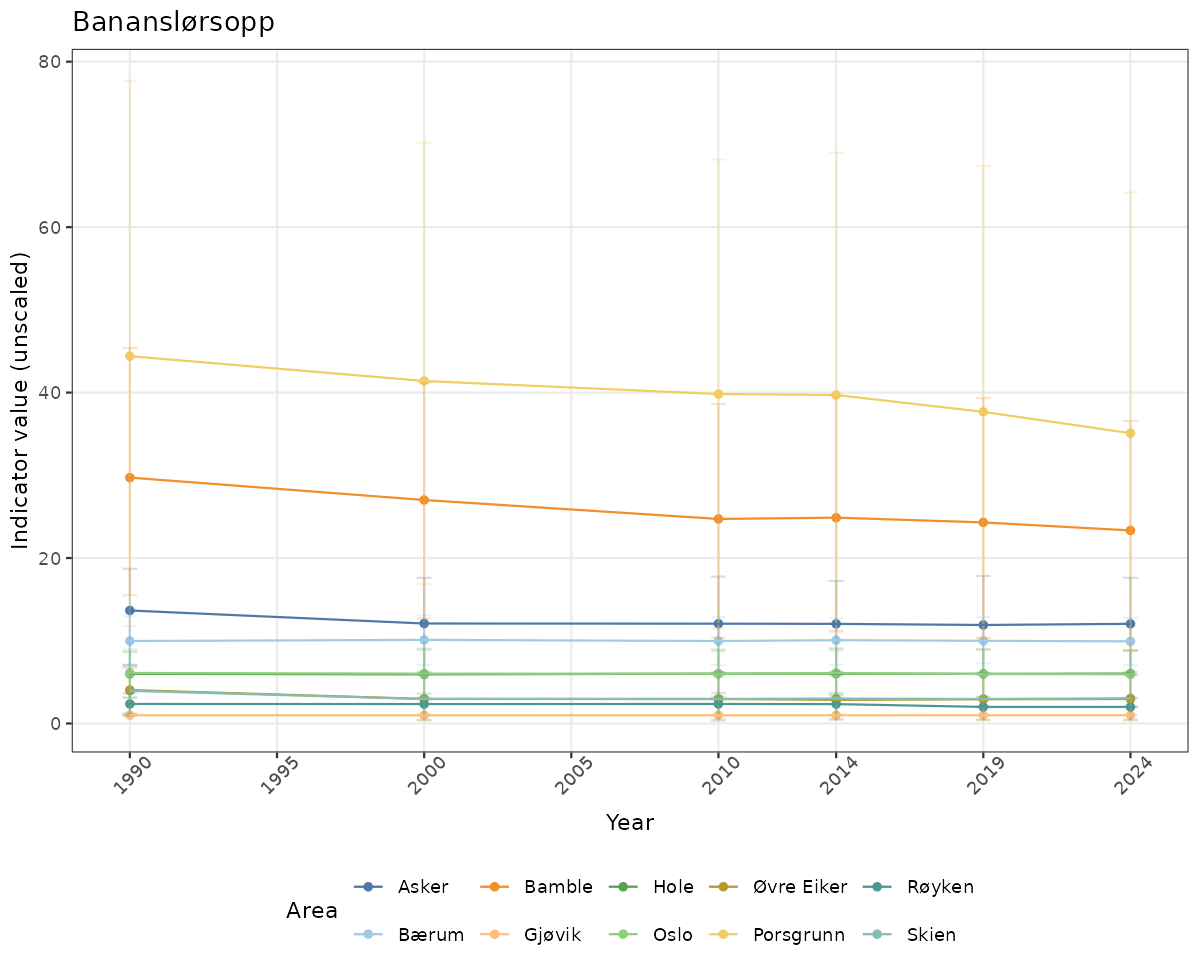
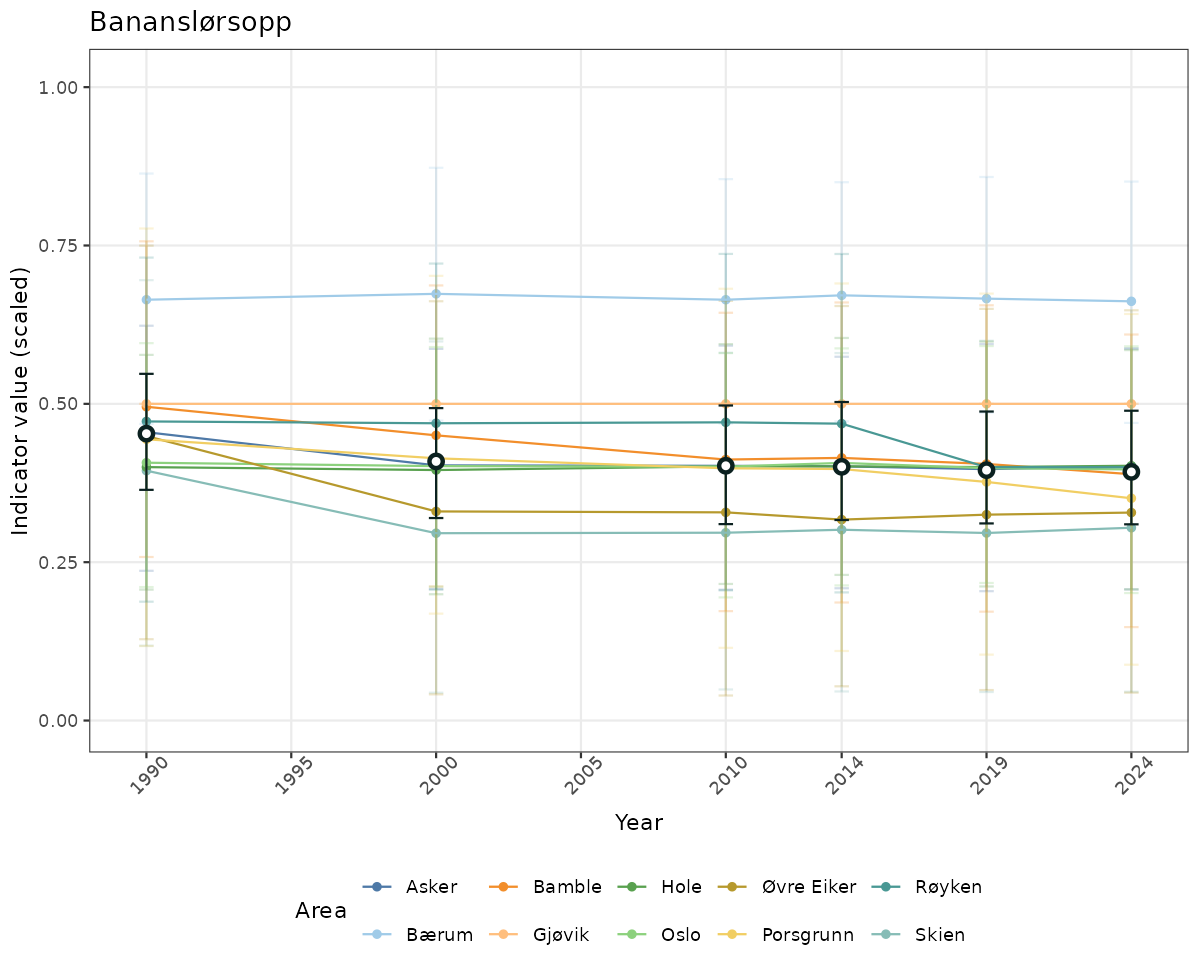
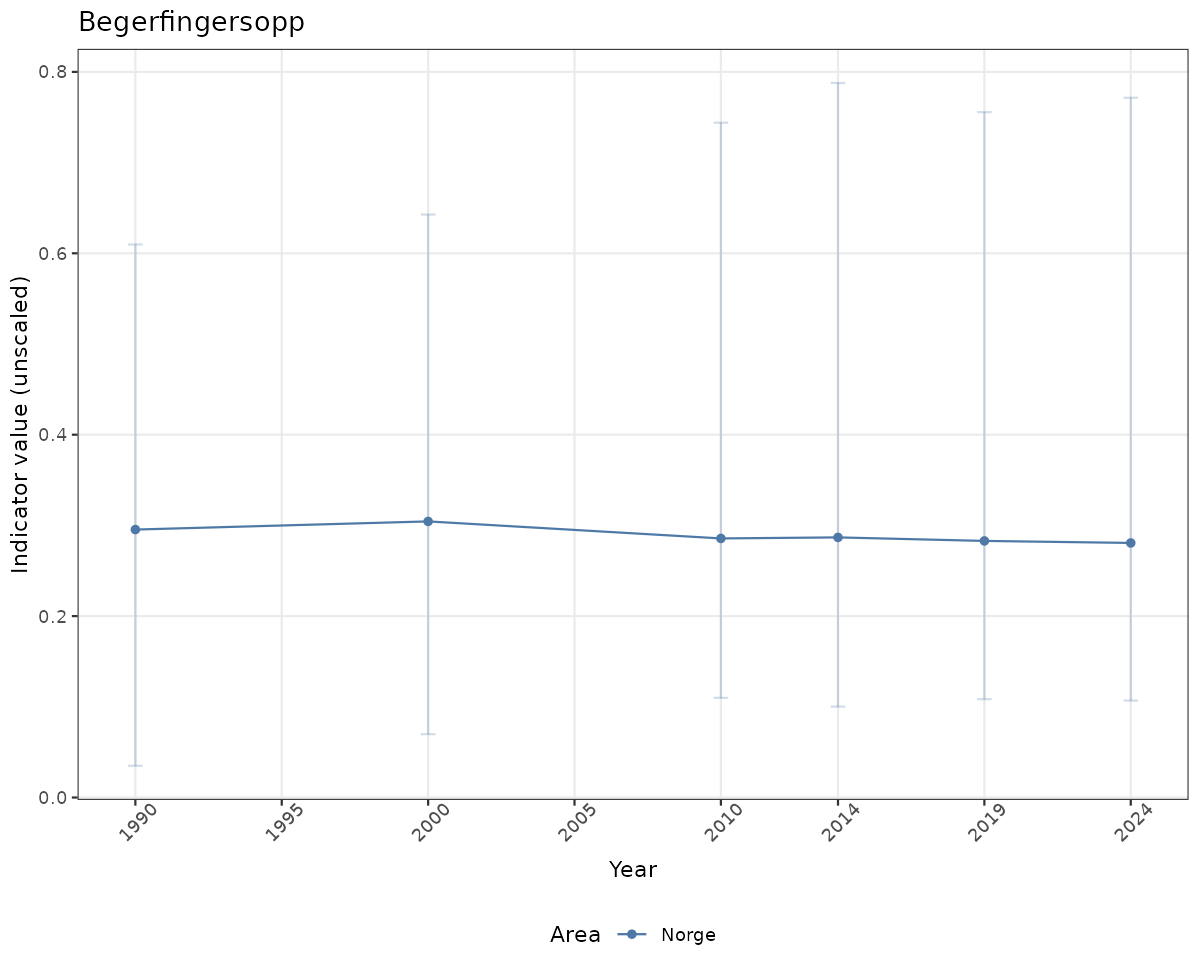
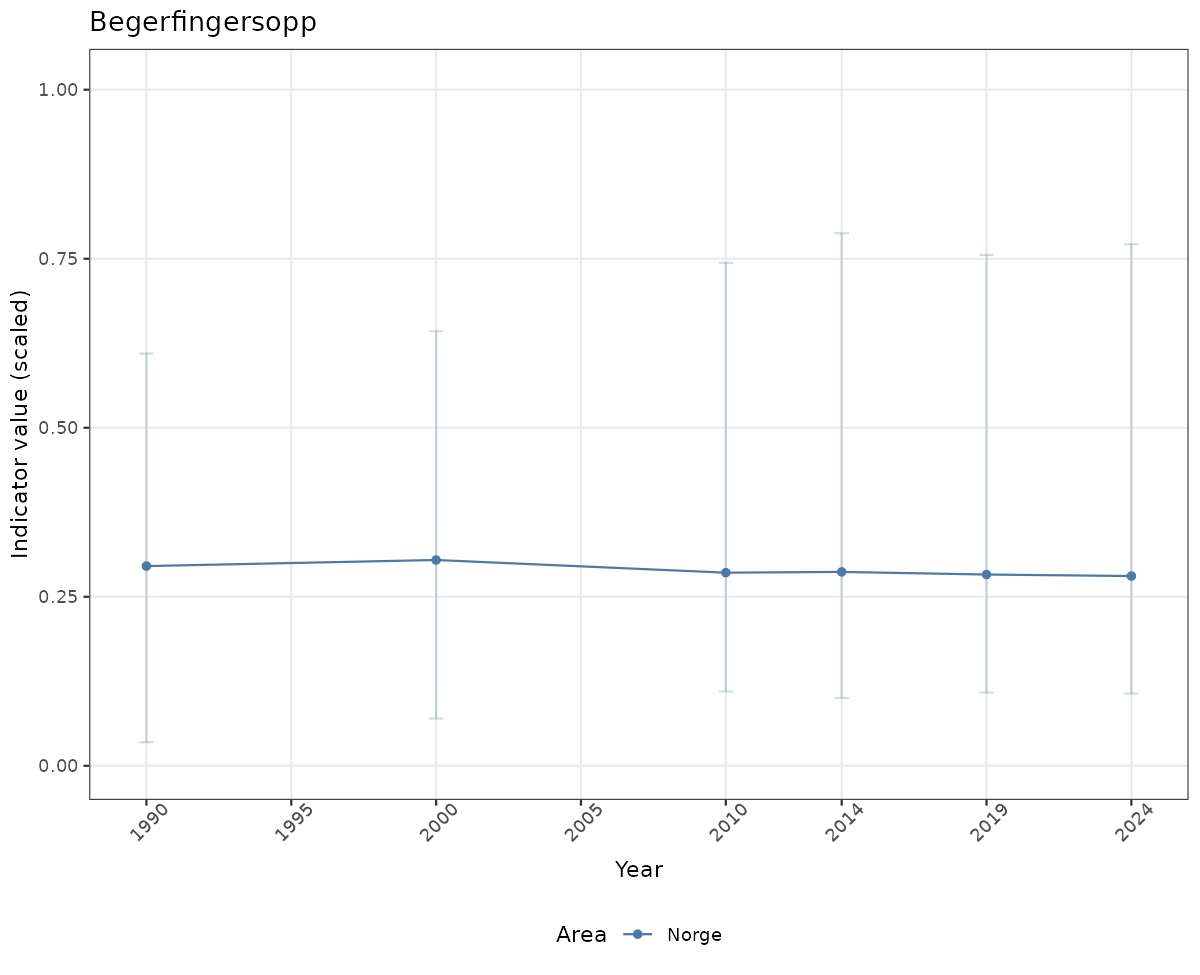
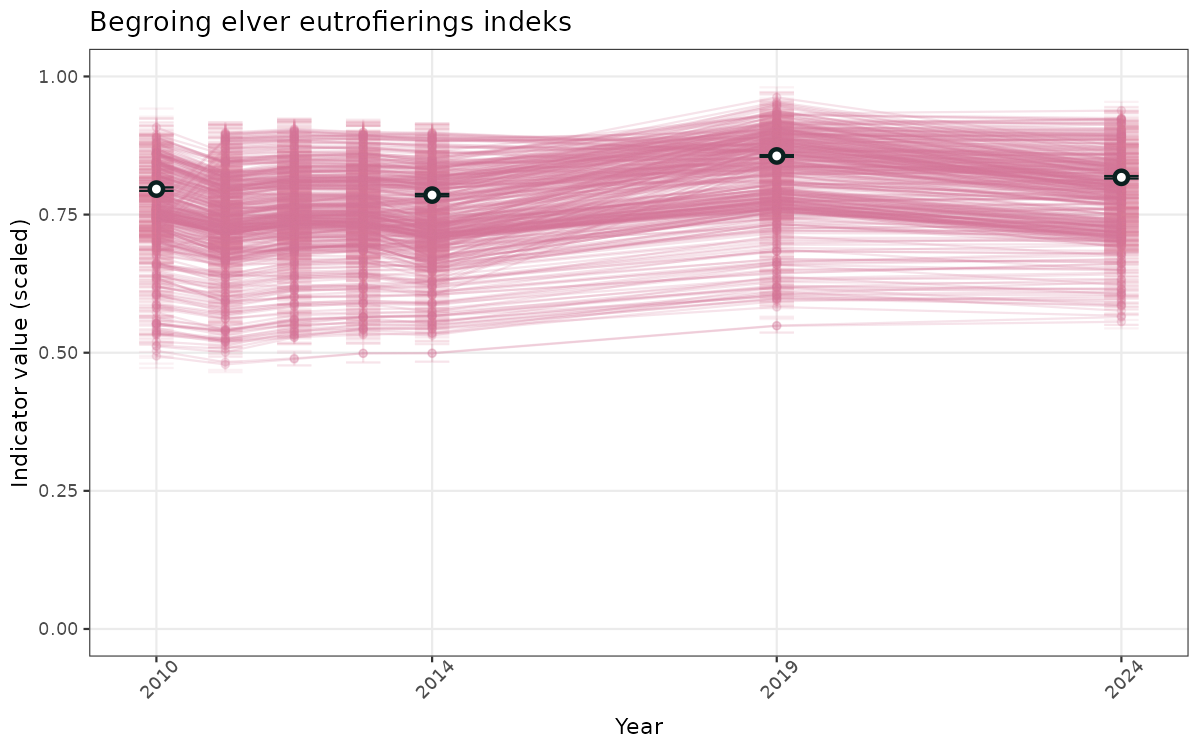
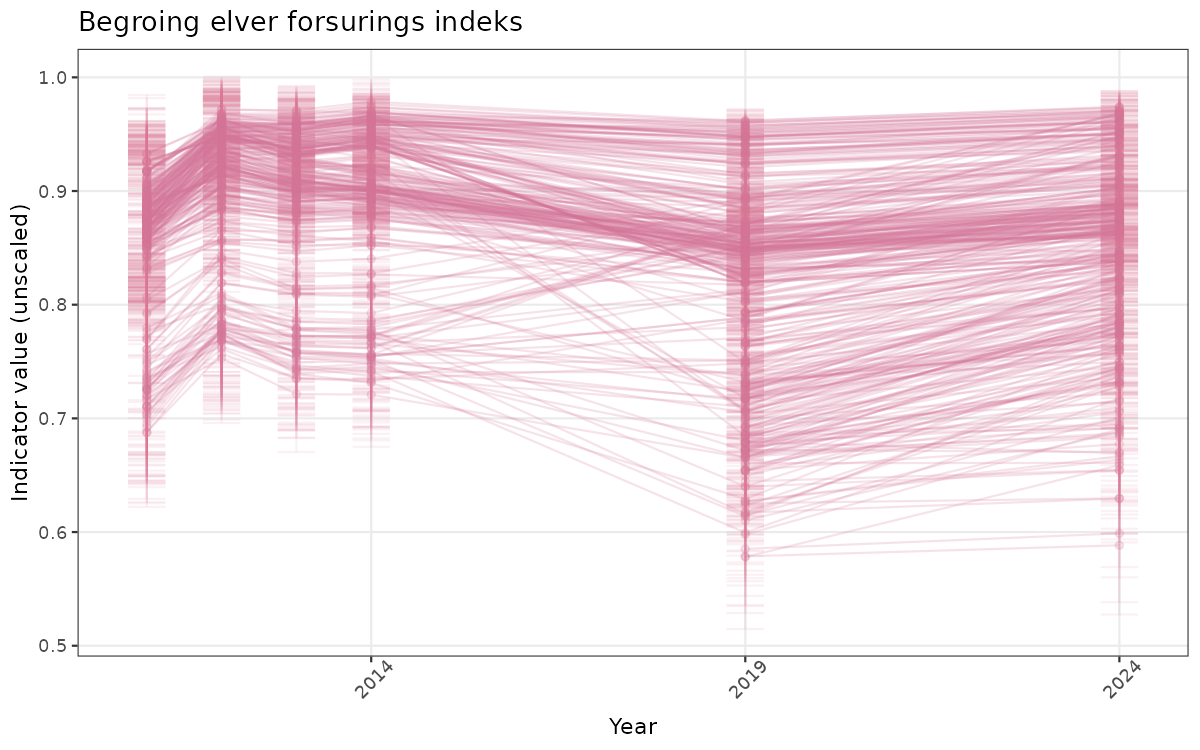
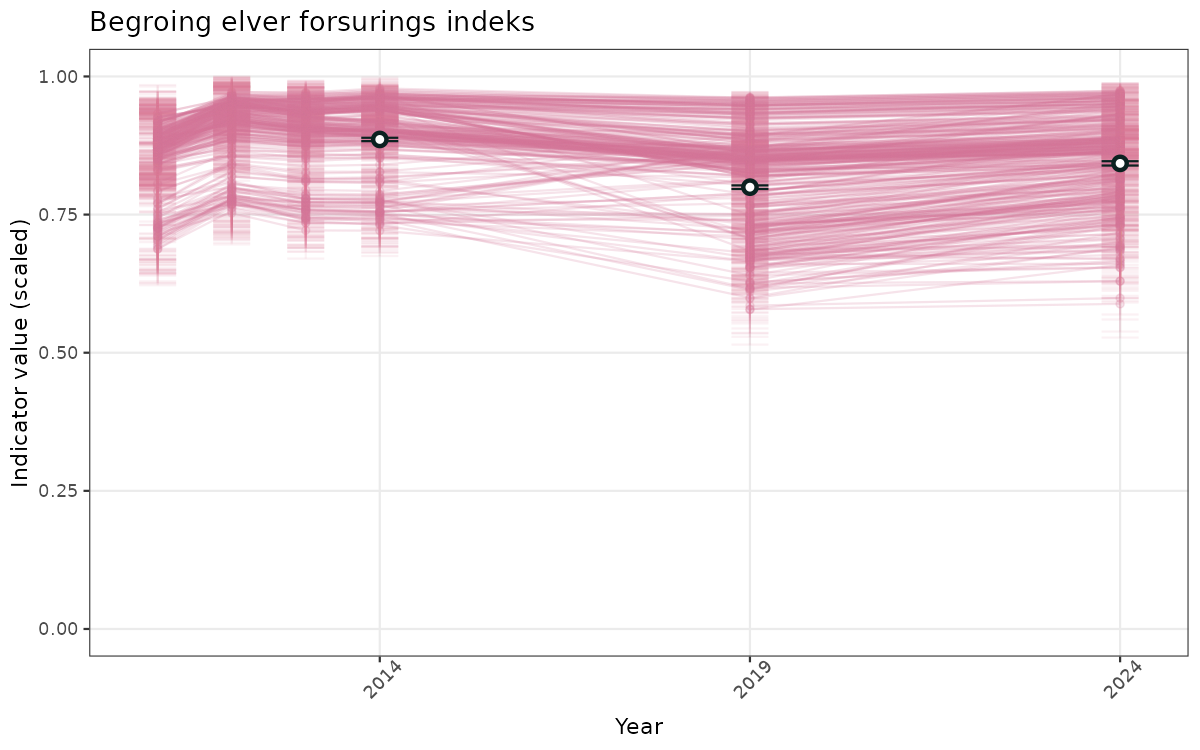
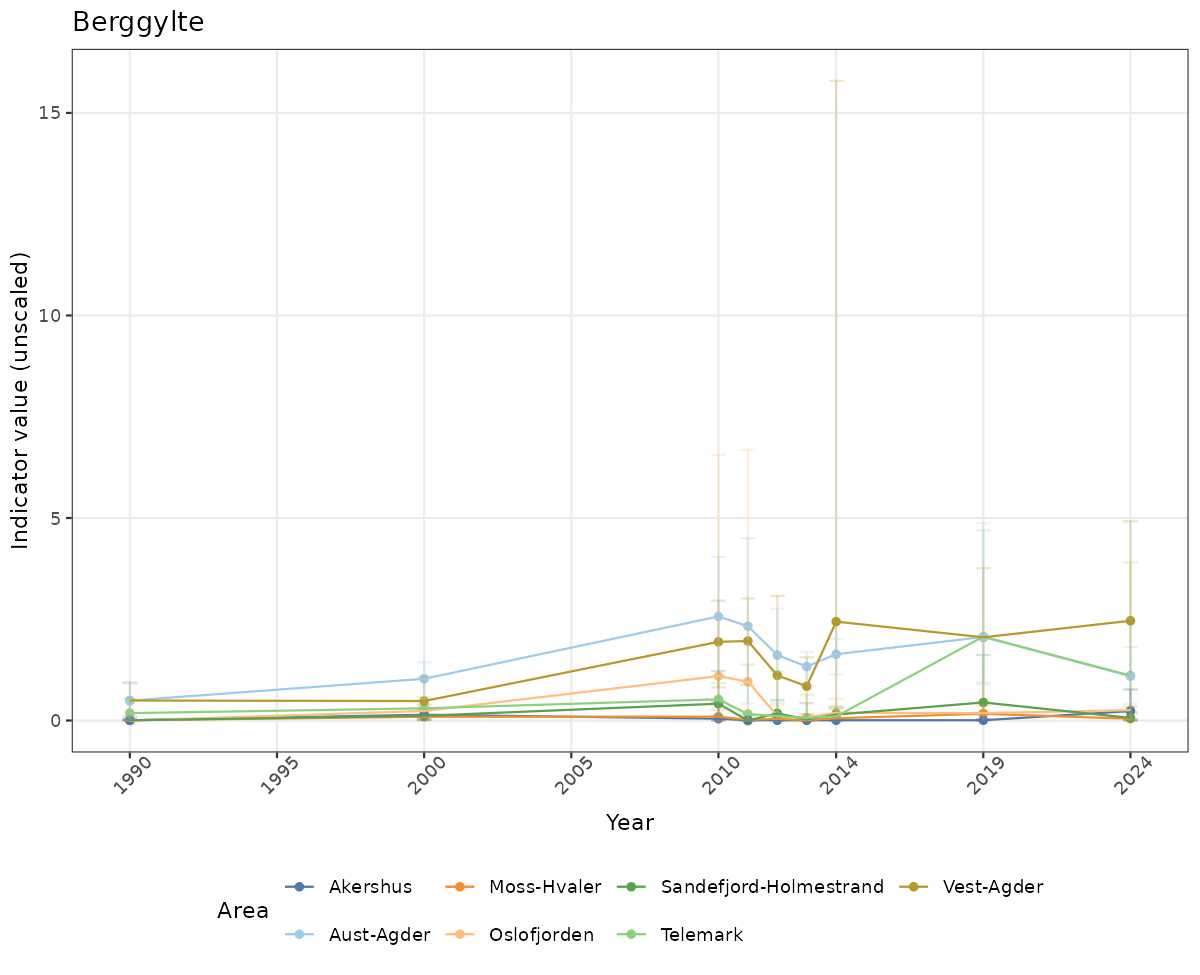
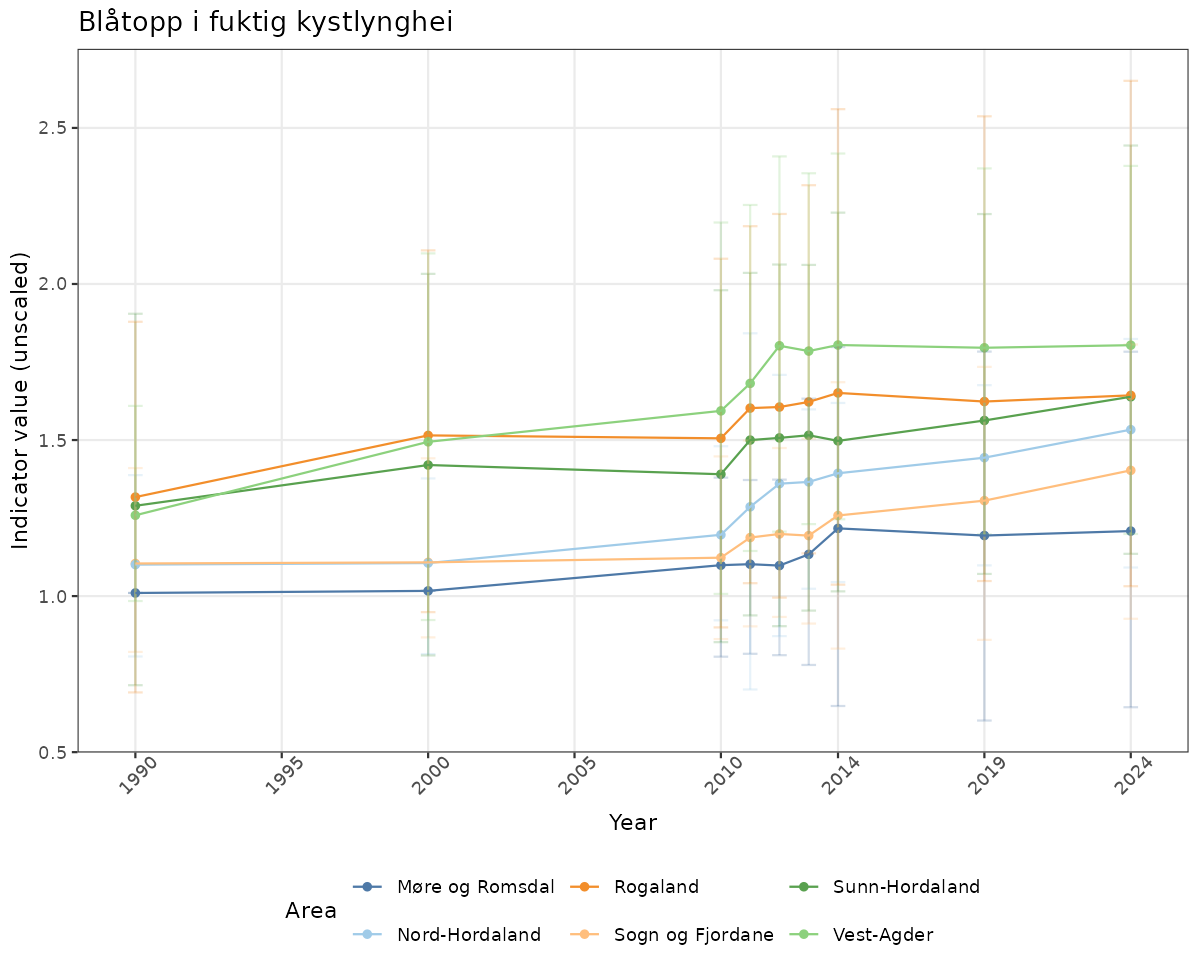
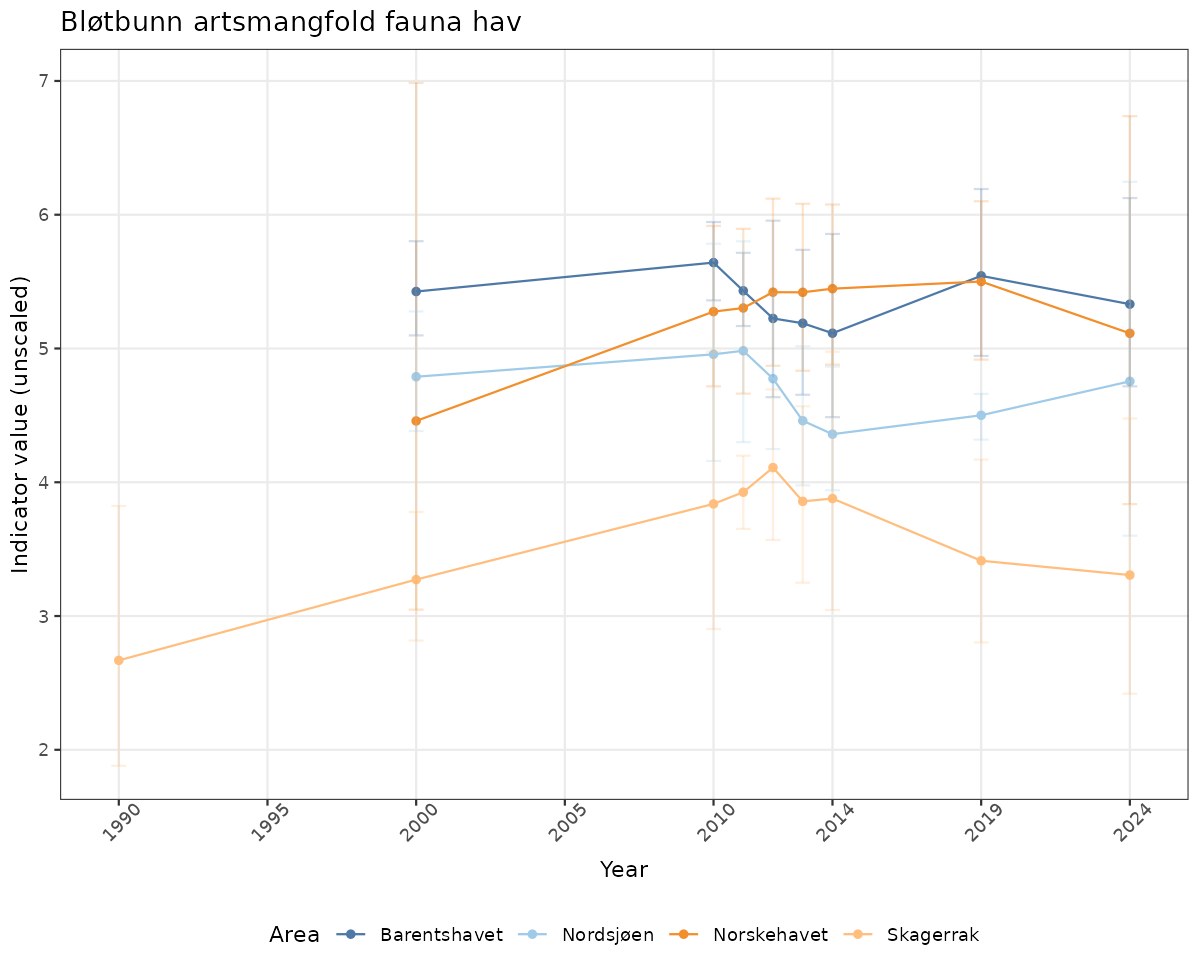
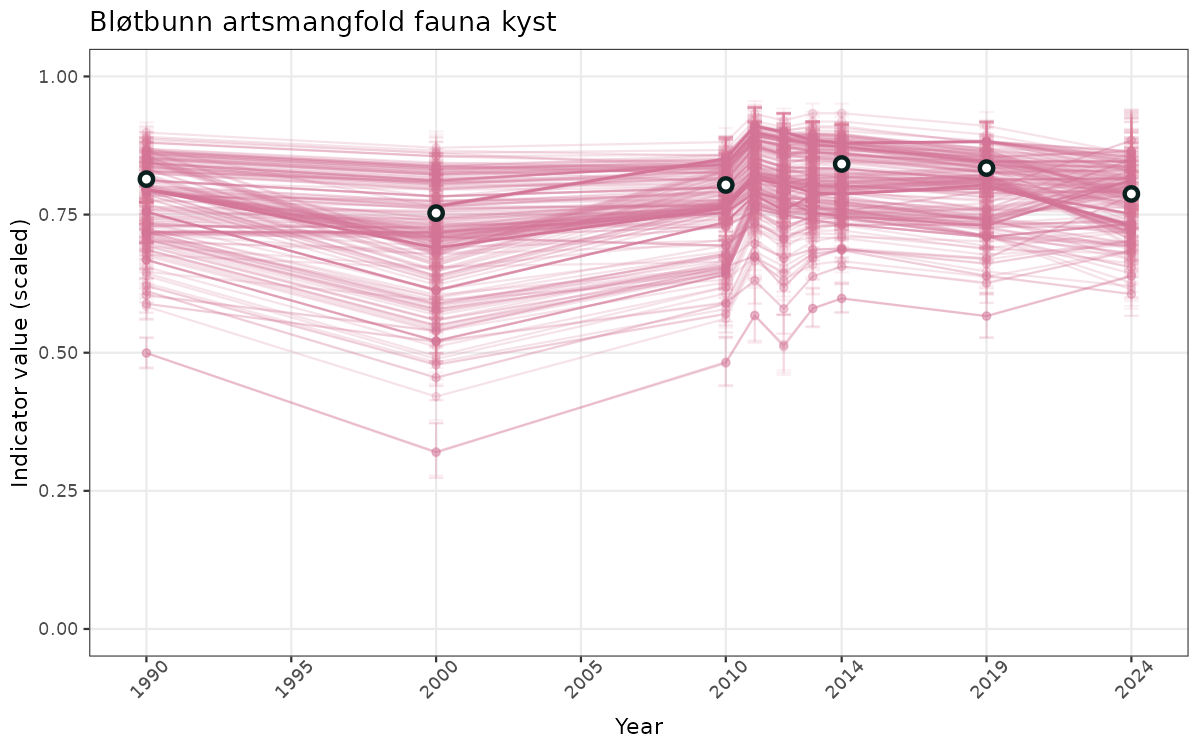
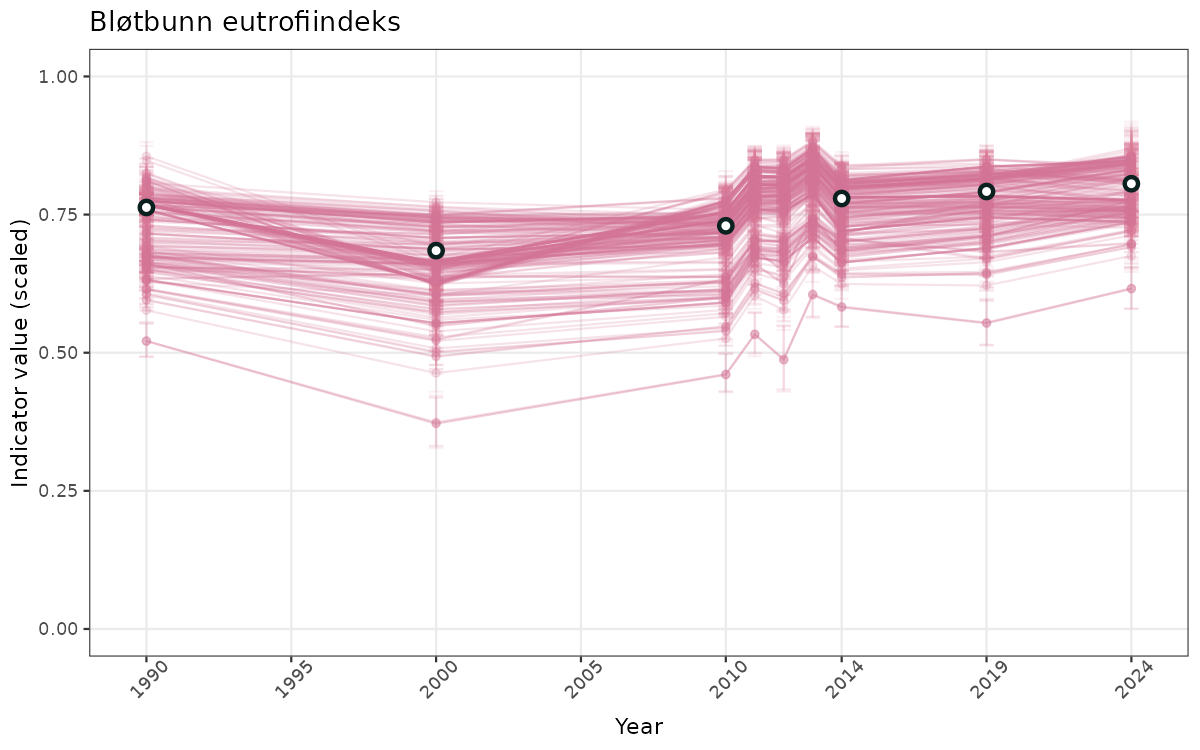
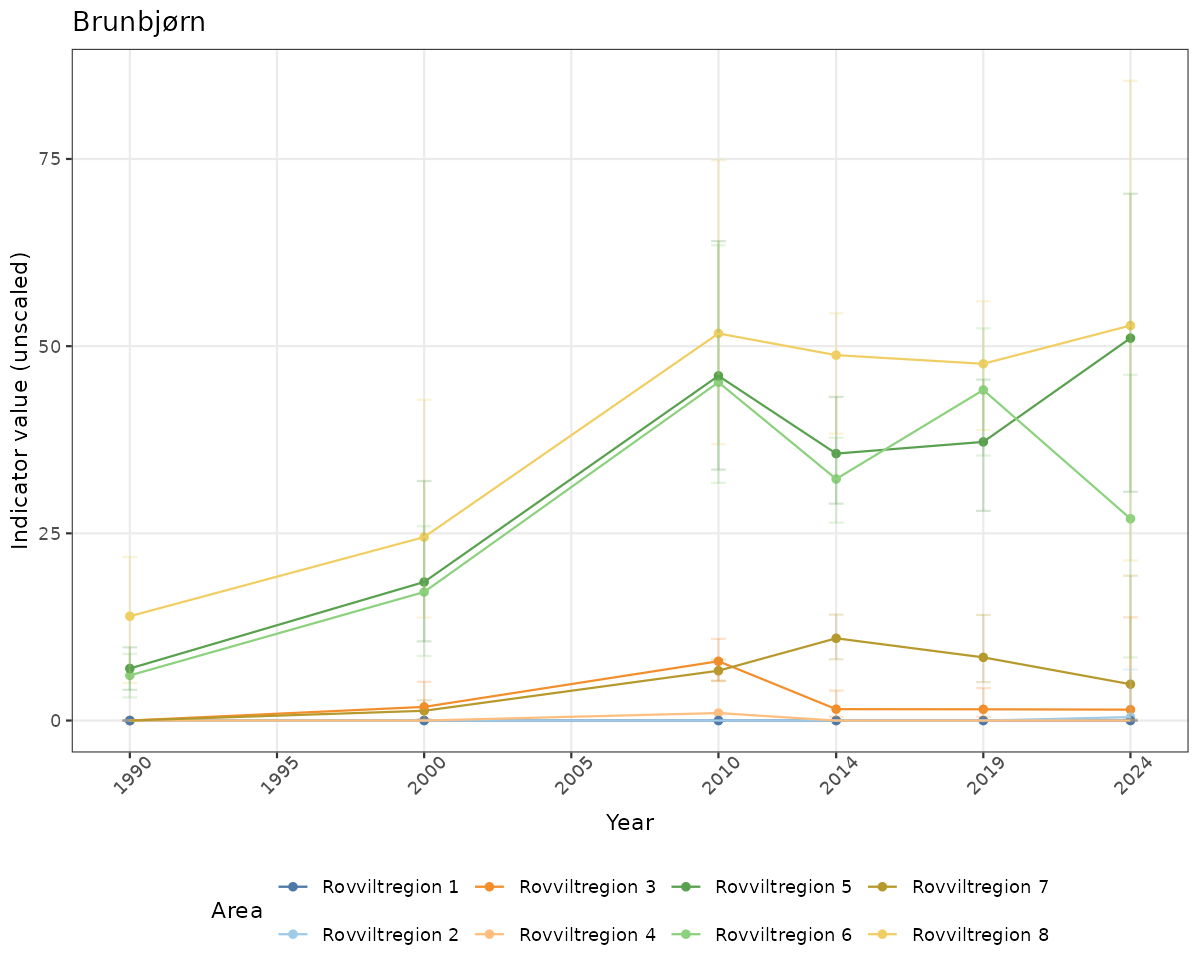
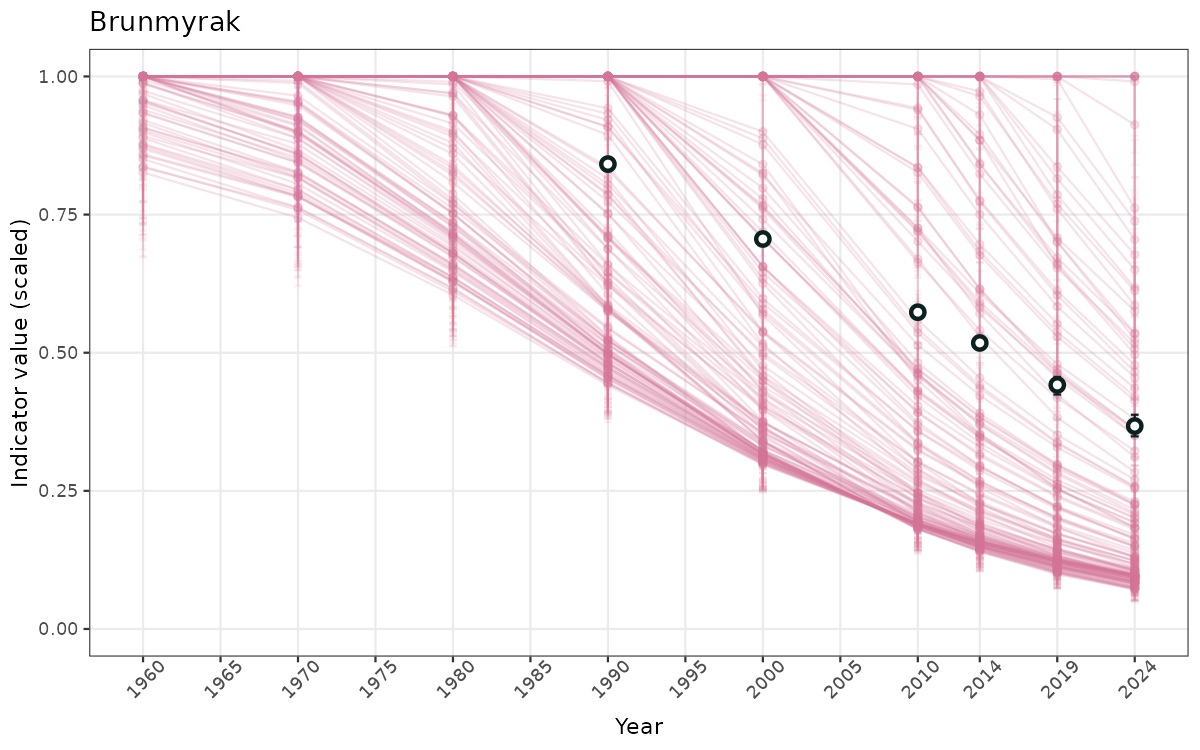
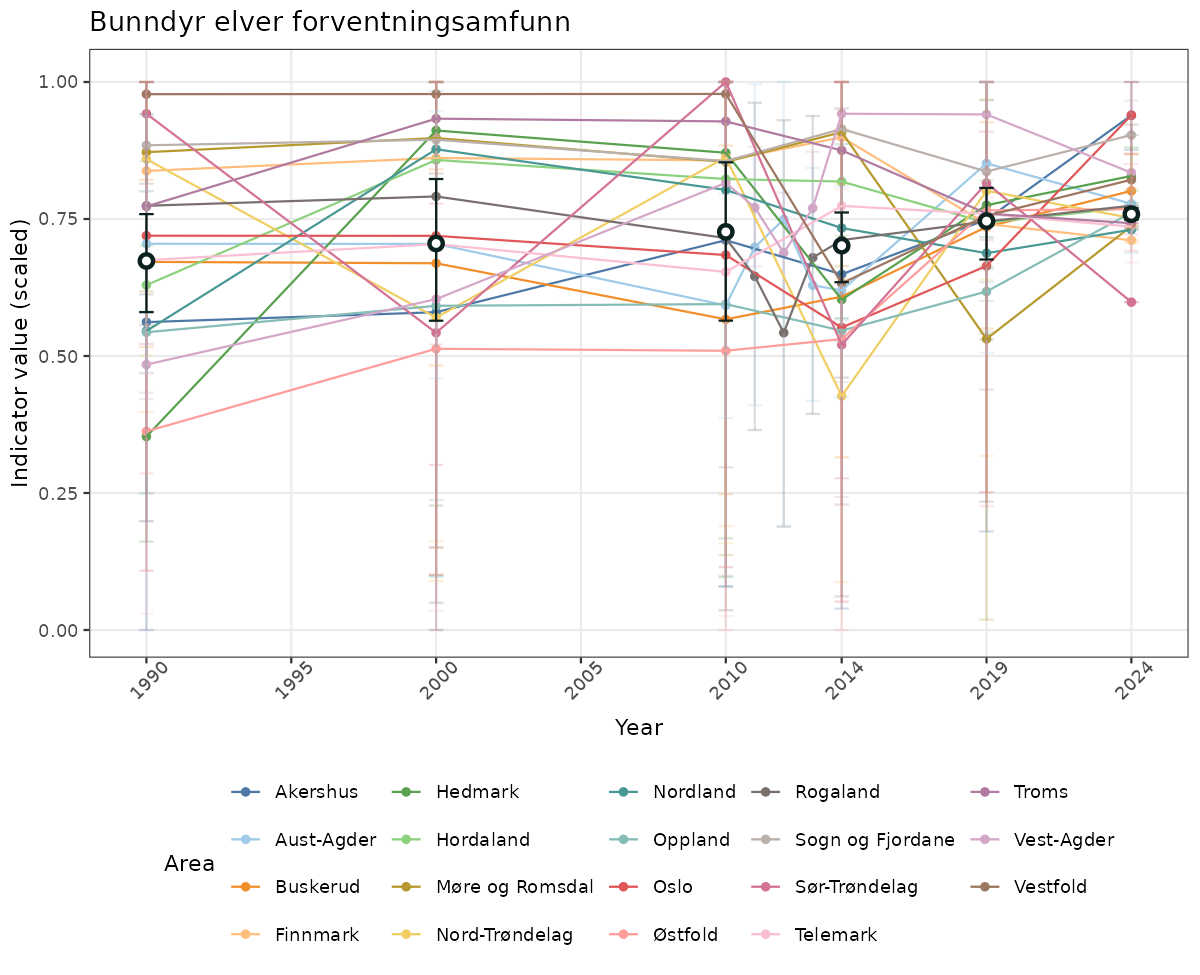
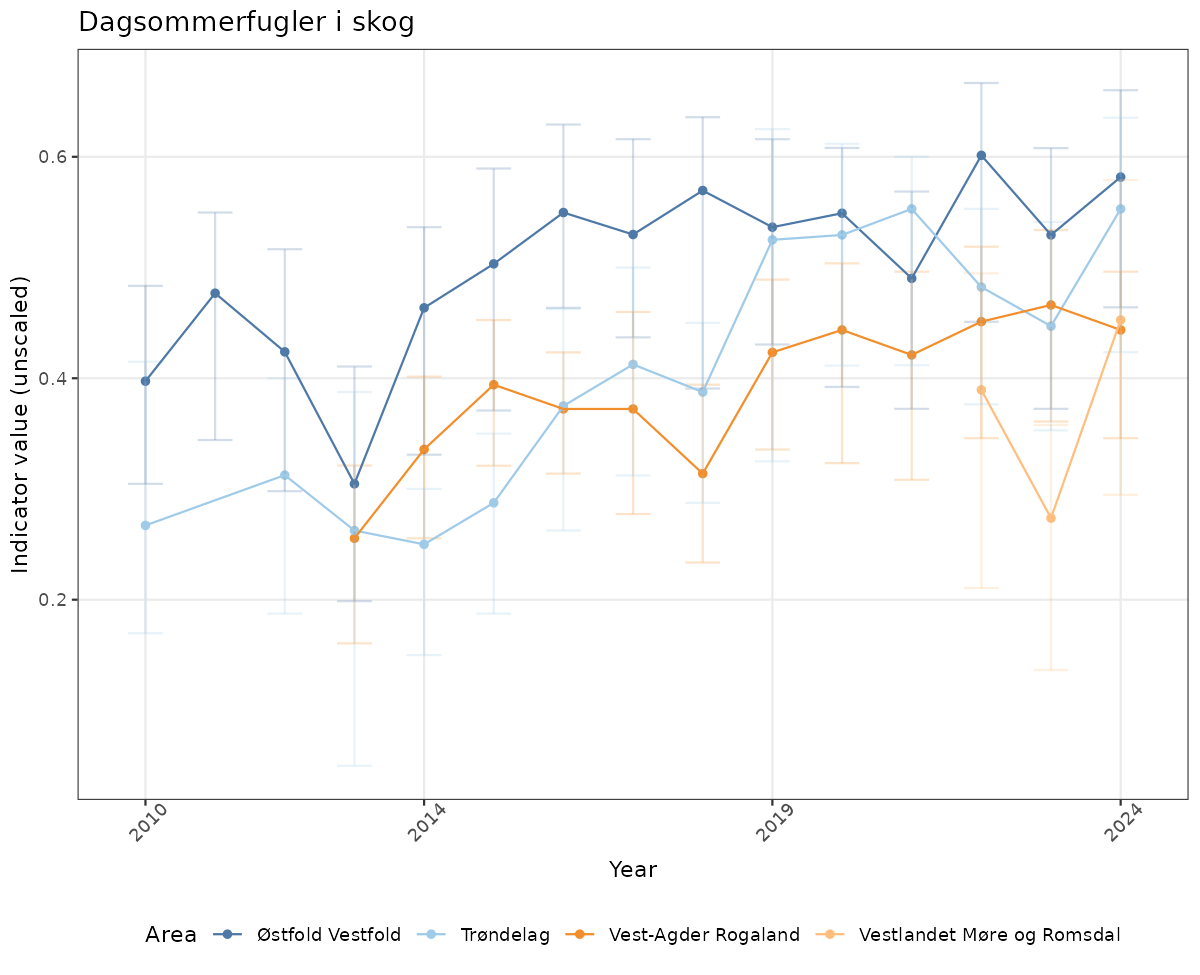
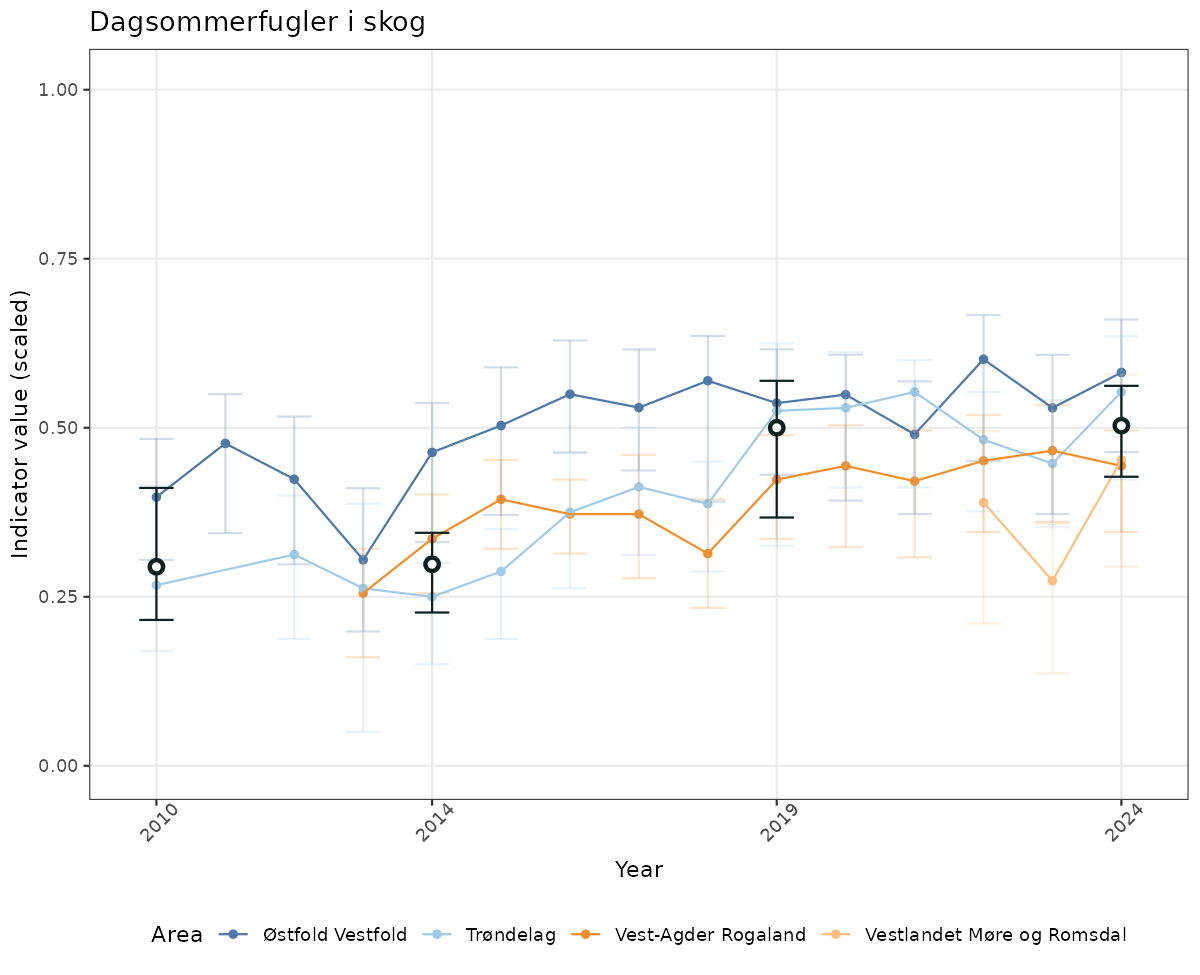
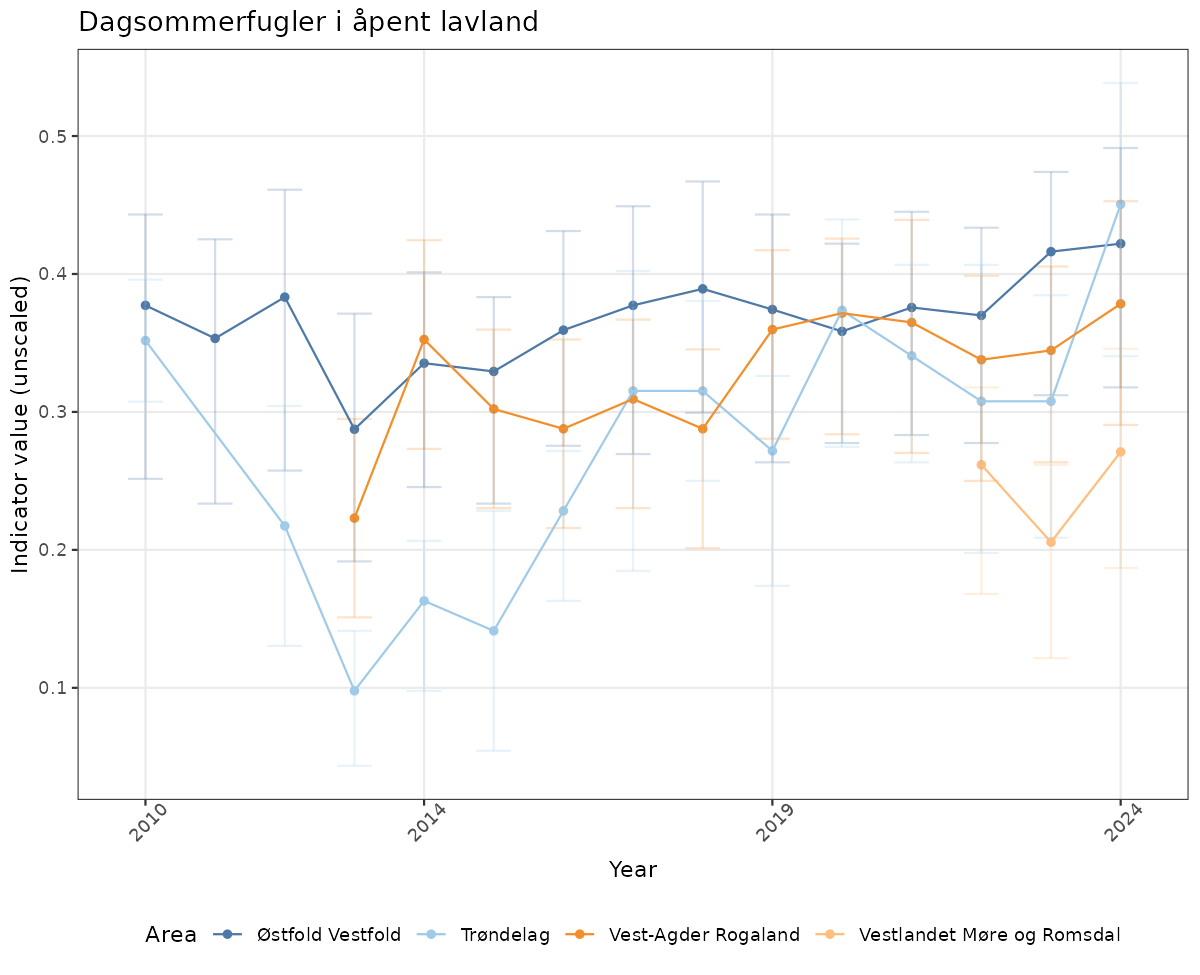
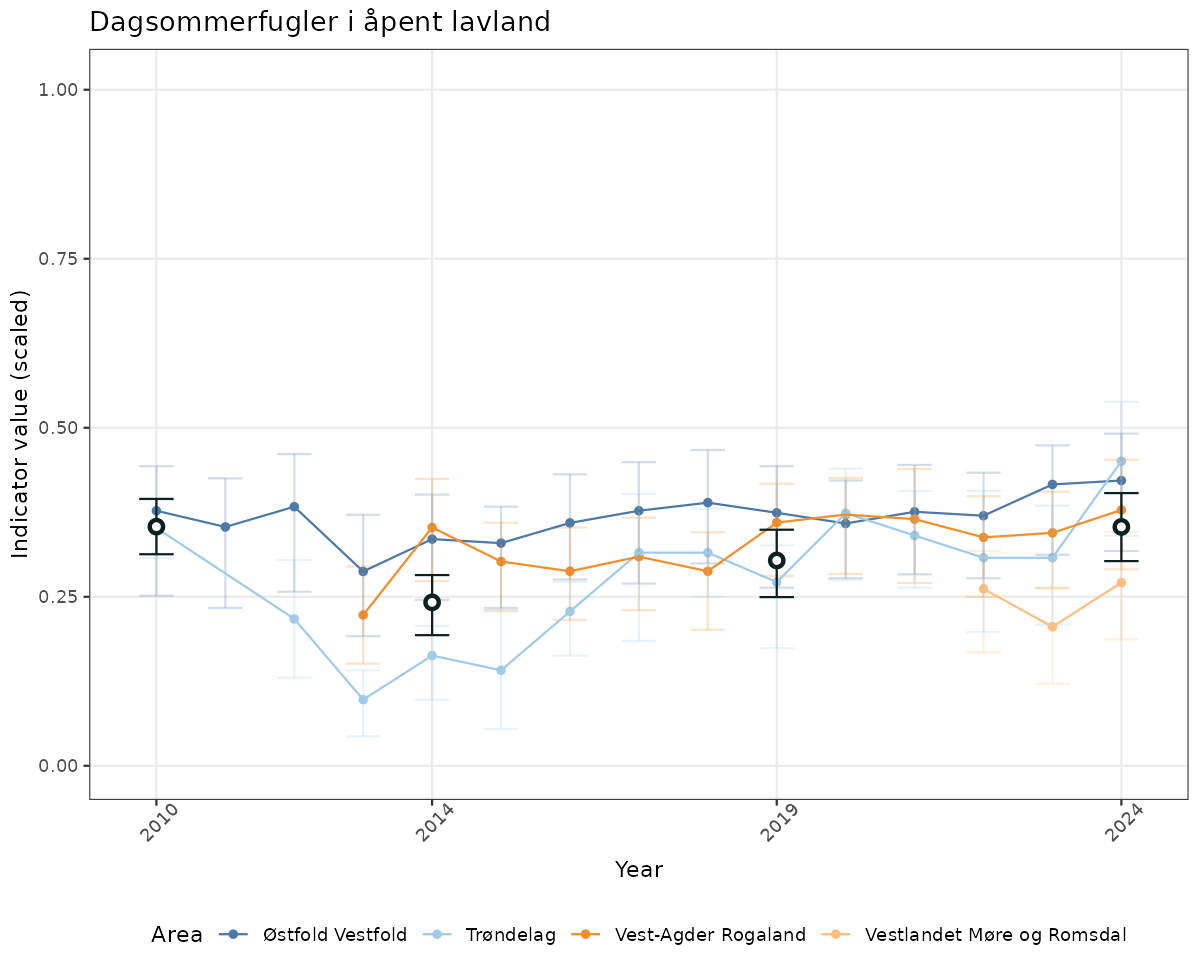
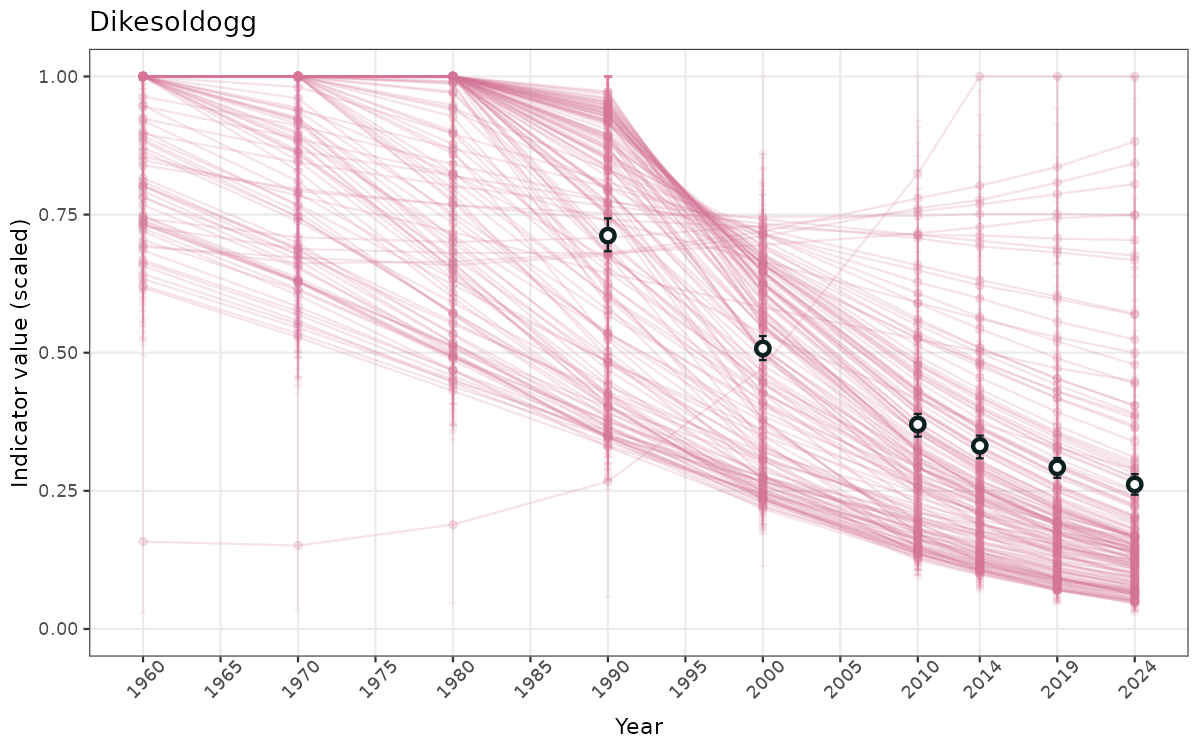
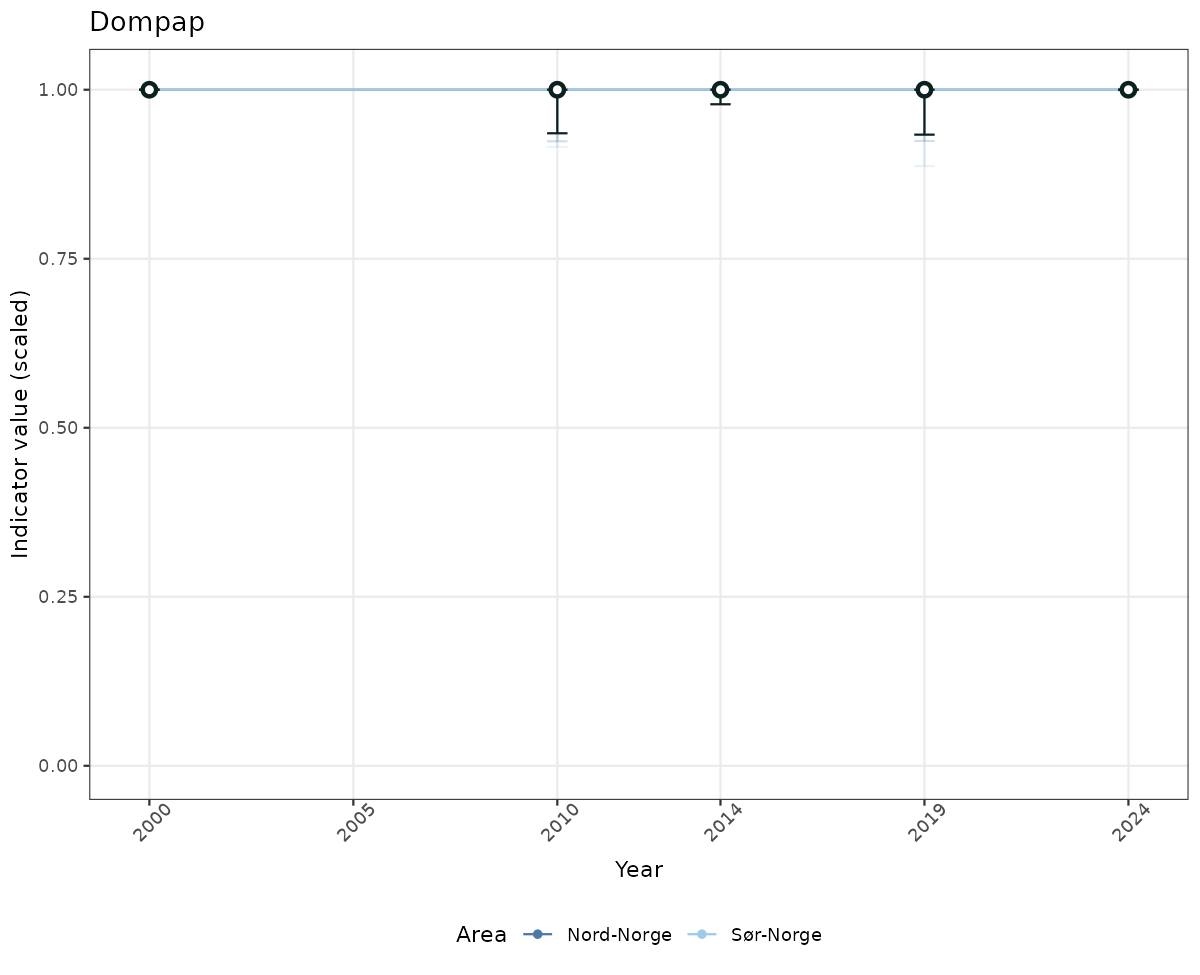
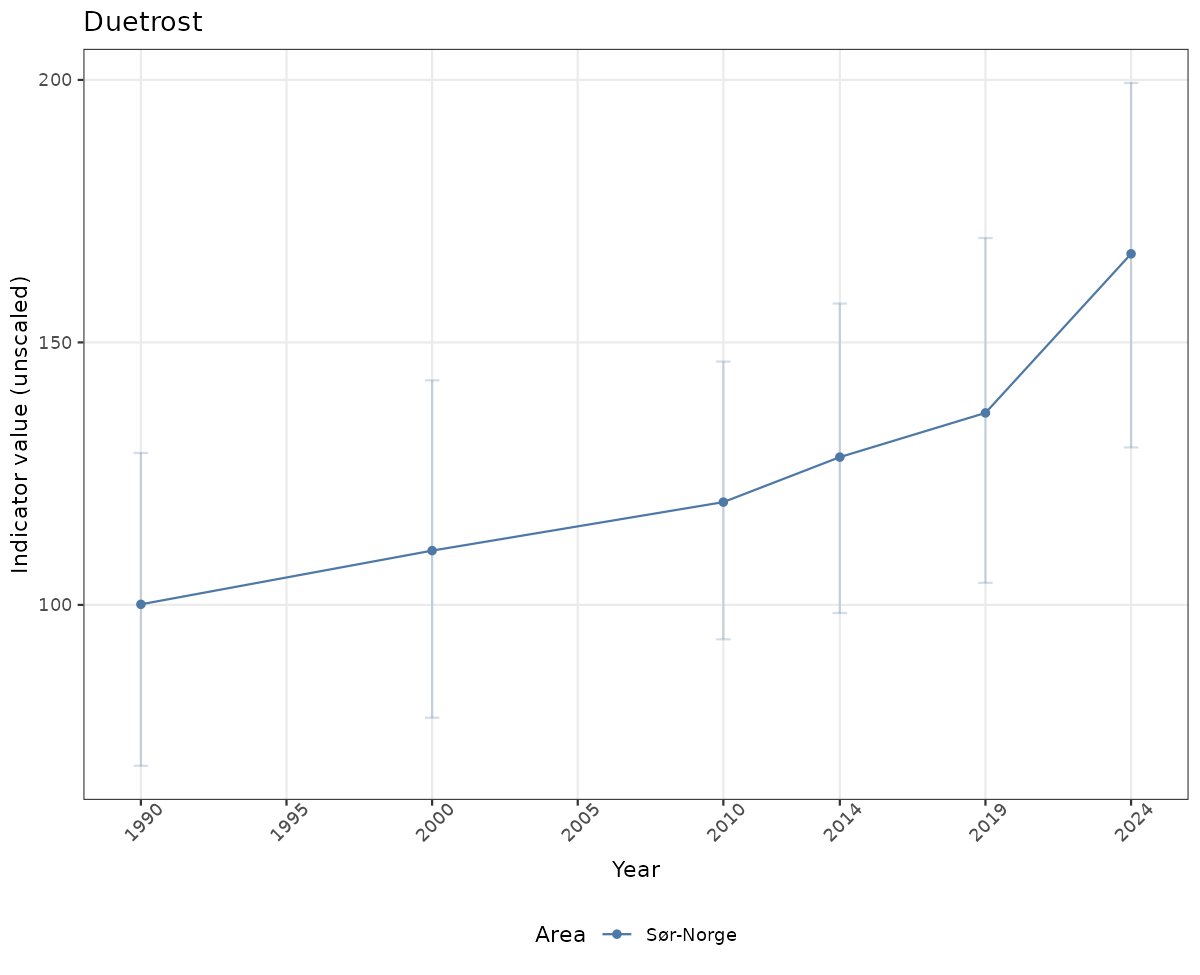
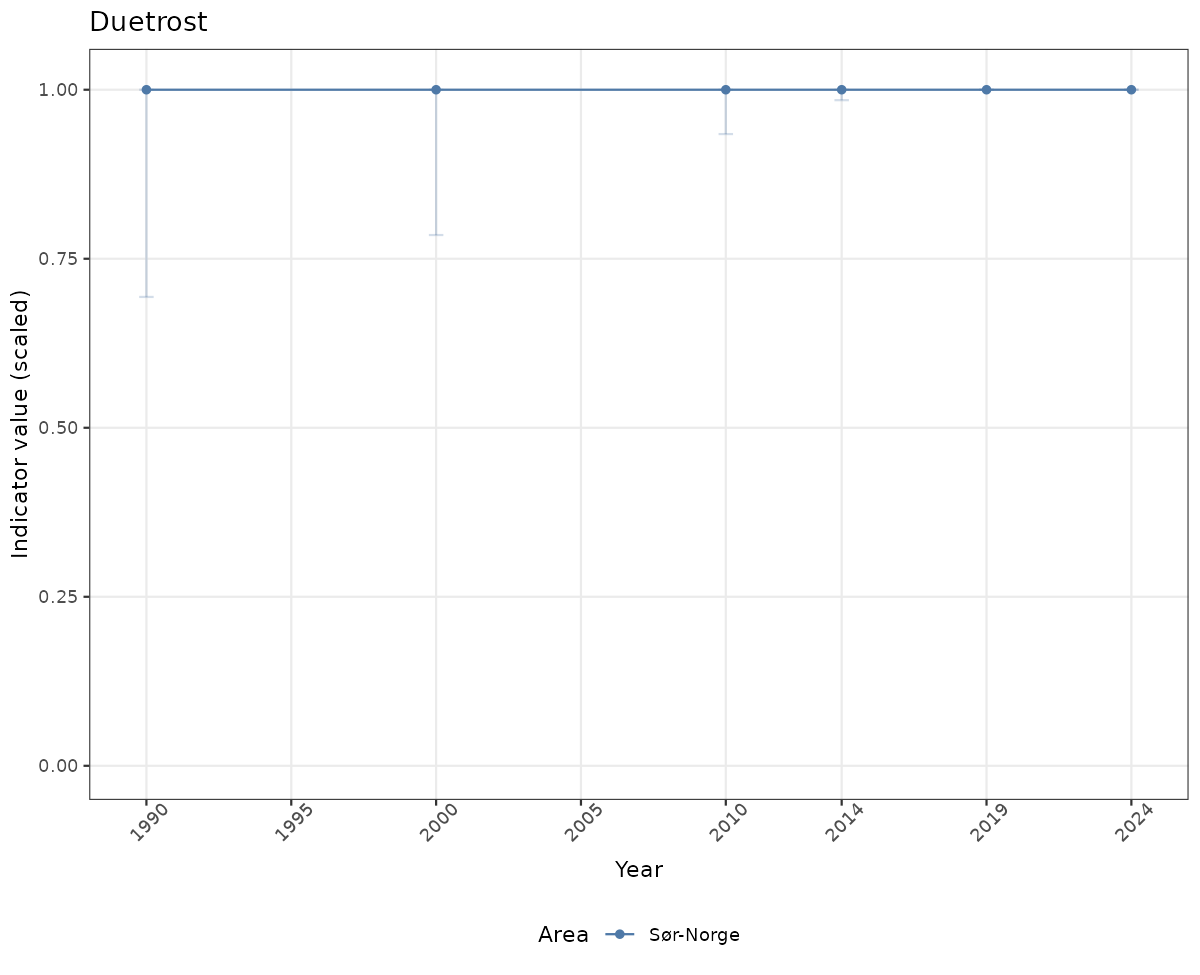
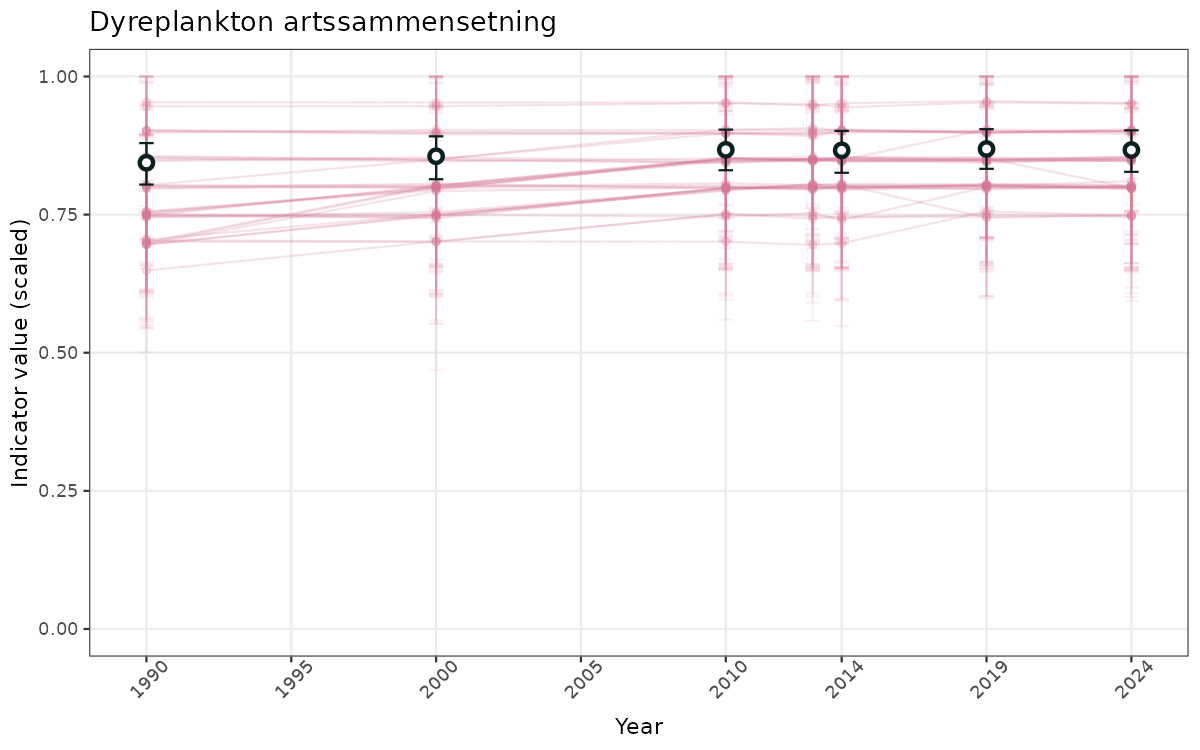
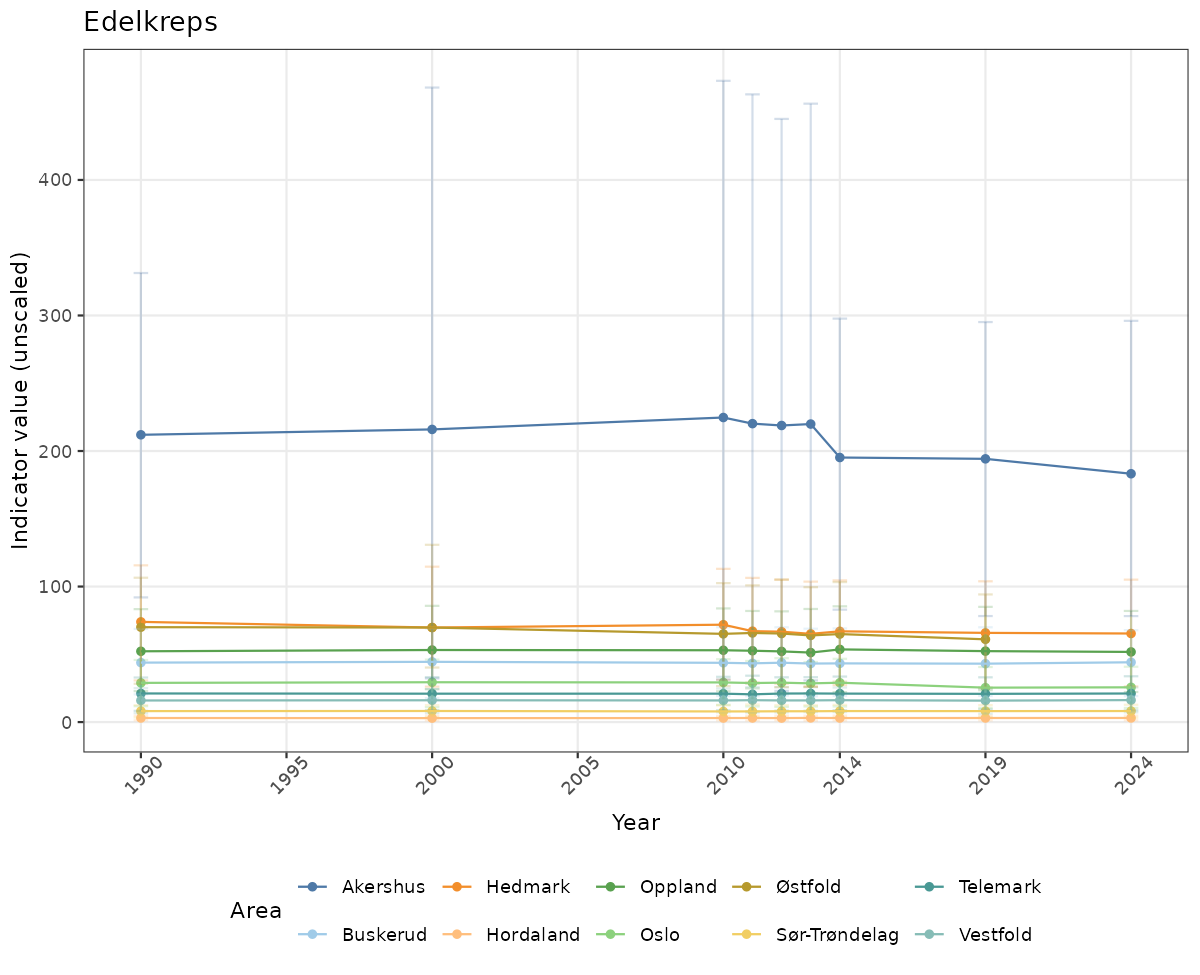
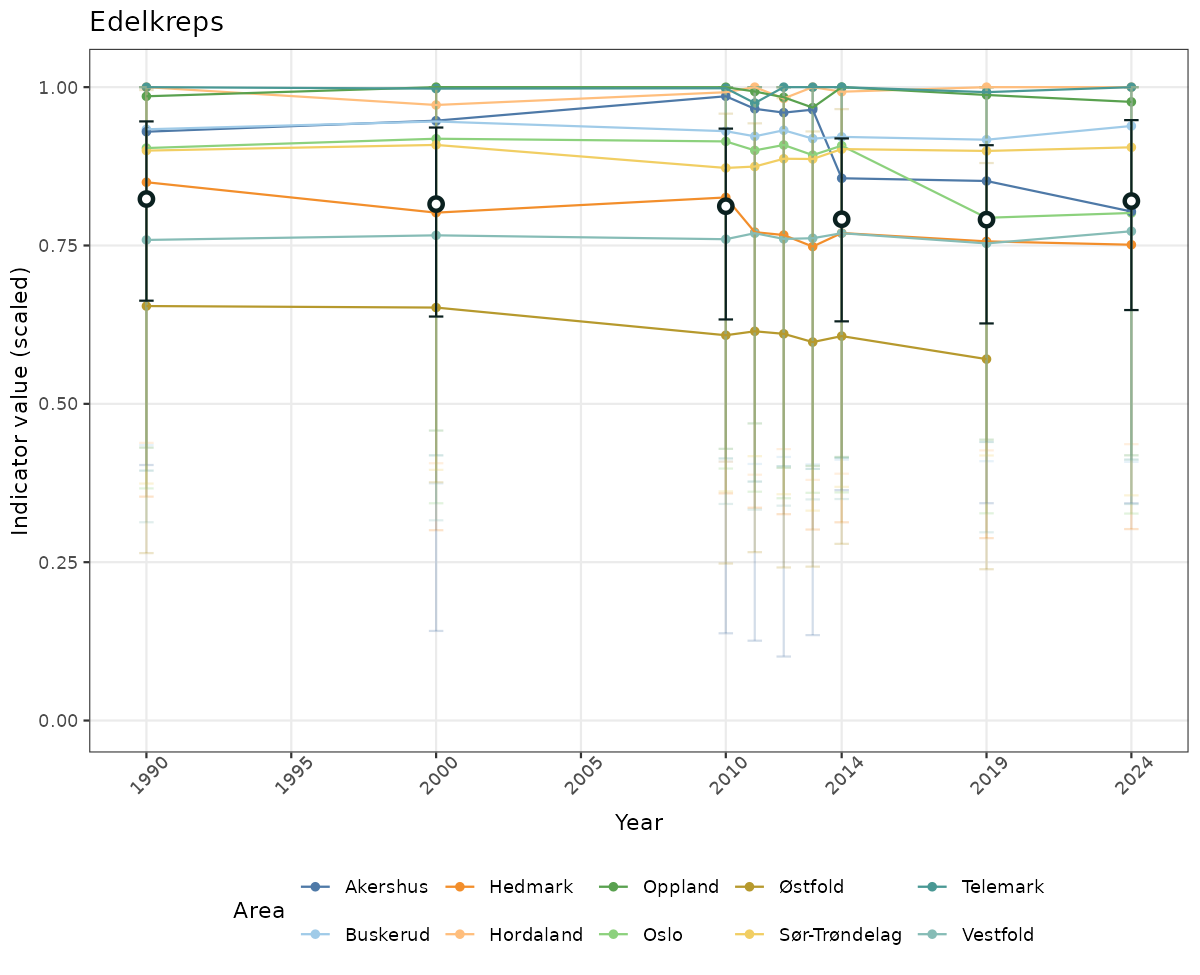
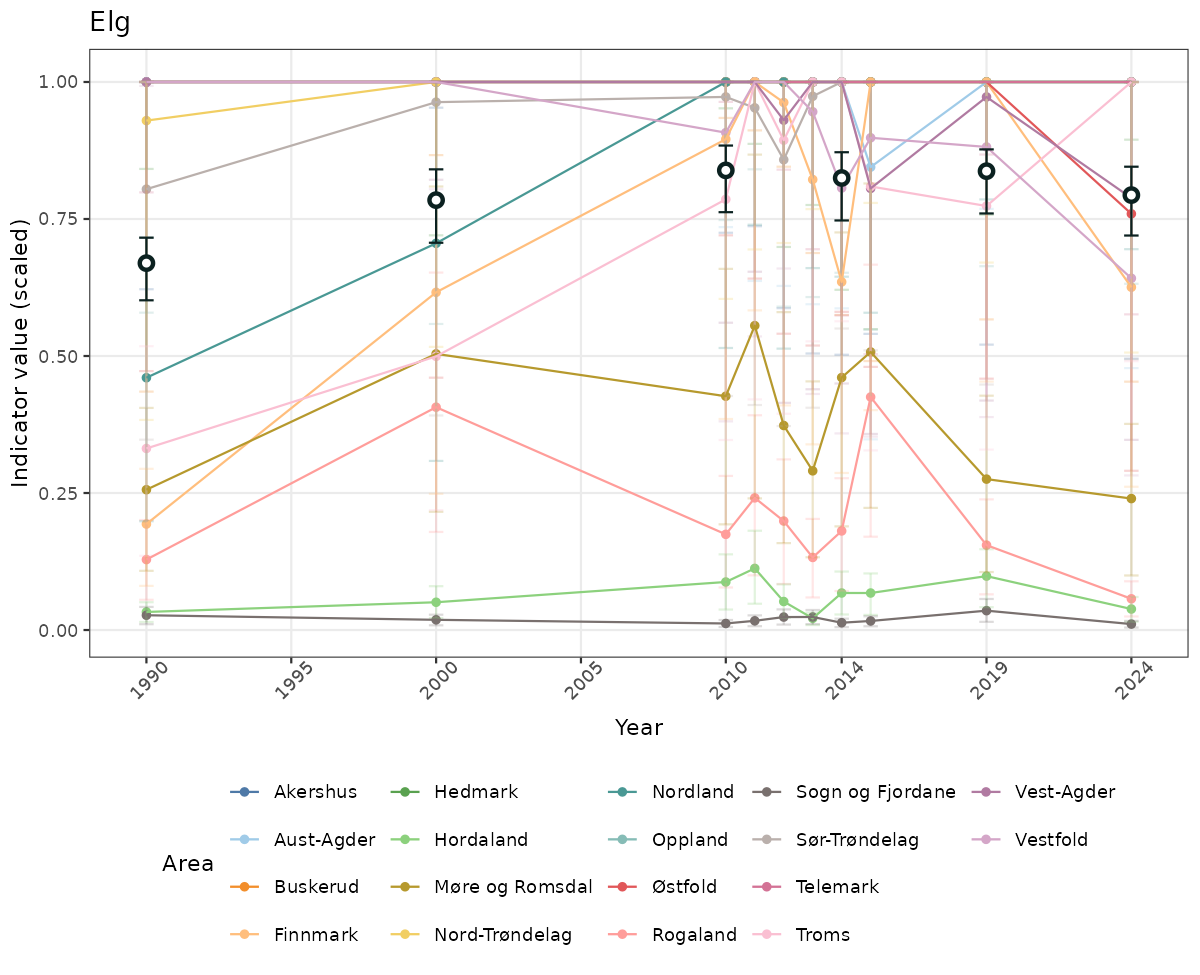
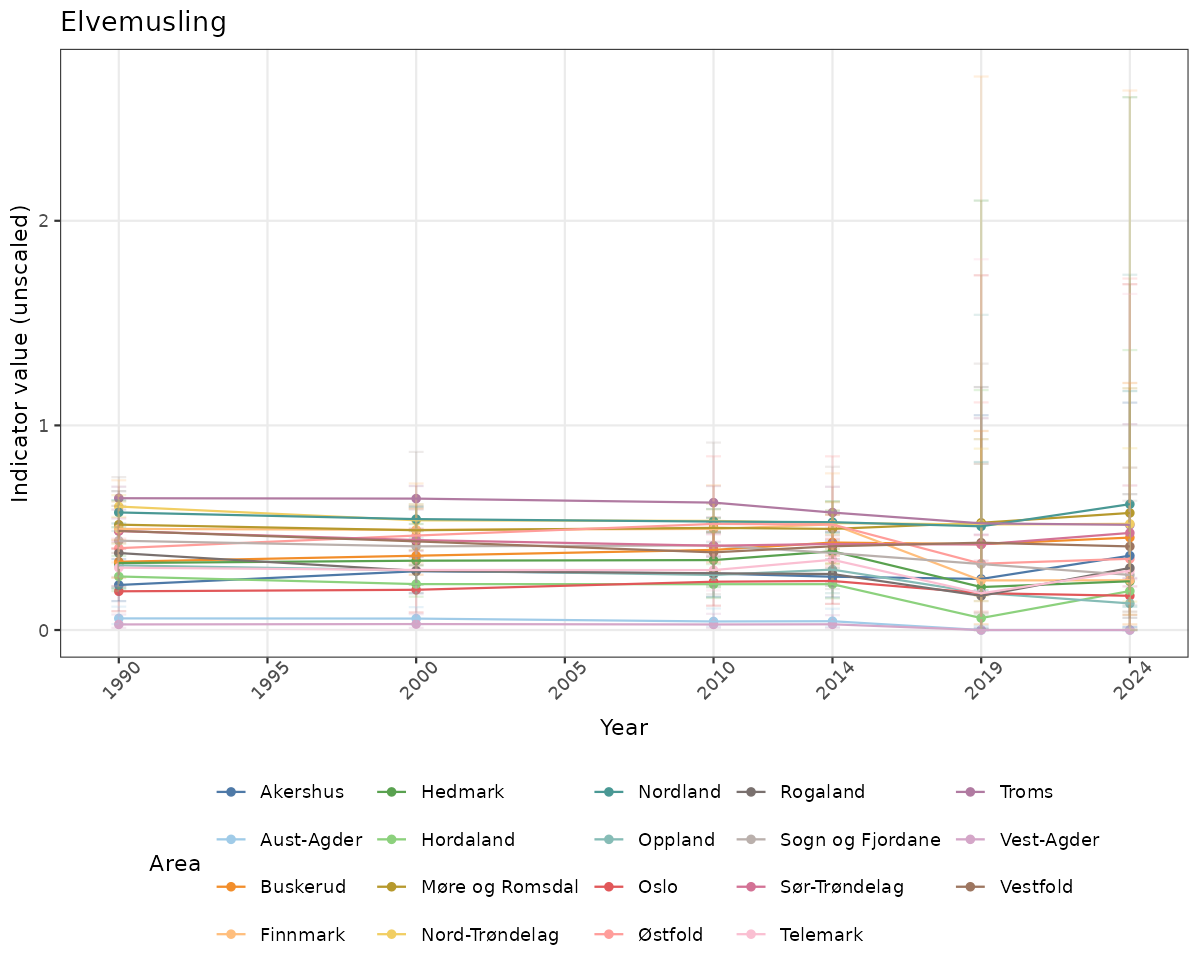
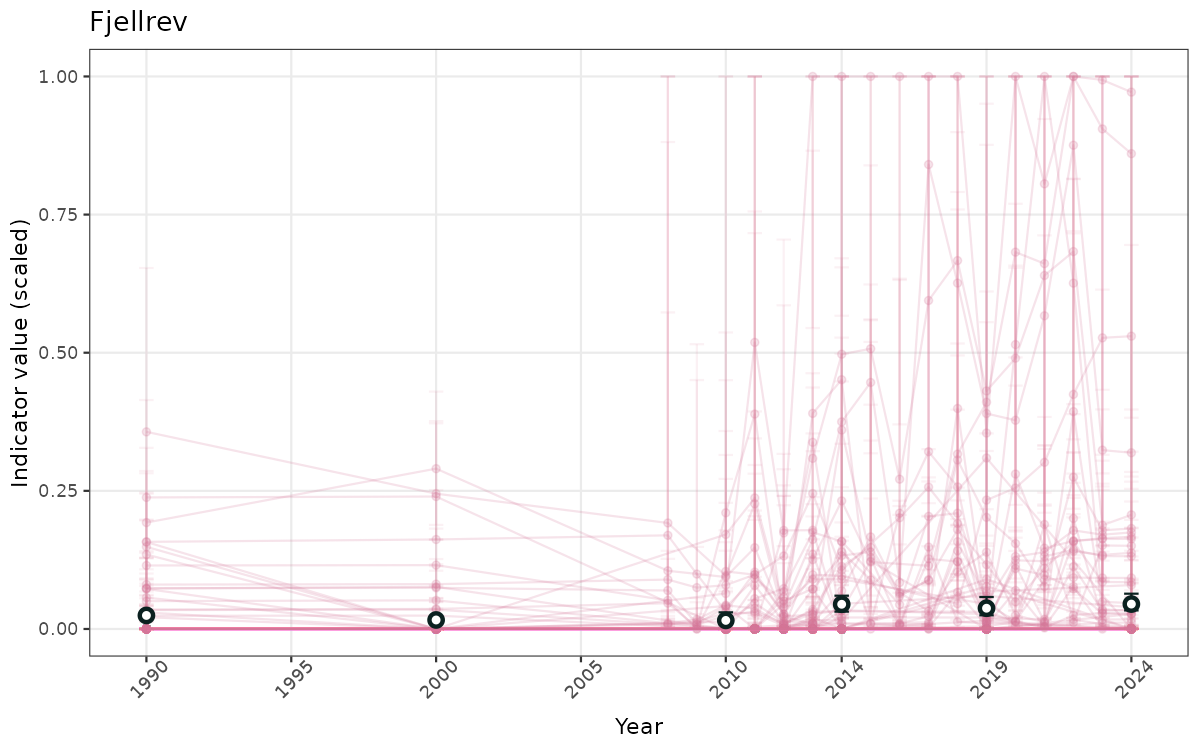
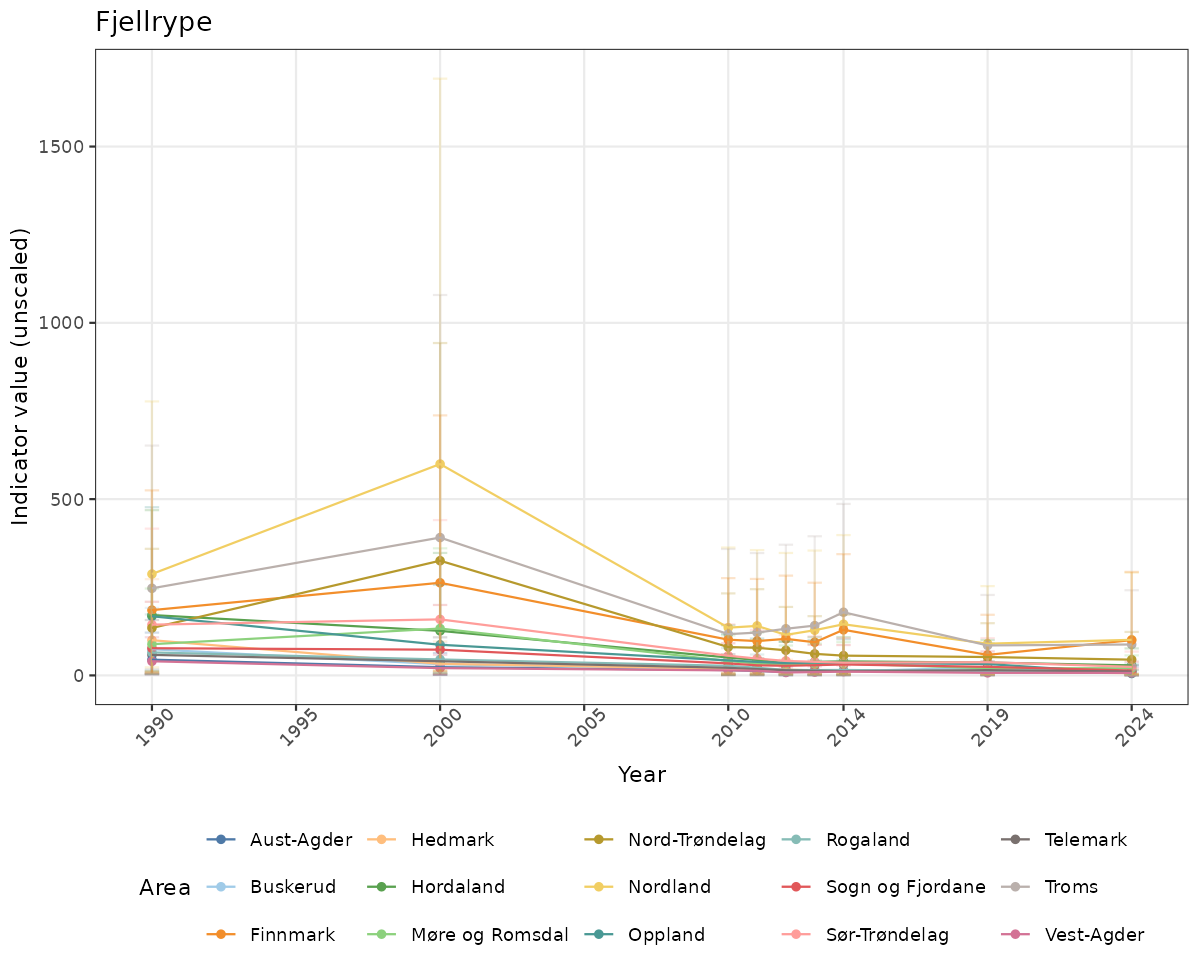
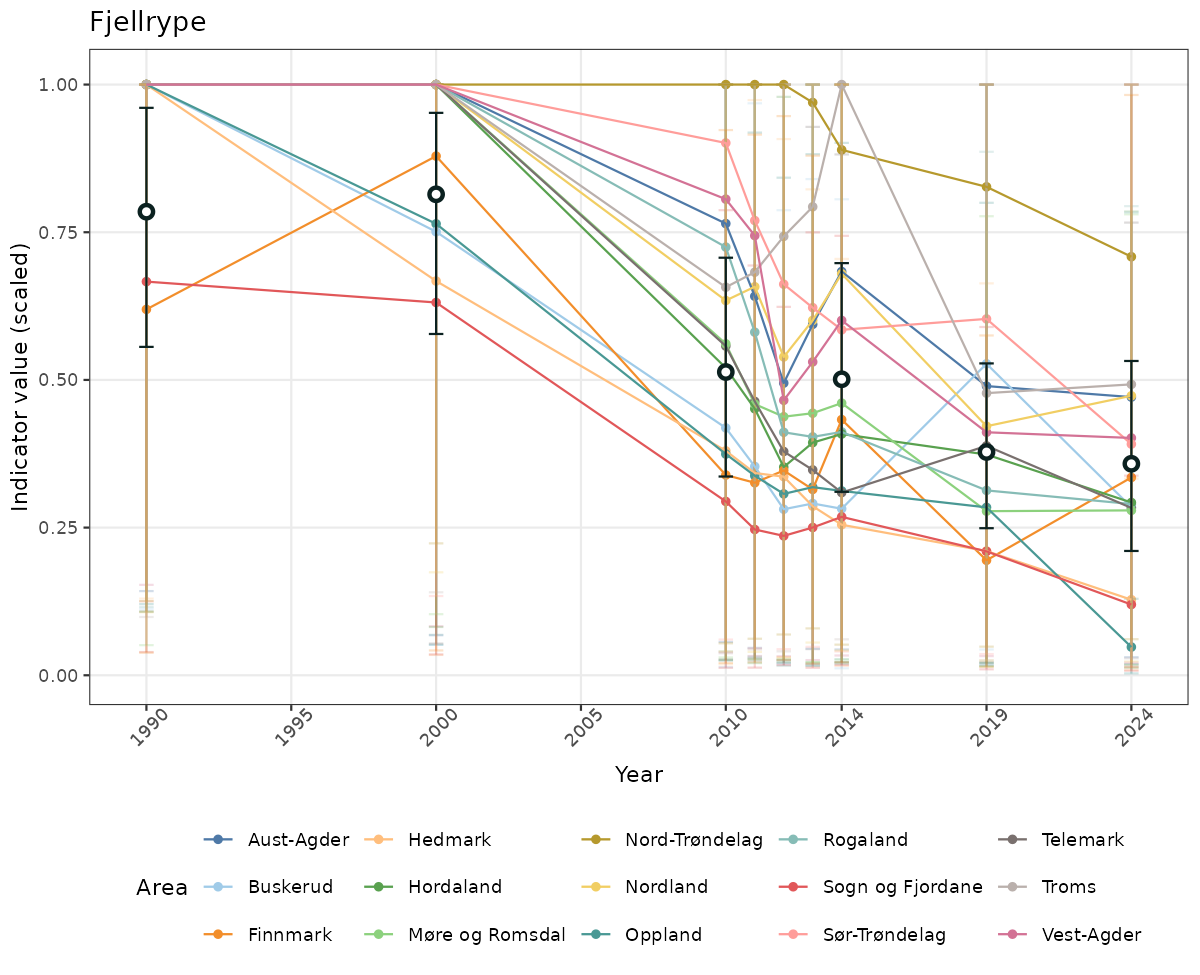
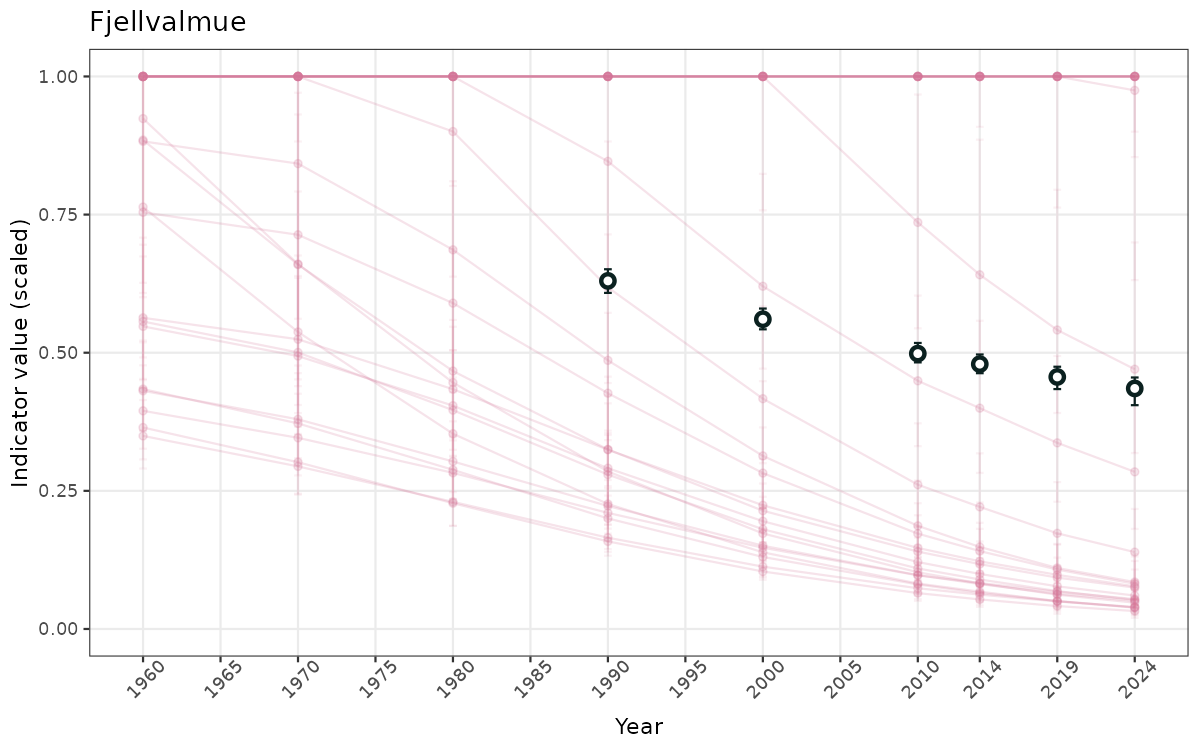
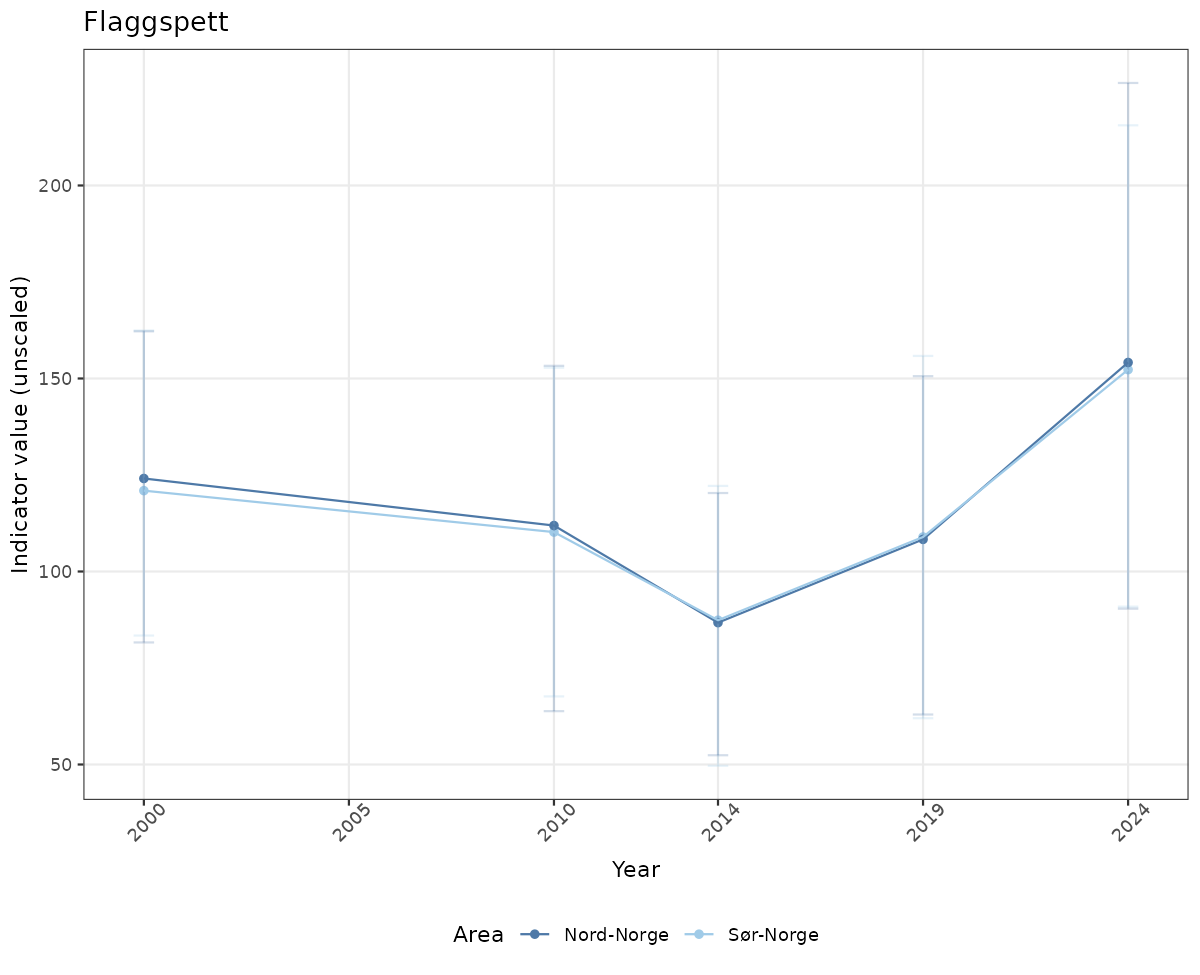
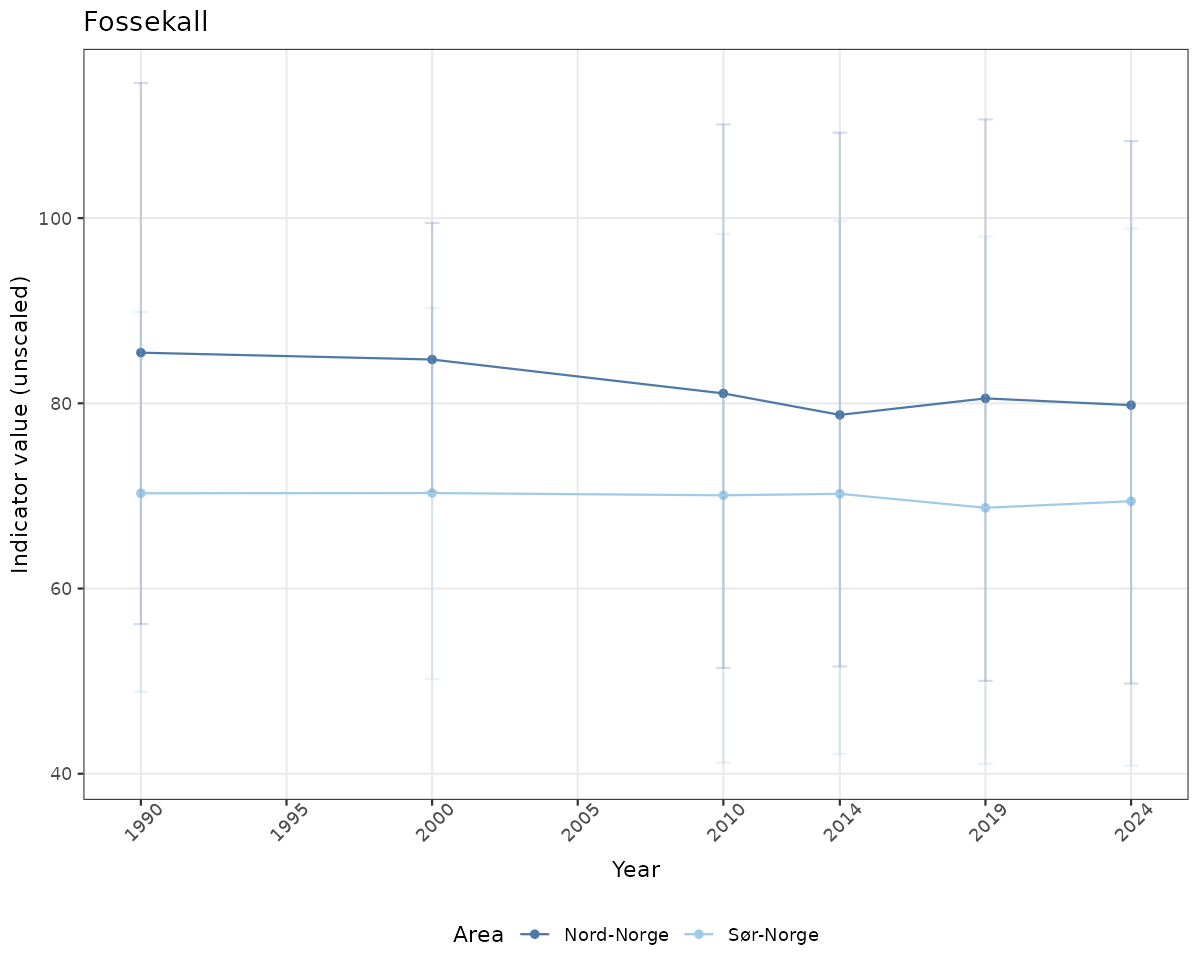
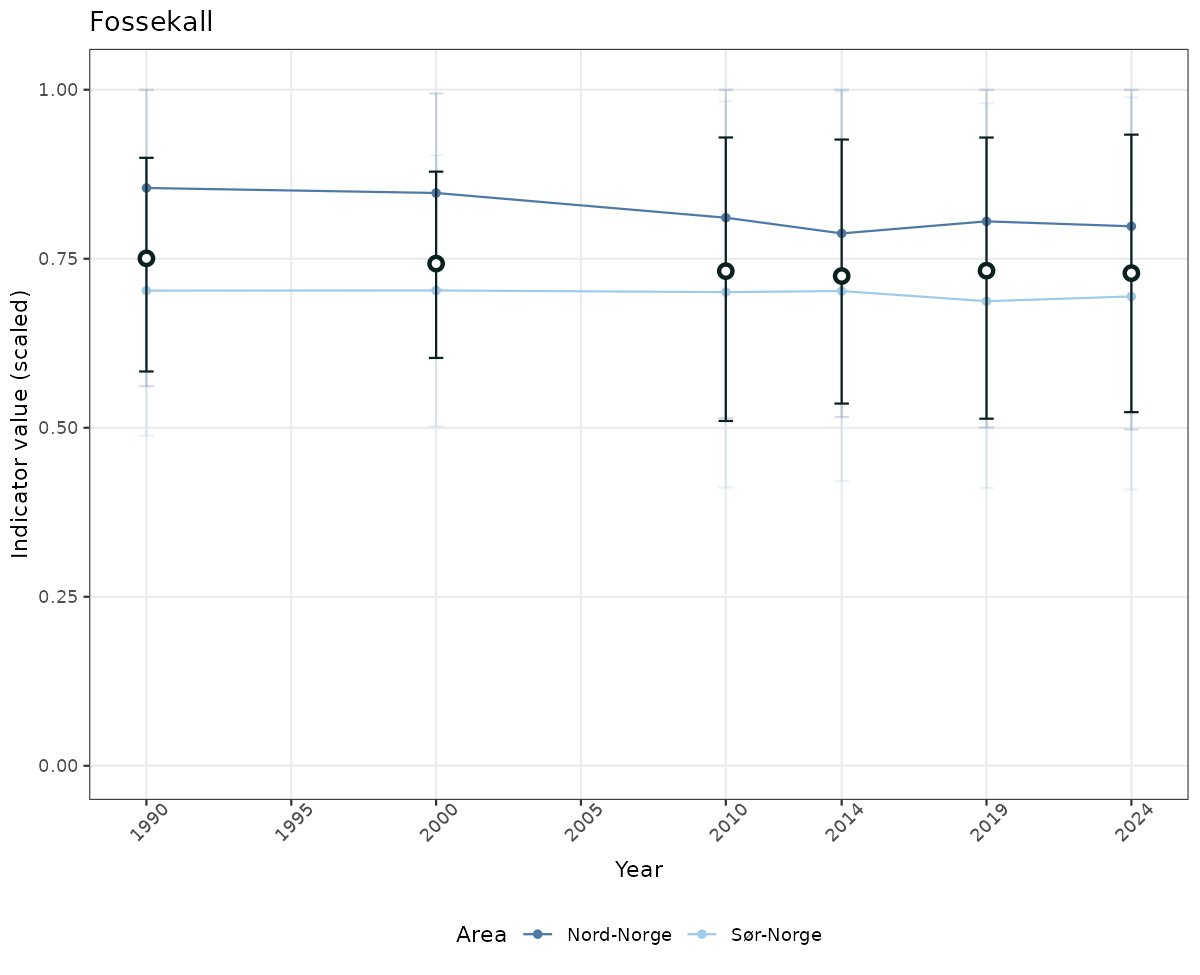
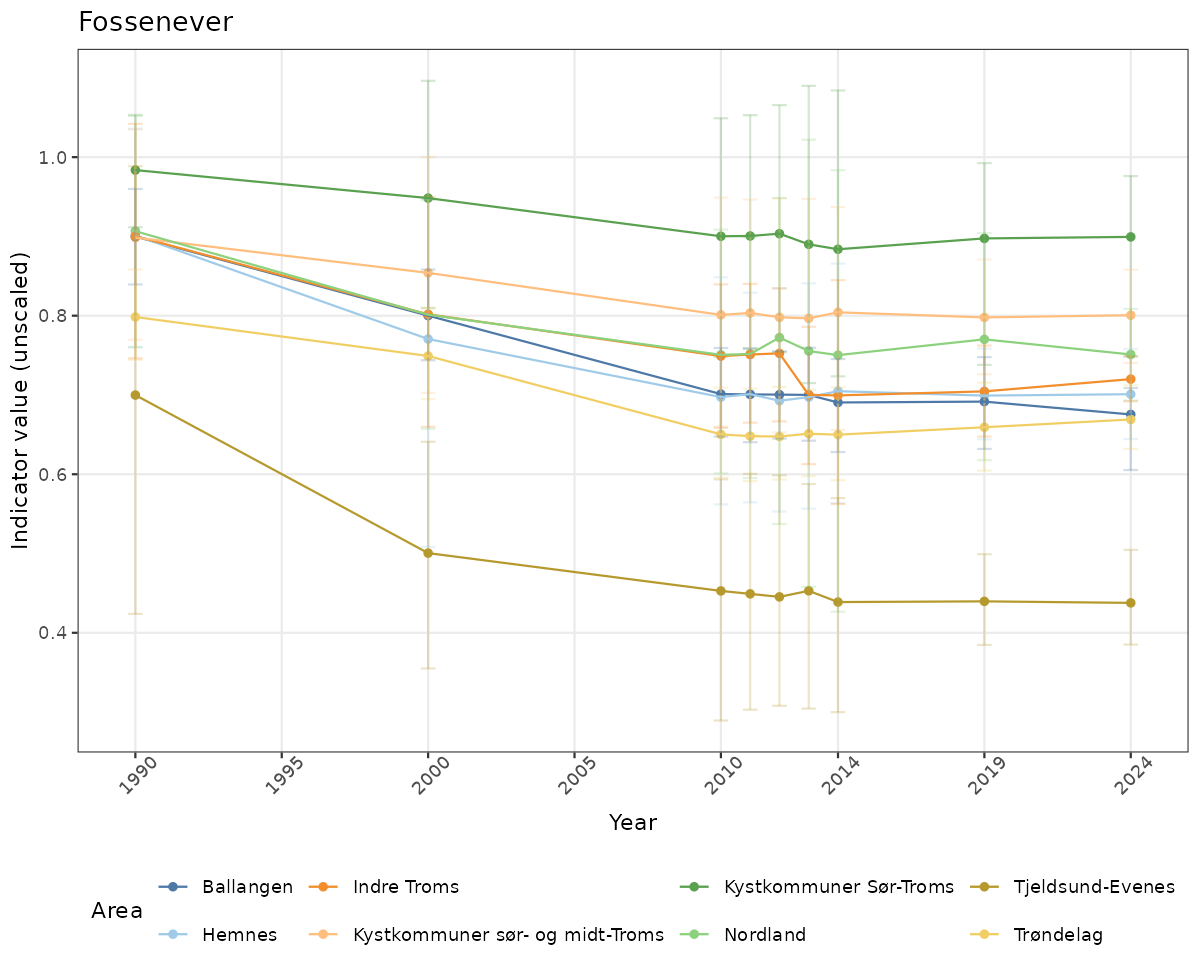
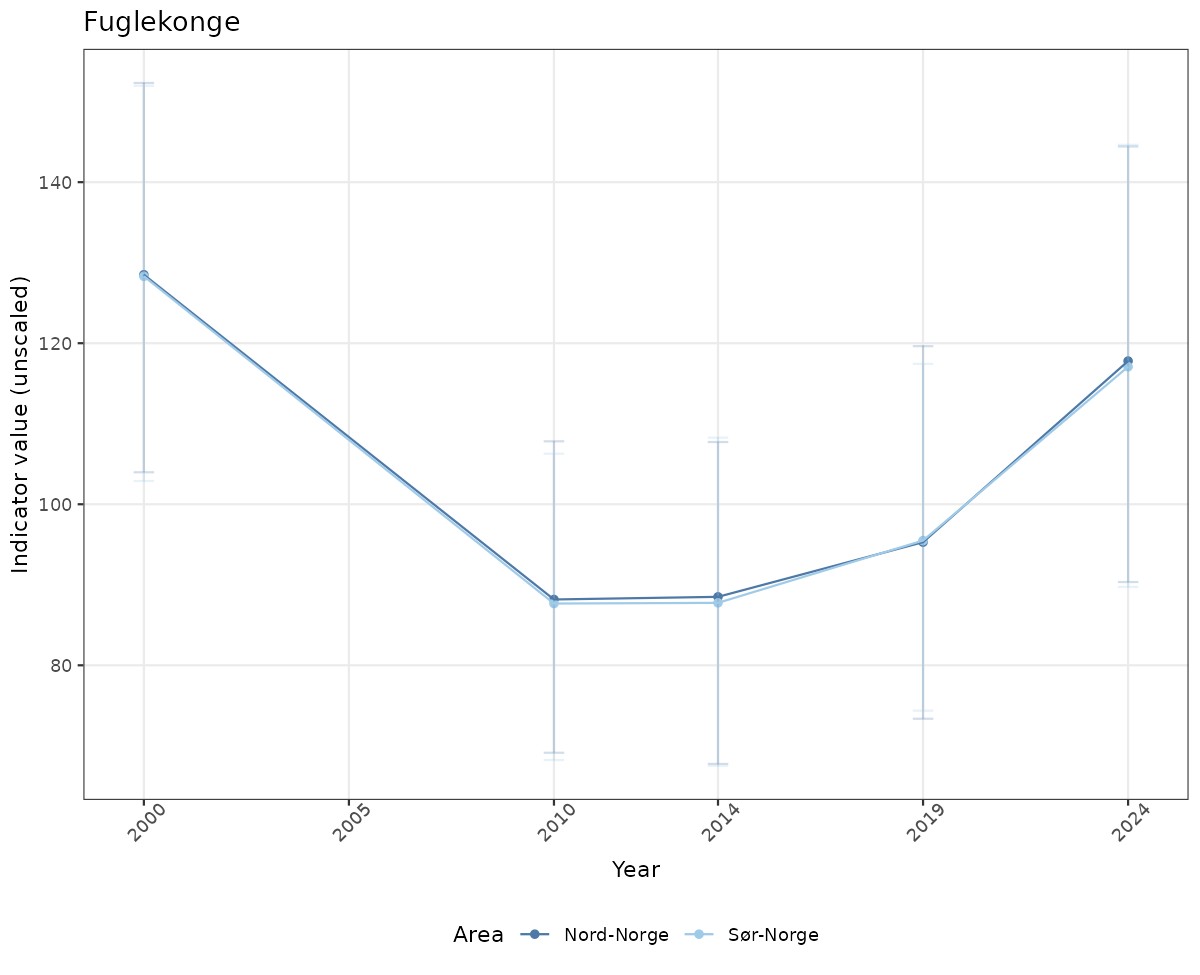
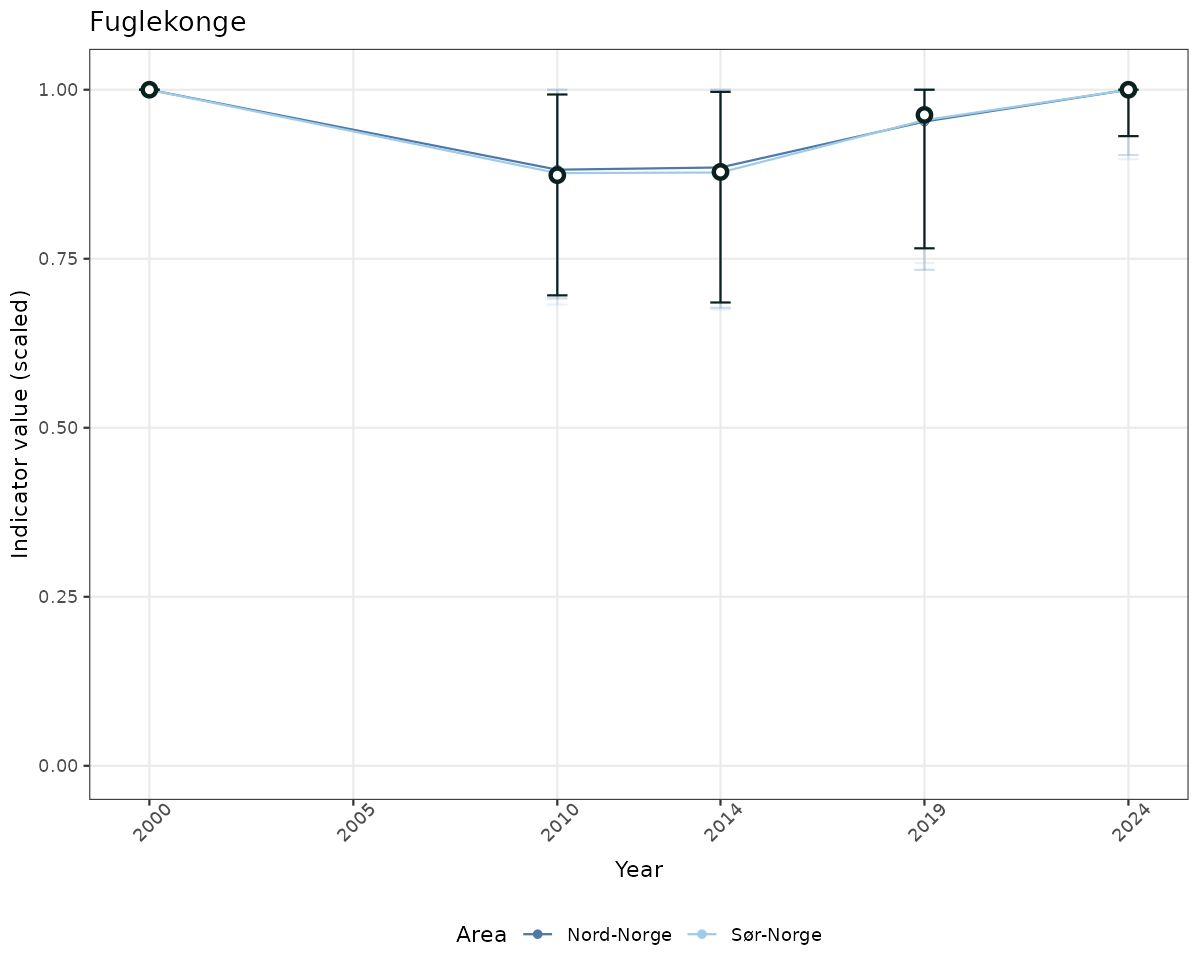
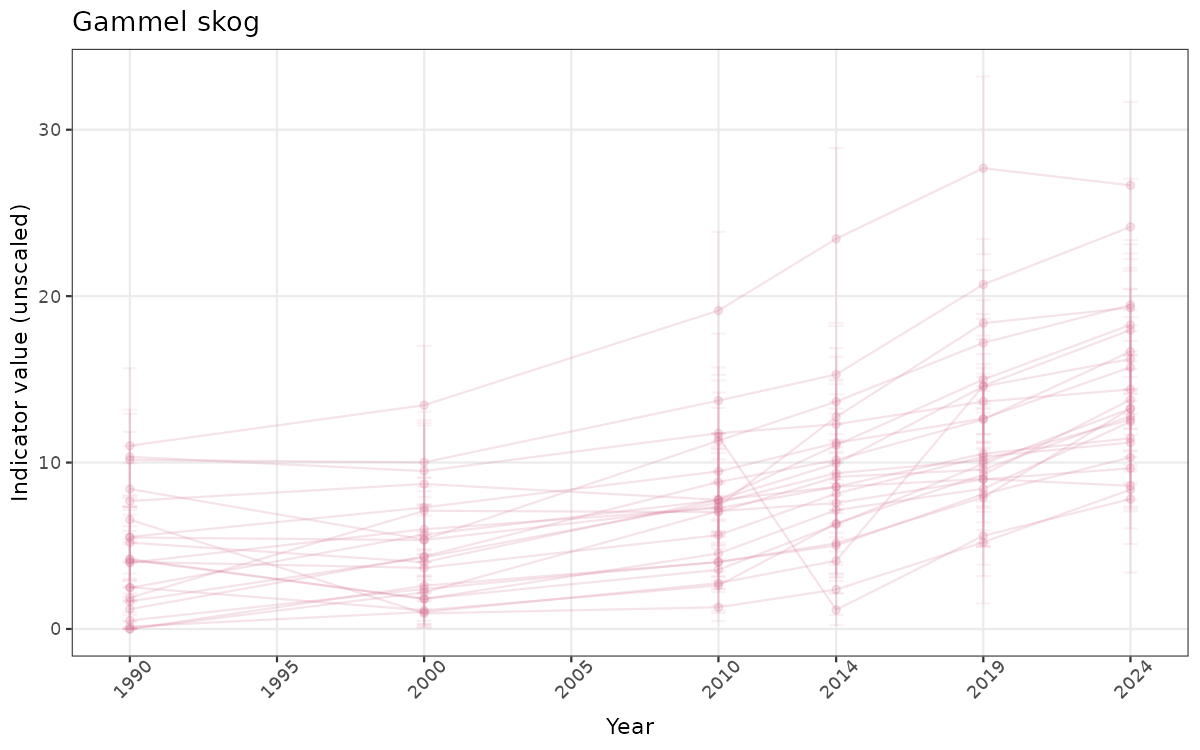
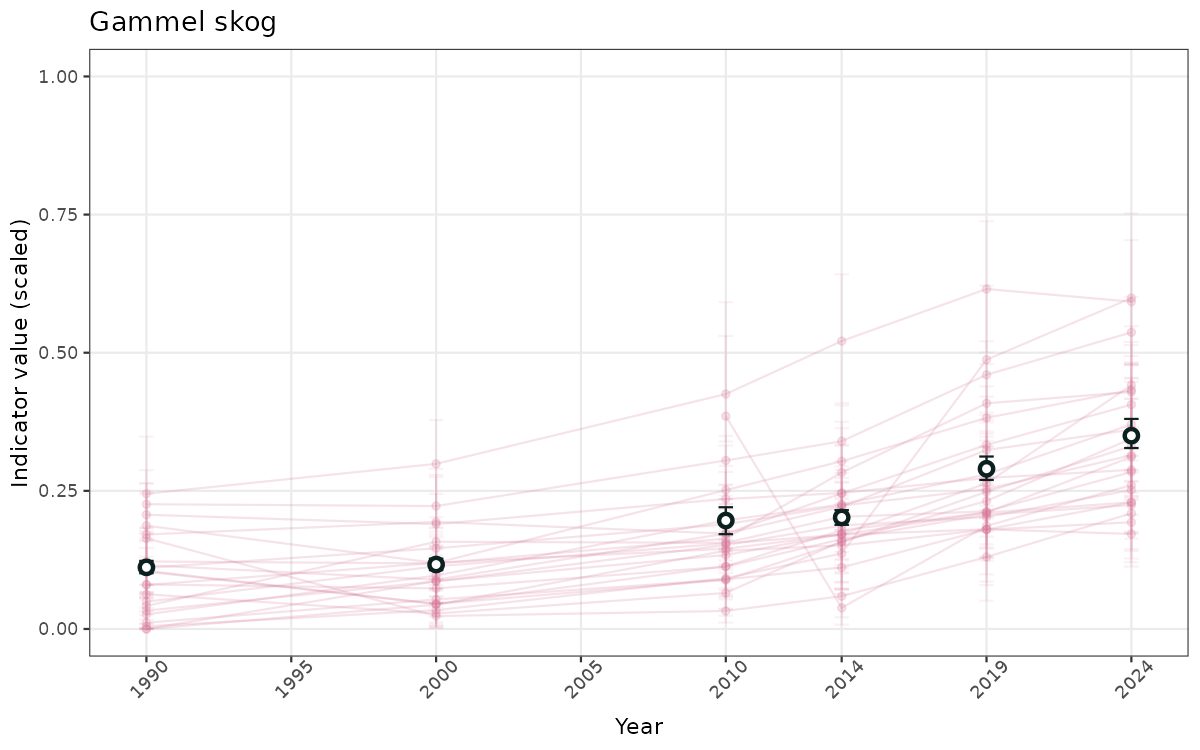
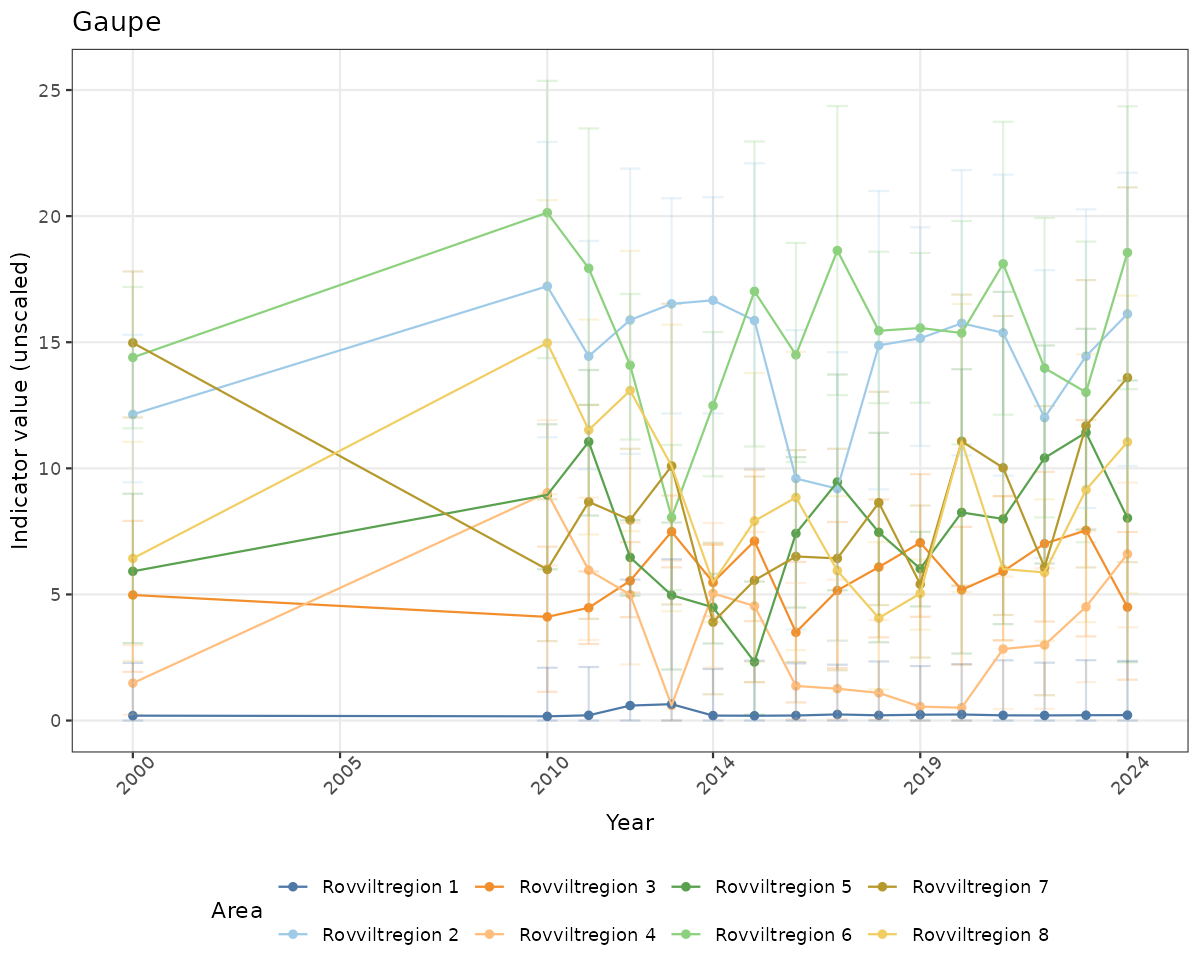
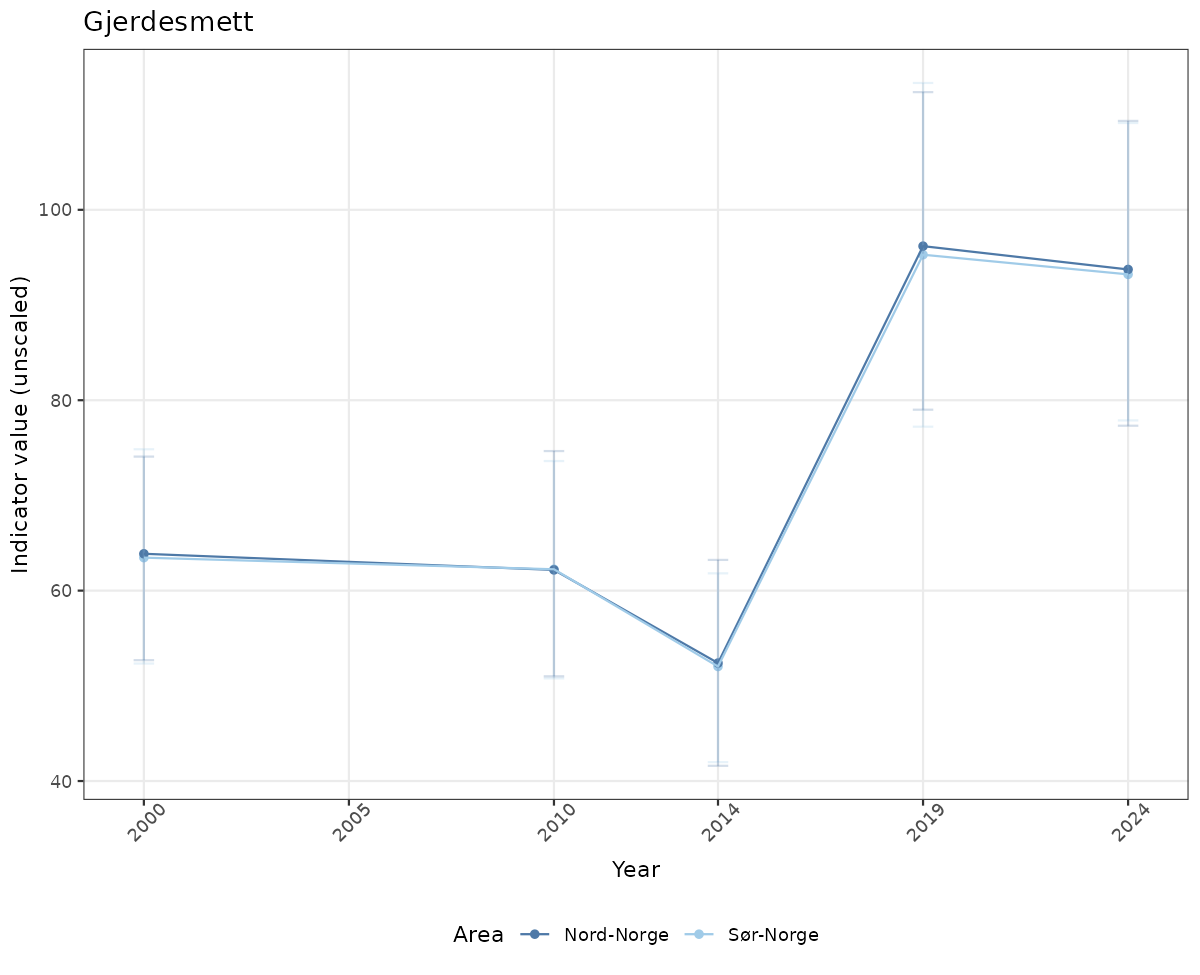
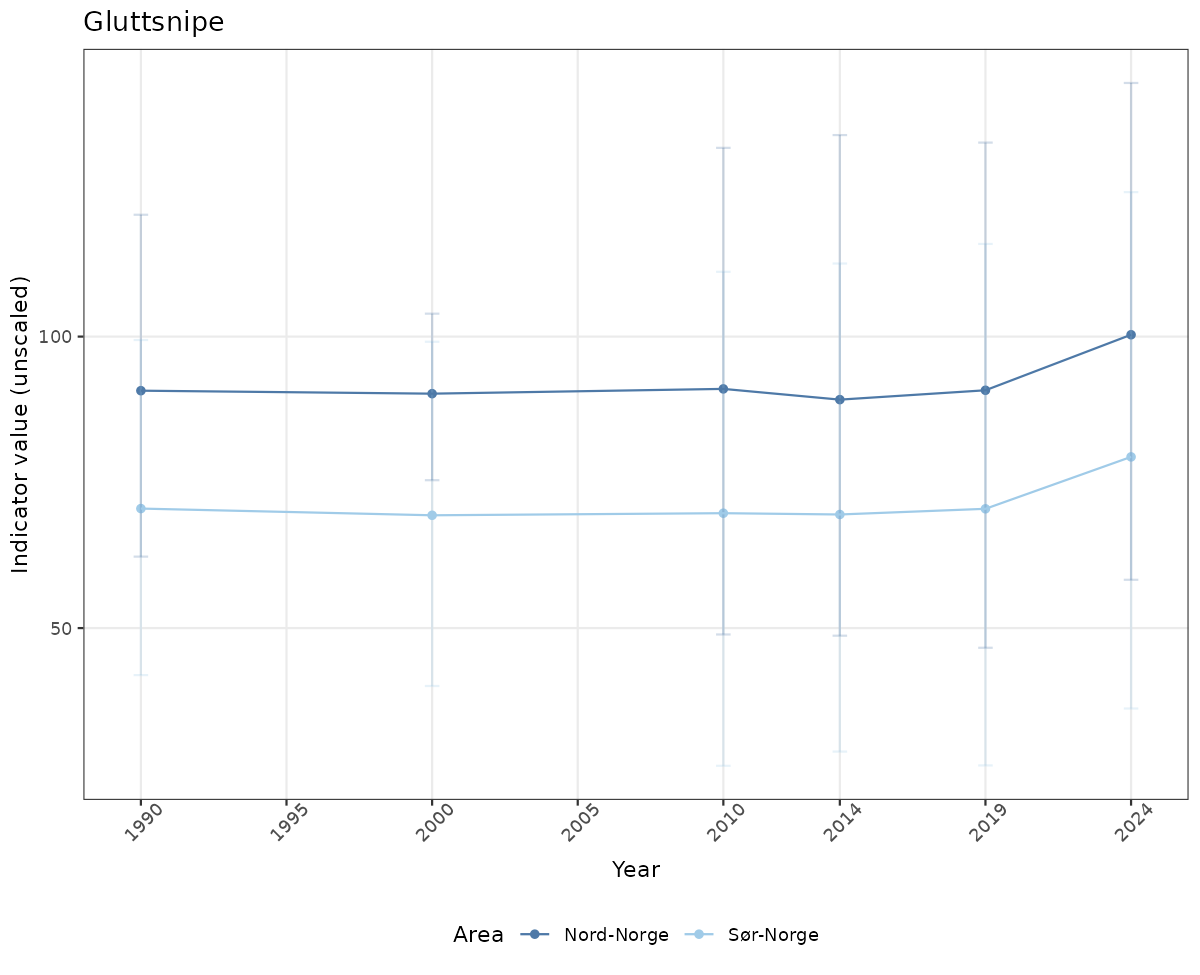
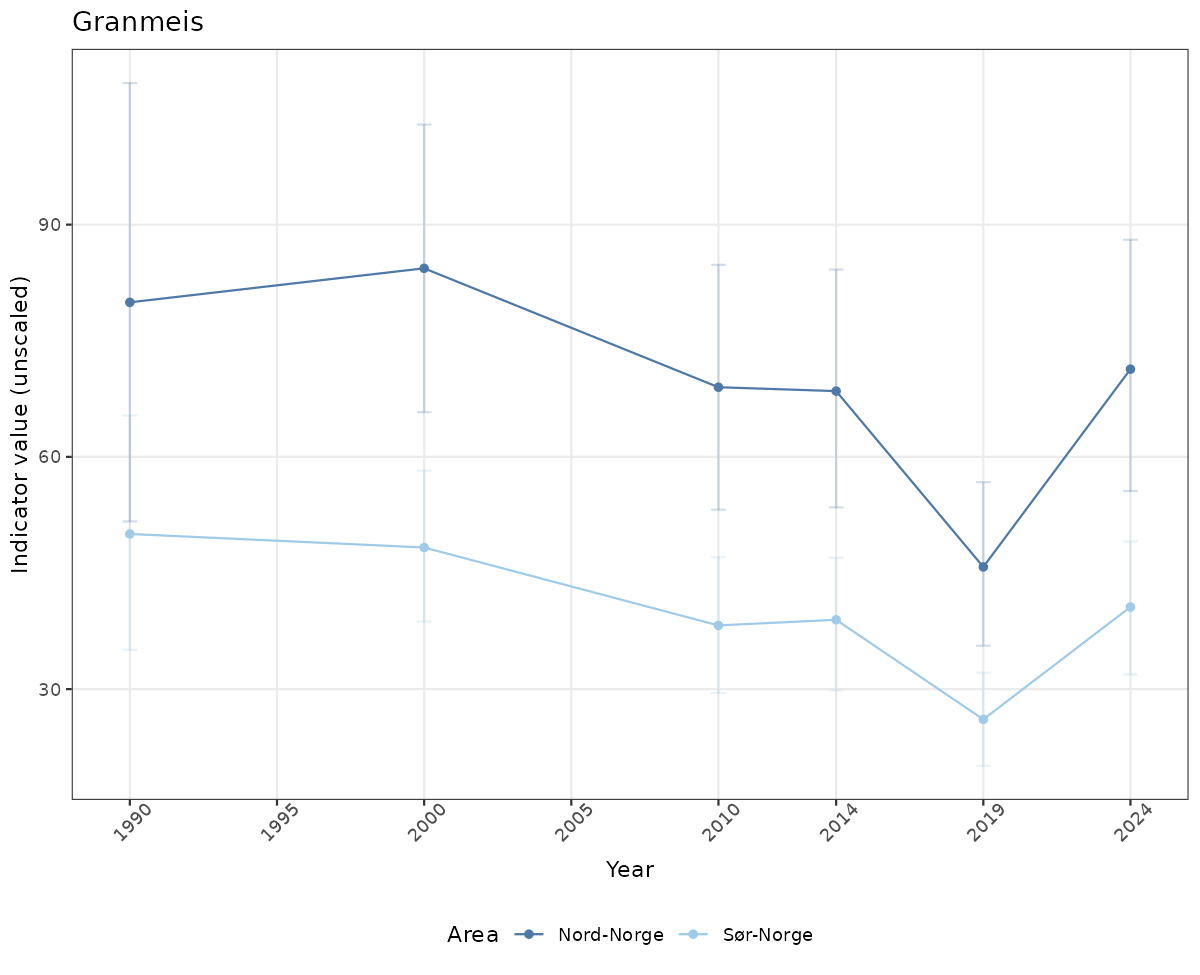
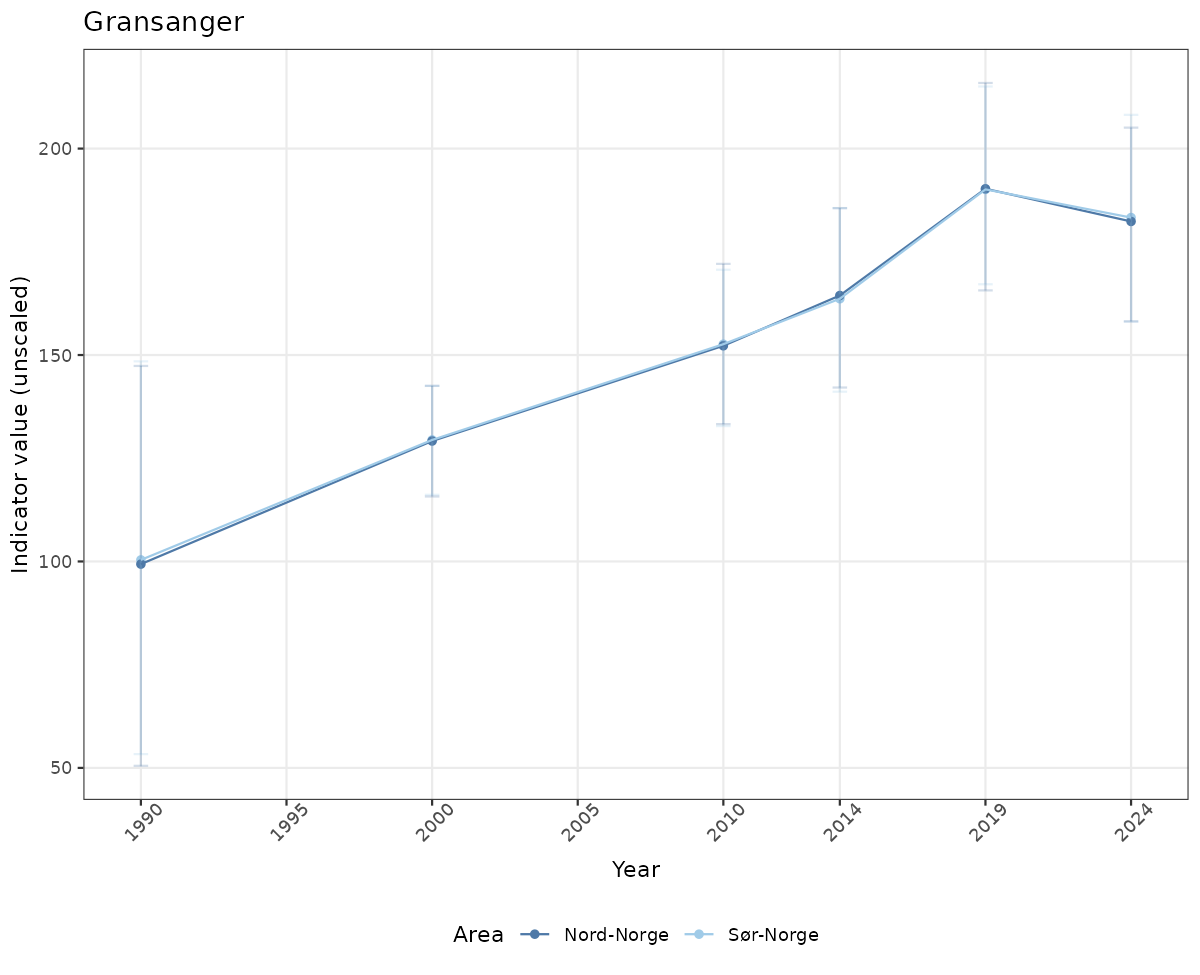
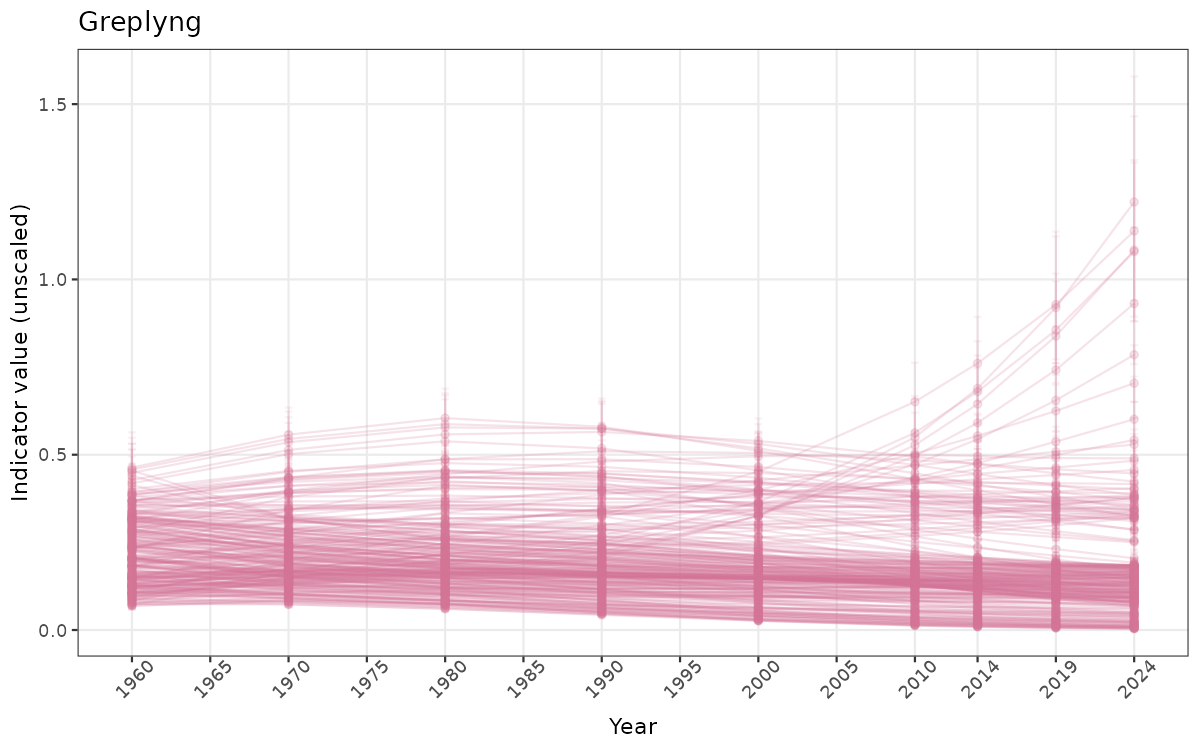
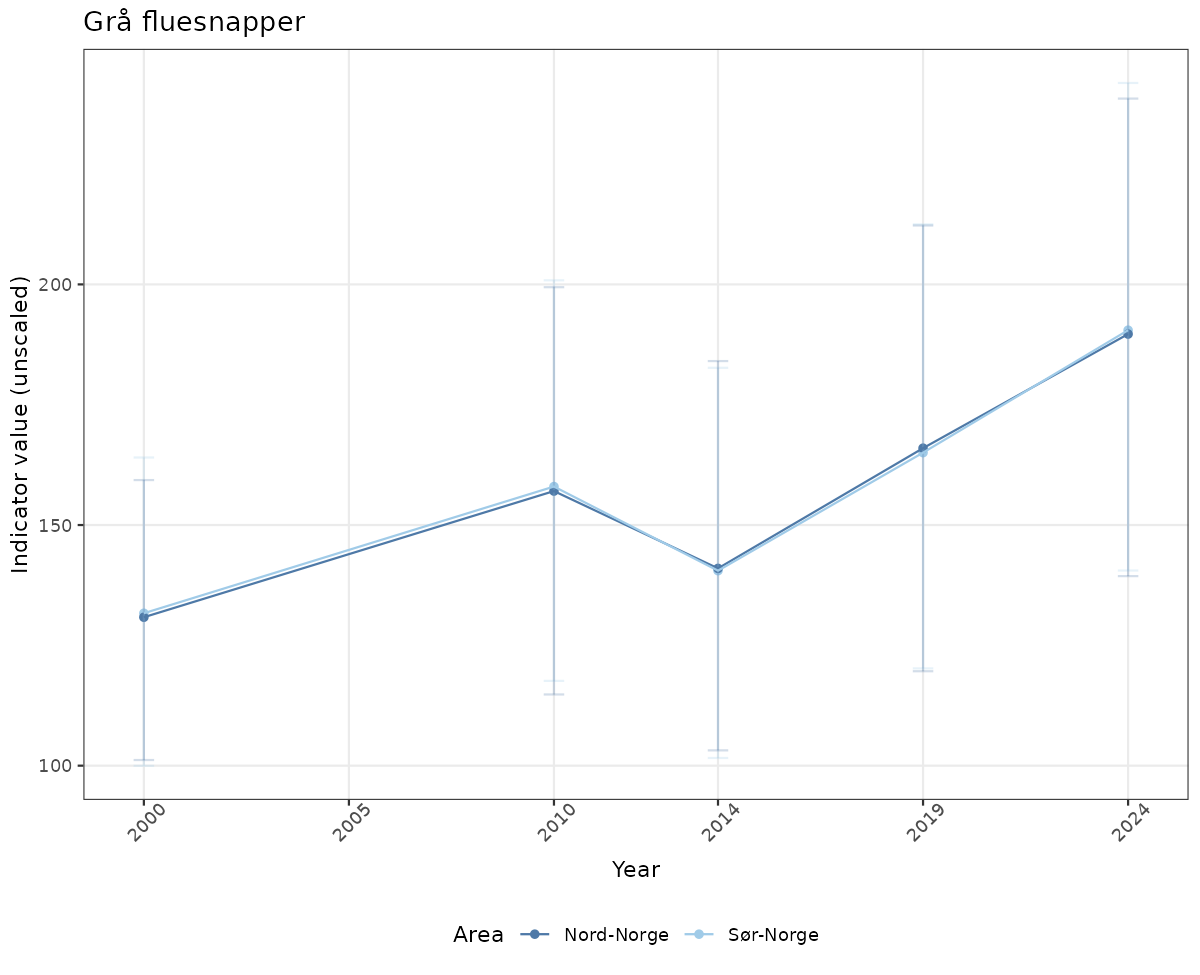
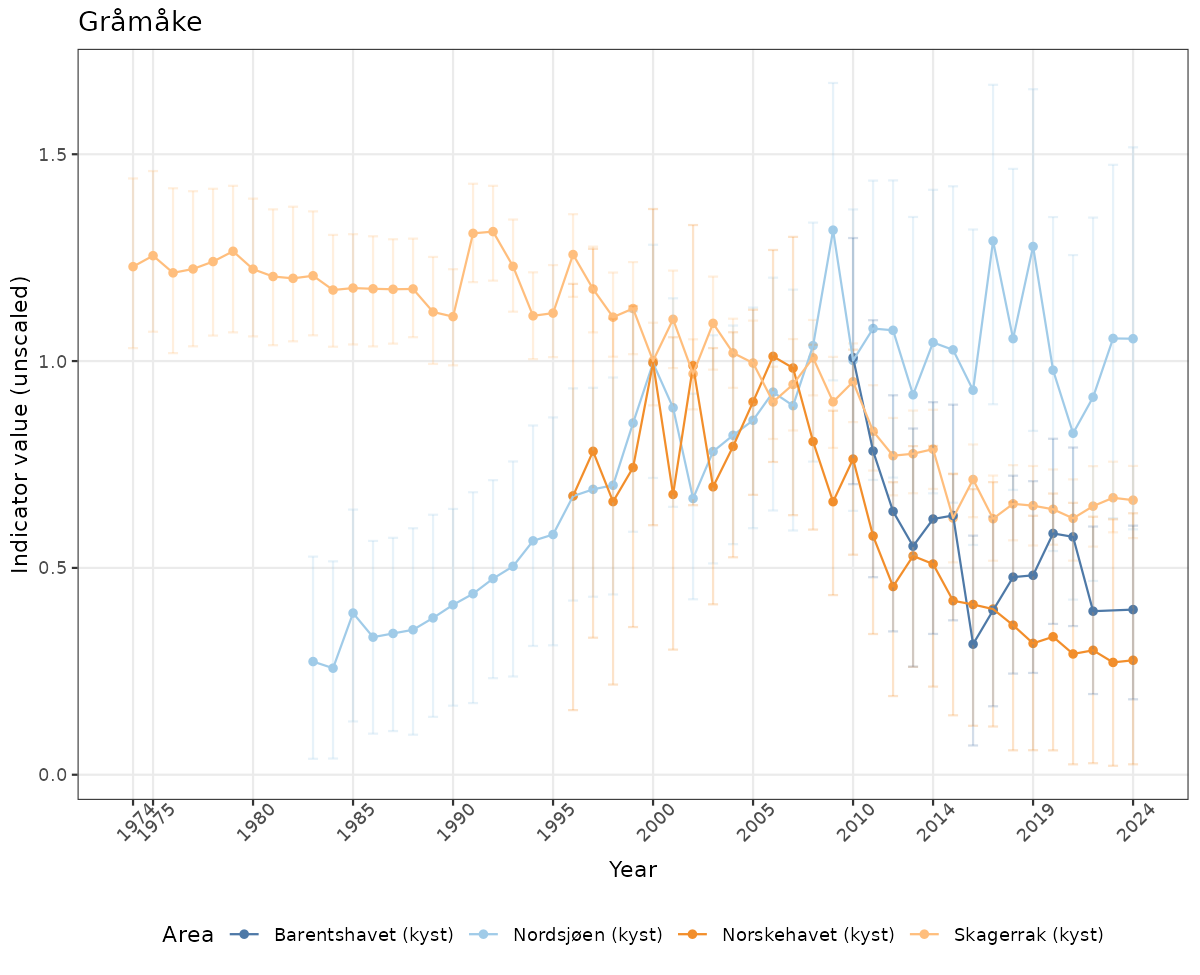
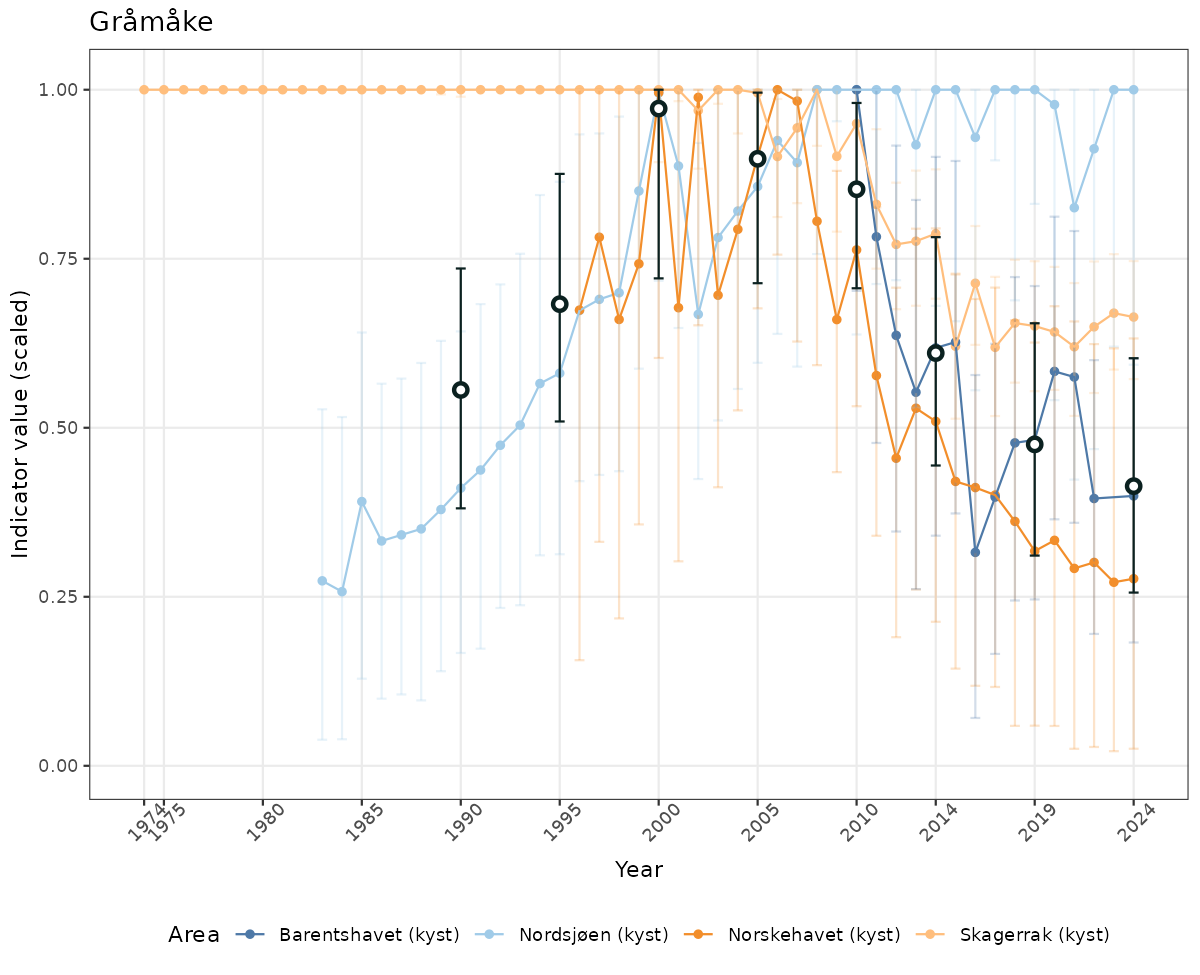
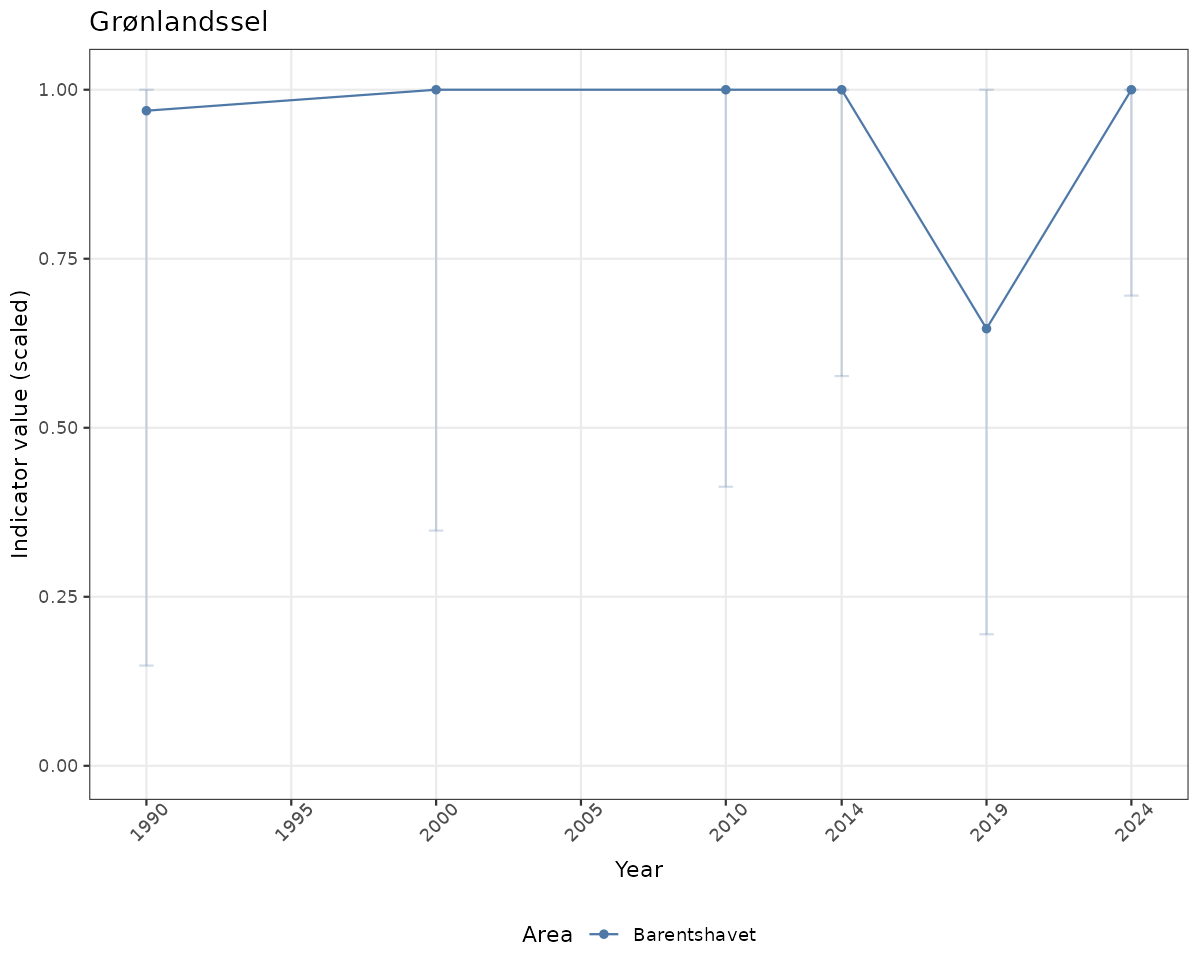
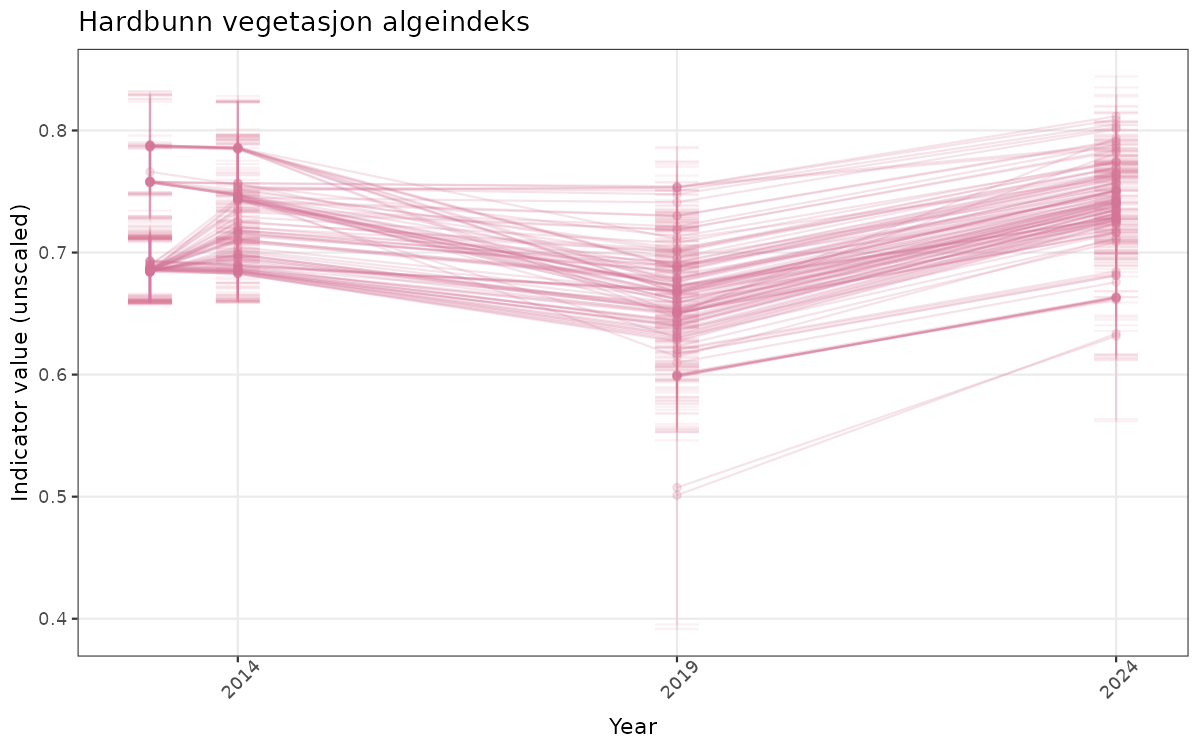
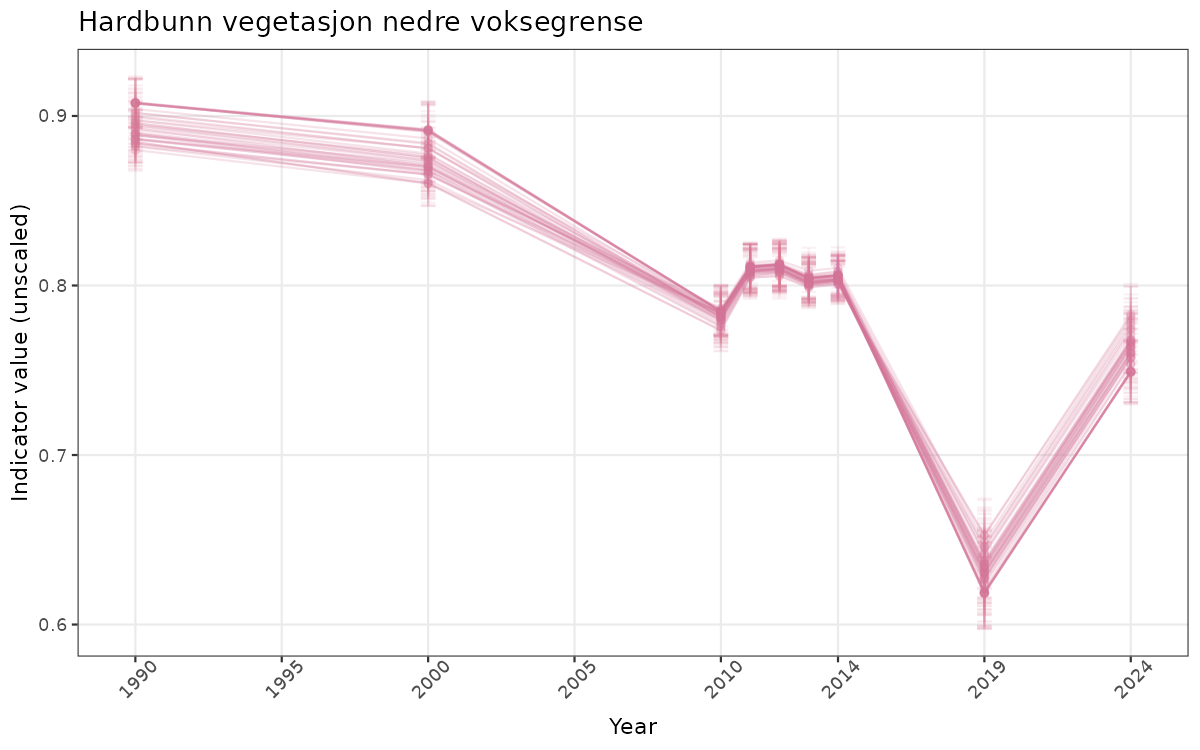
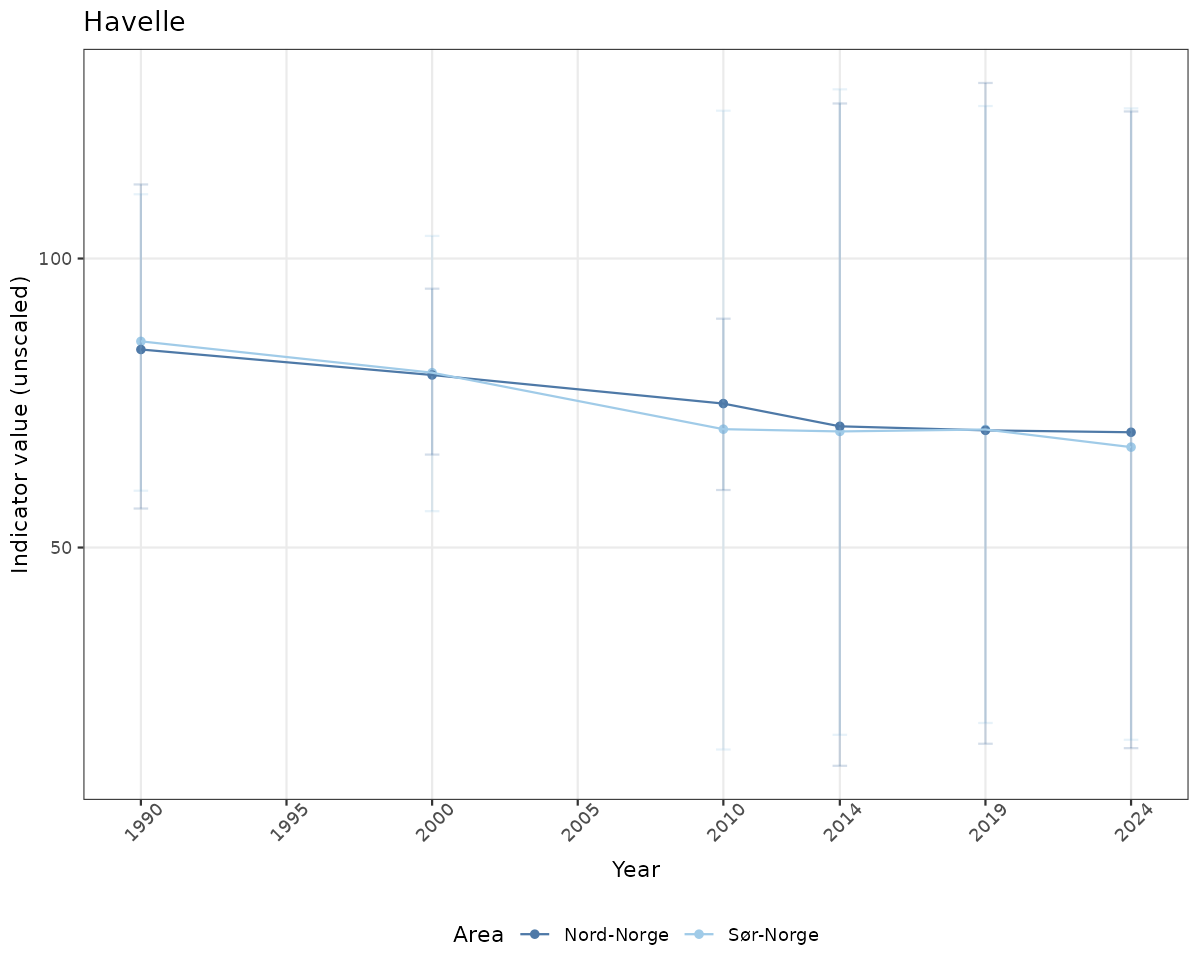
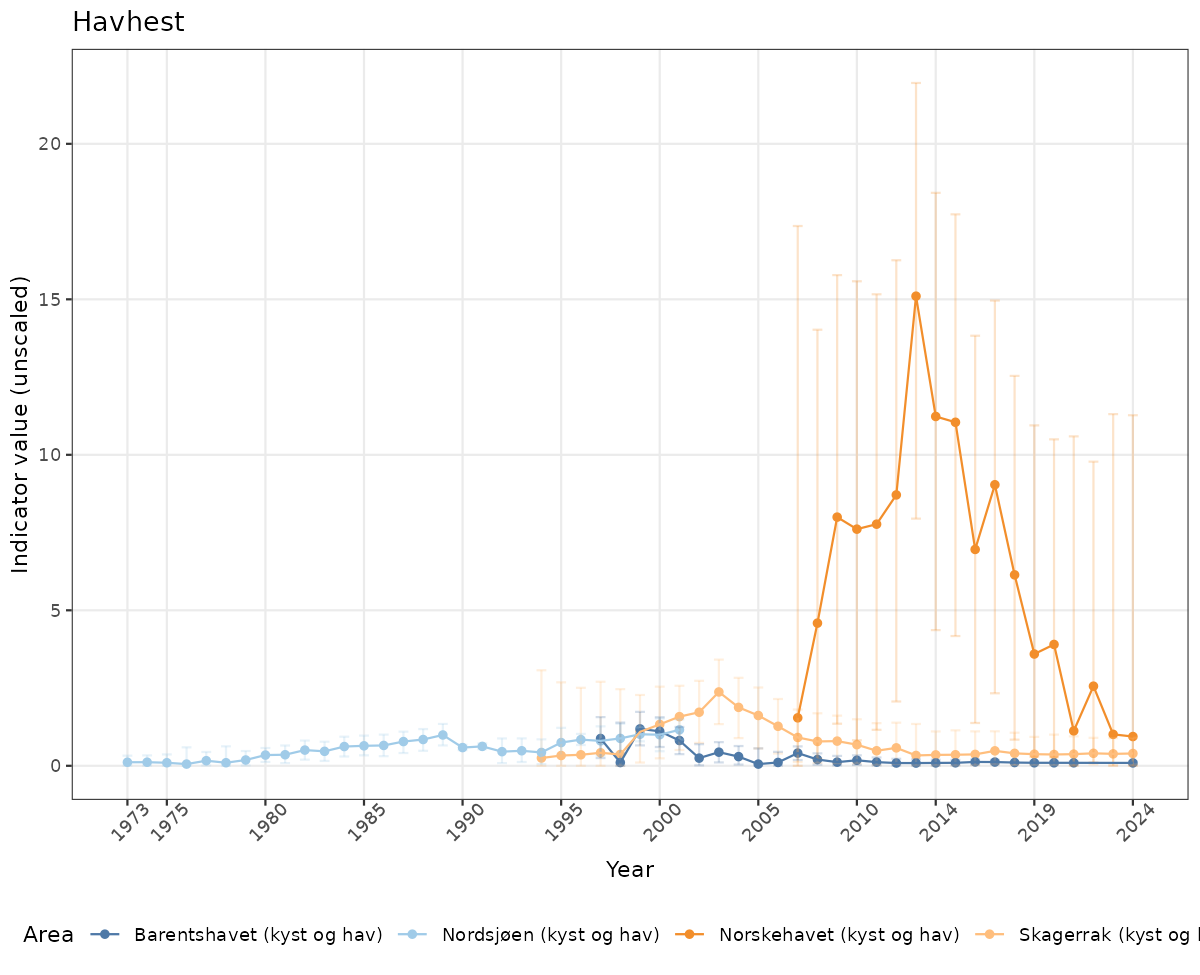
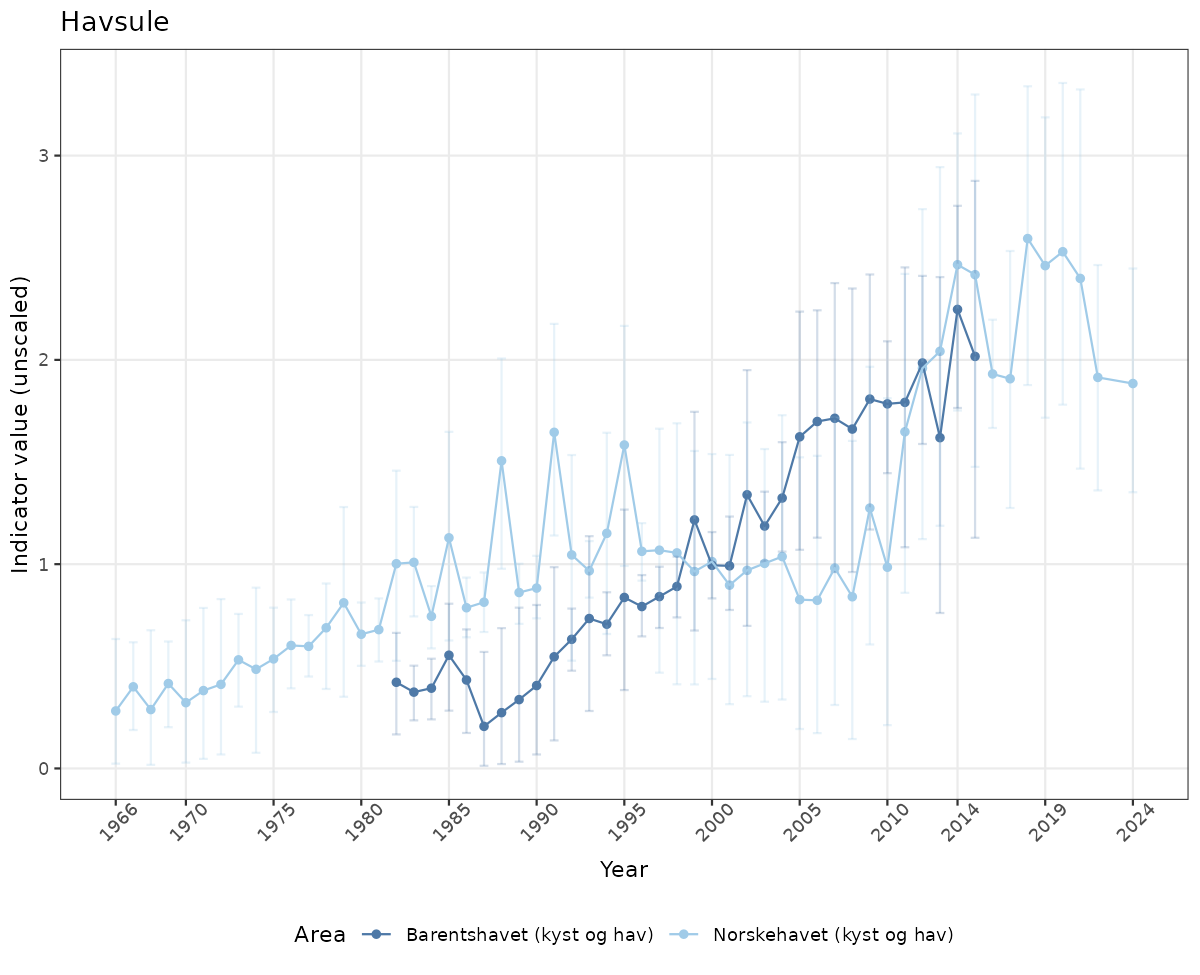
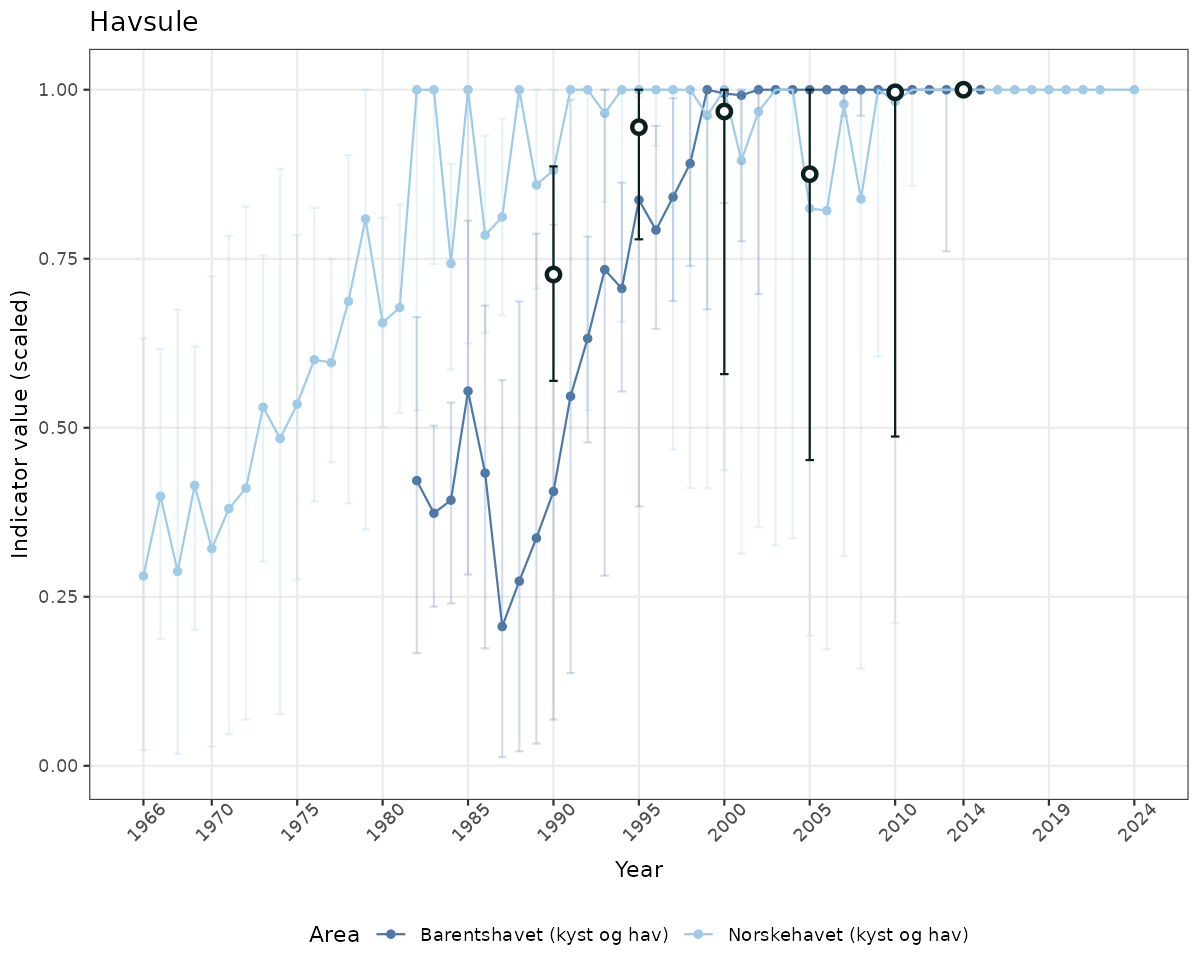
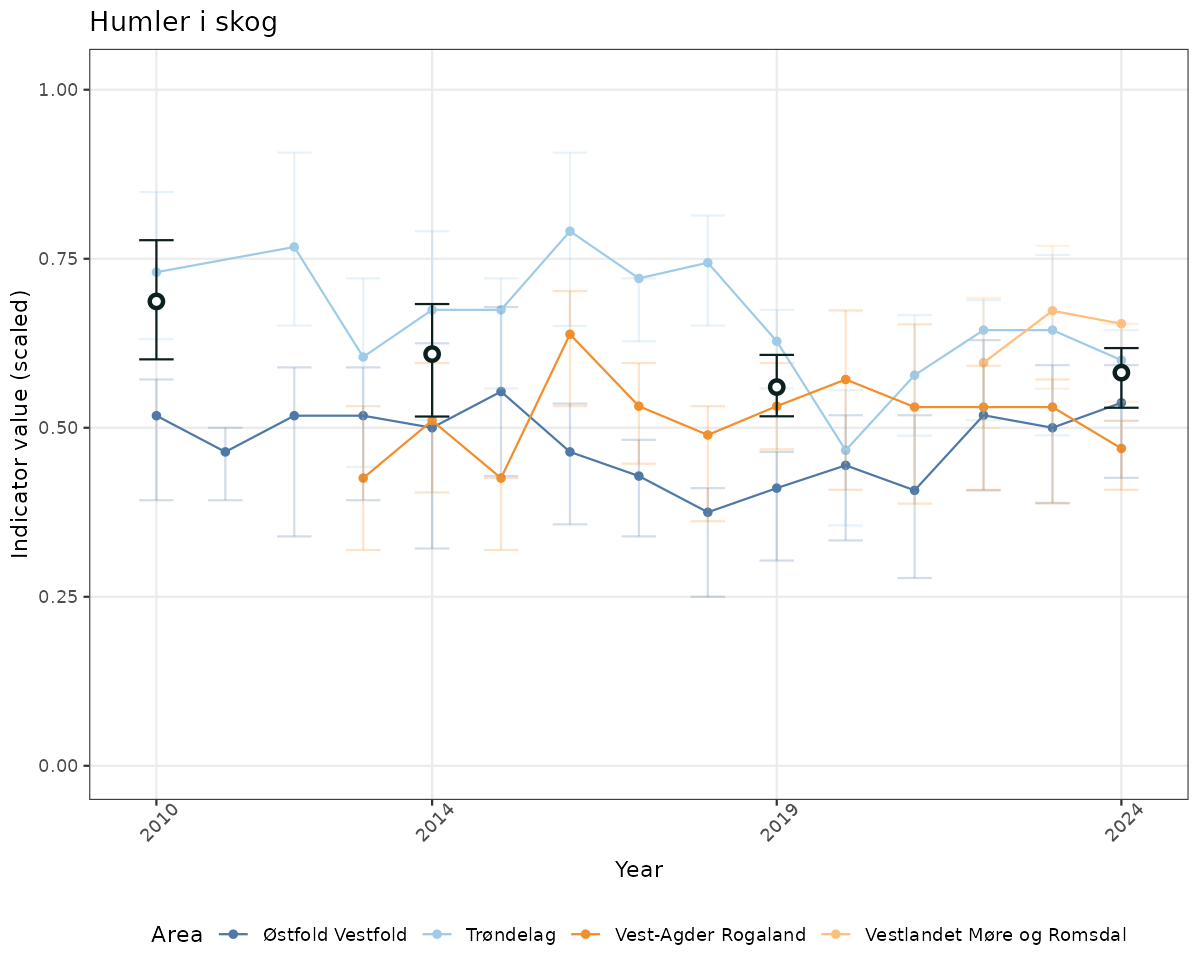
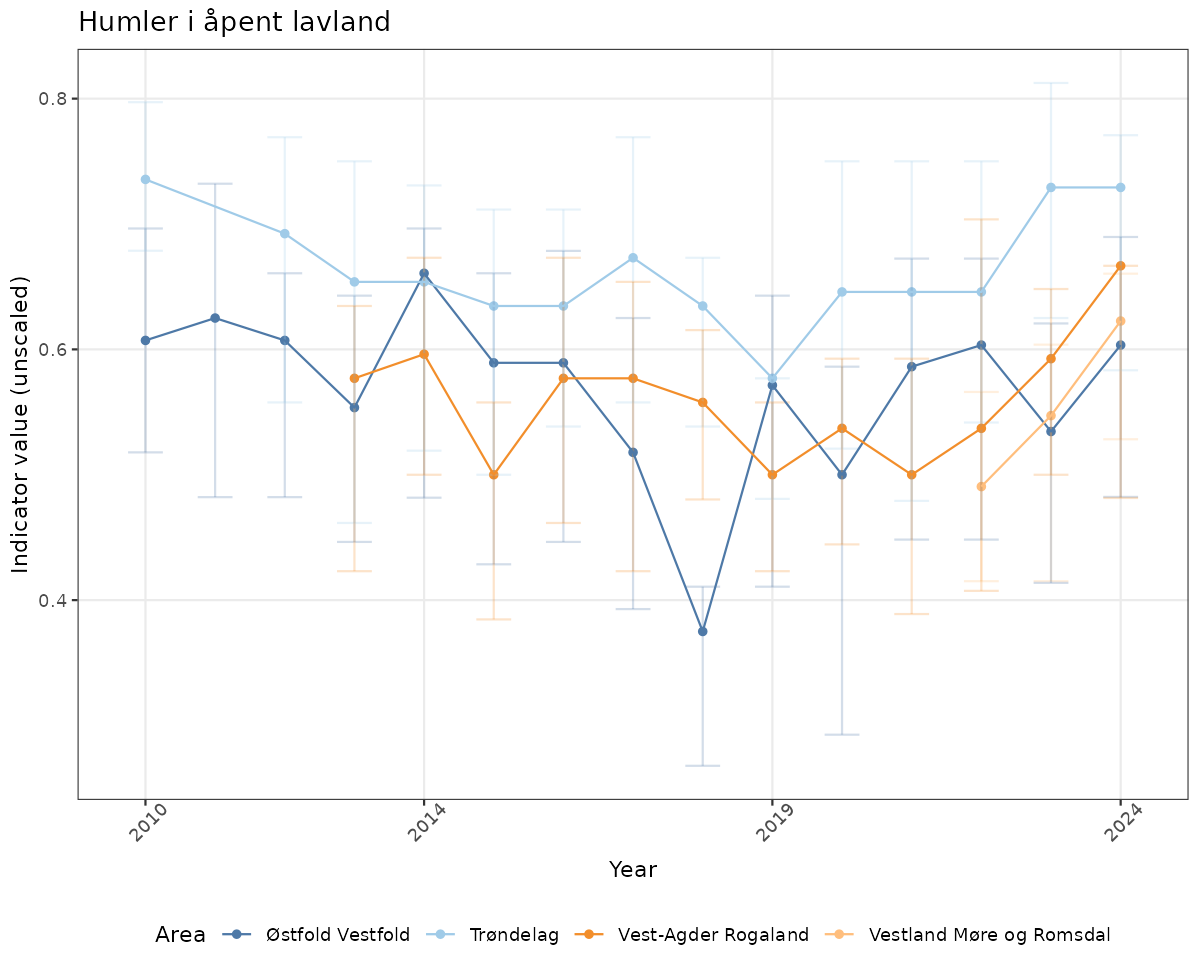
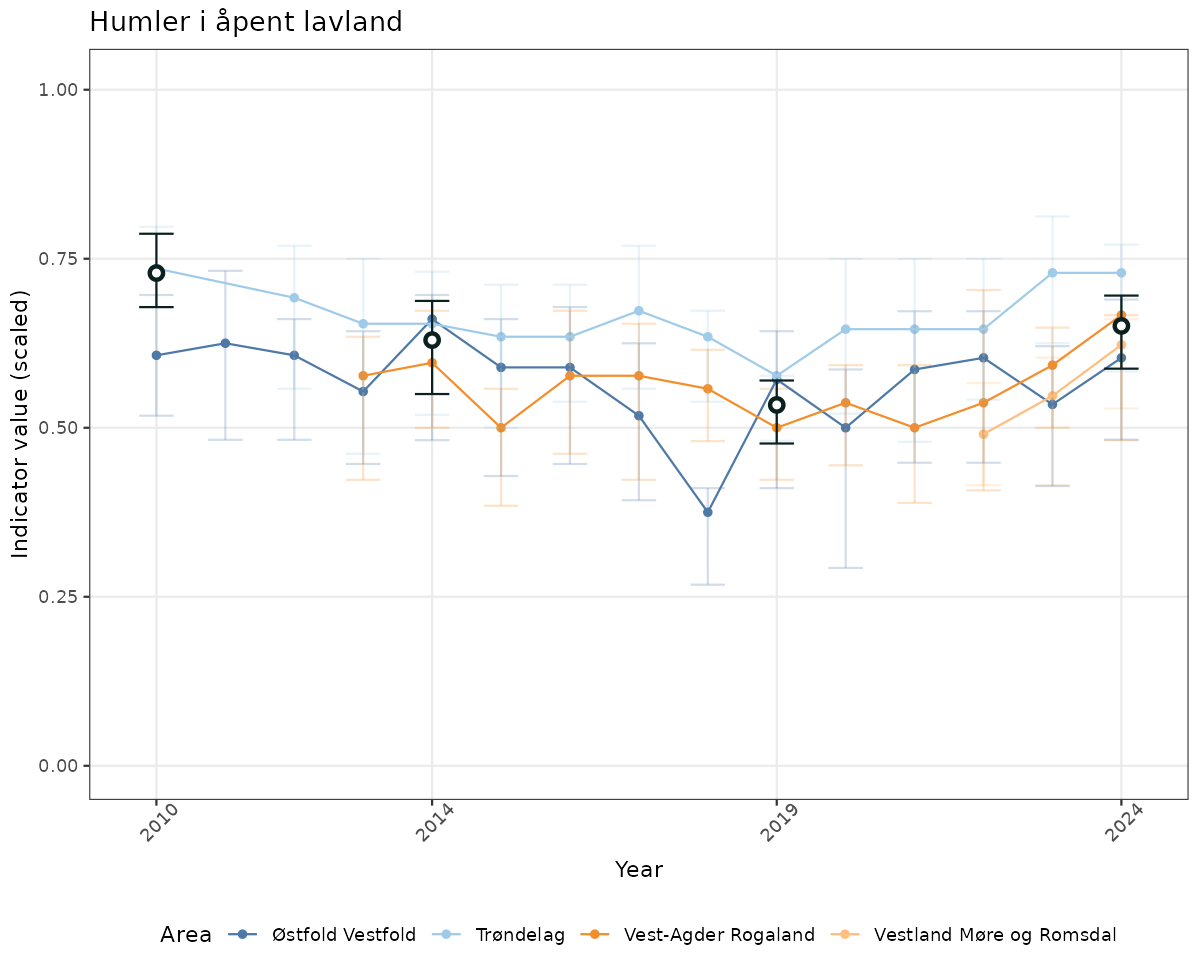
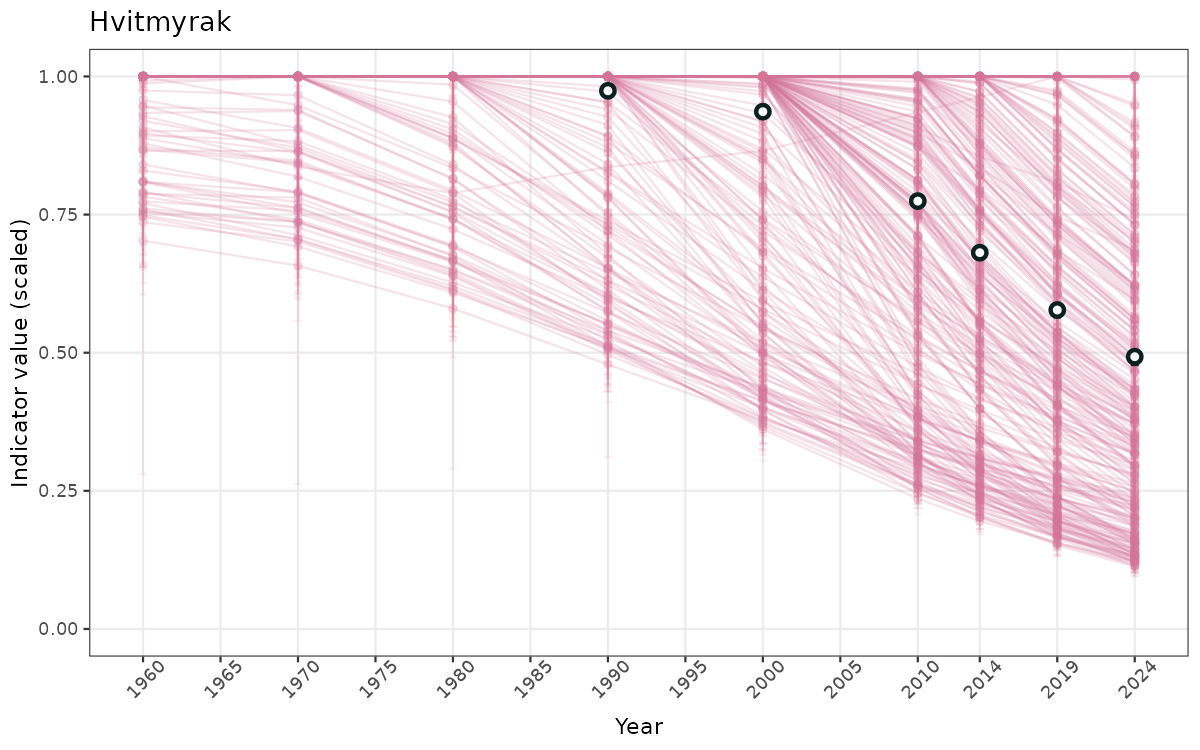
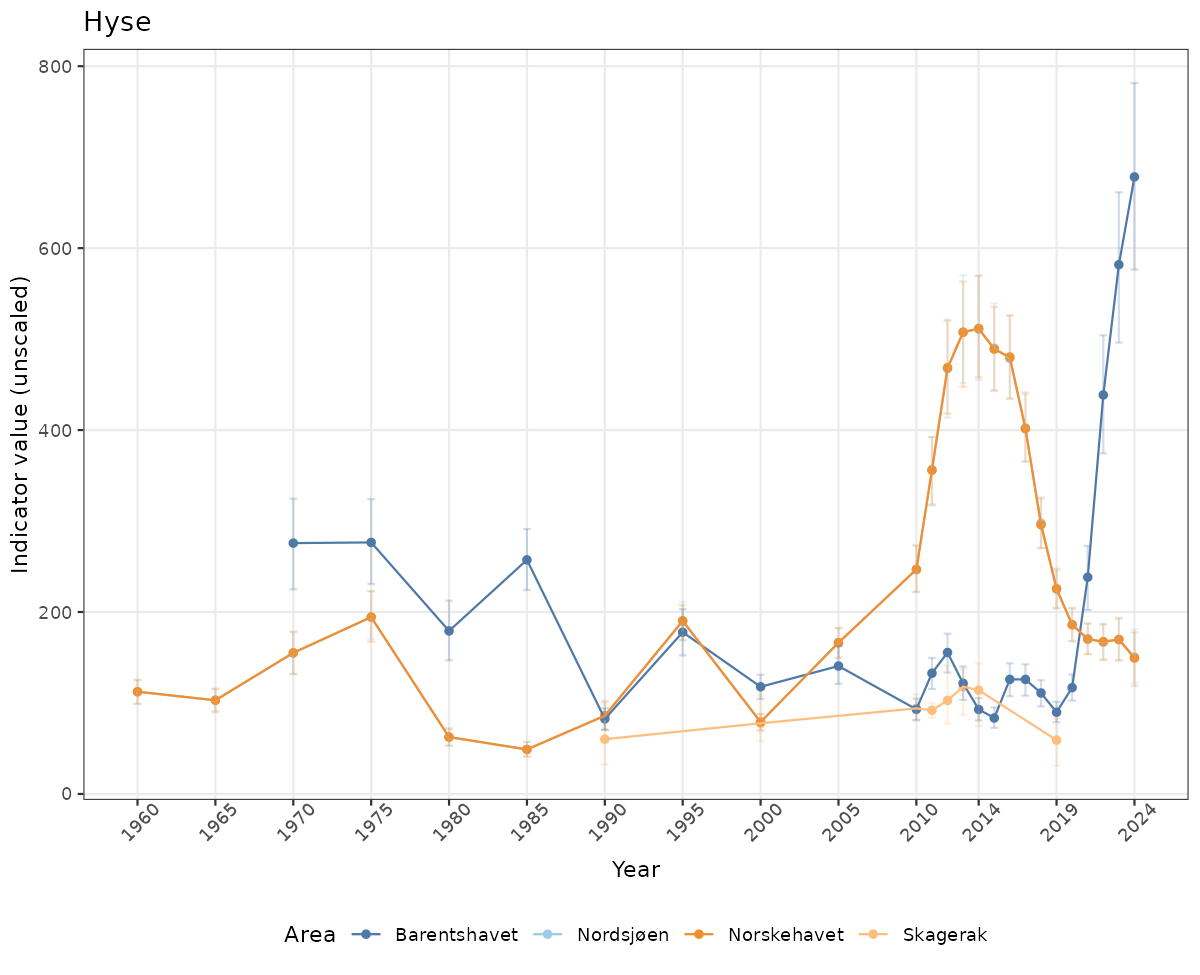
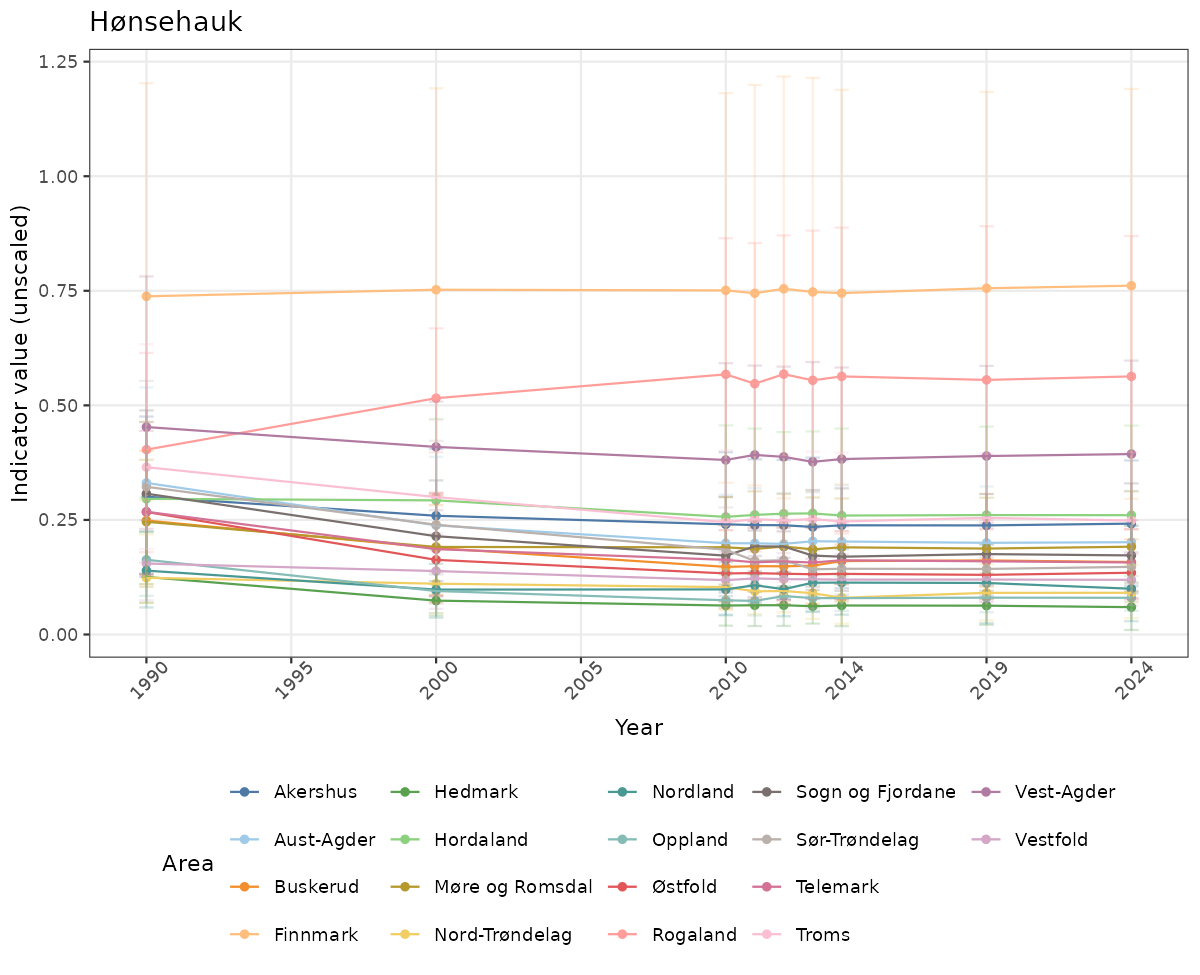
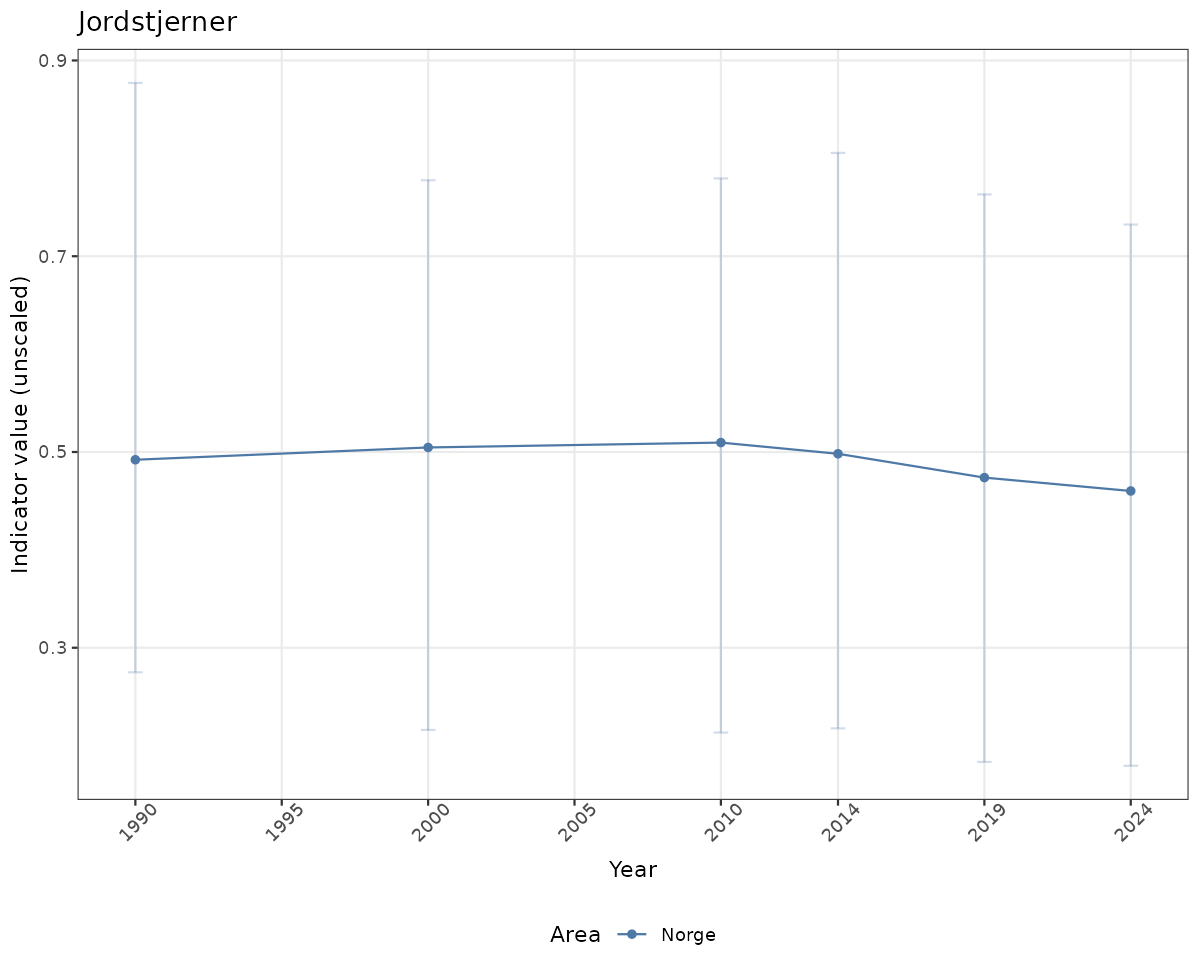
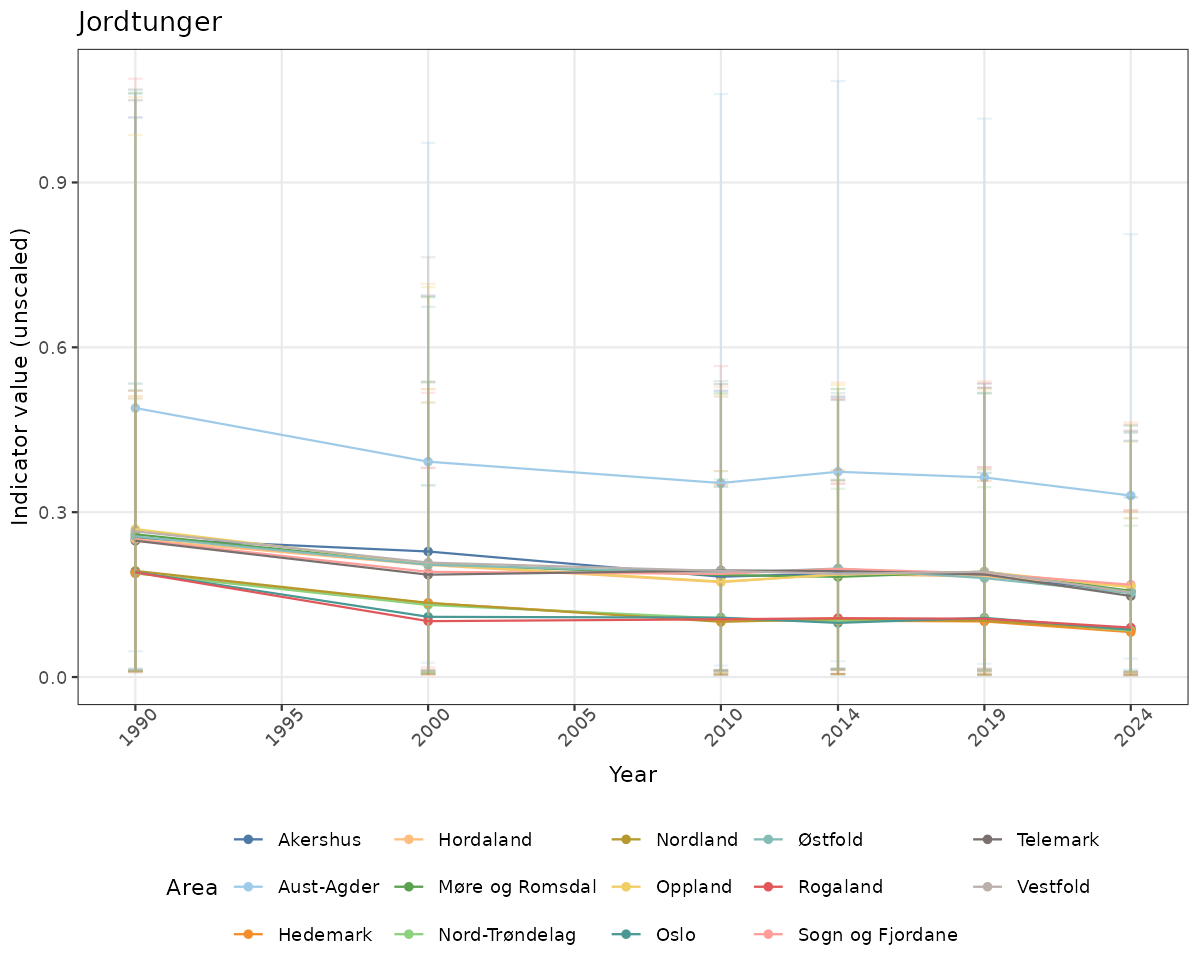
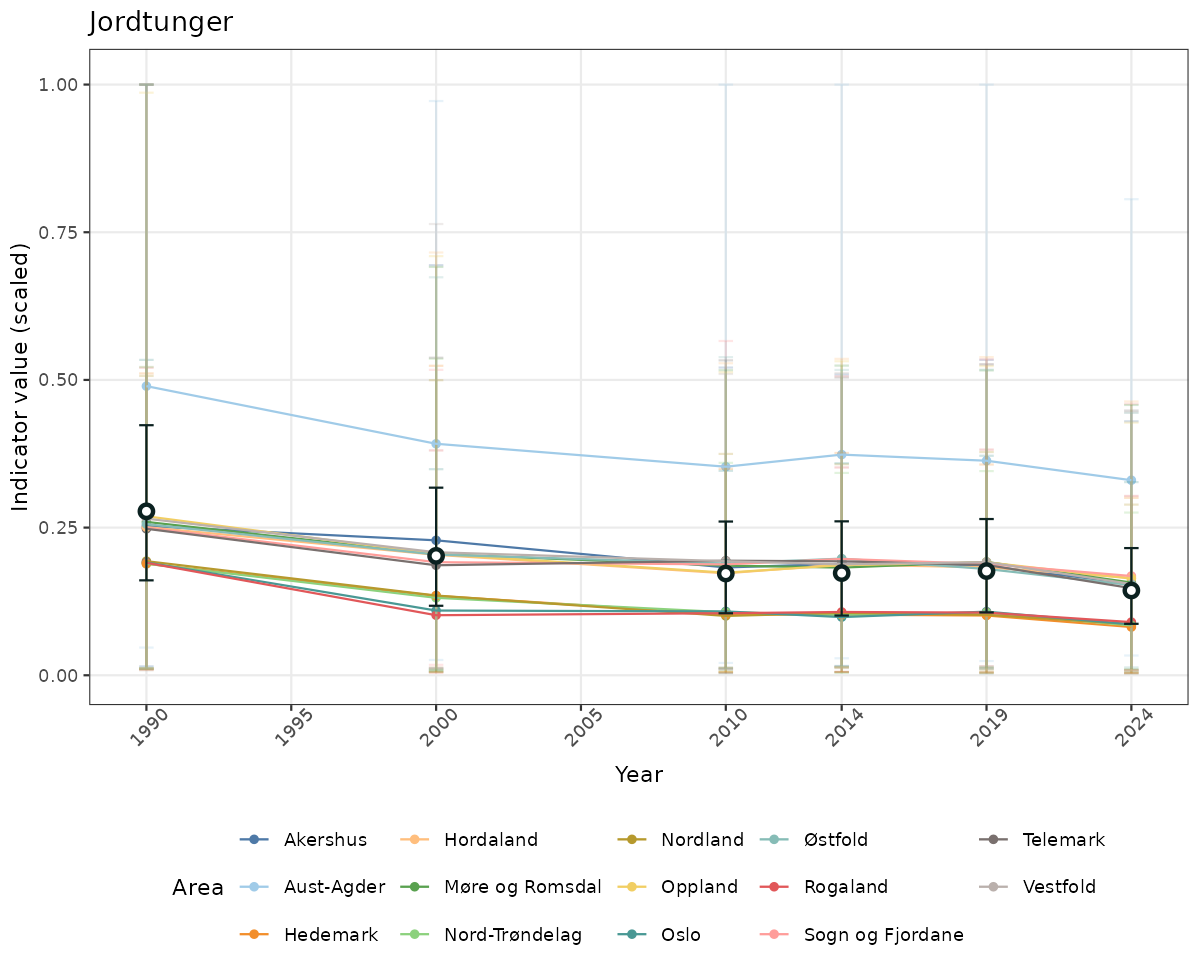
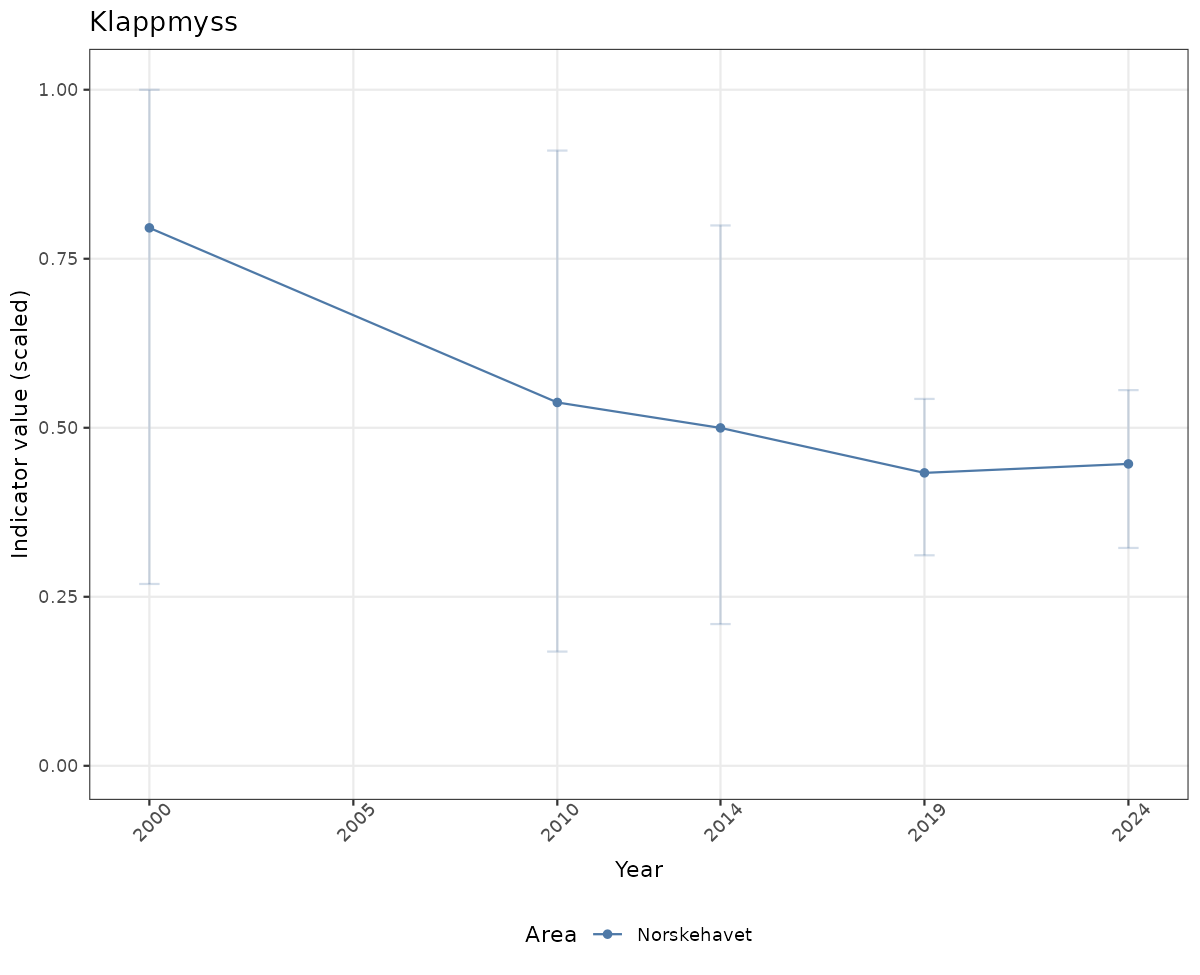
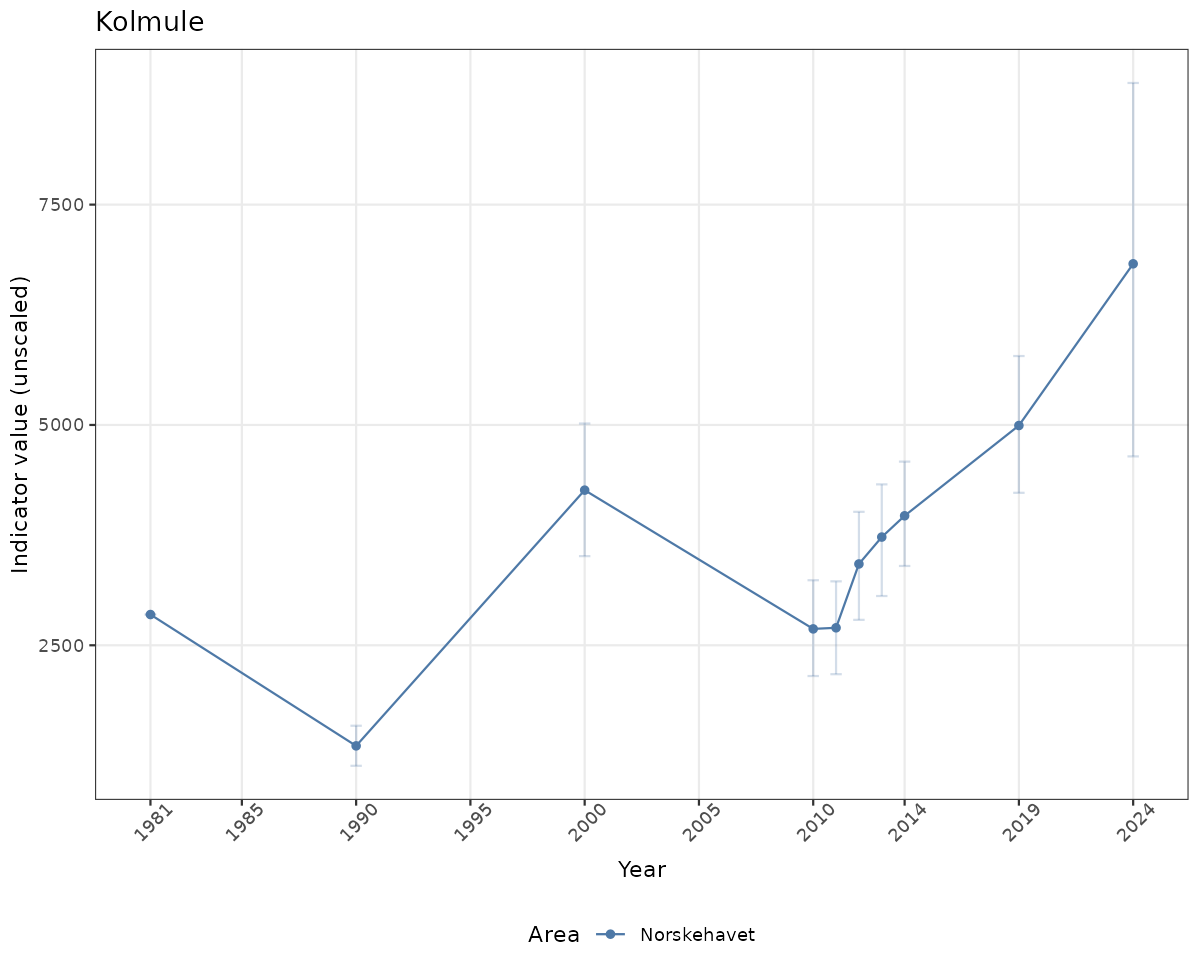
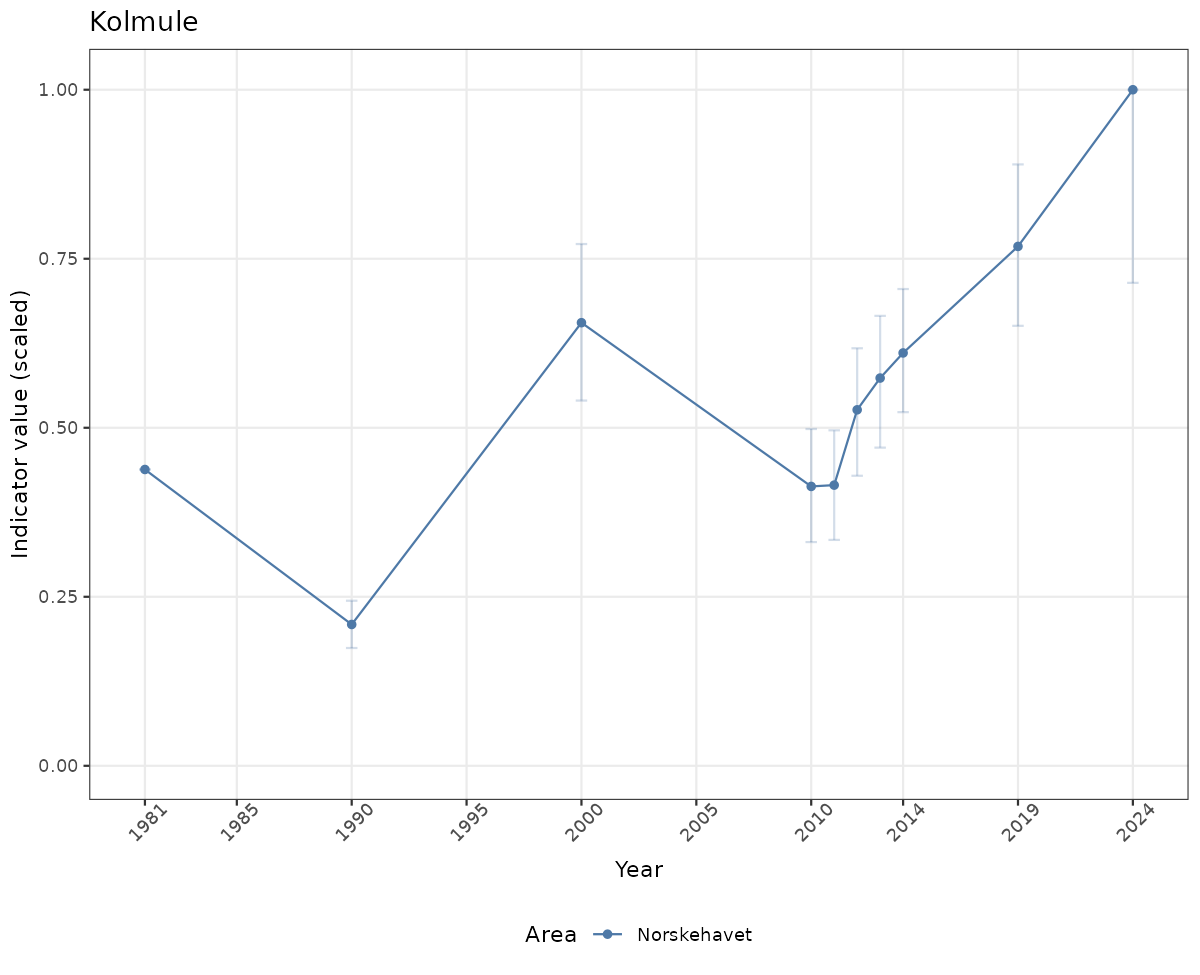
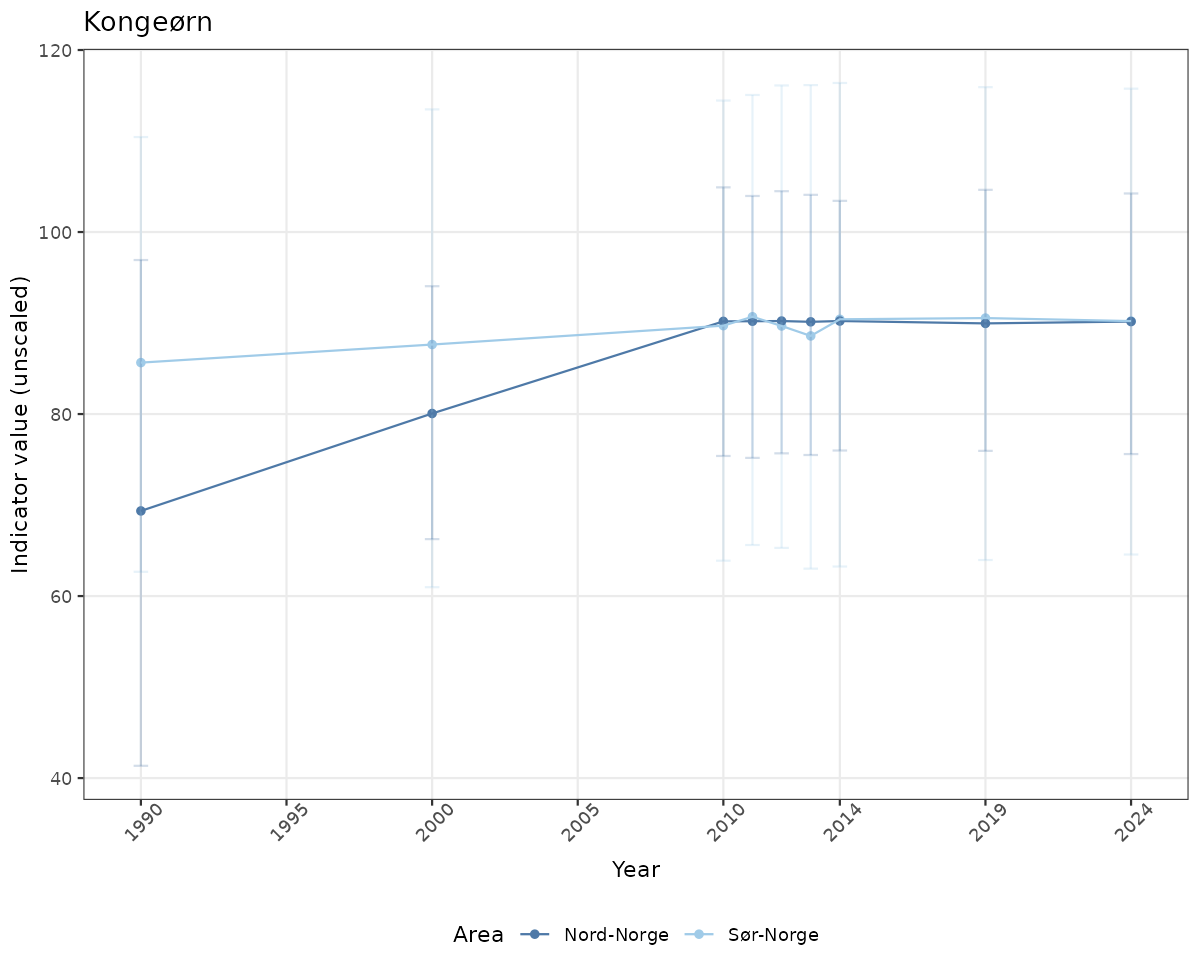
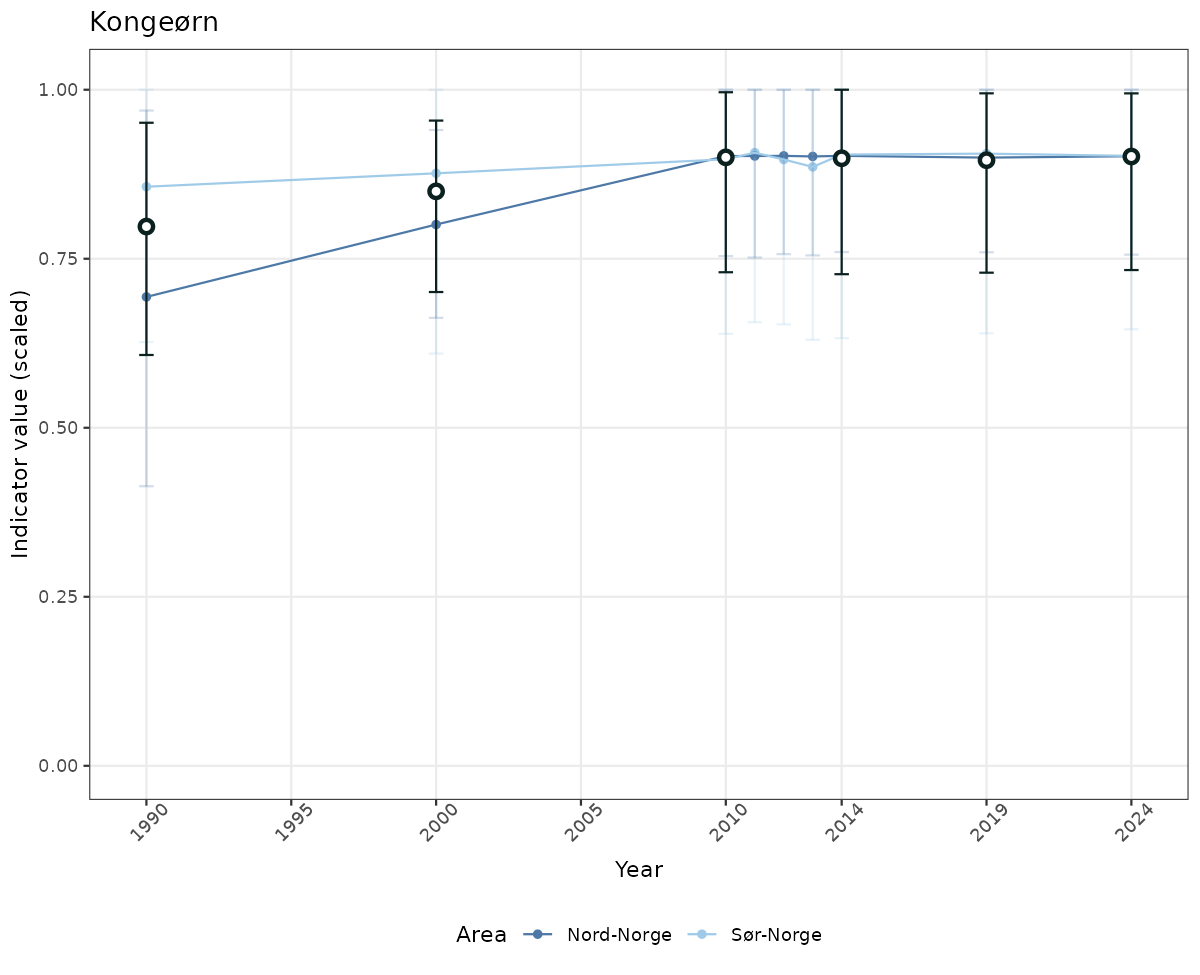
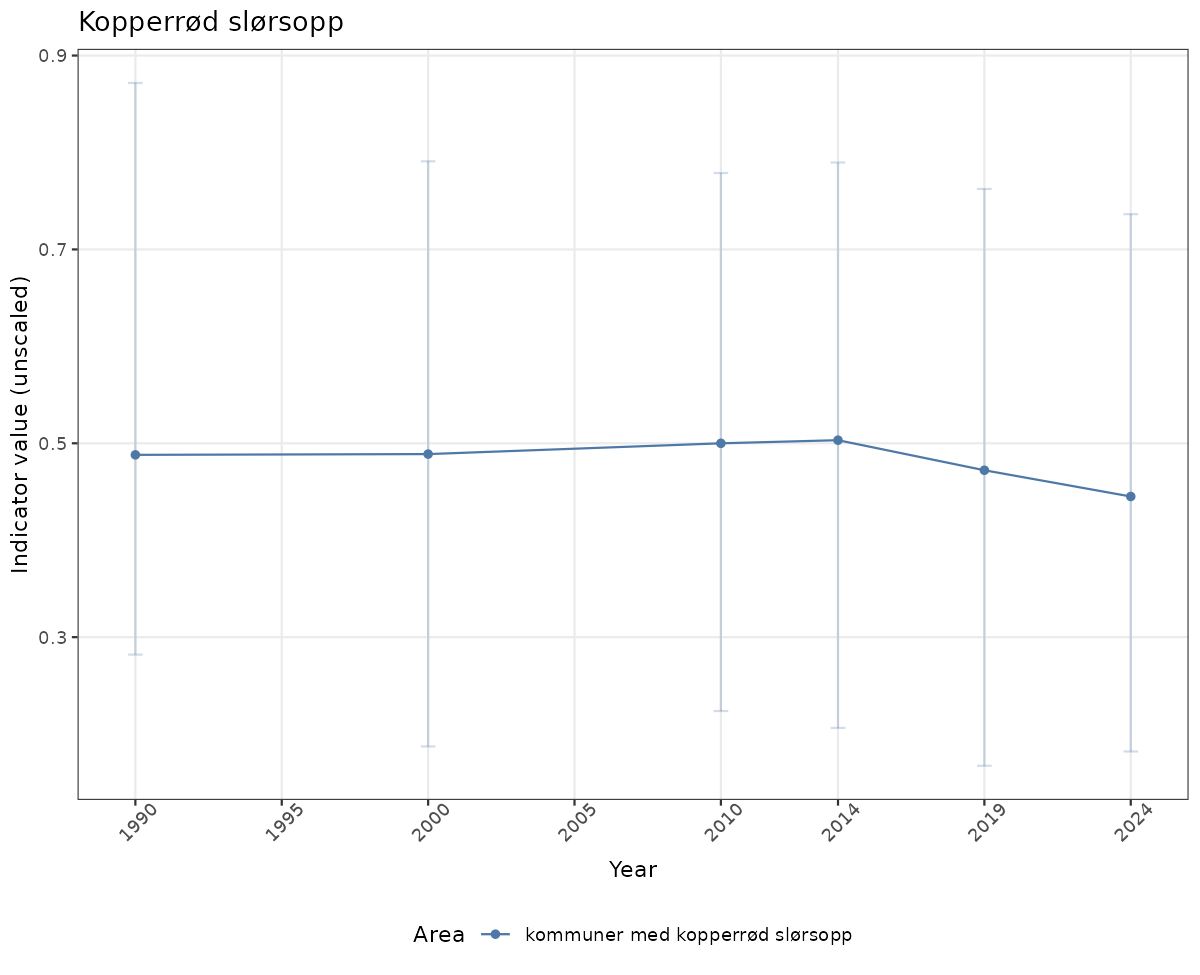
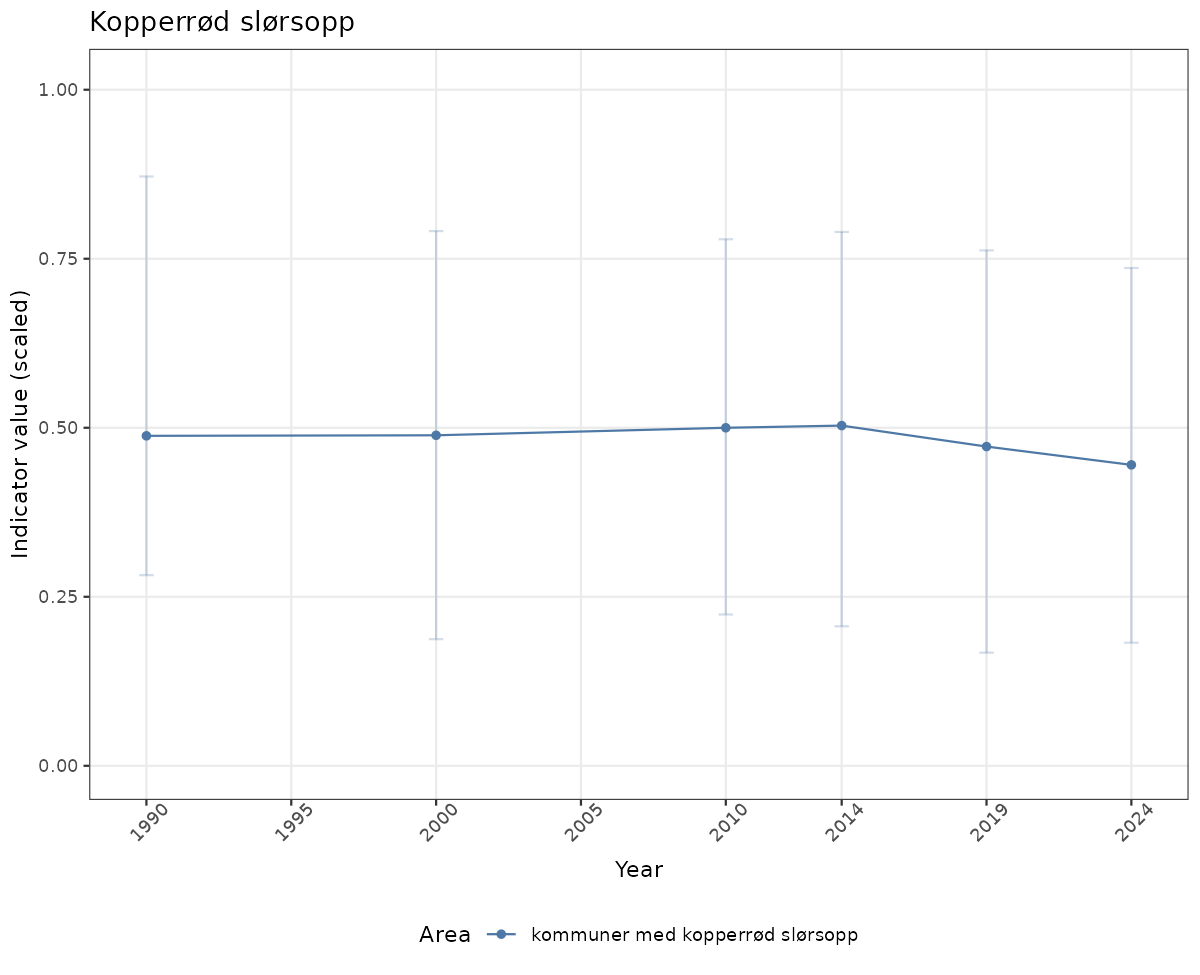
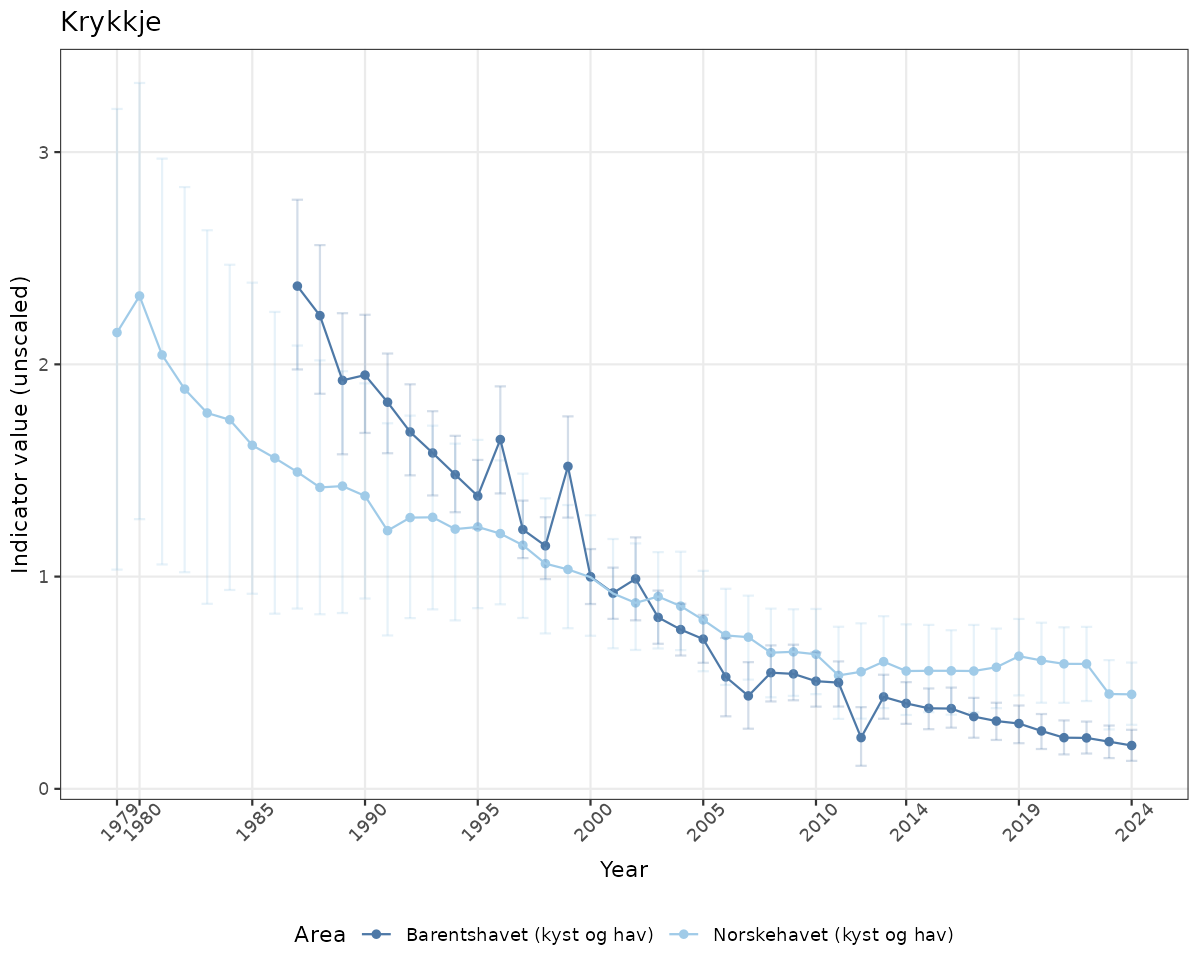
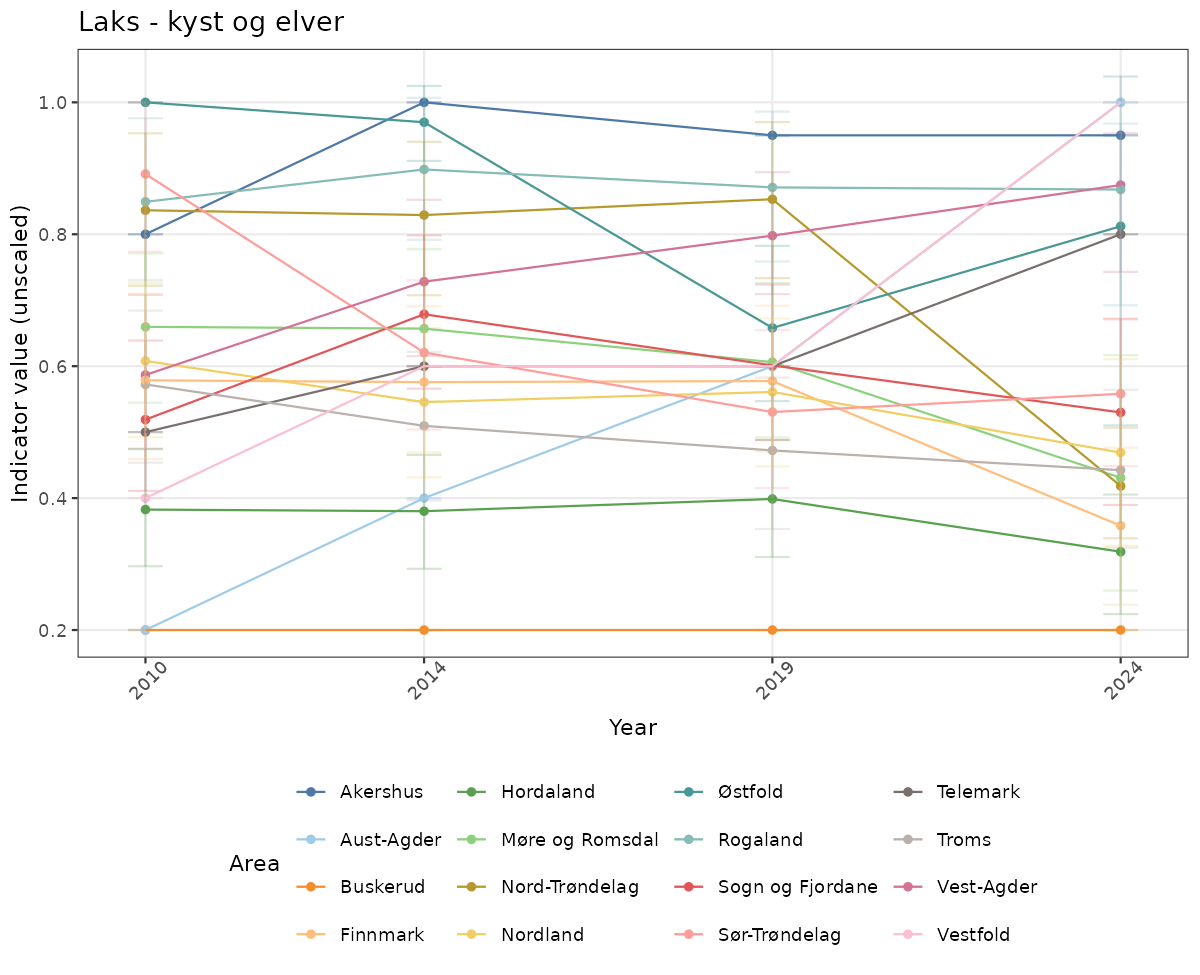
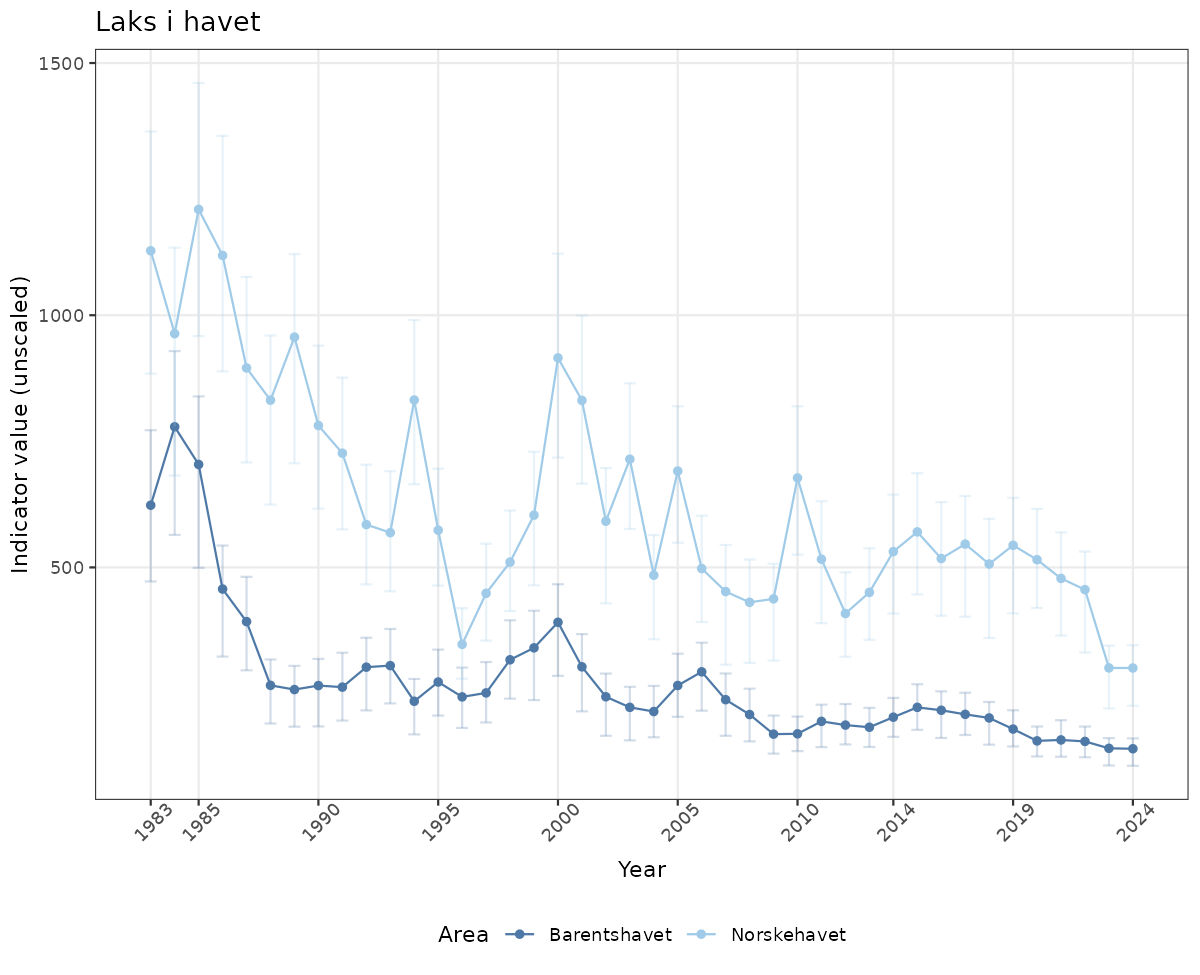
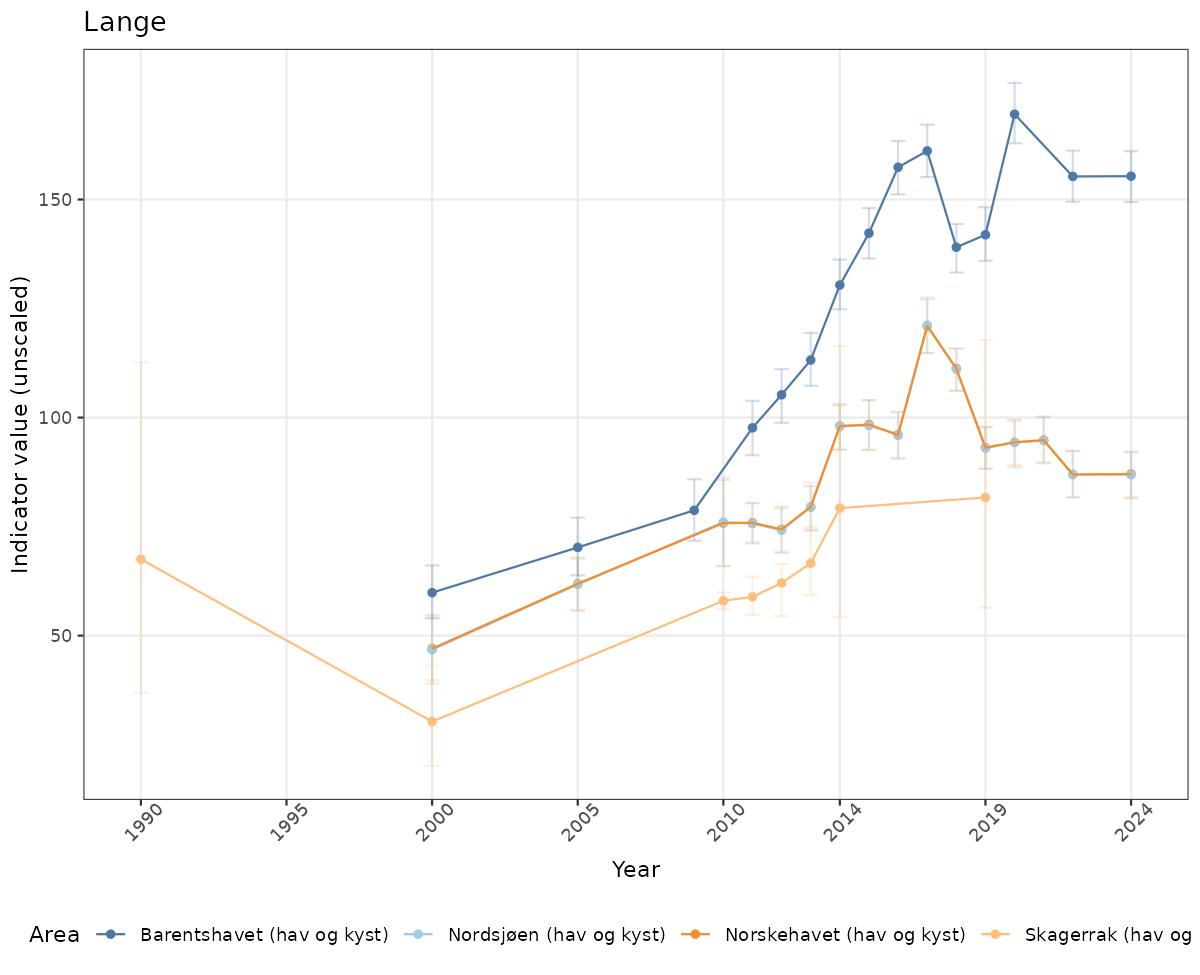
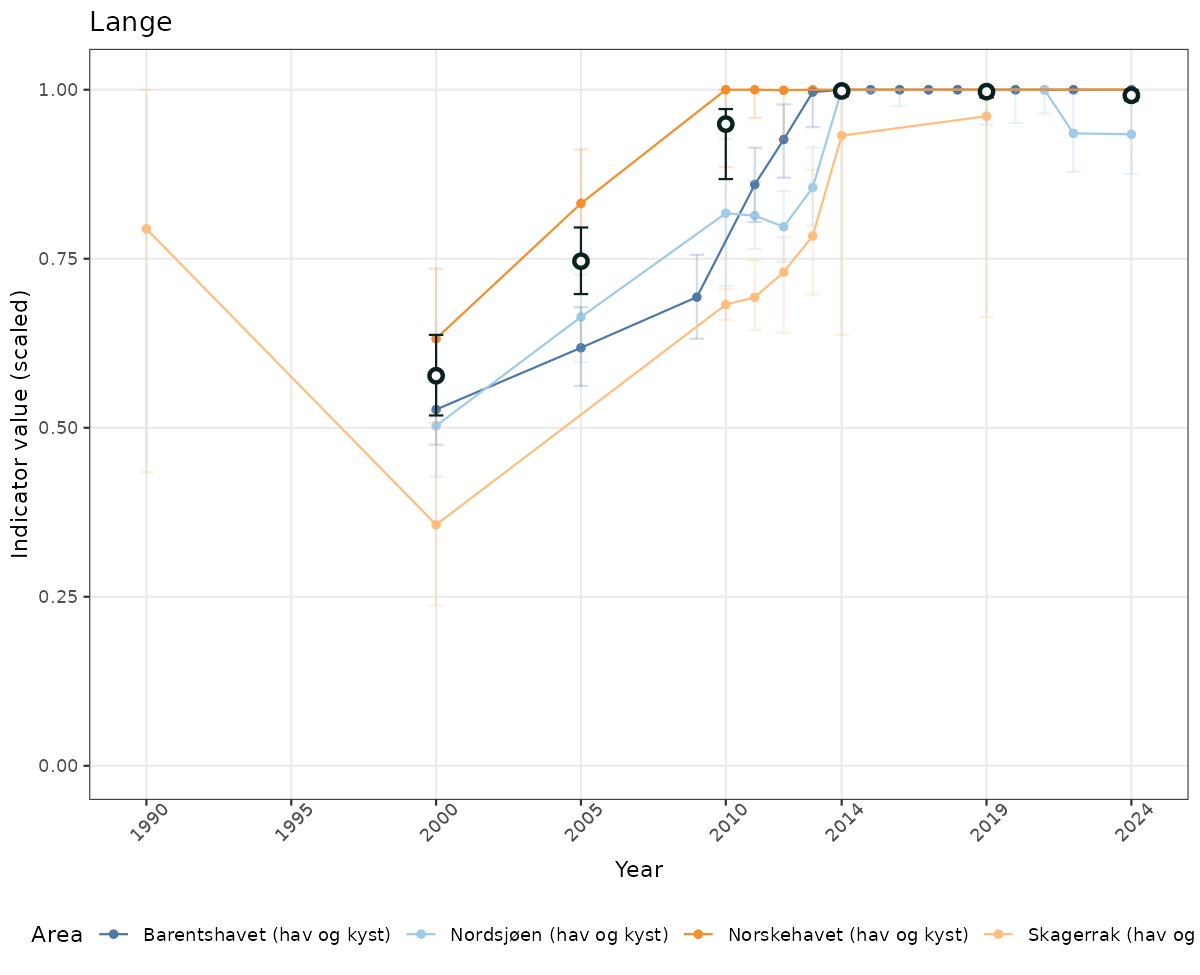
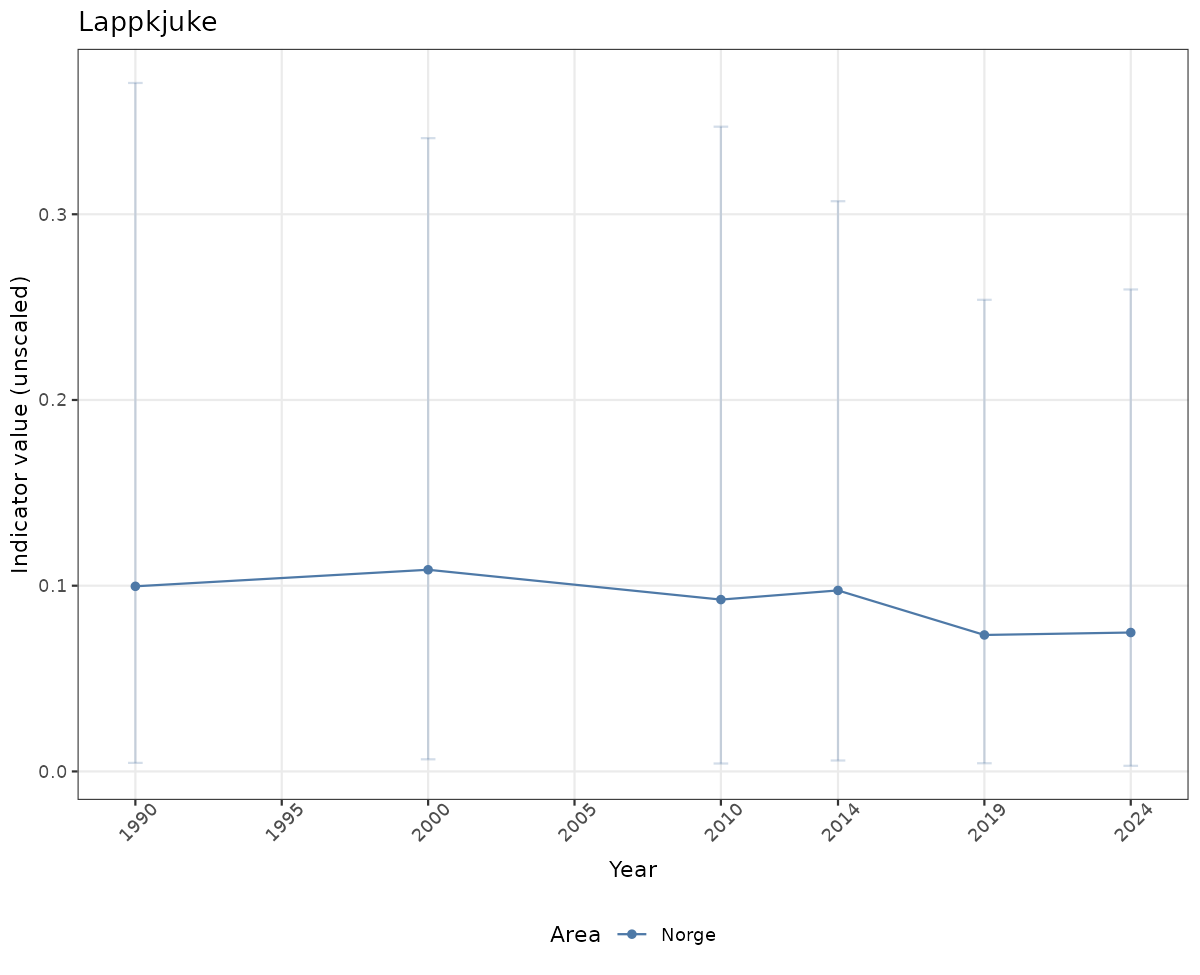
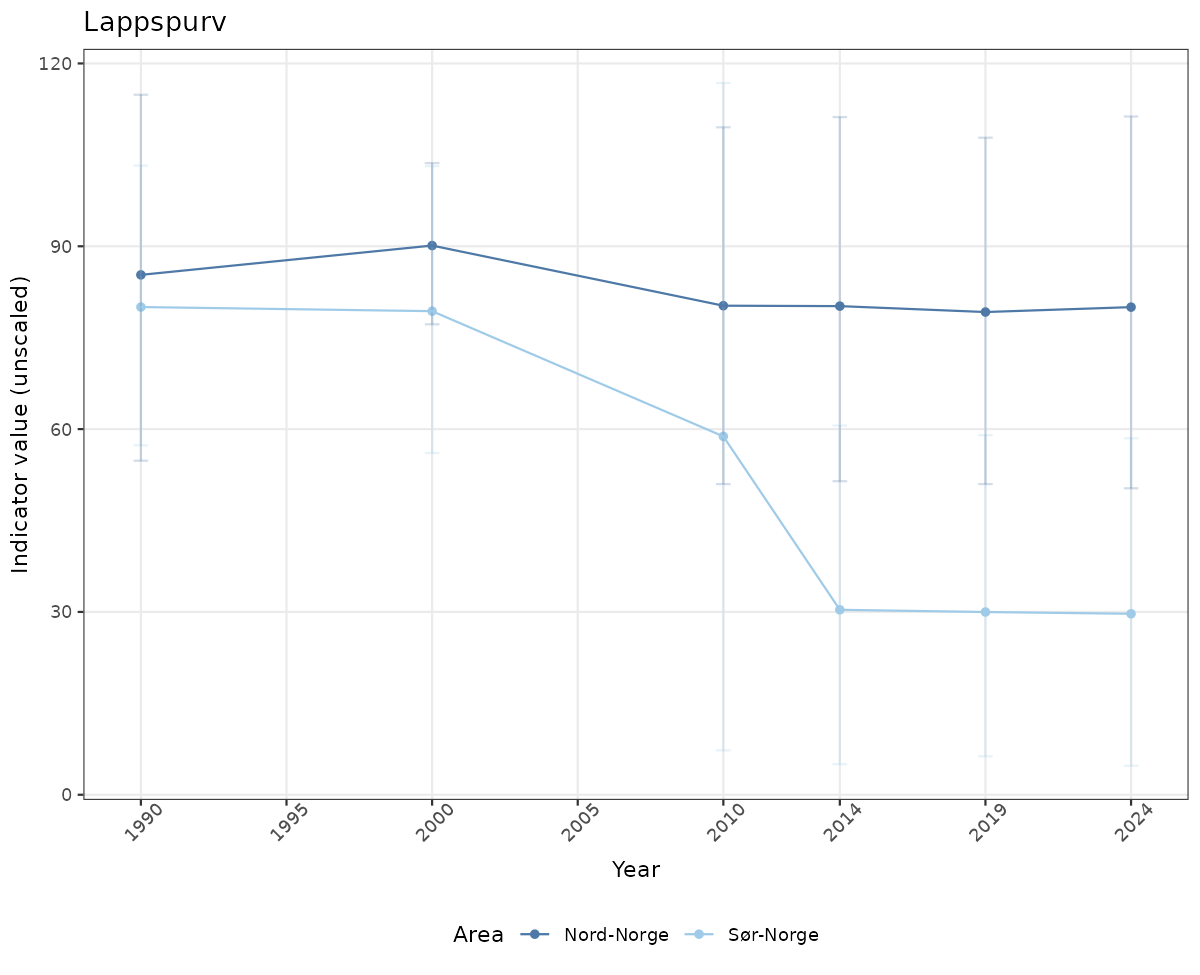
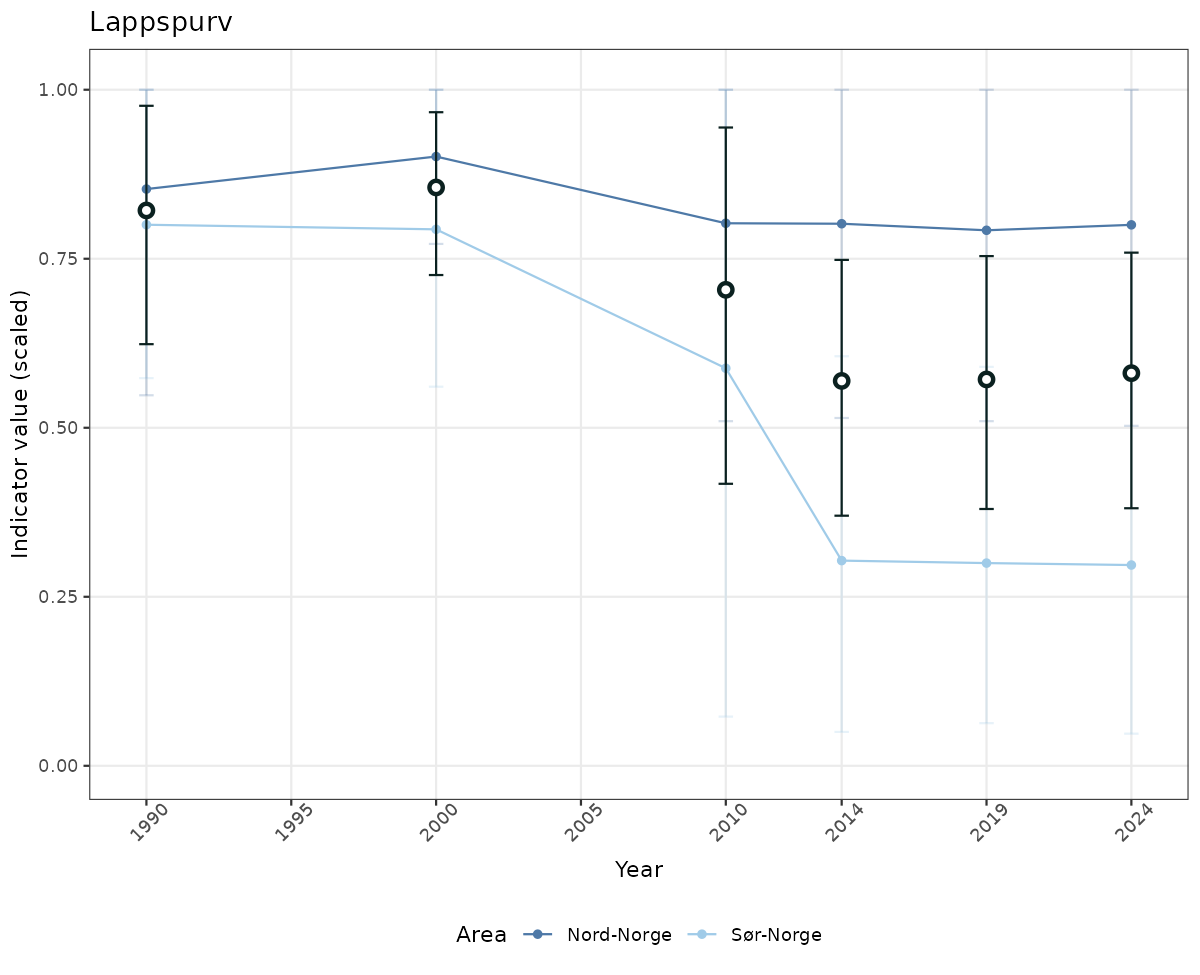
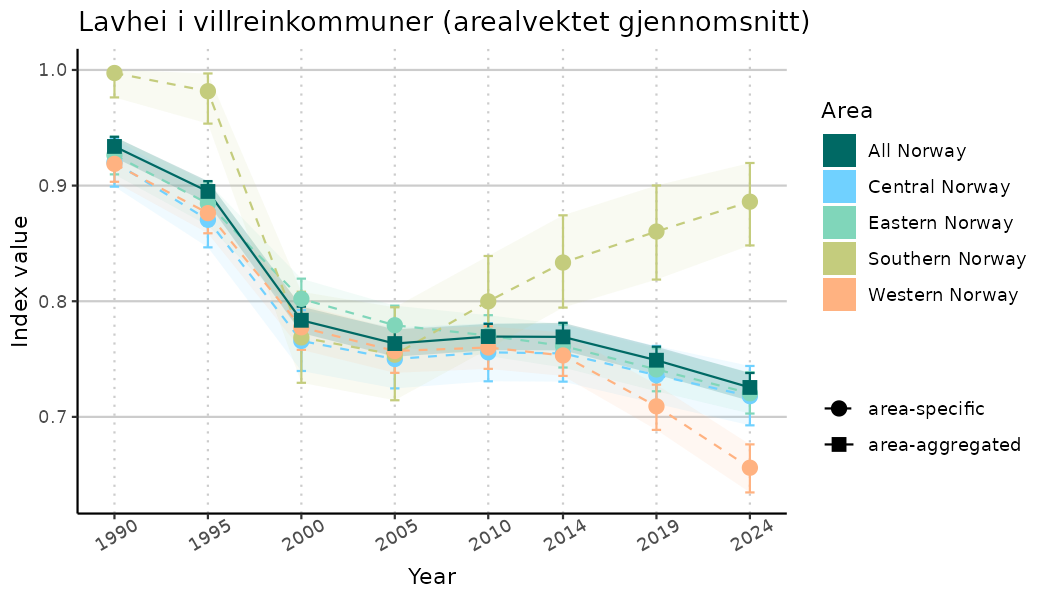
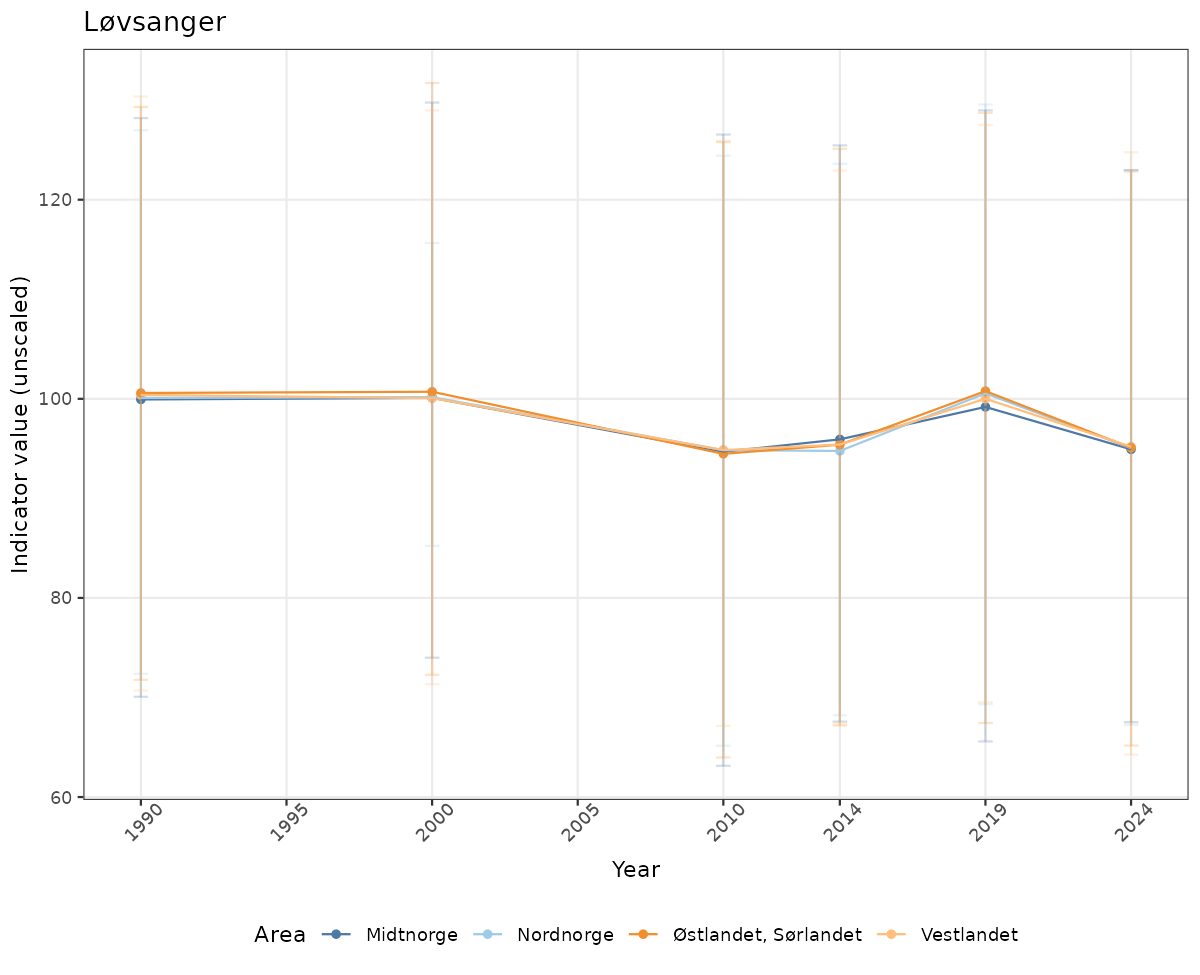
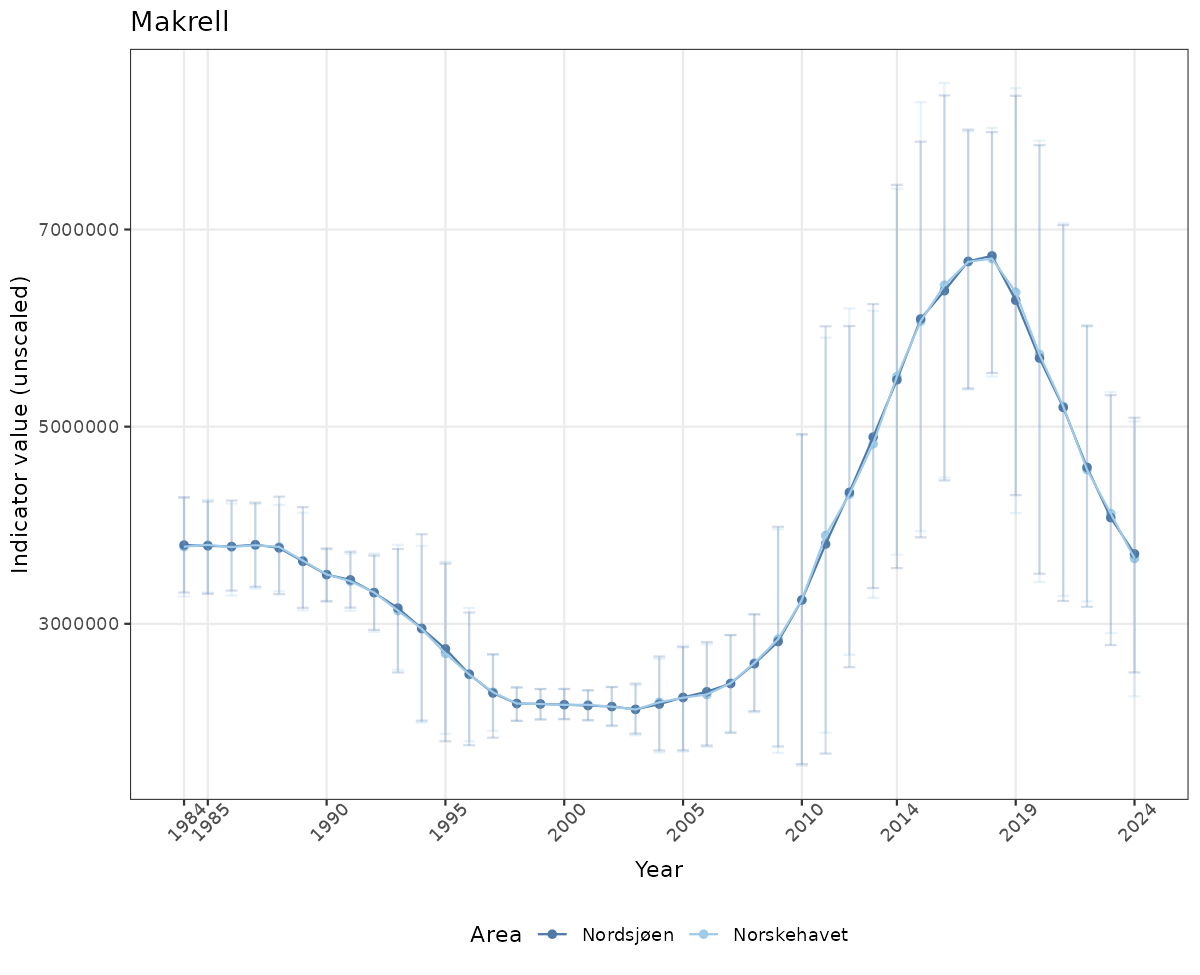
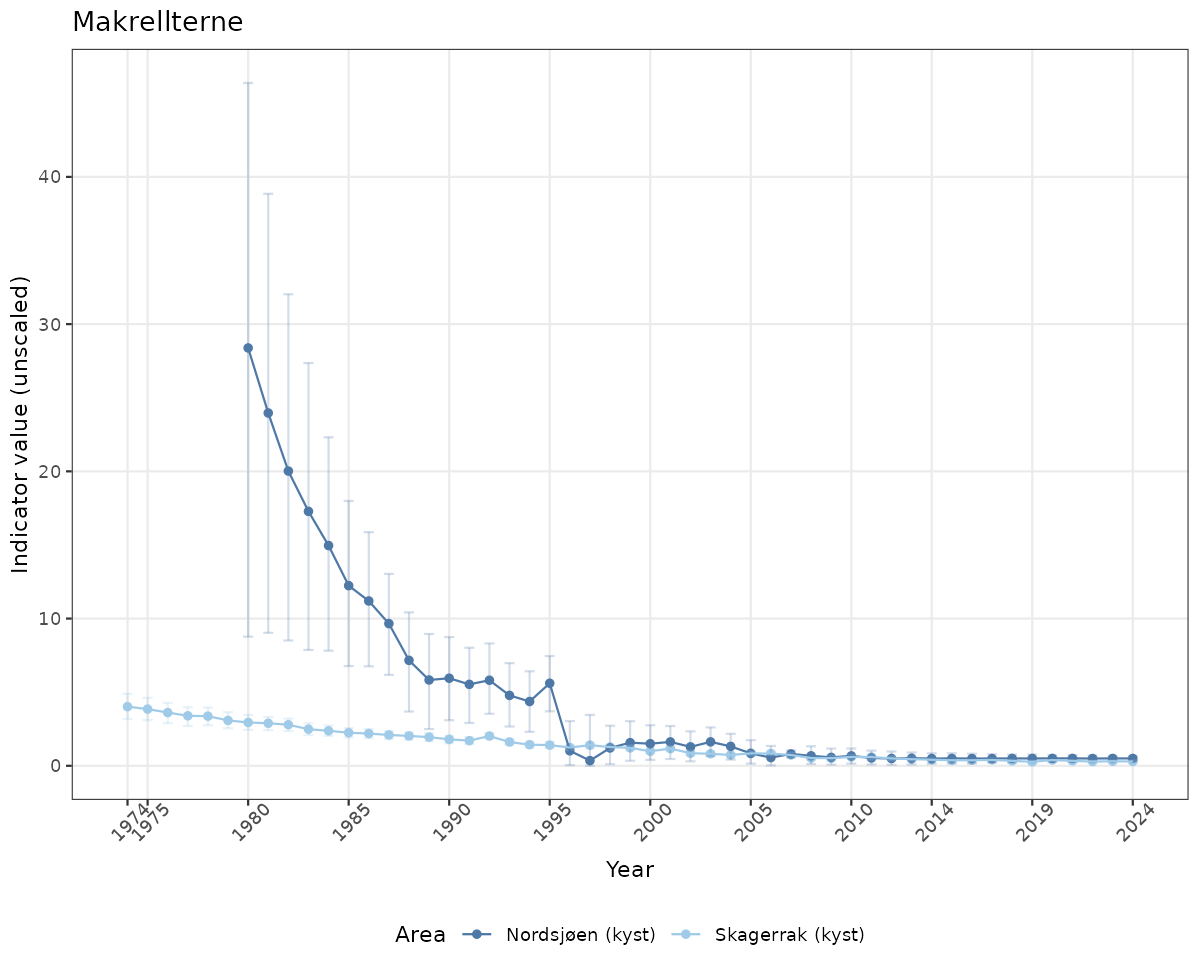
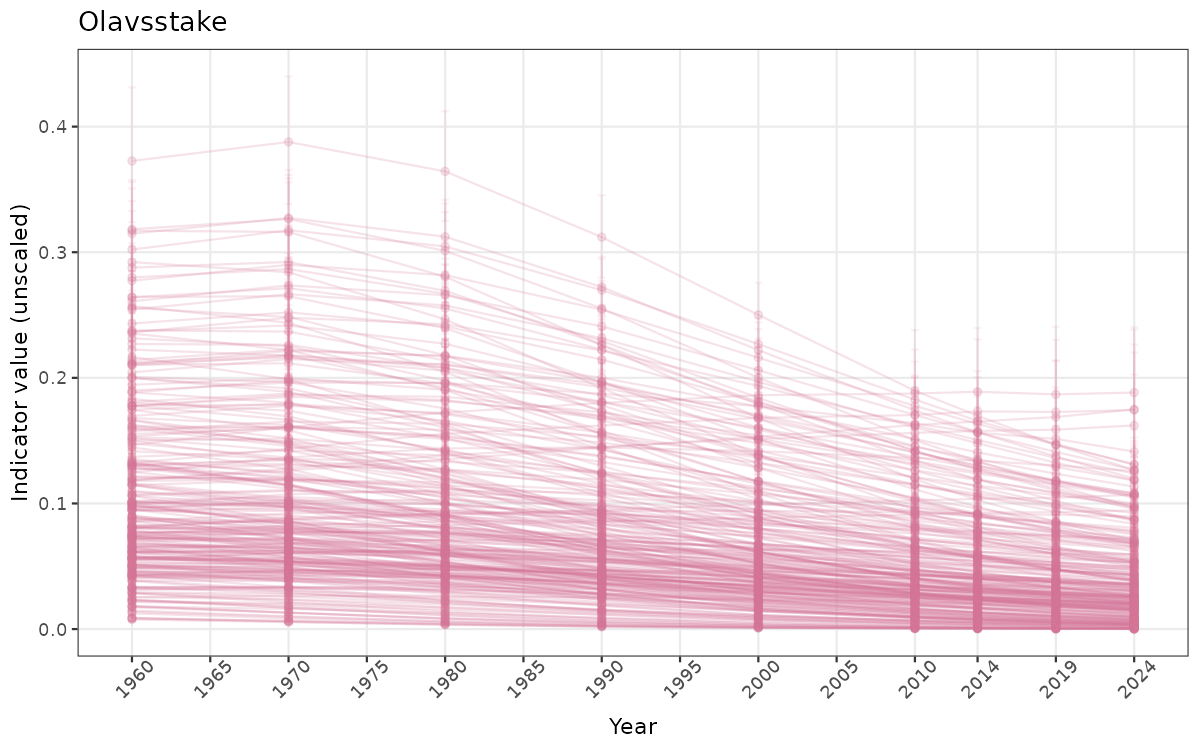
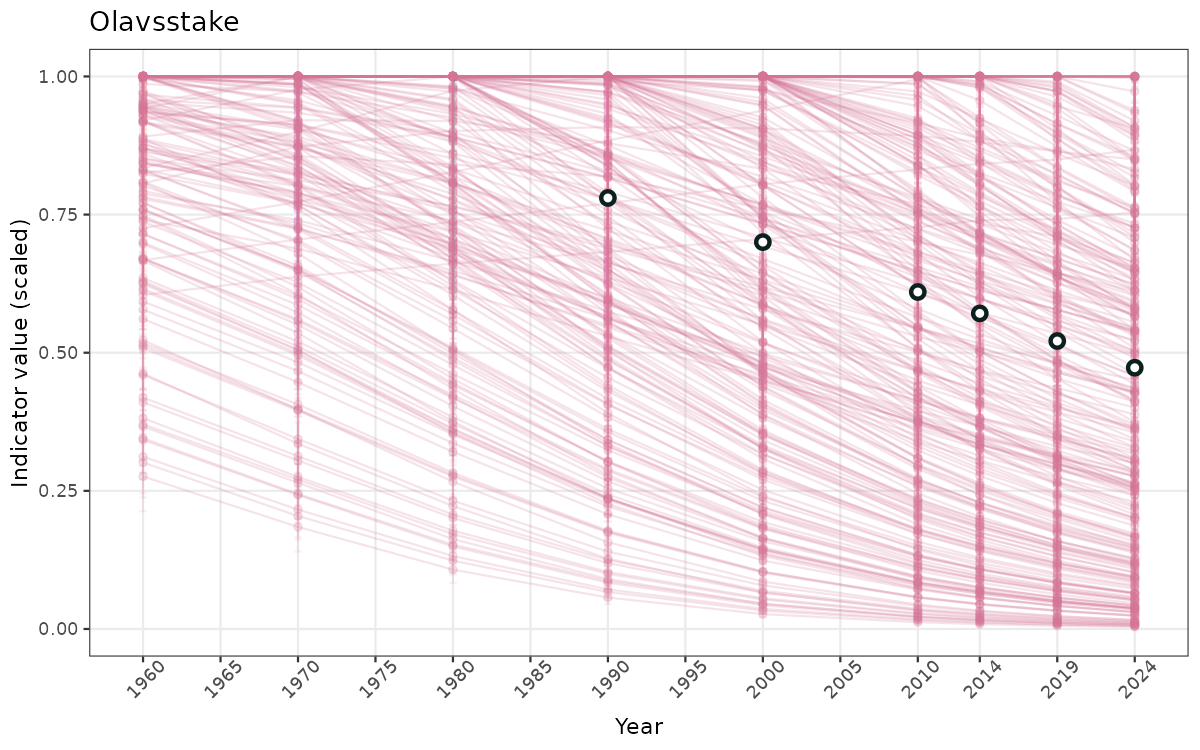
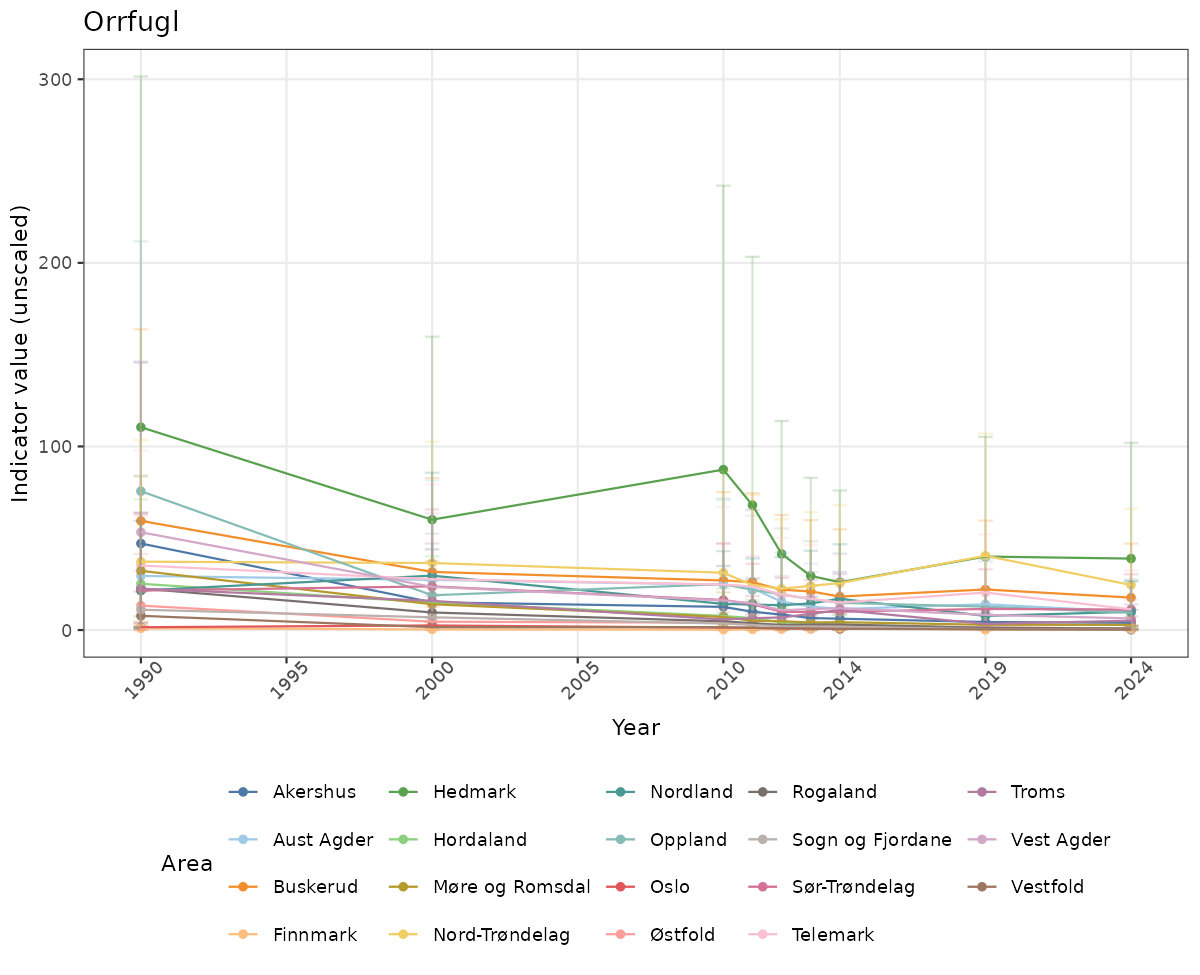
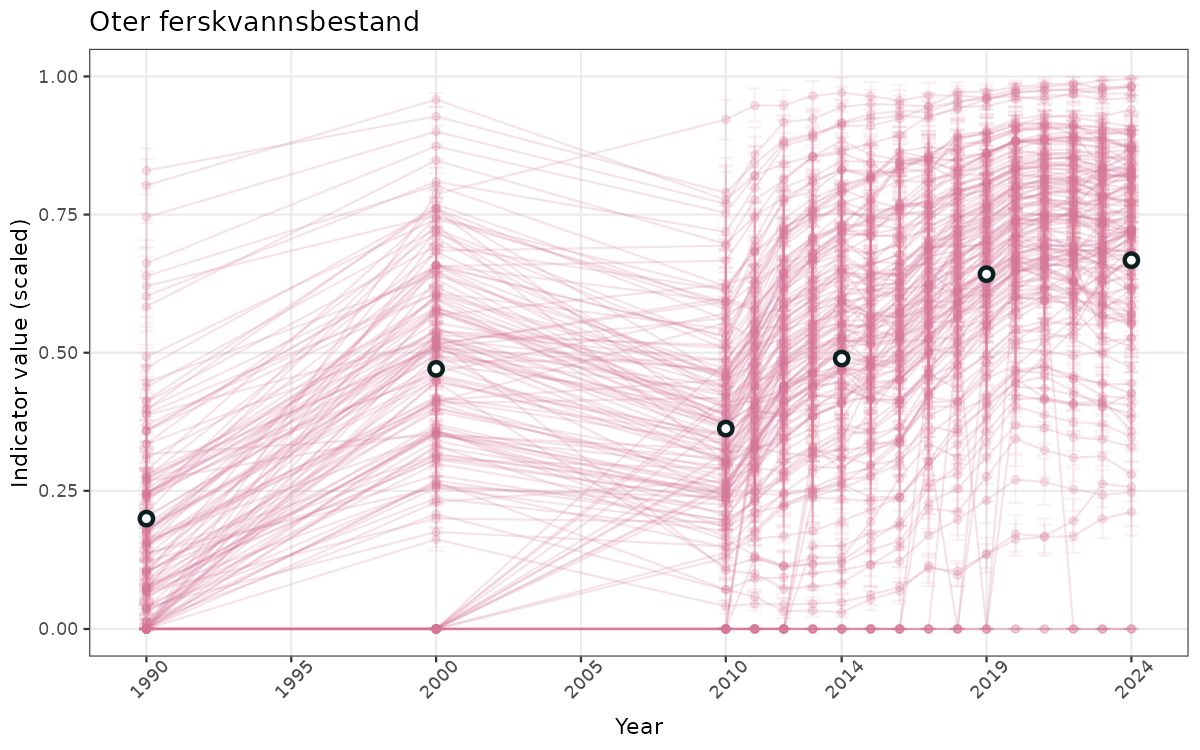
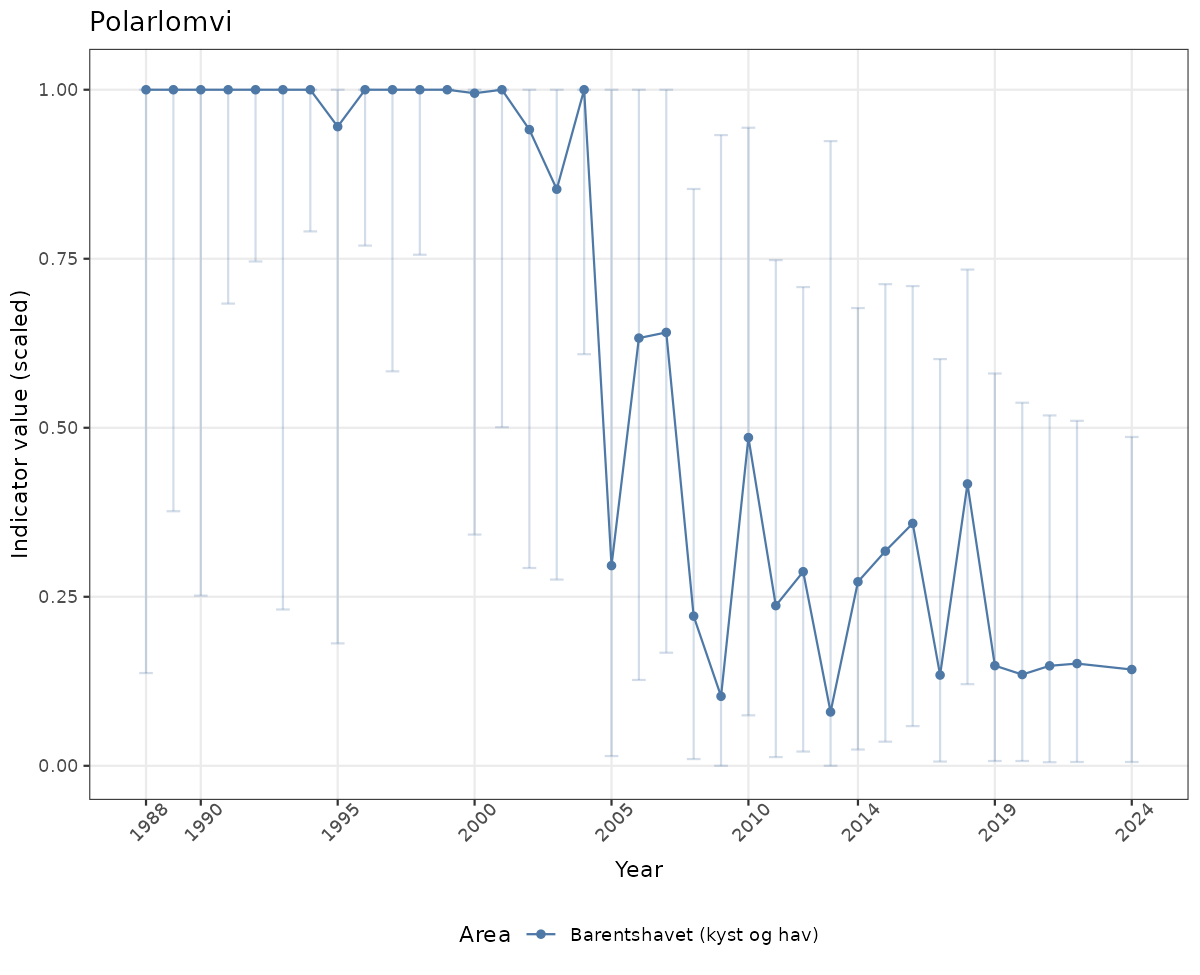
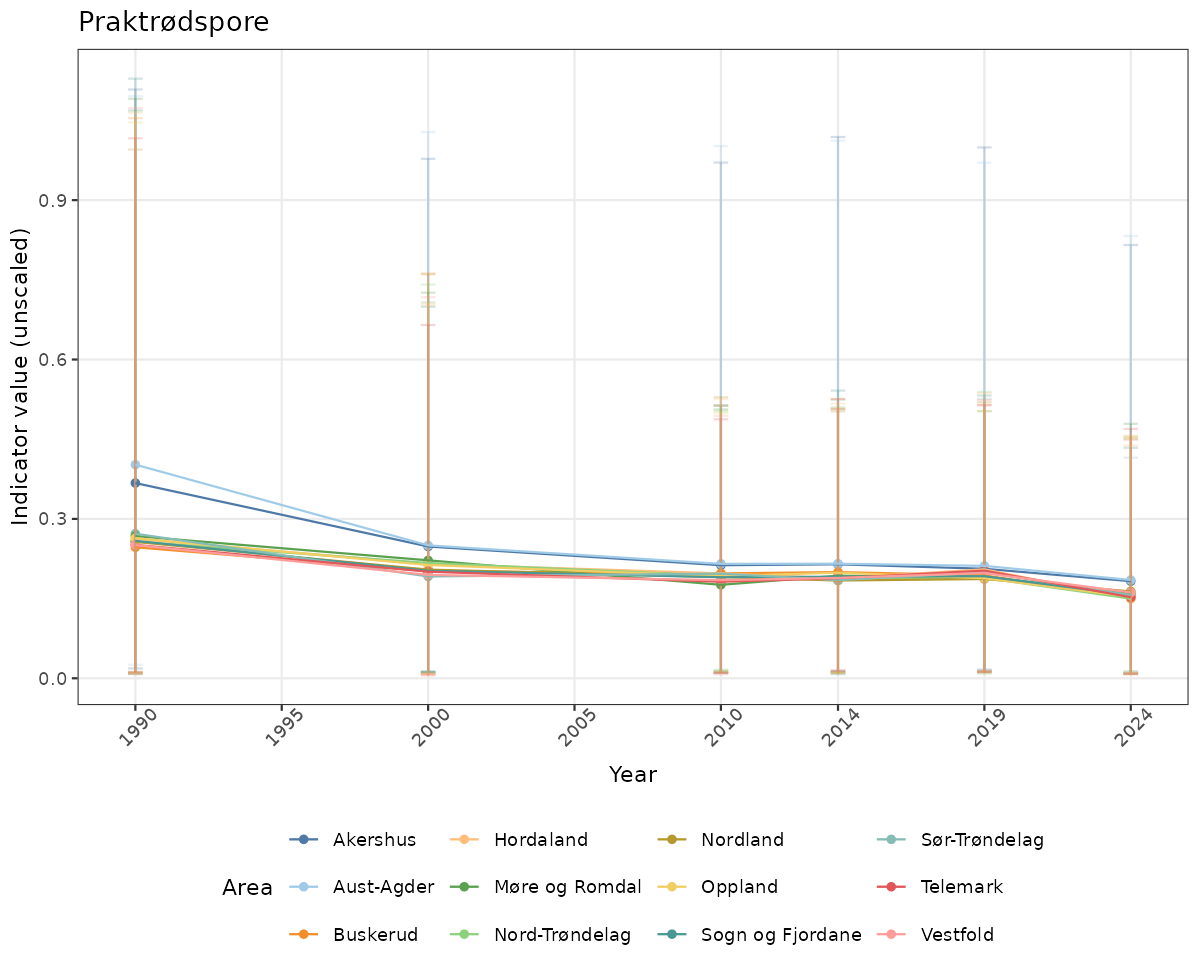
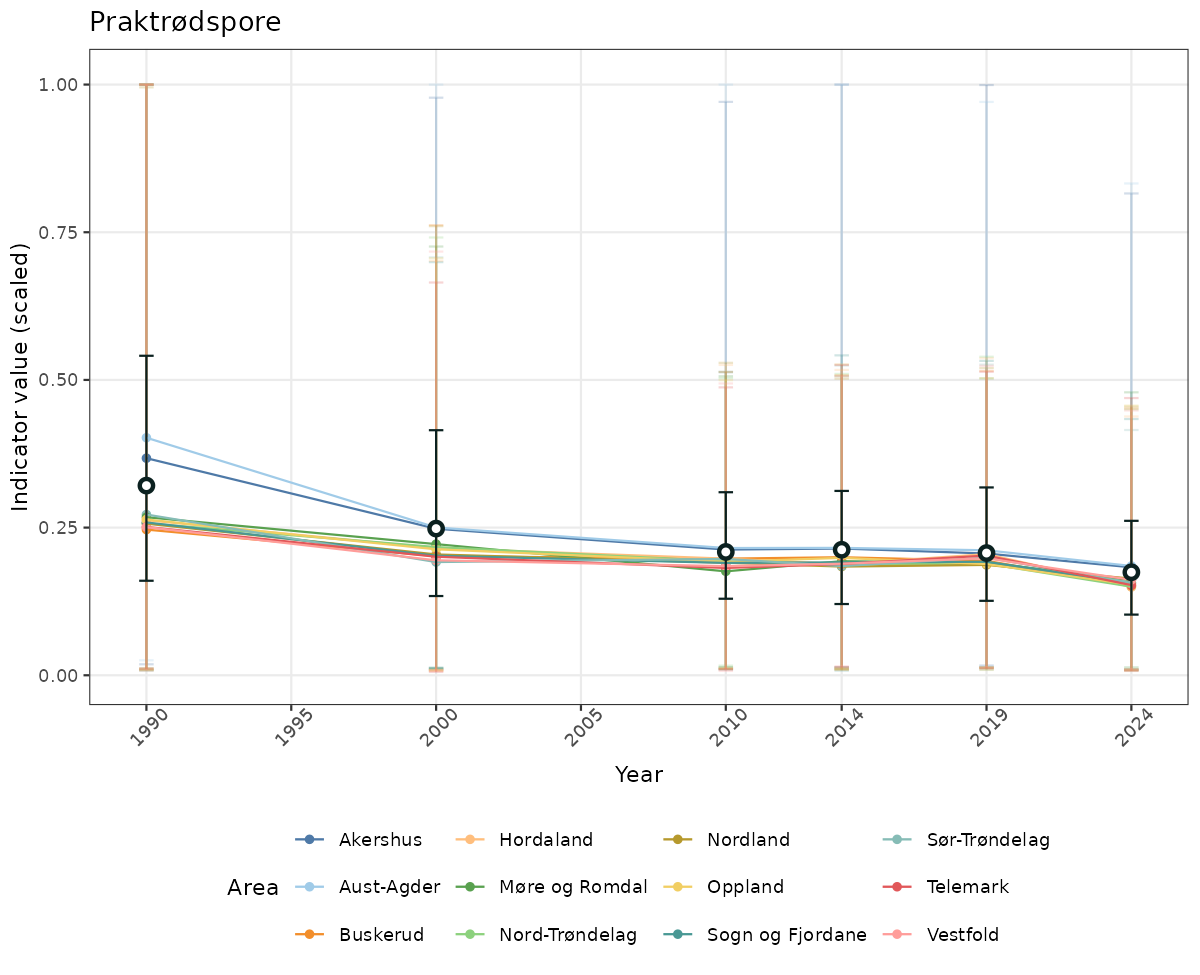
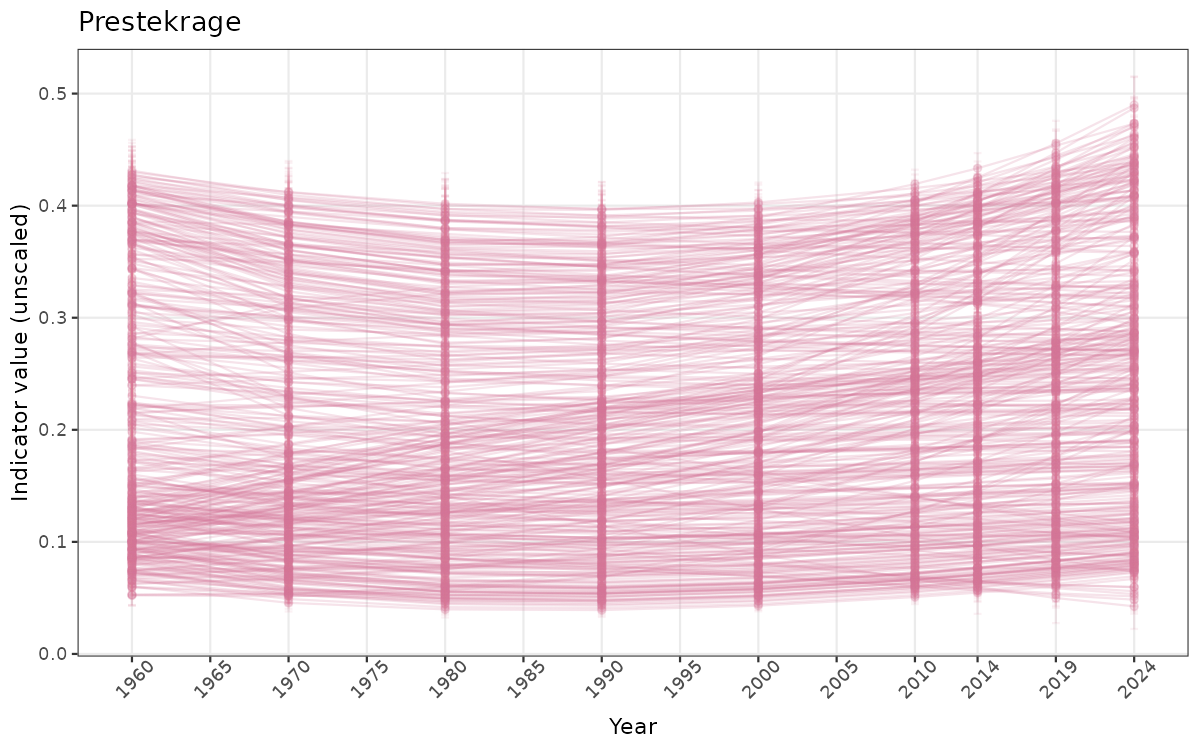
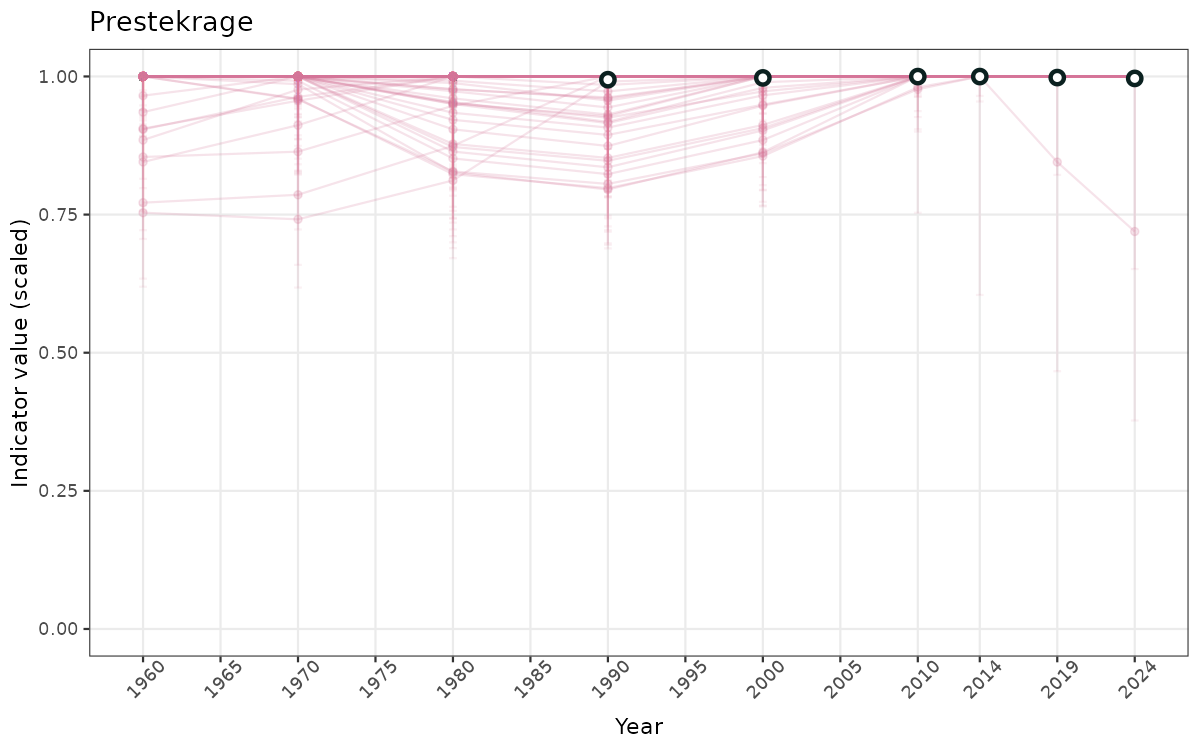
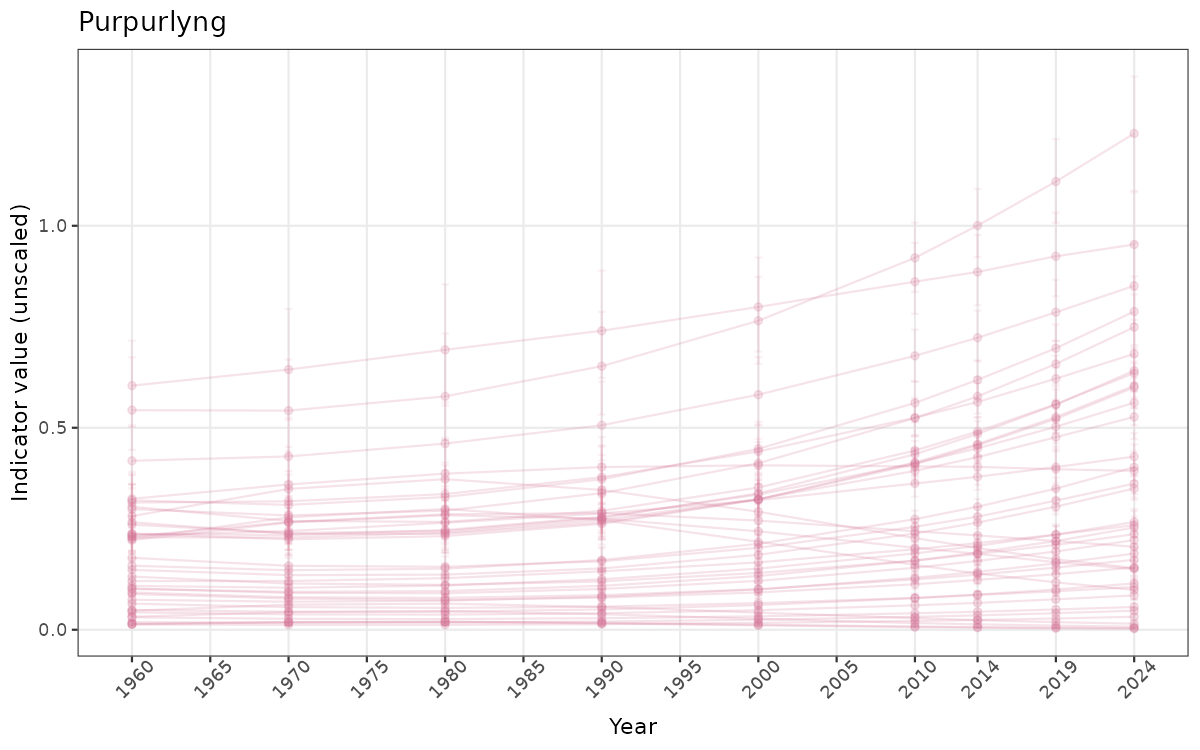
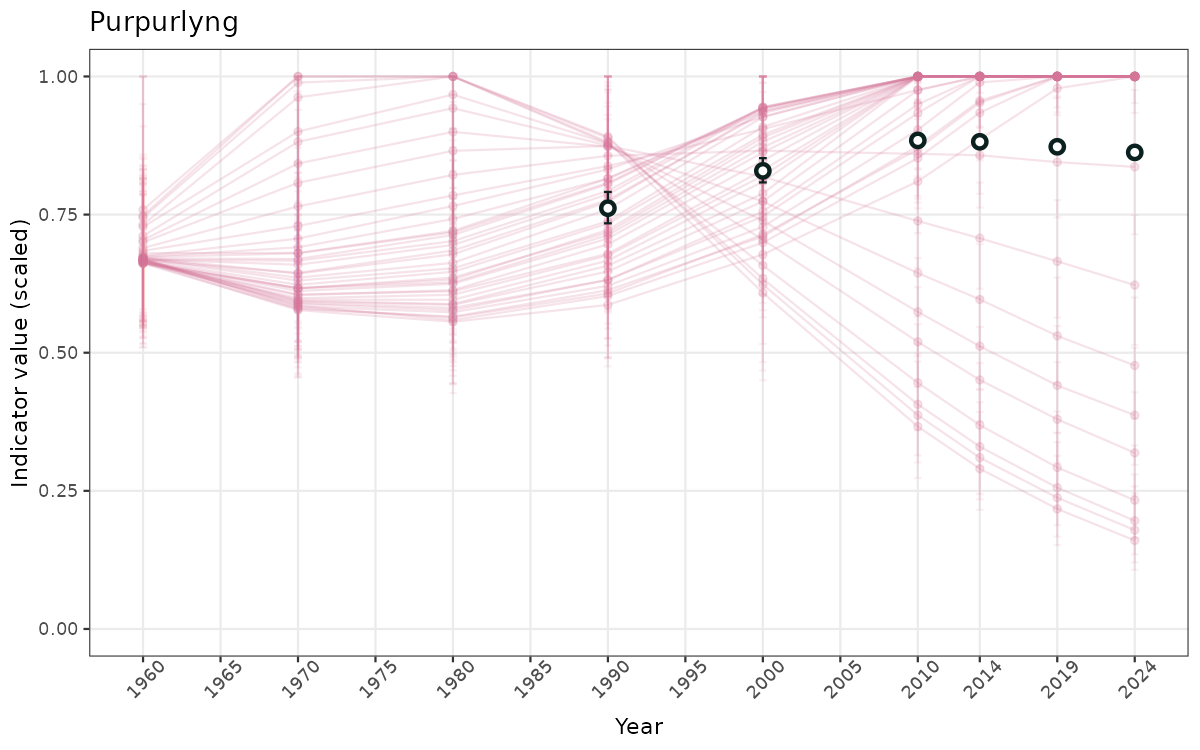
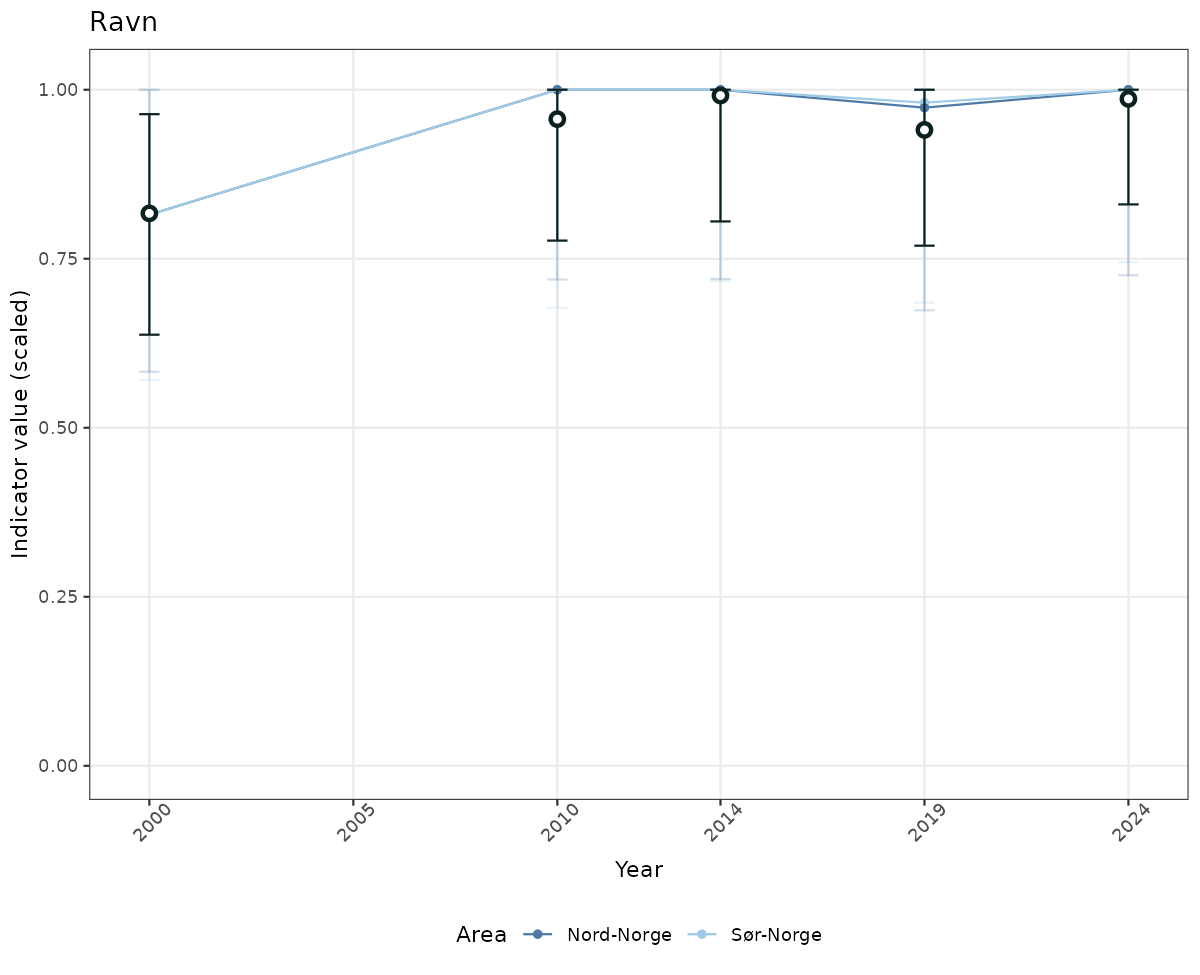
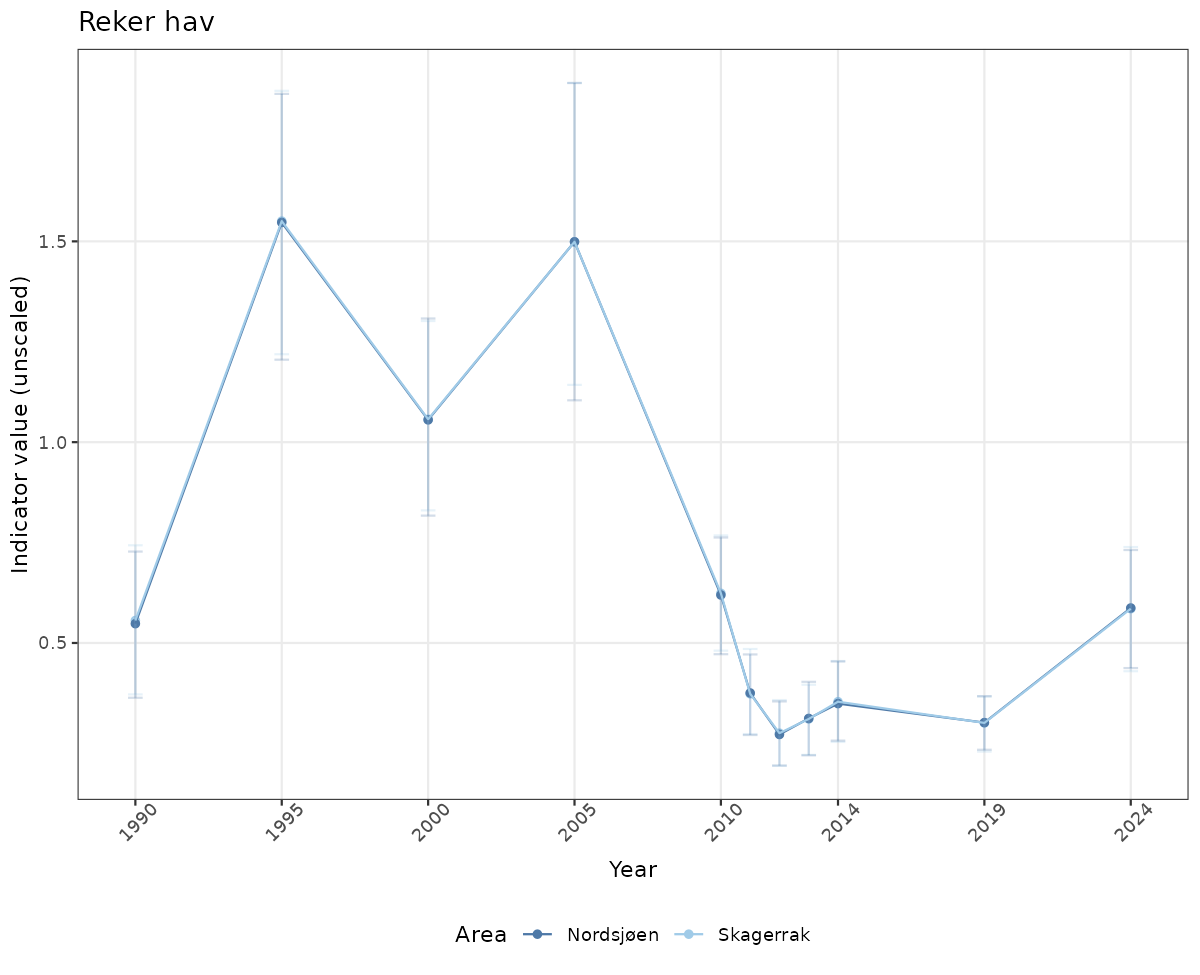
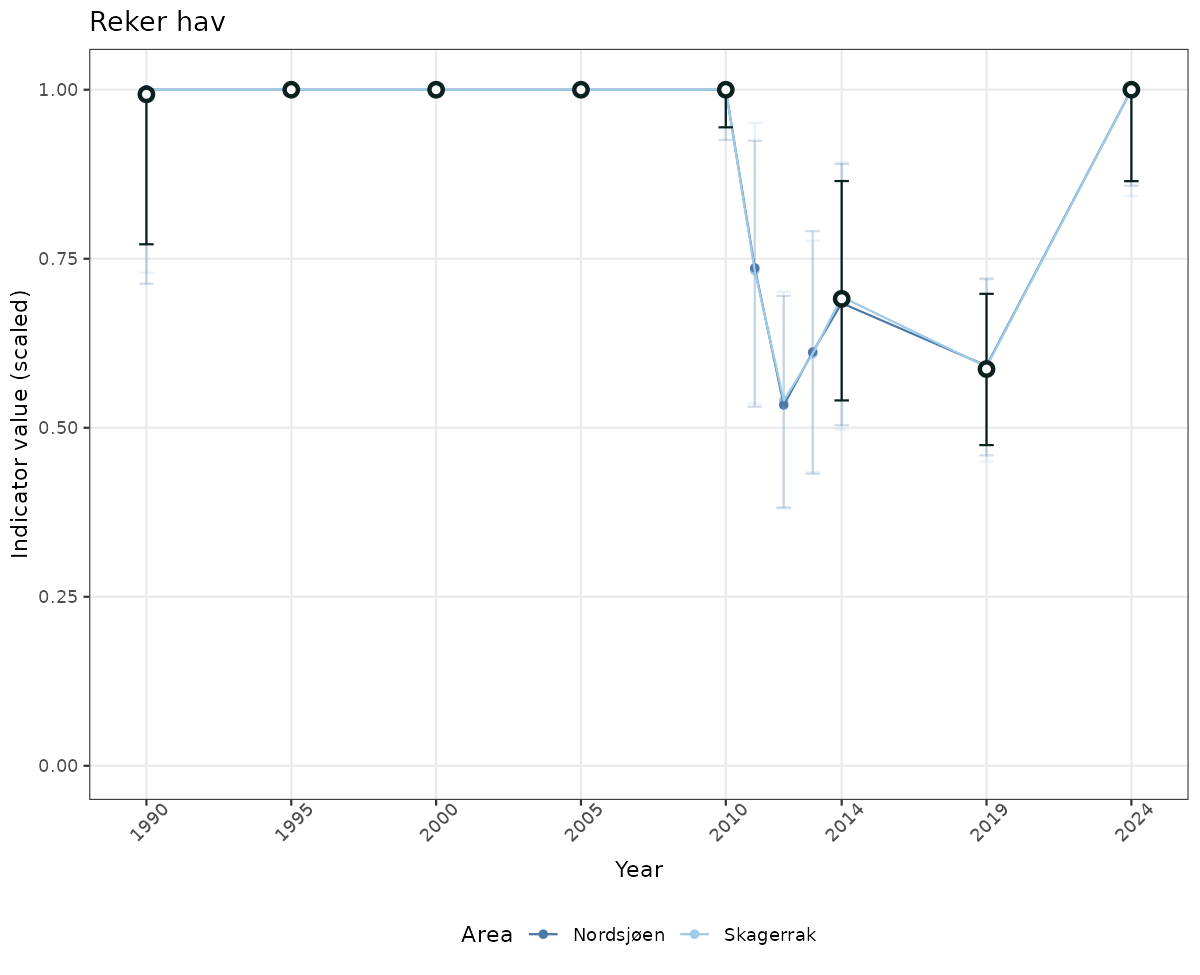
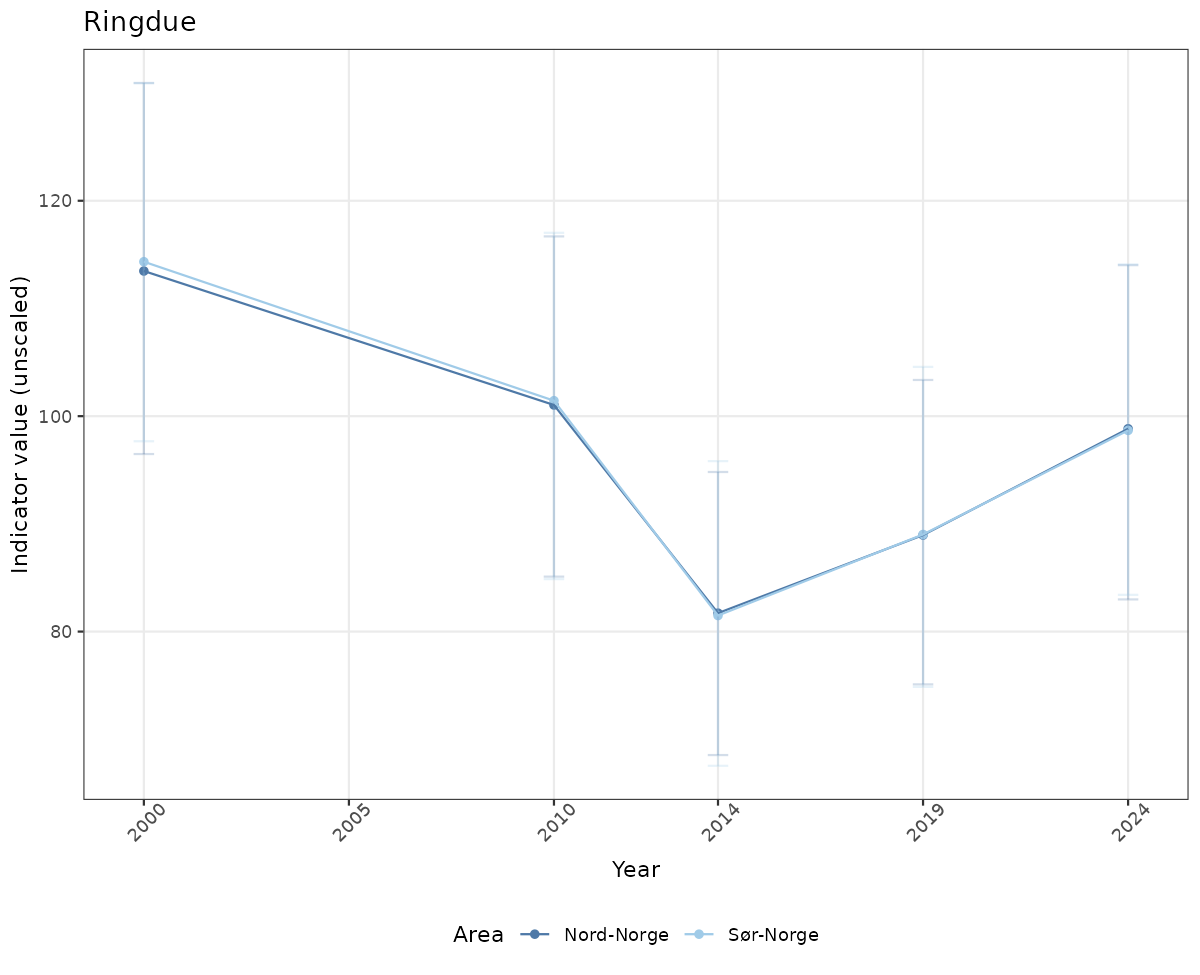
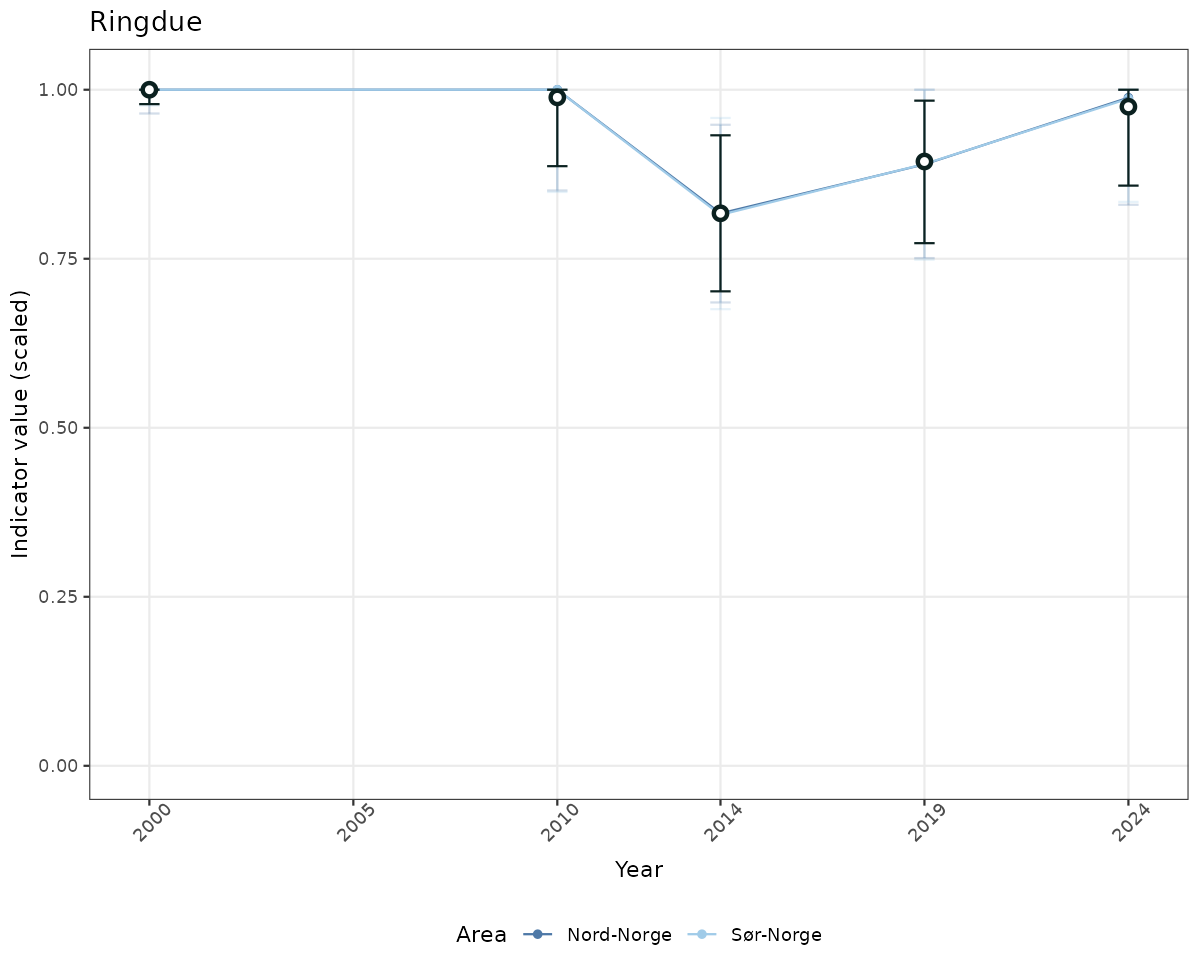
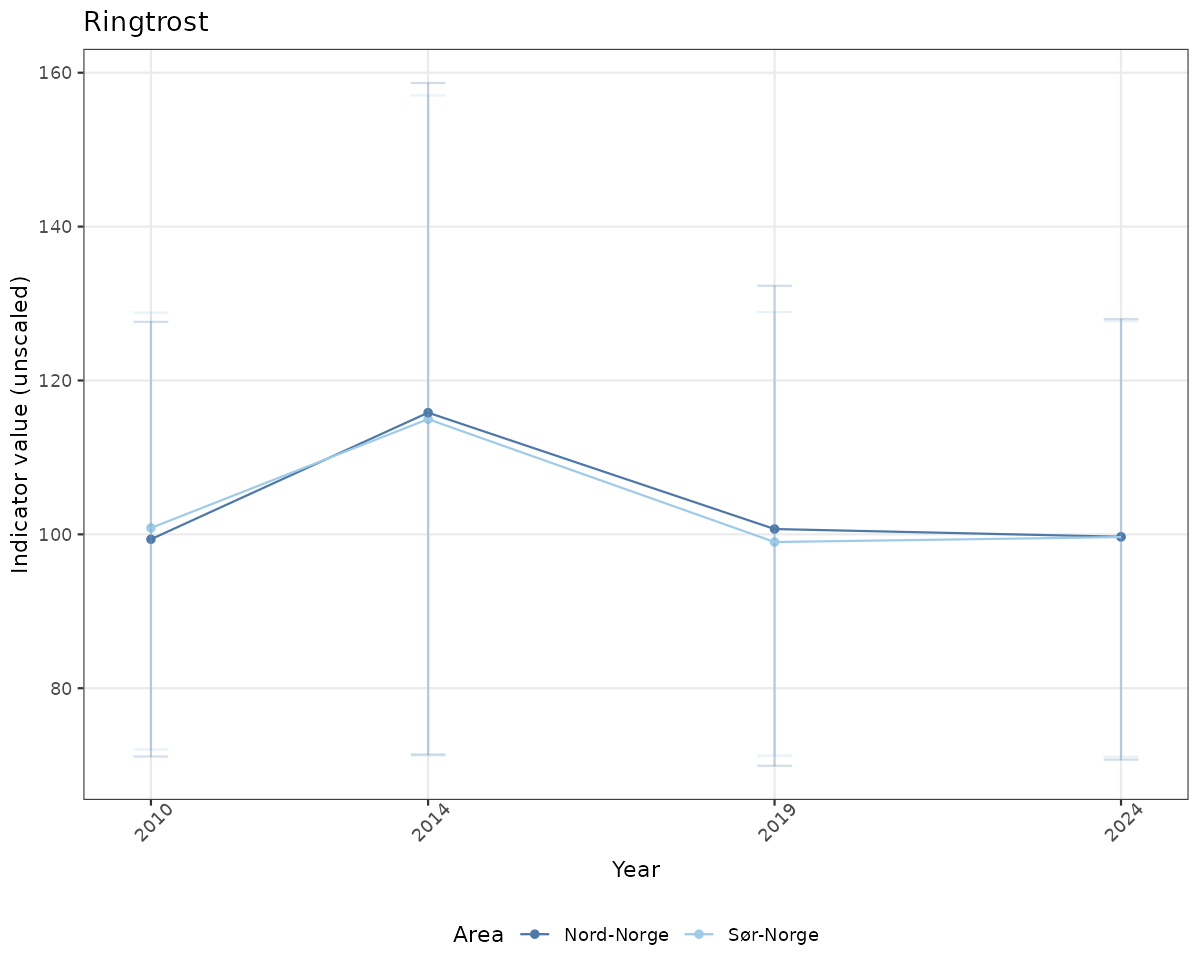
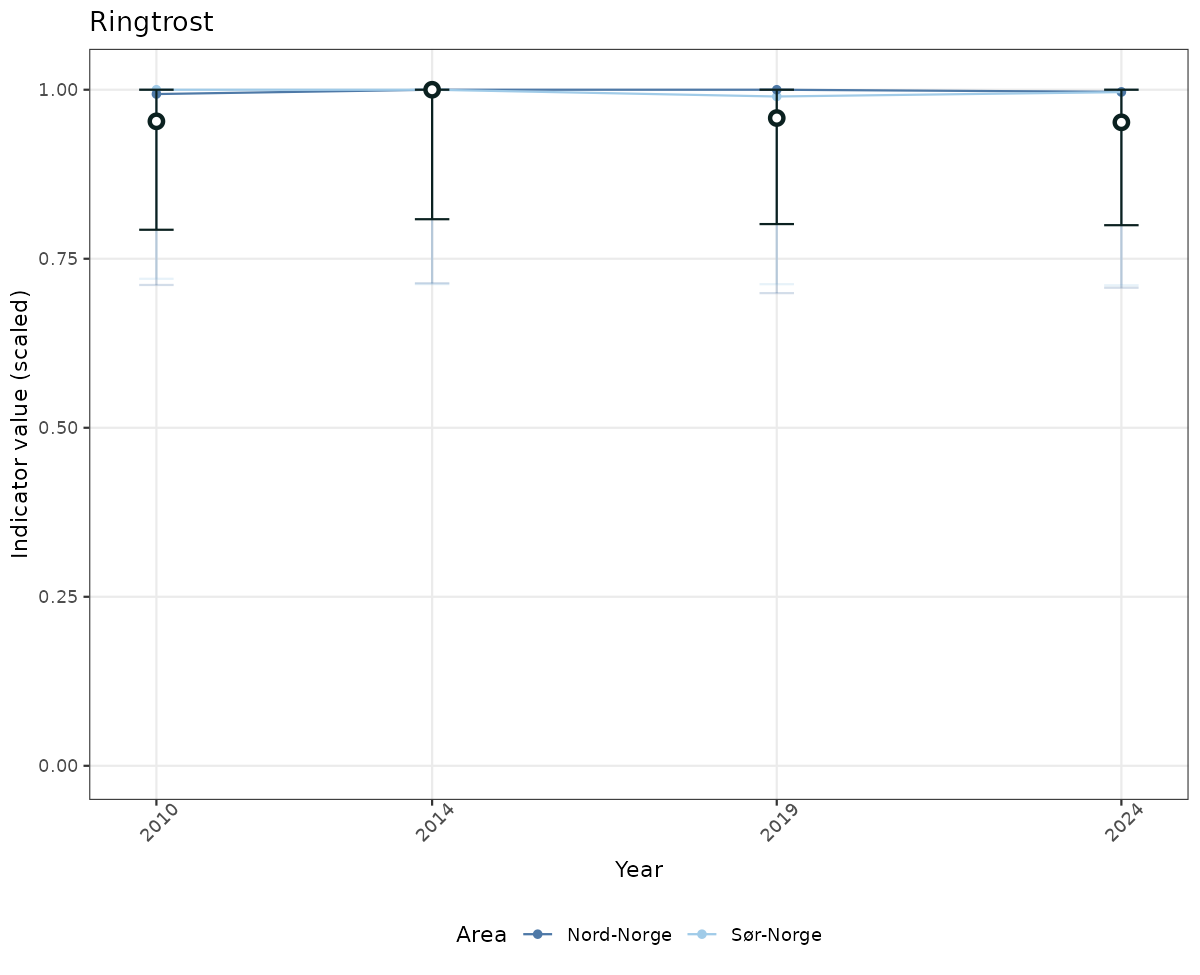
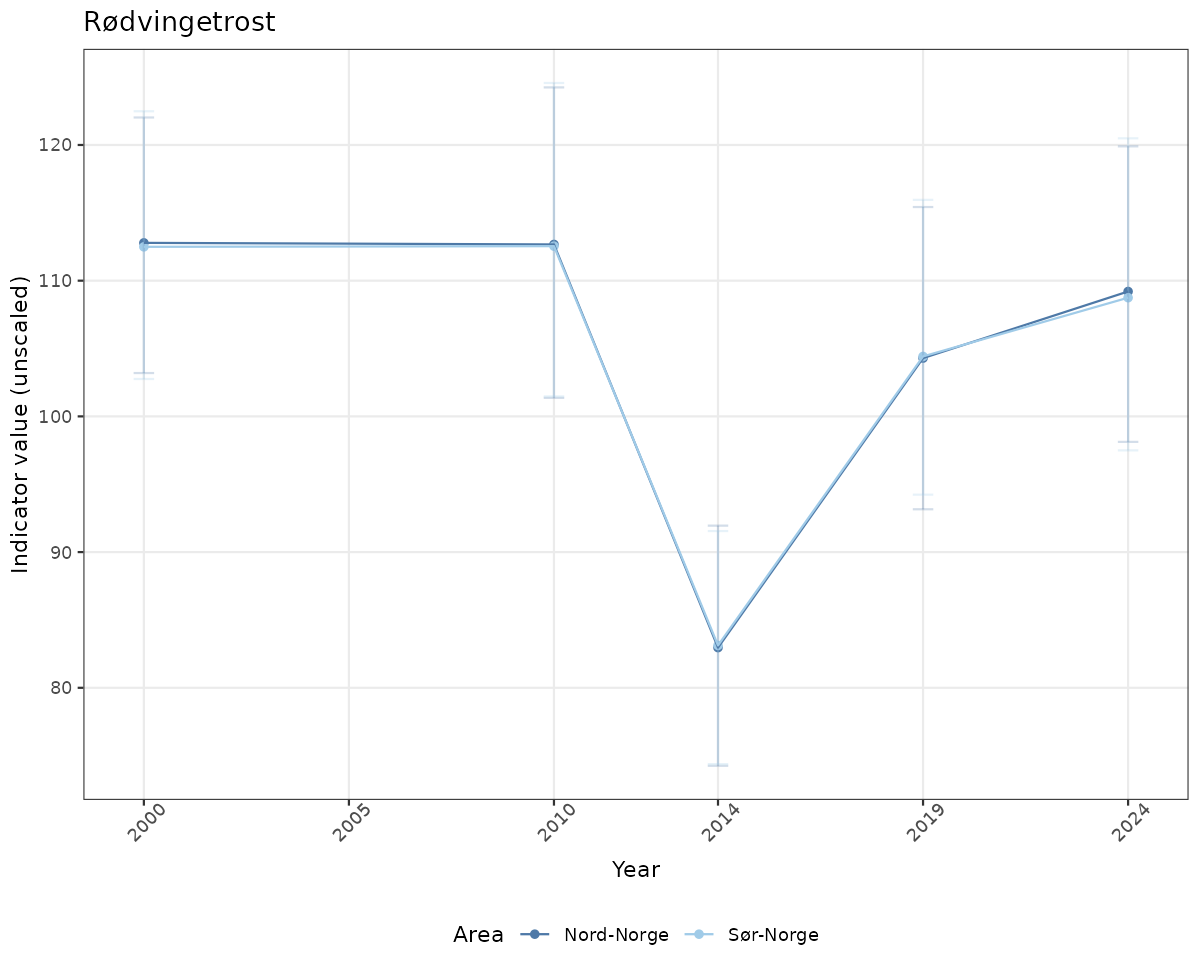
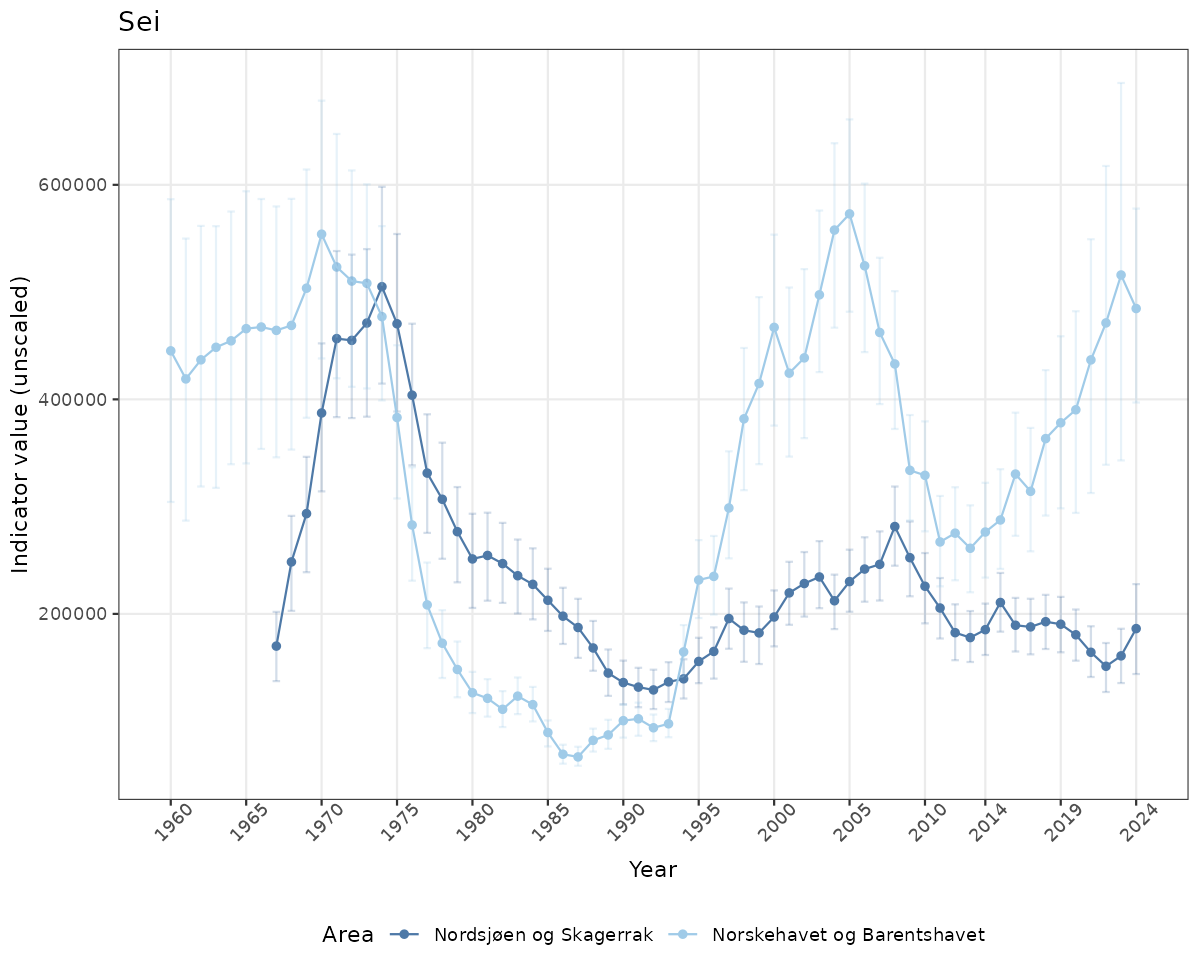
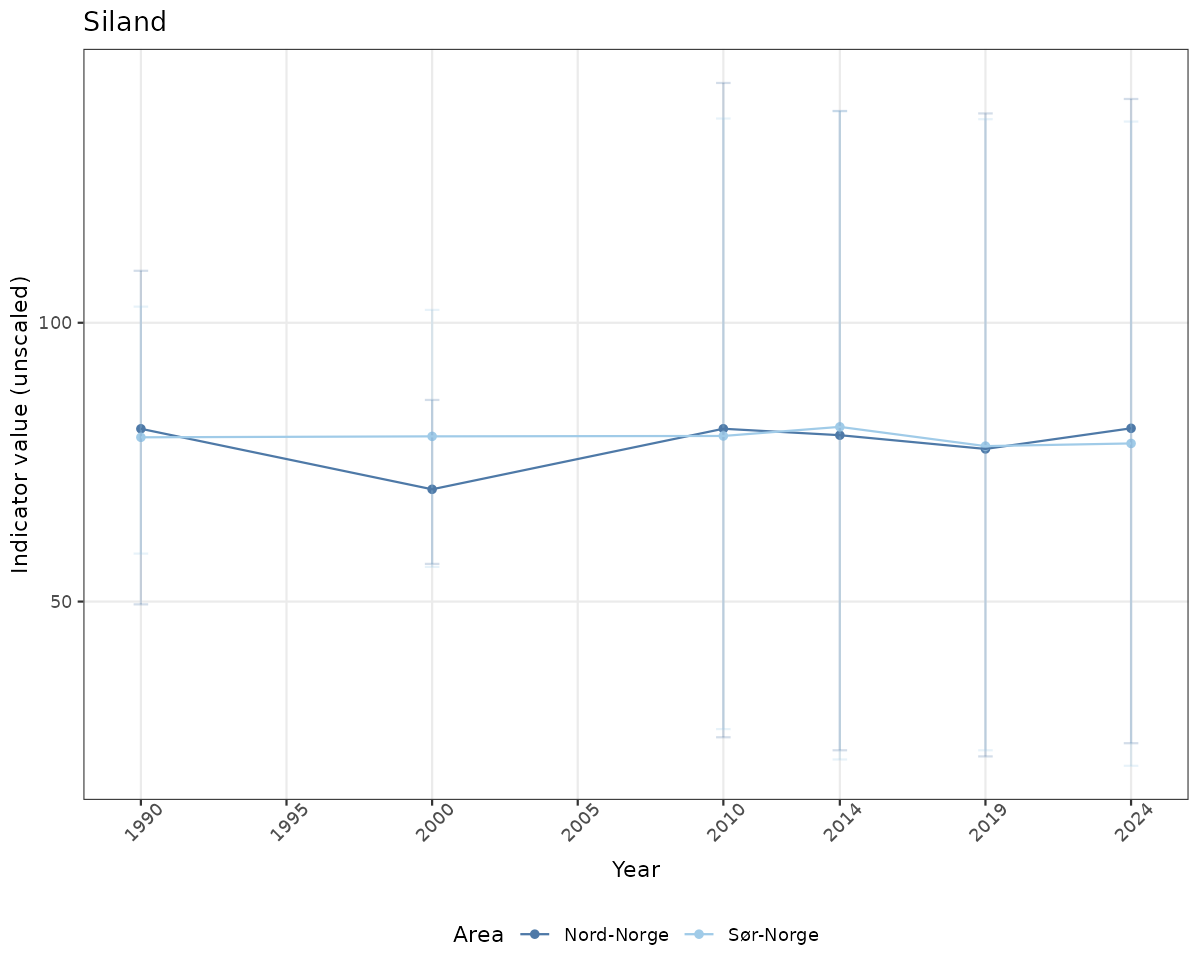
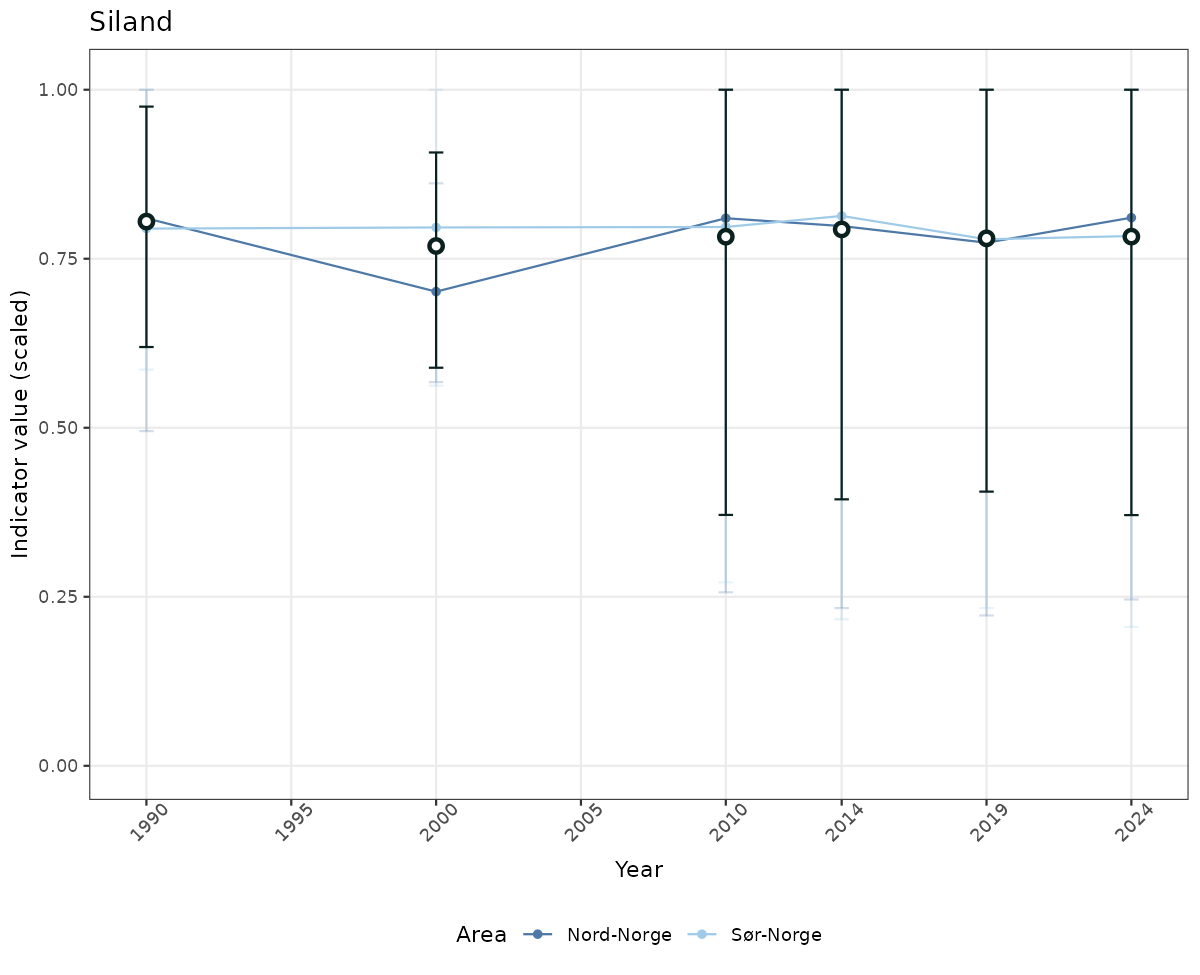
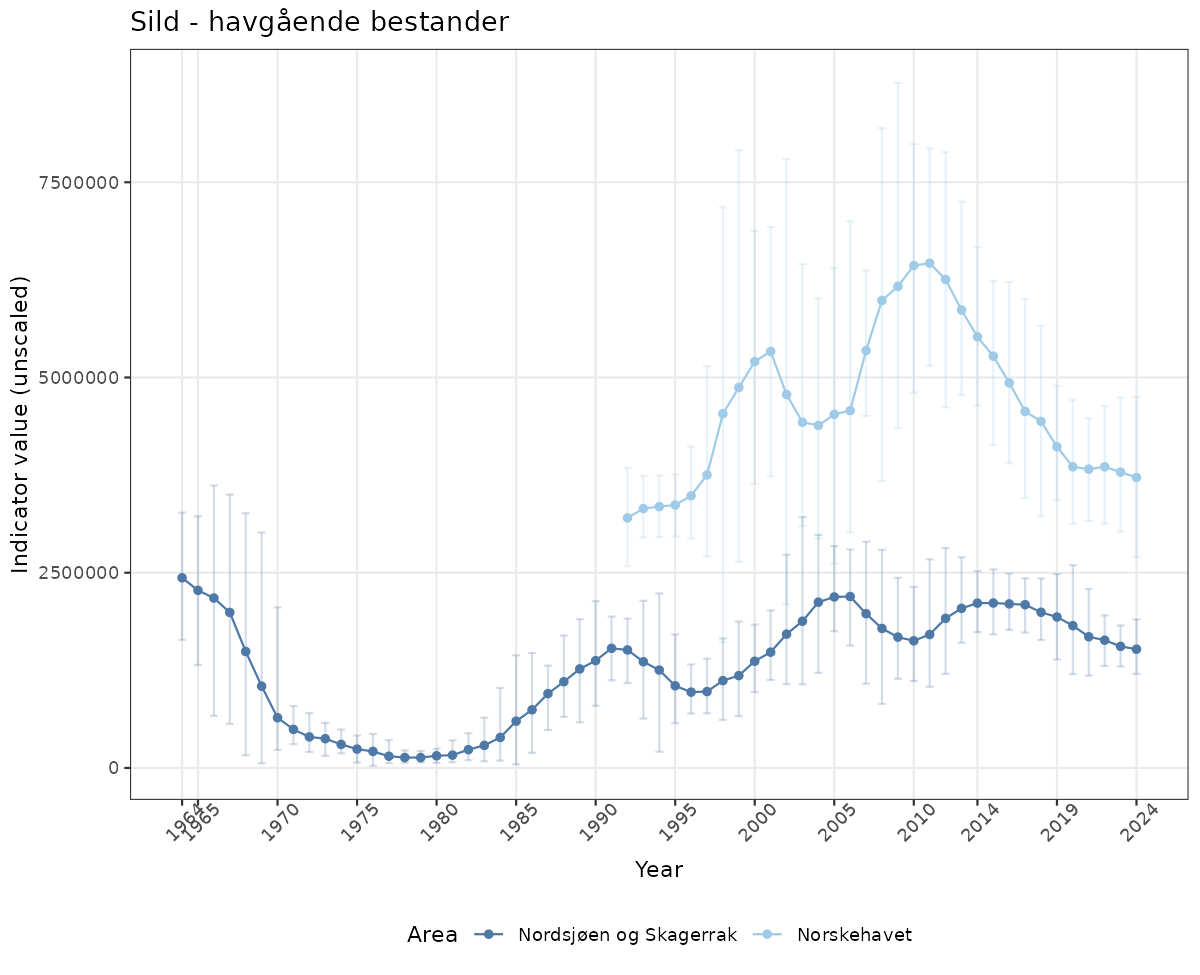
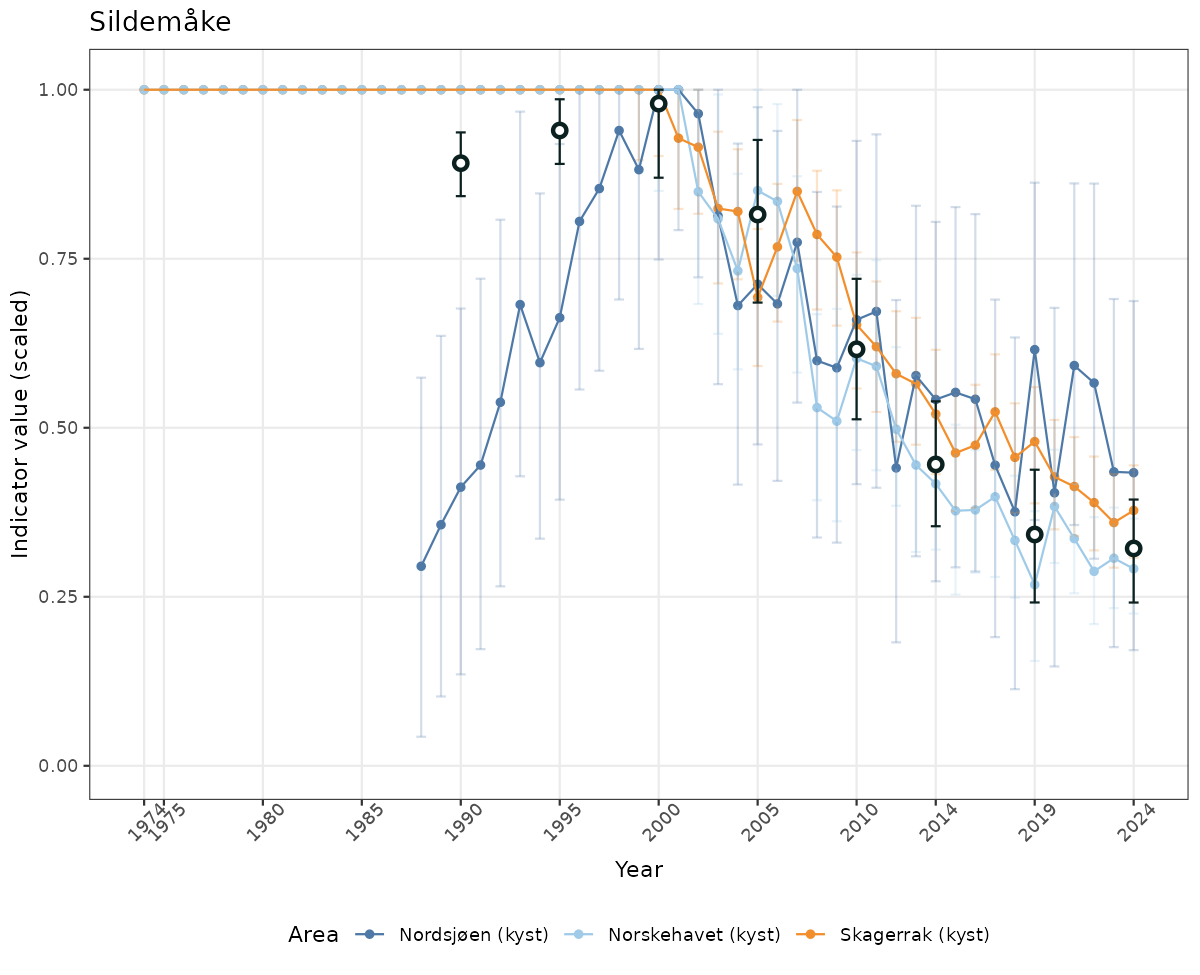
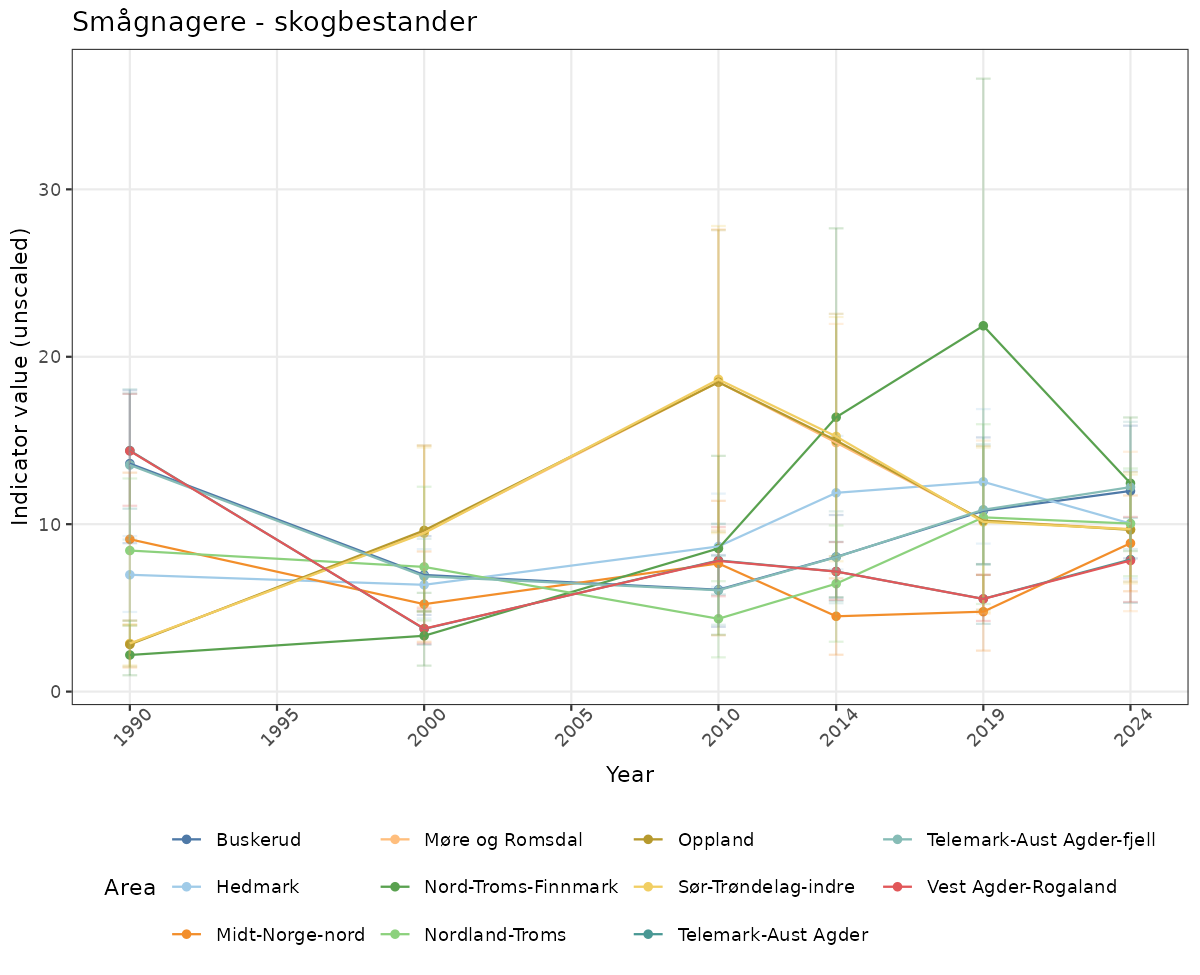
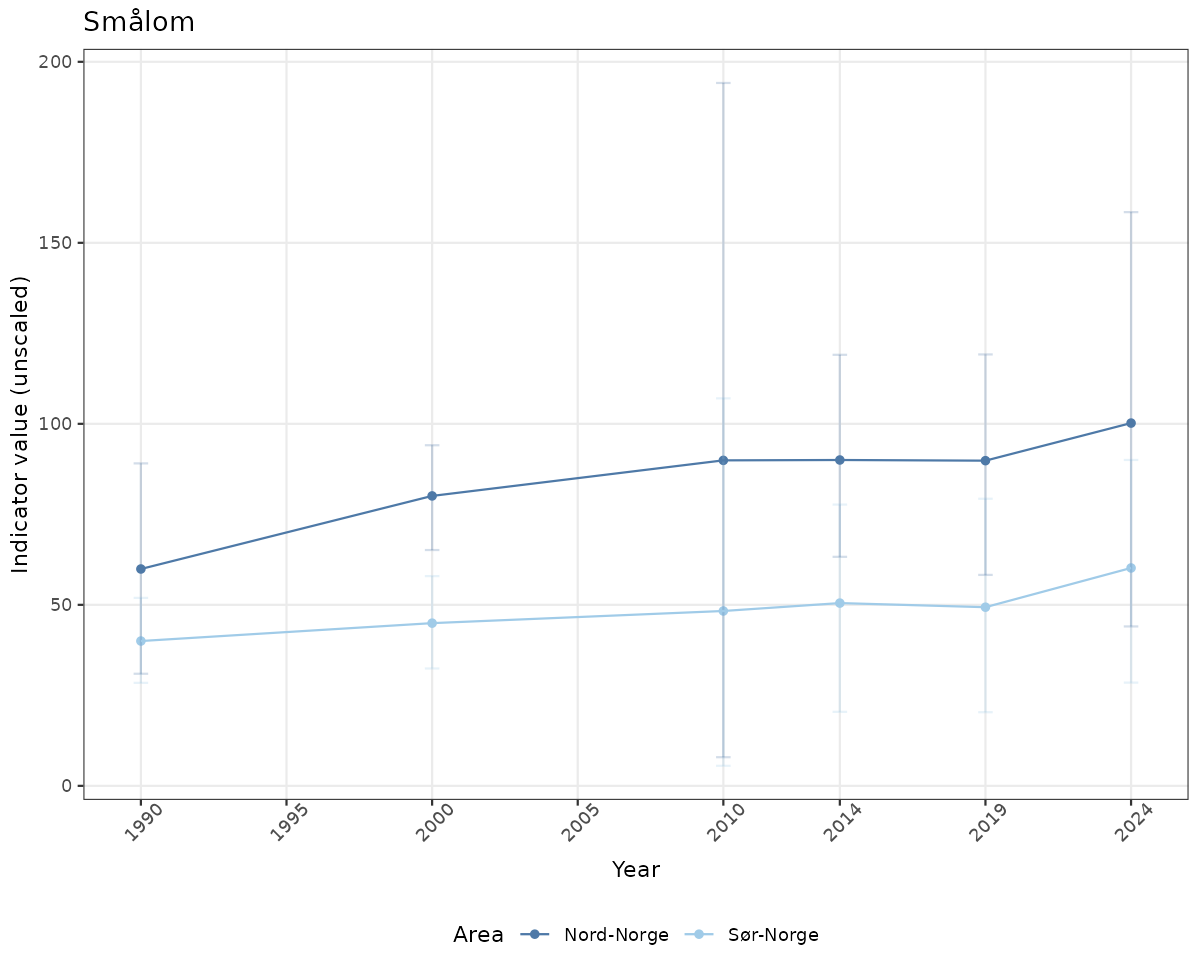
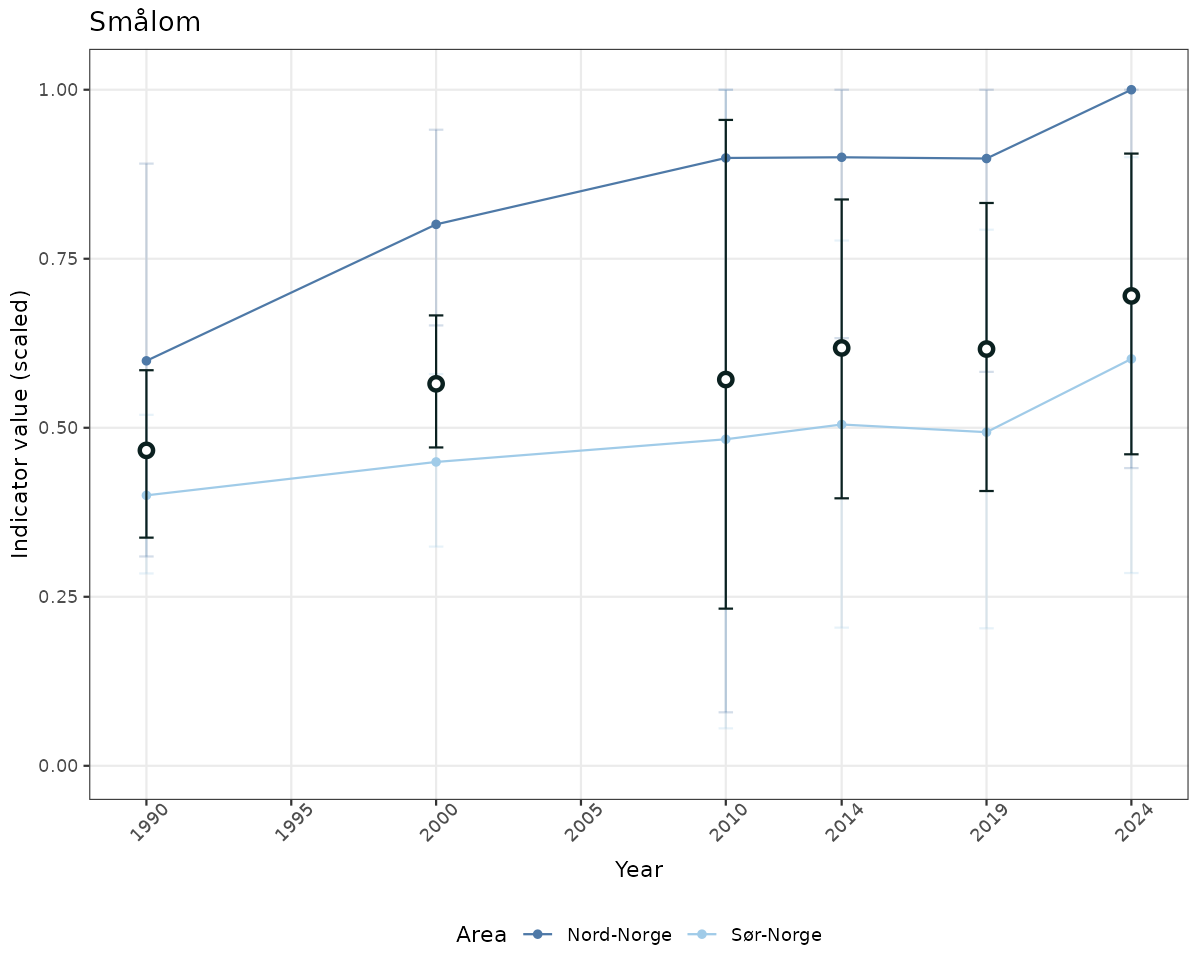
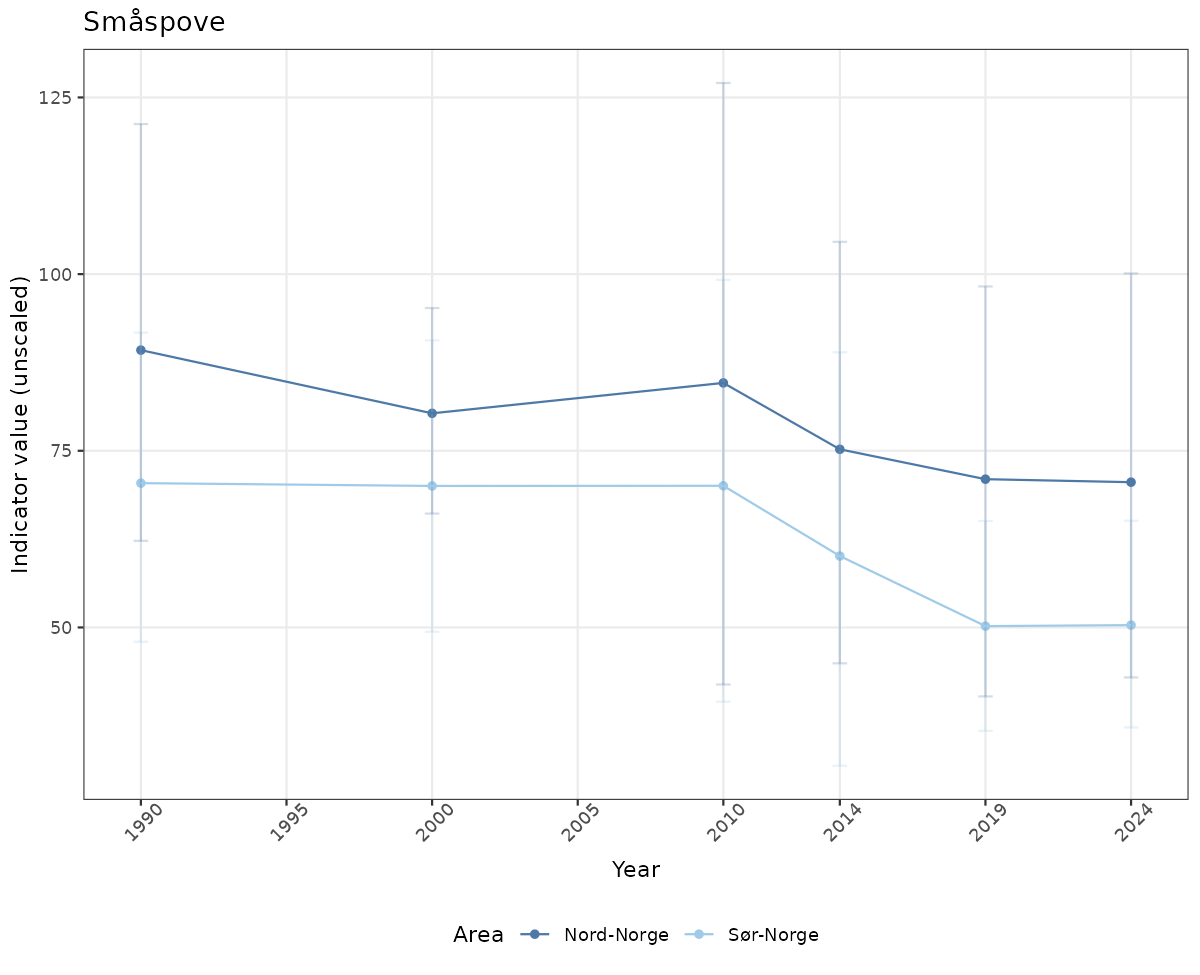
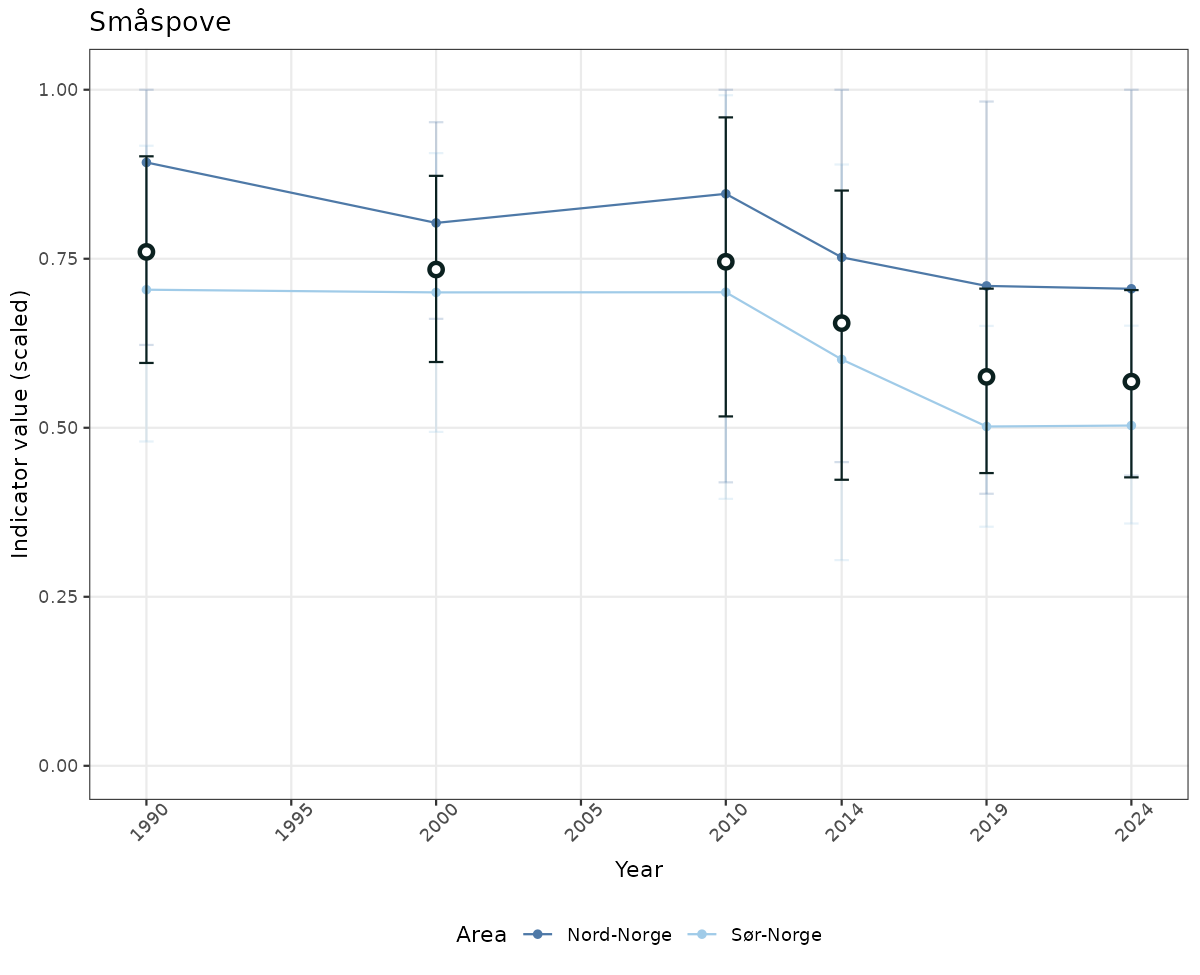
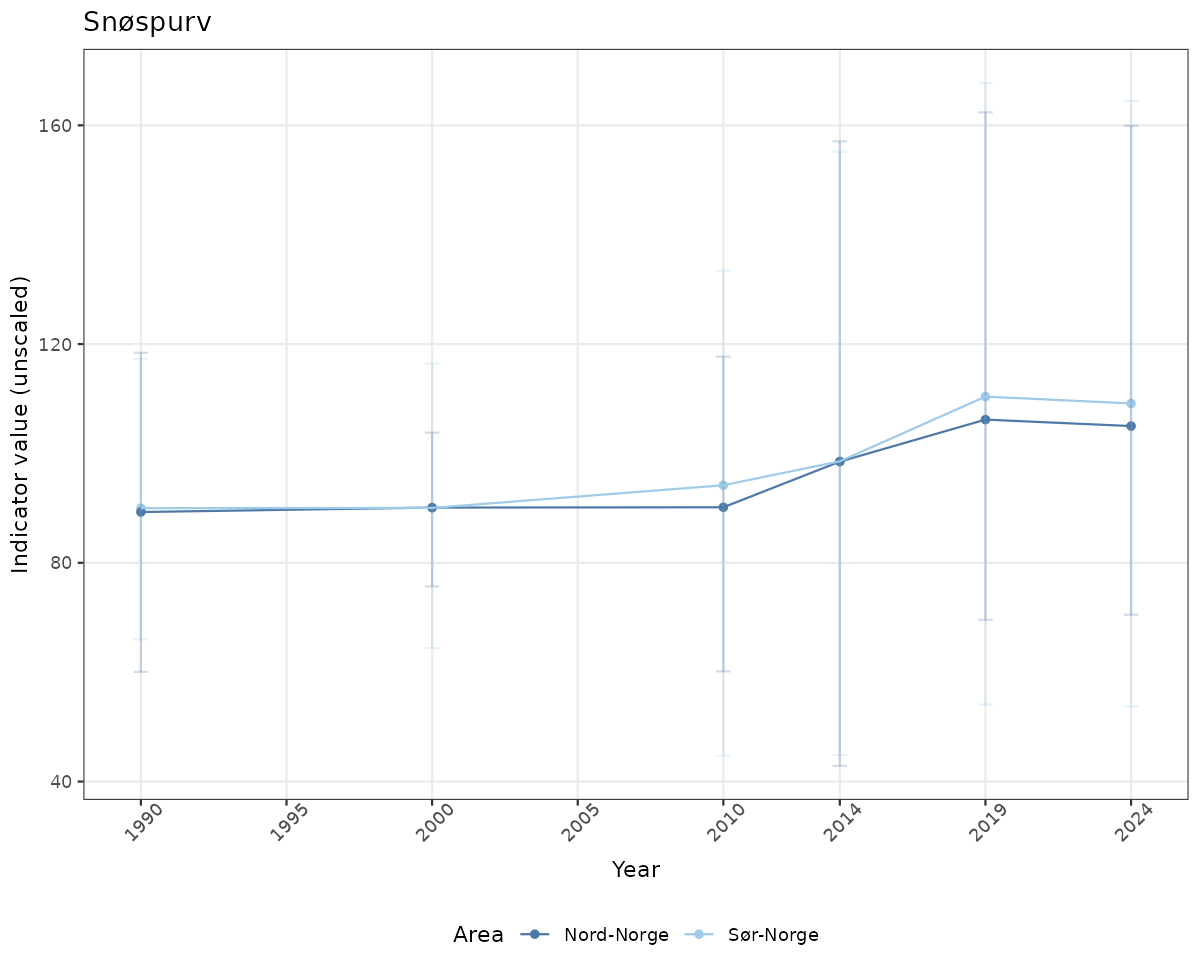
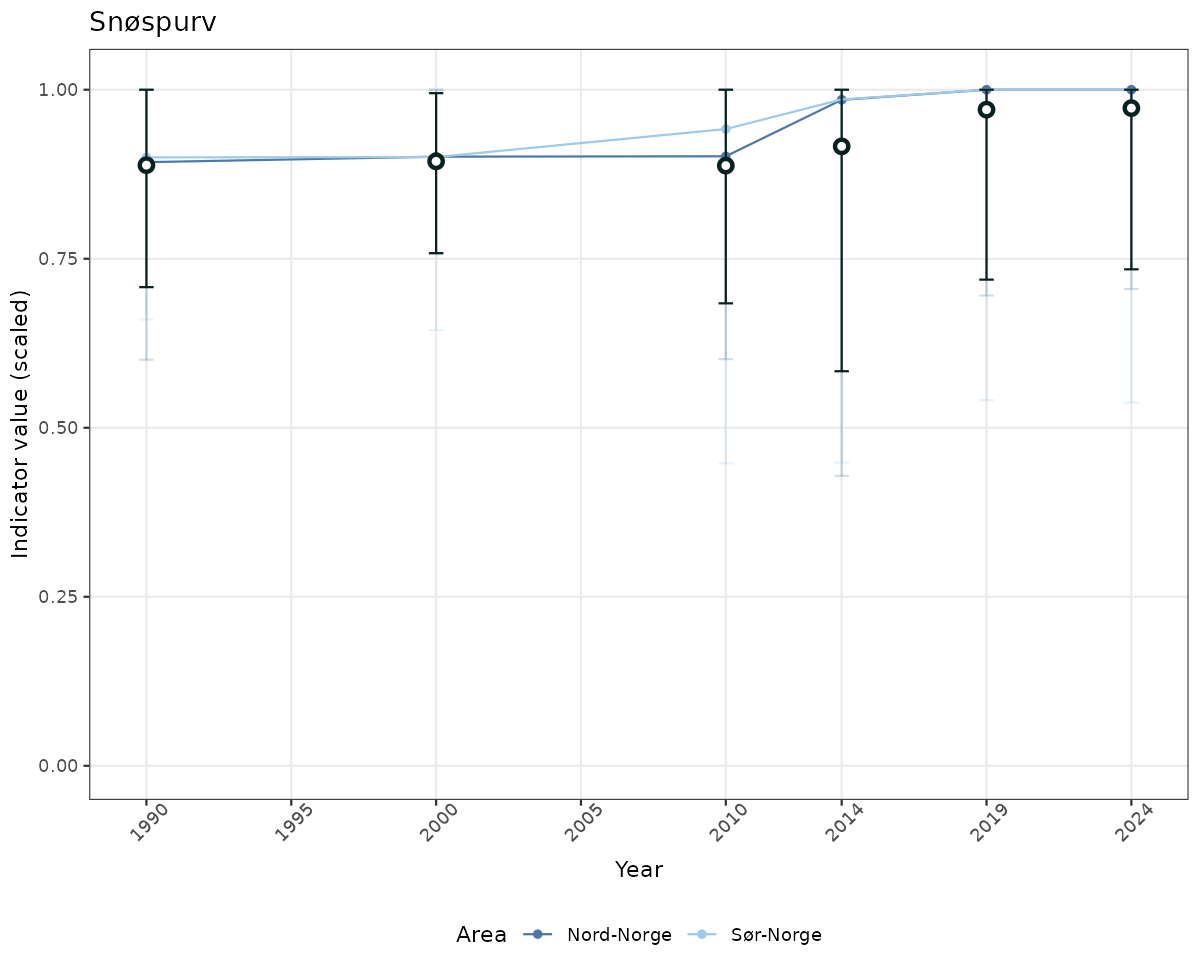
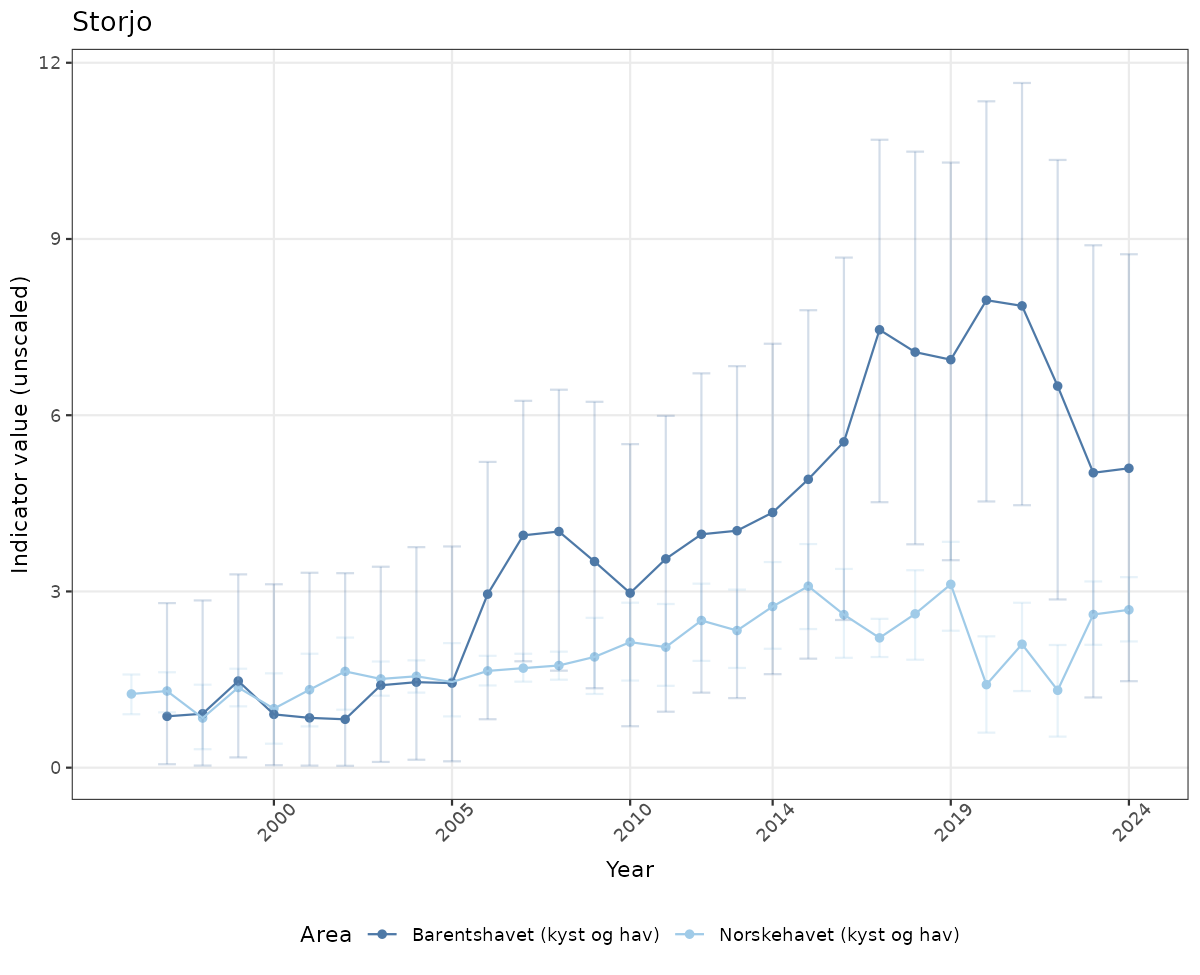
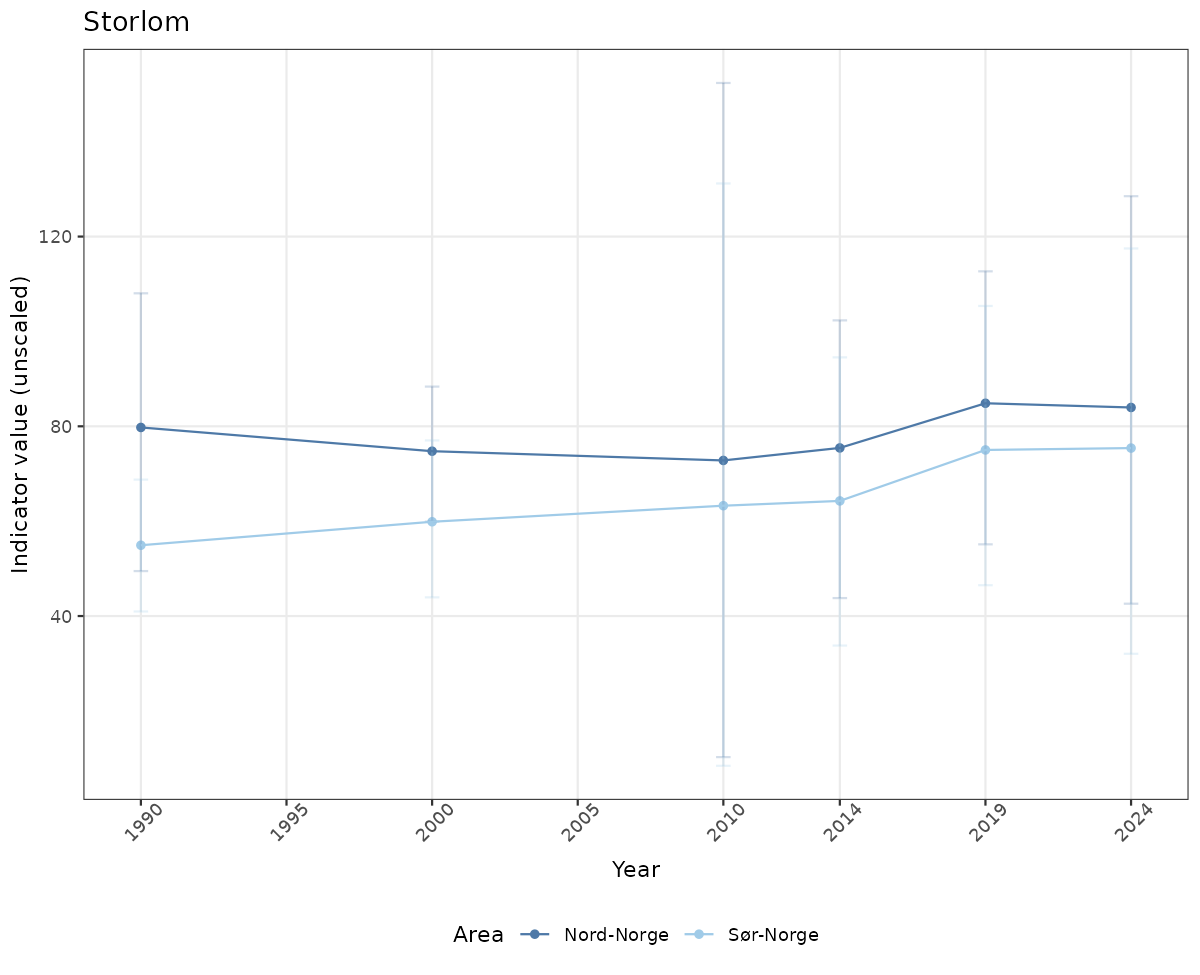
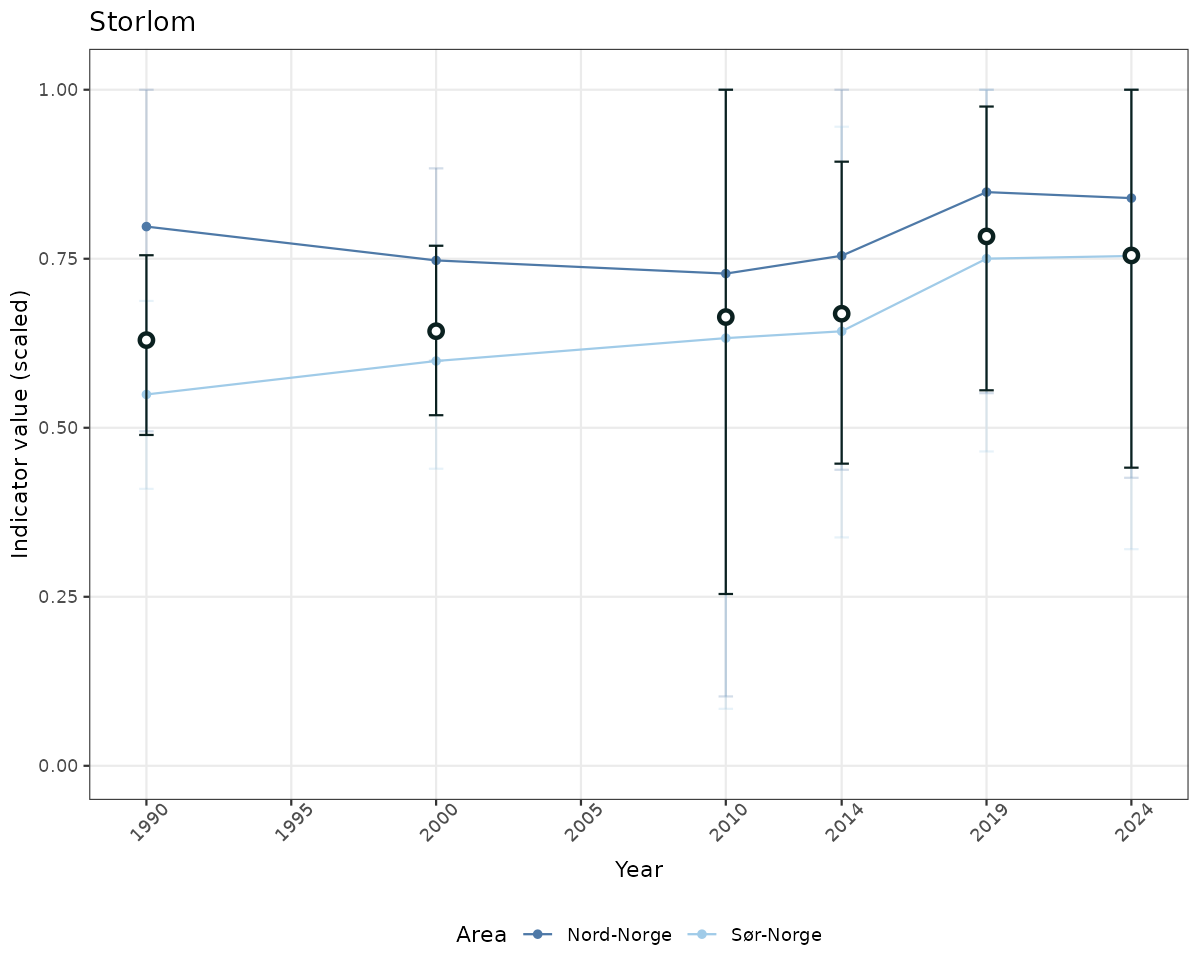
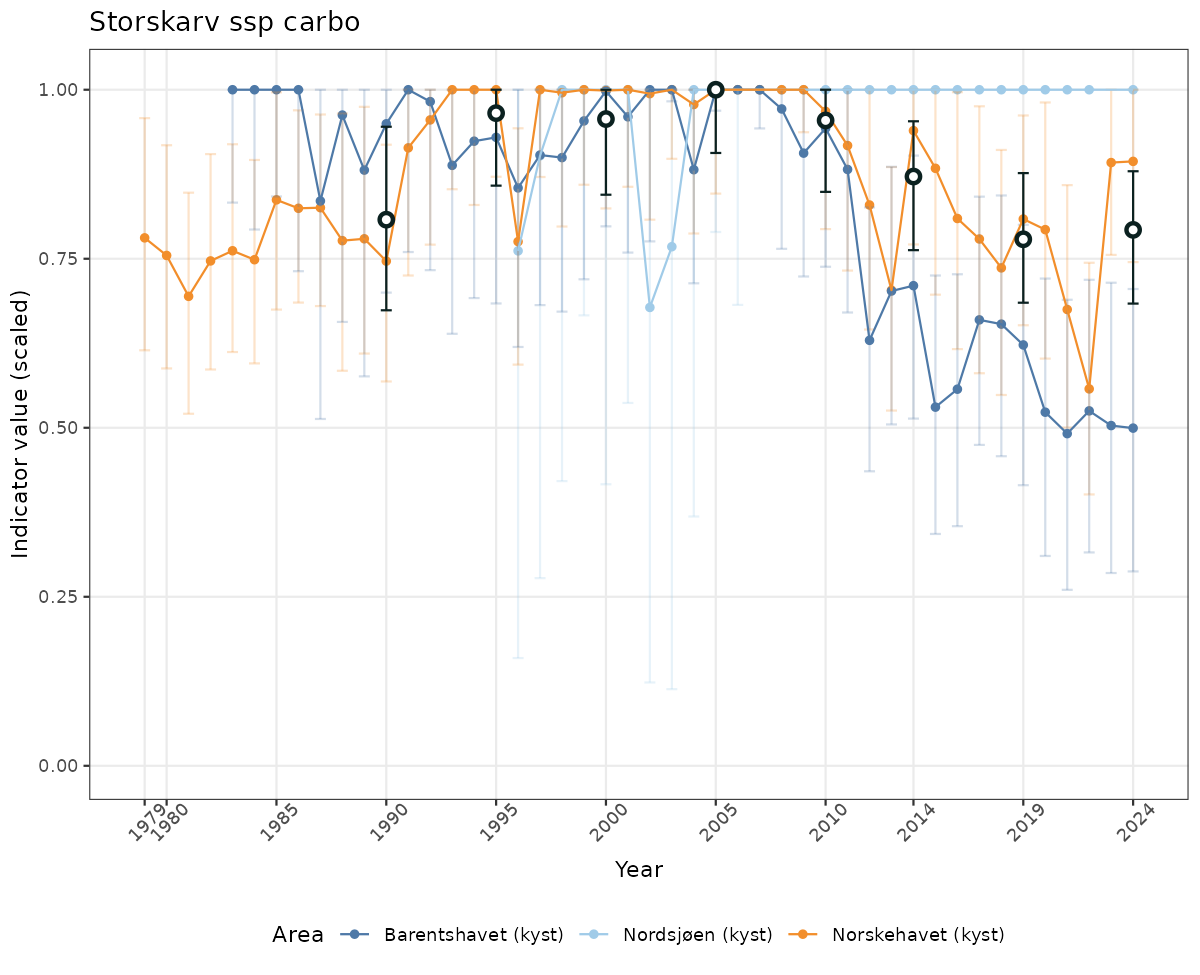
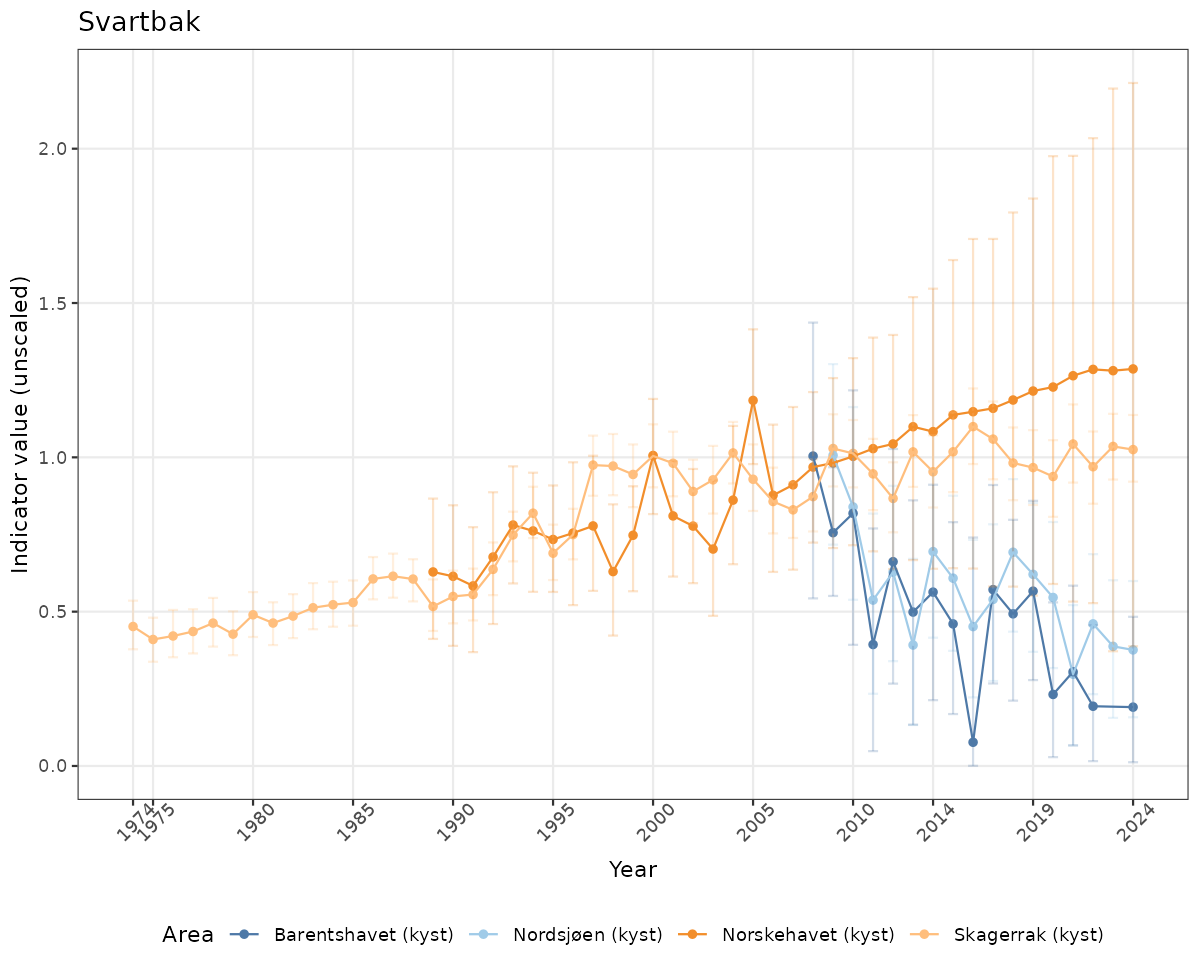
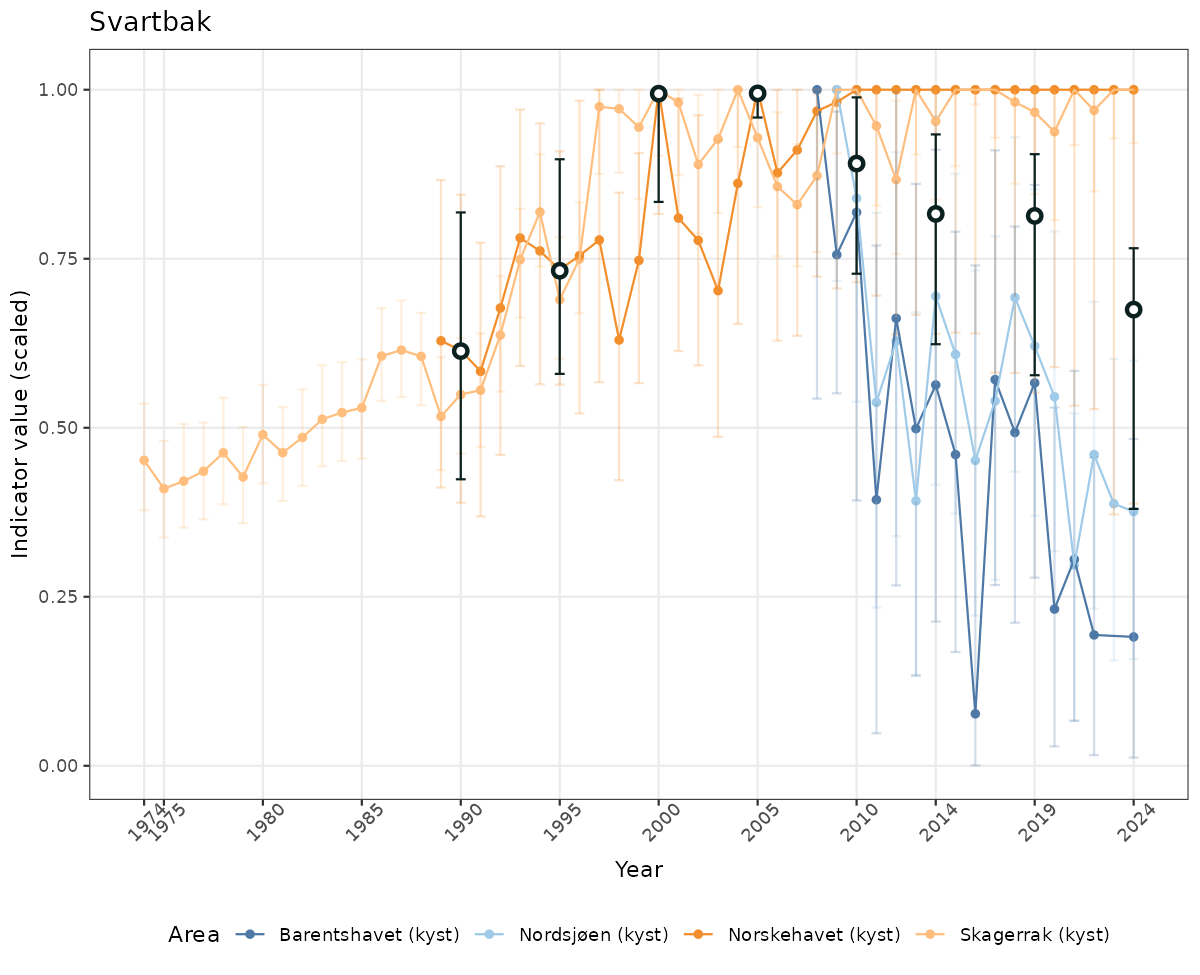
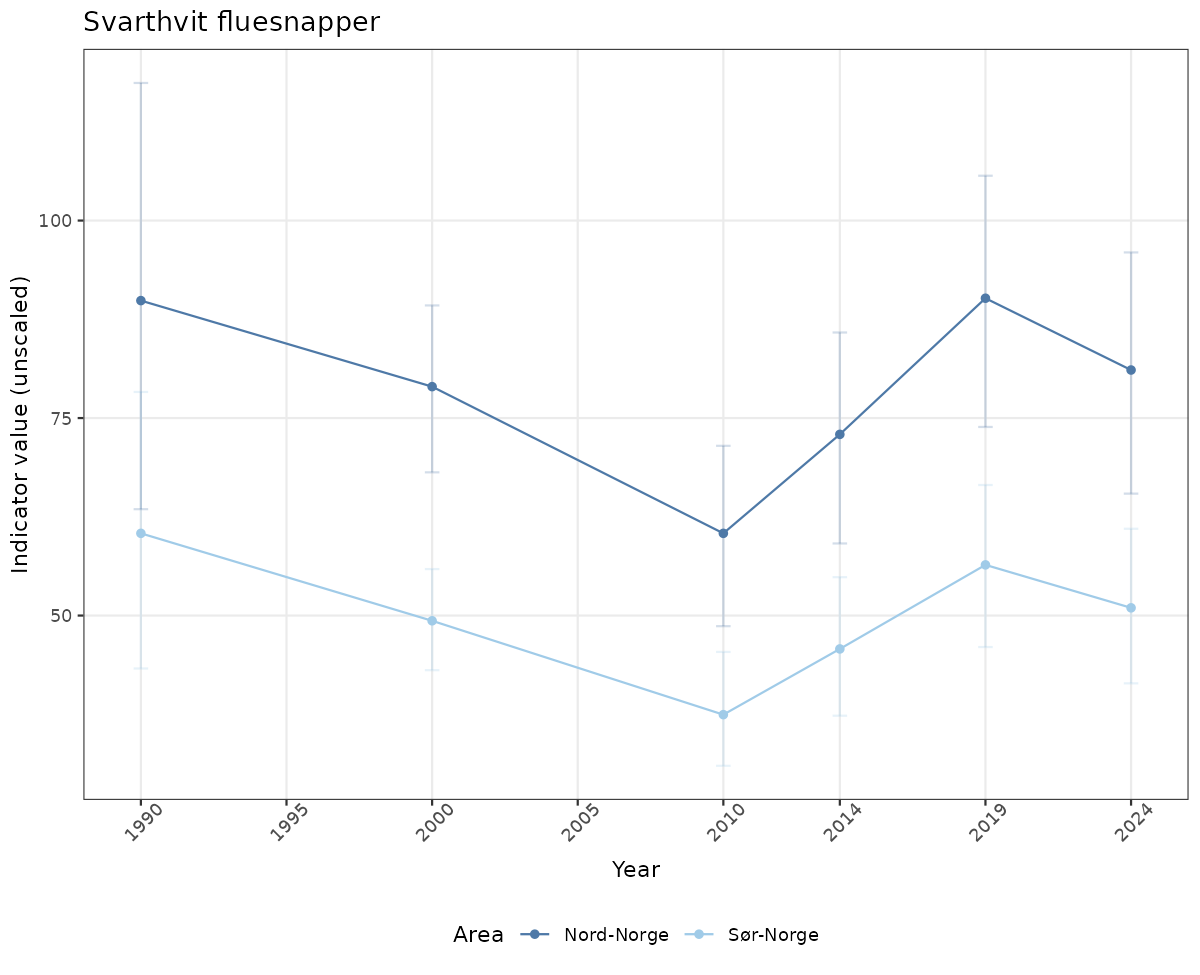
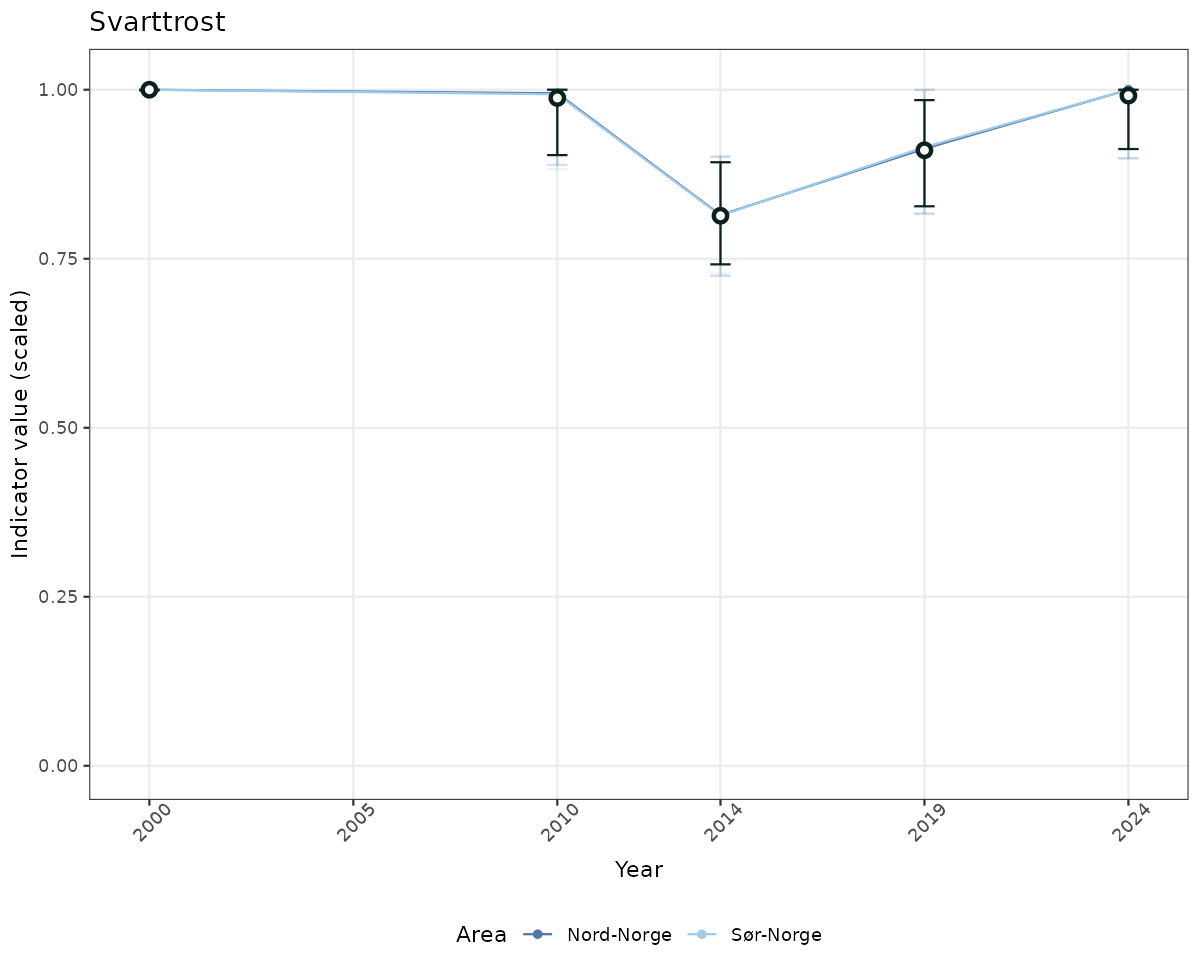
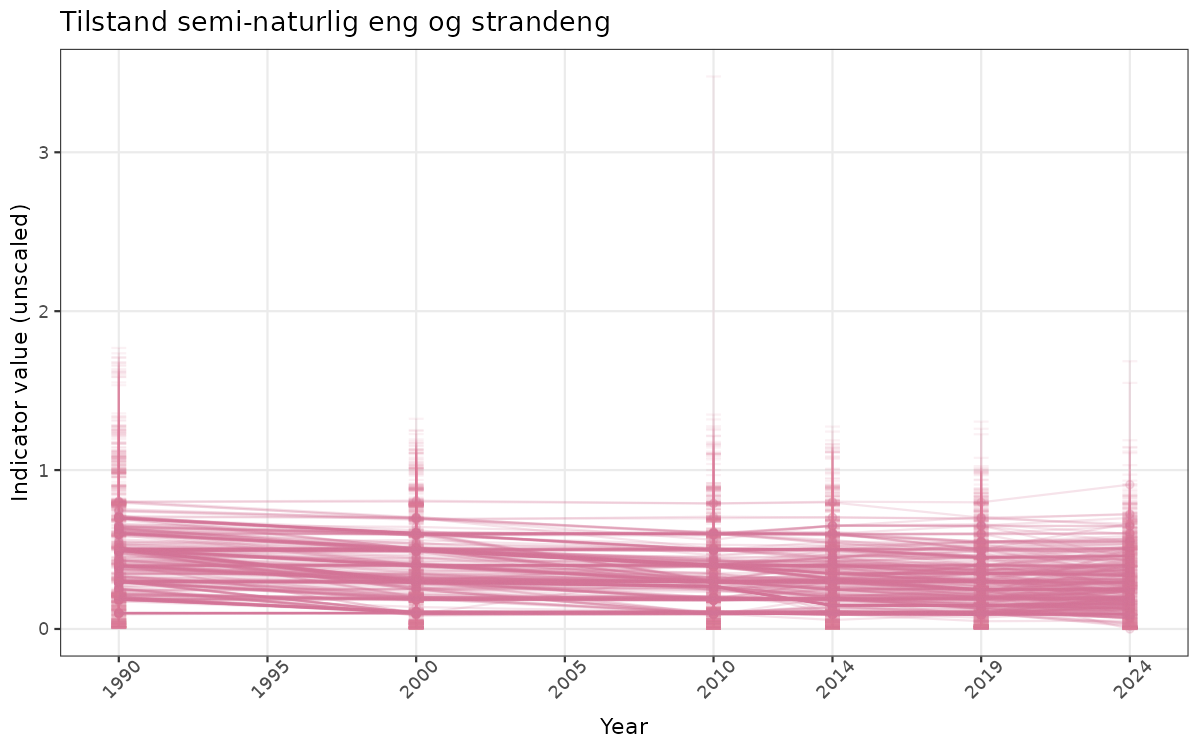
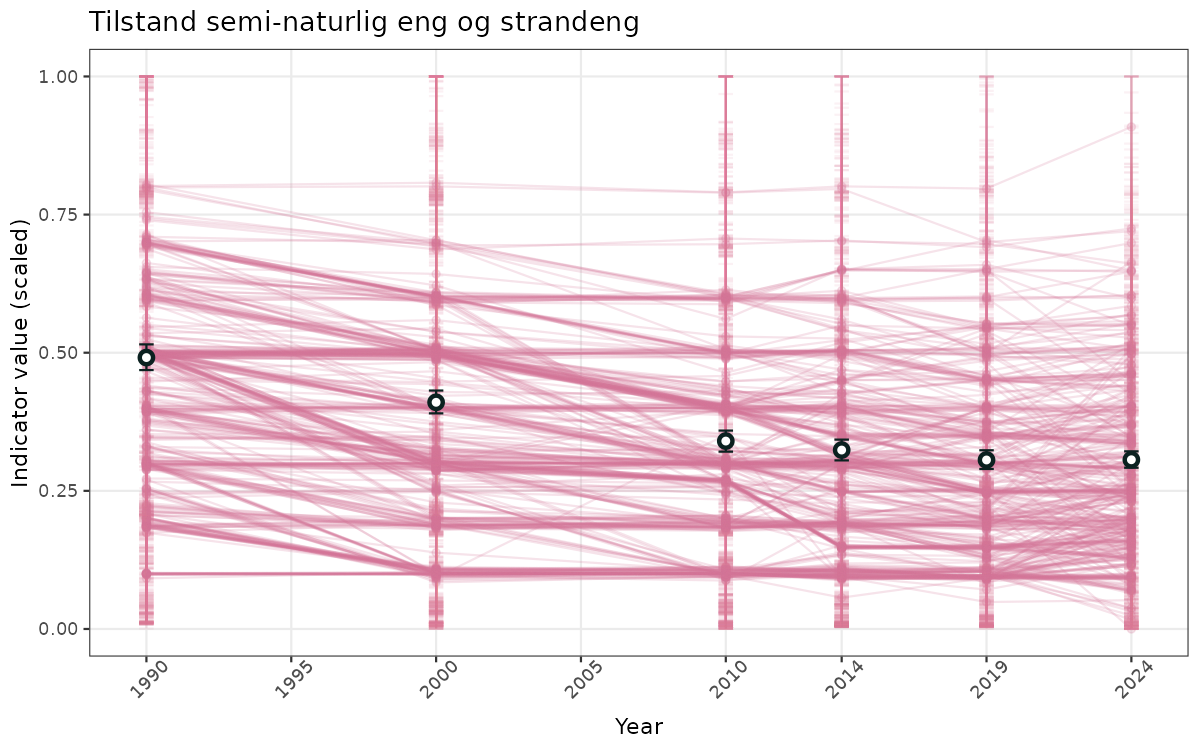
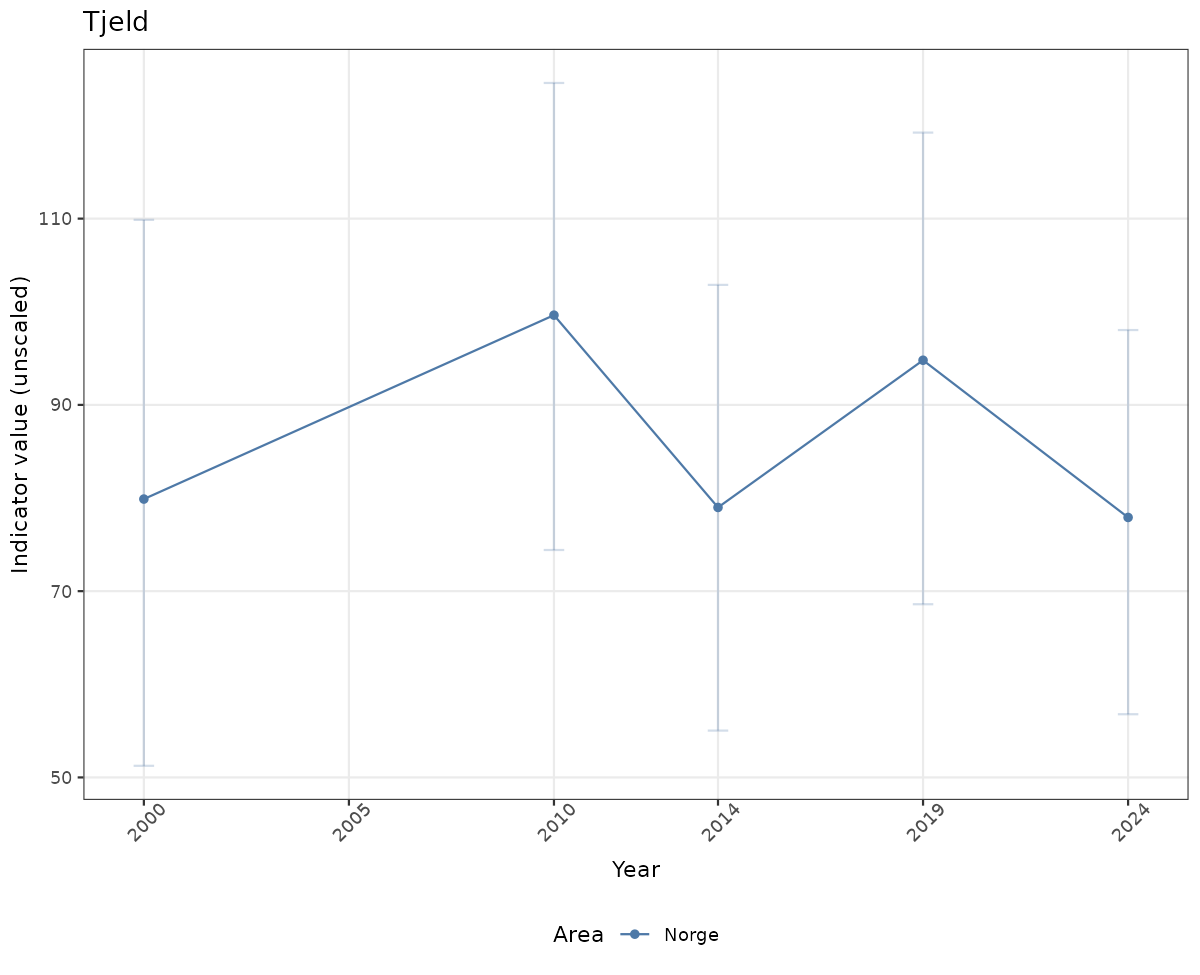
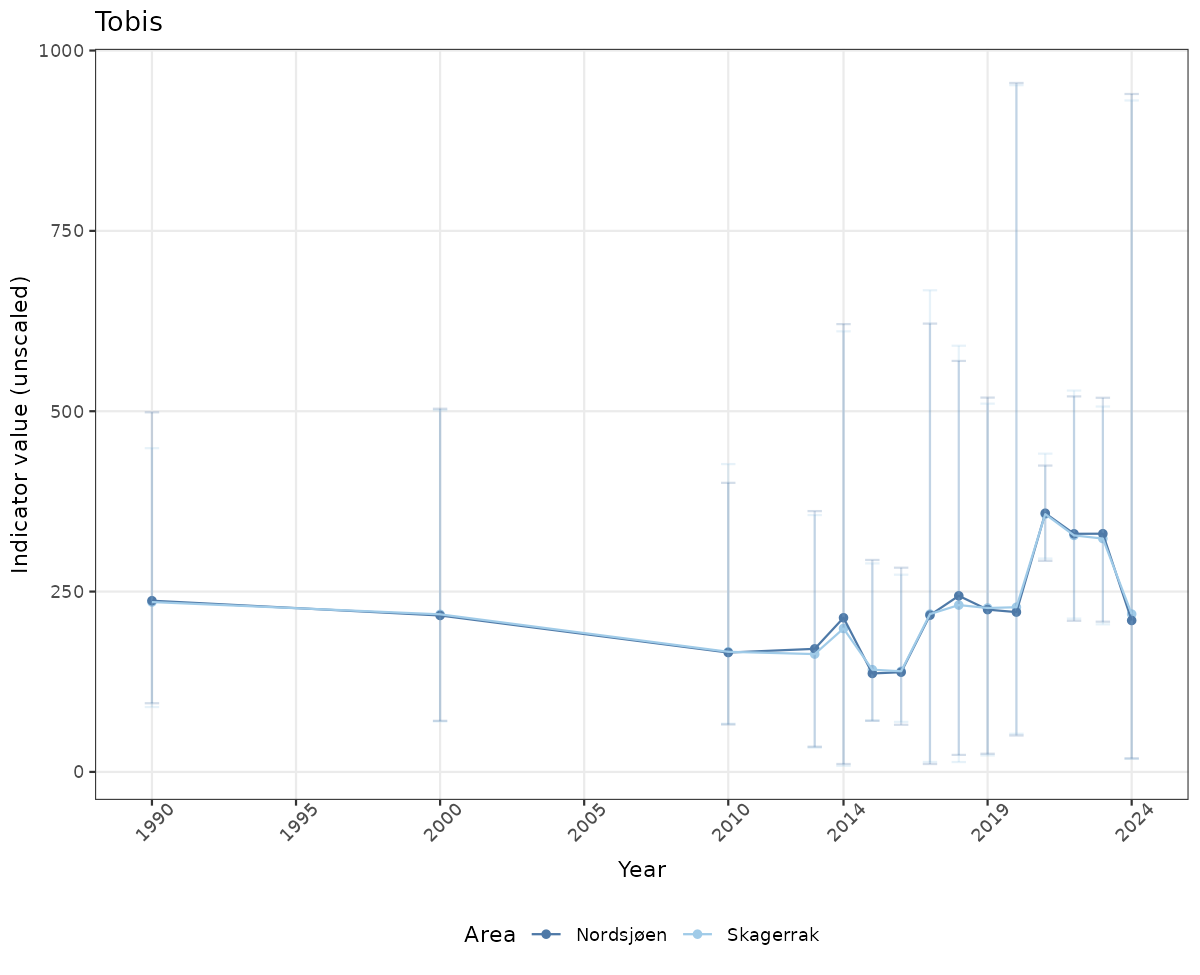
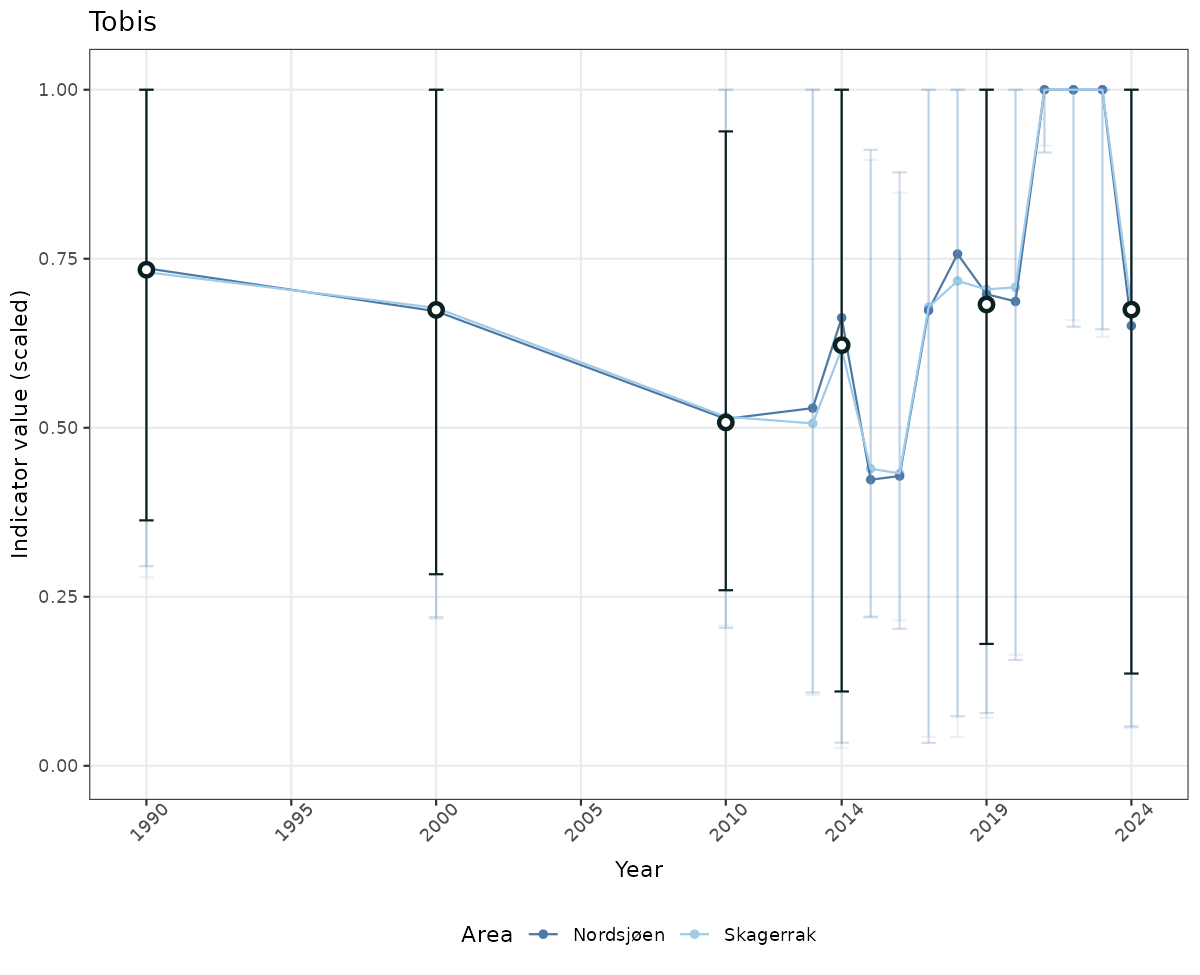
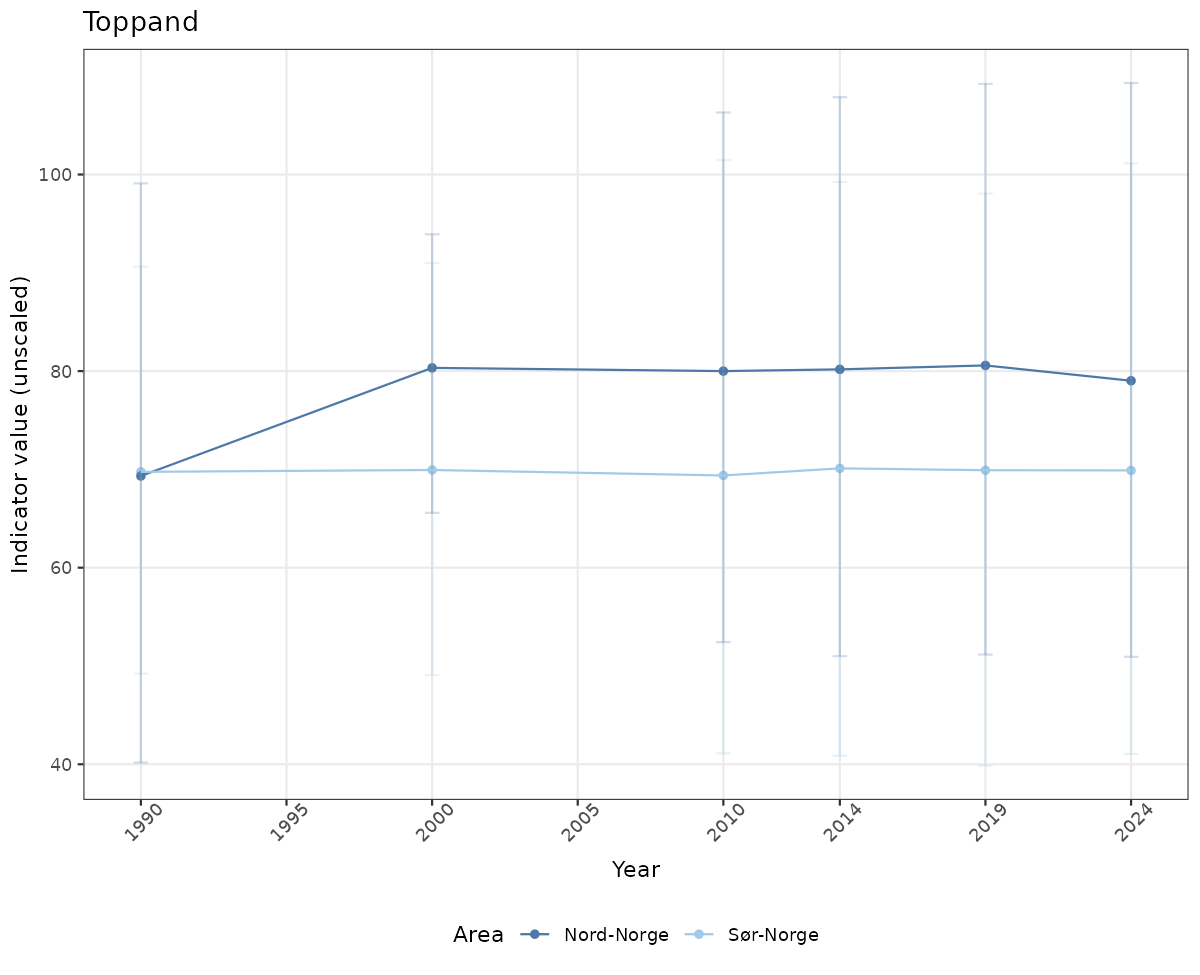
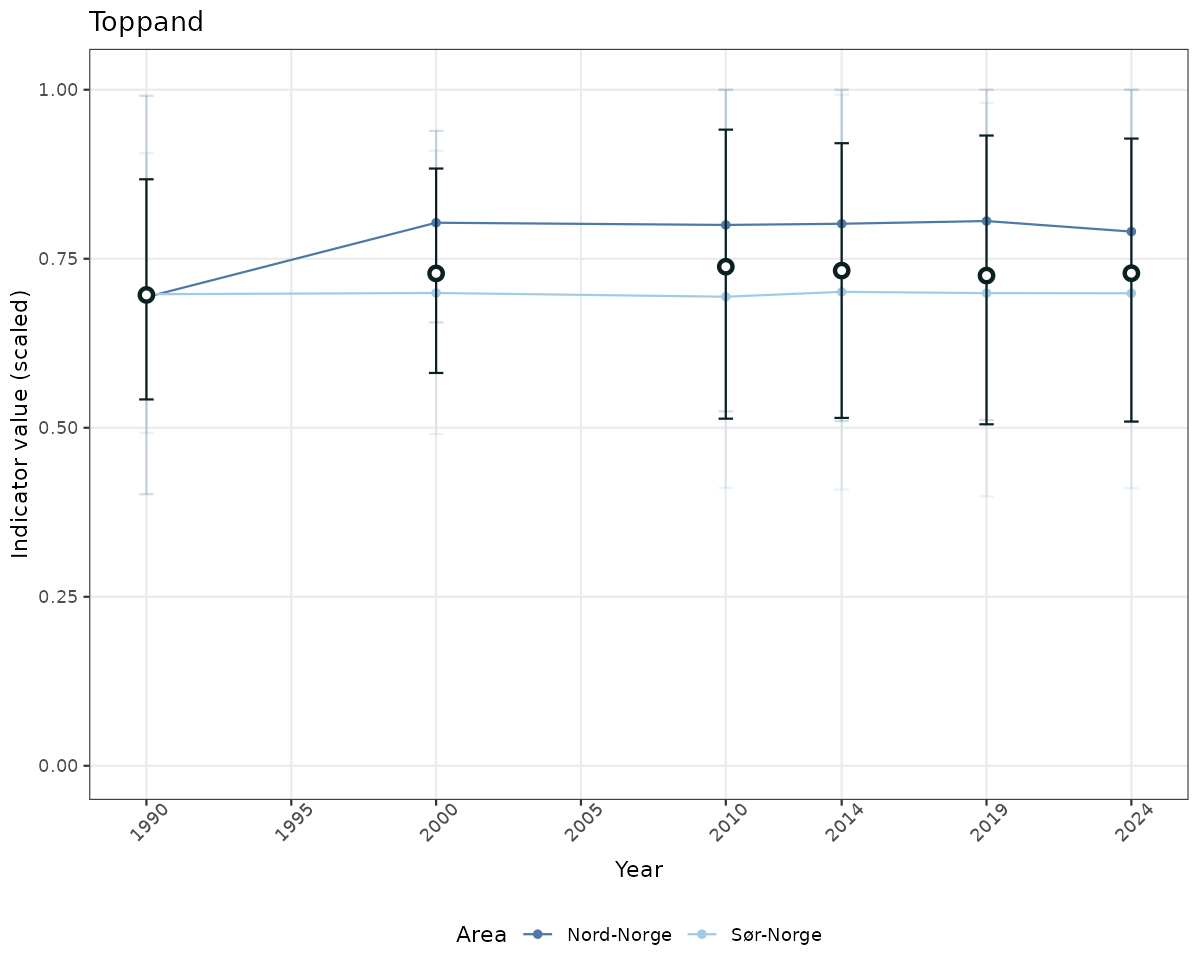
- Time-series plots of medians and 95% confidence intervals (featuring lines for different areas)
- Year-specific maps of a) medians and b) uncertainty as relative standard deviation
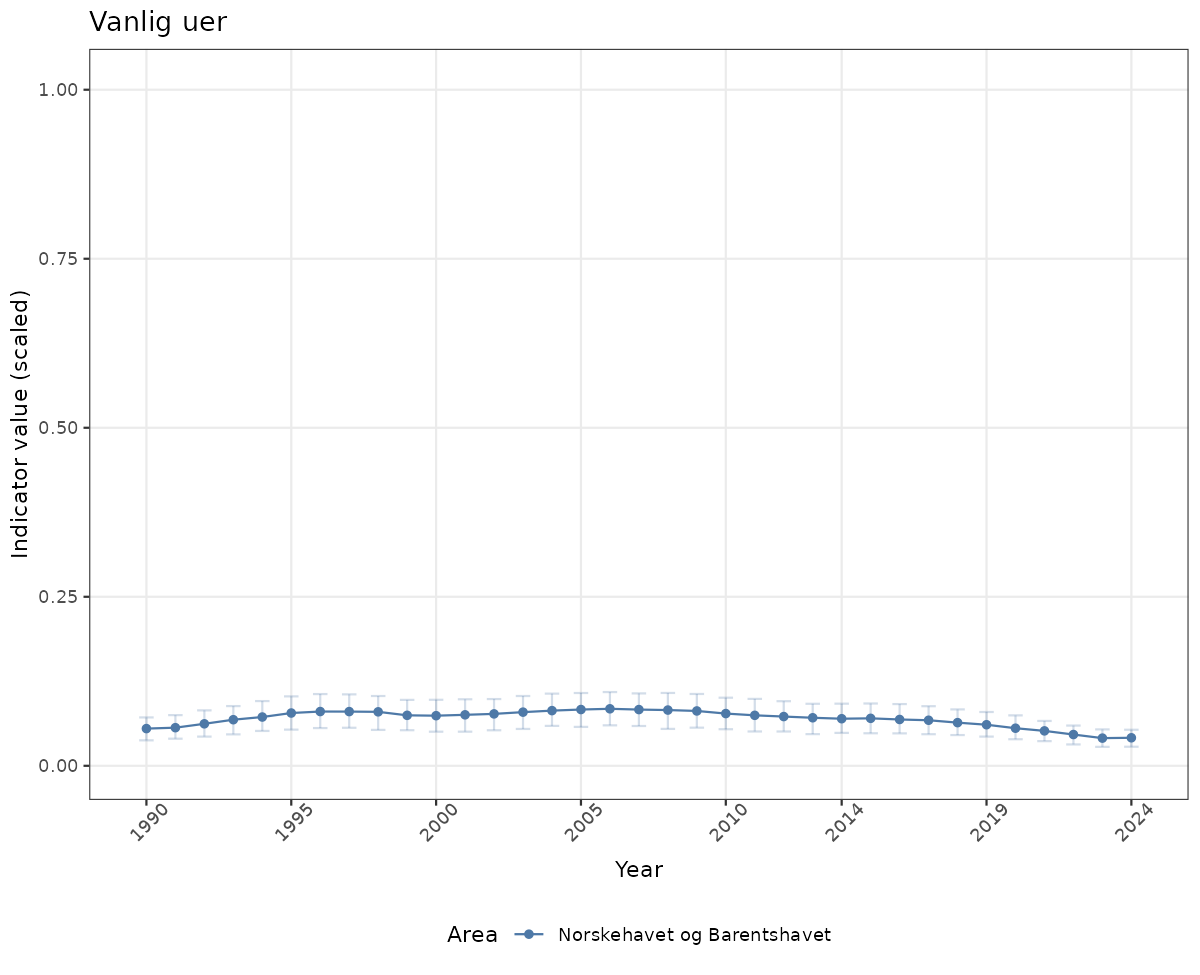
Both visualizations will be done for the unscaled (raw) and scaled indicator values.
Below, we present the corresponding visualizations in indicator-specific sub-chapters. There are a few things to note:
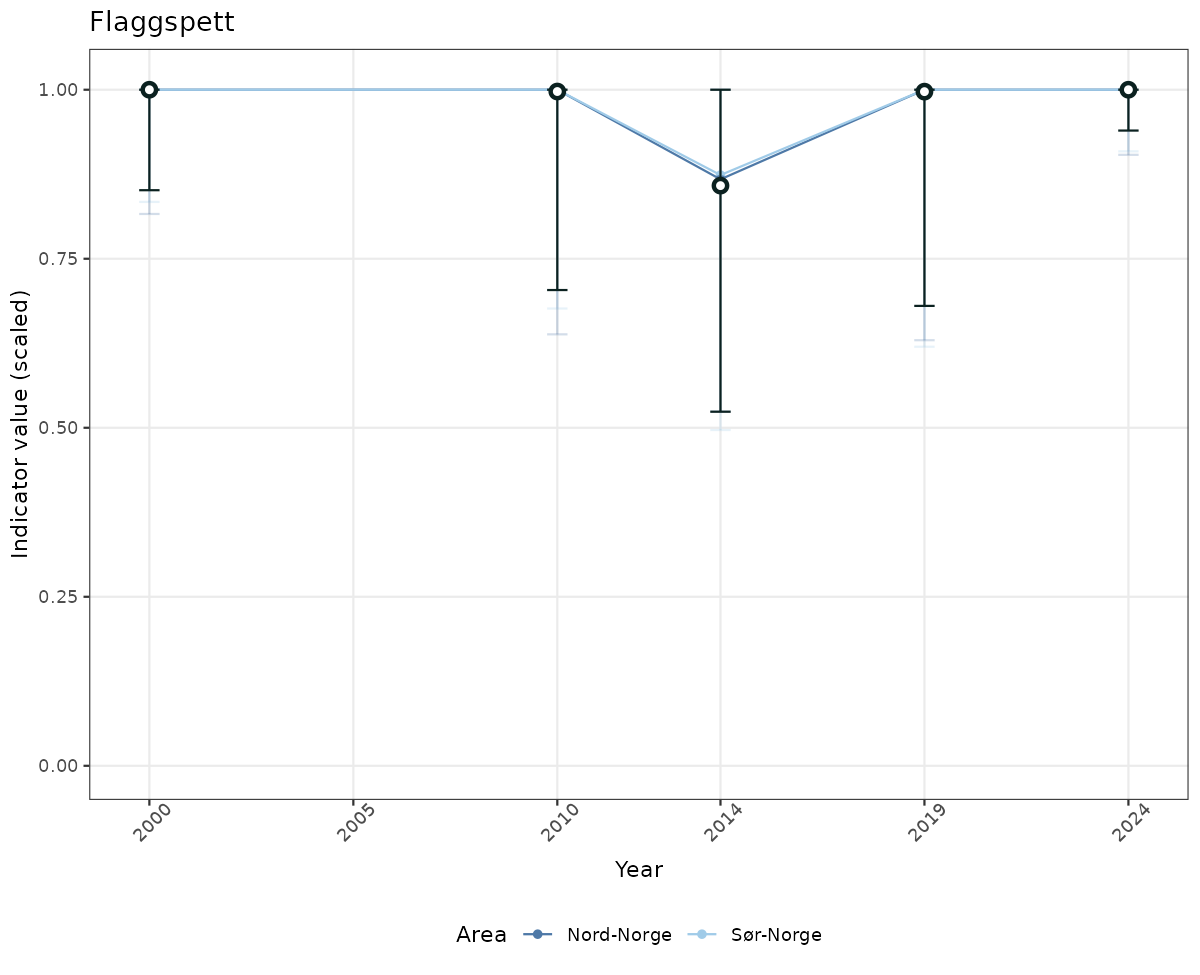
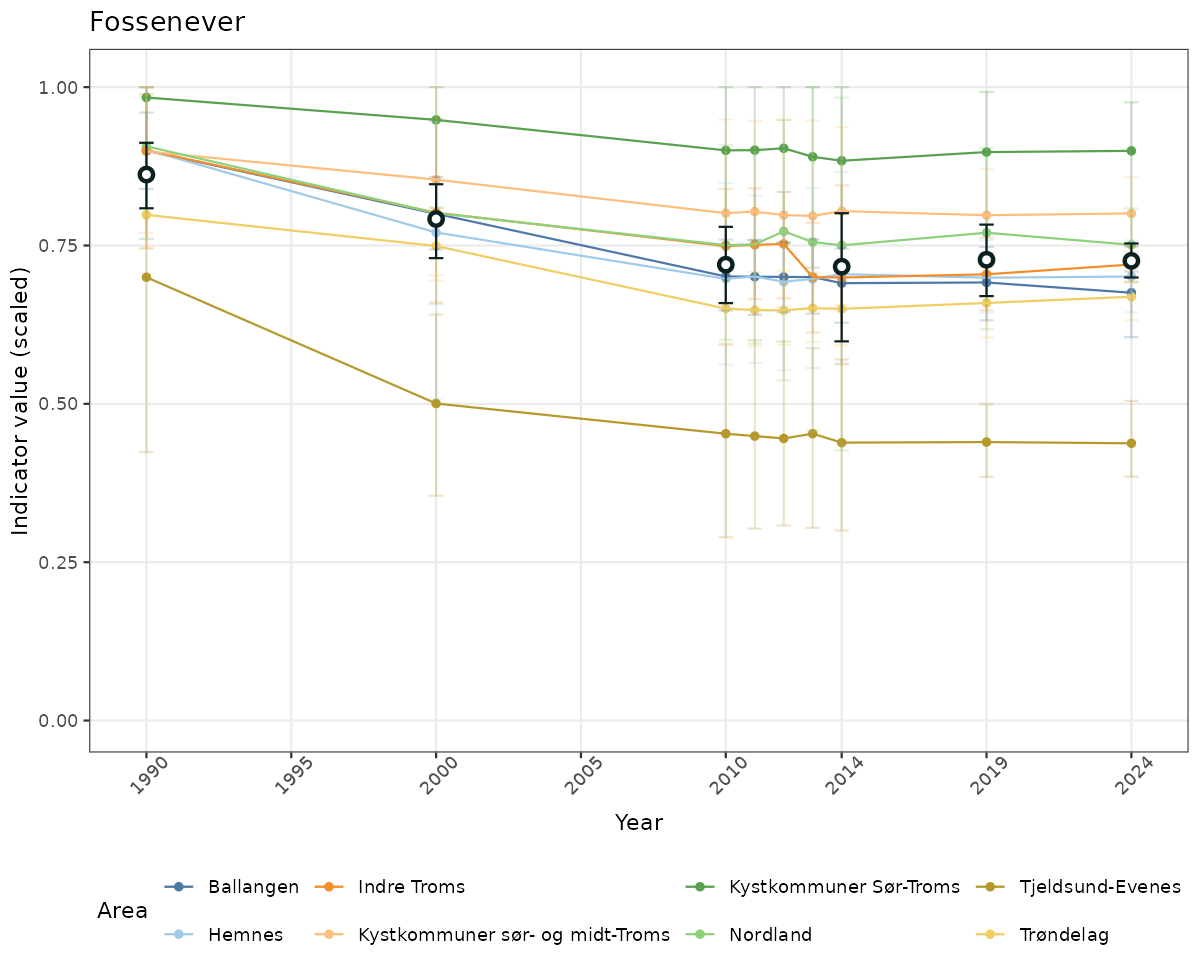
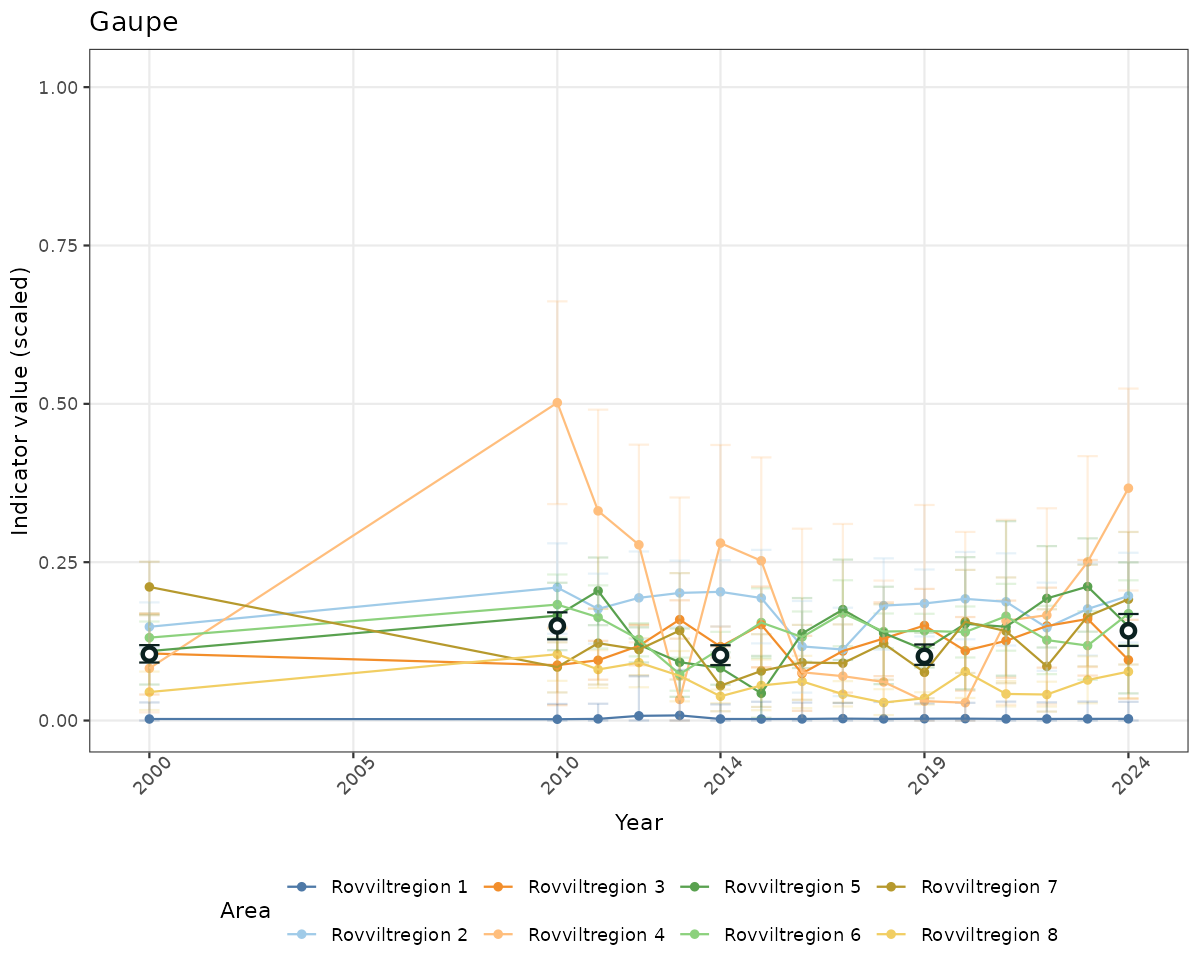
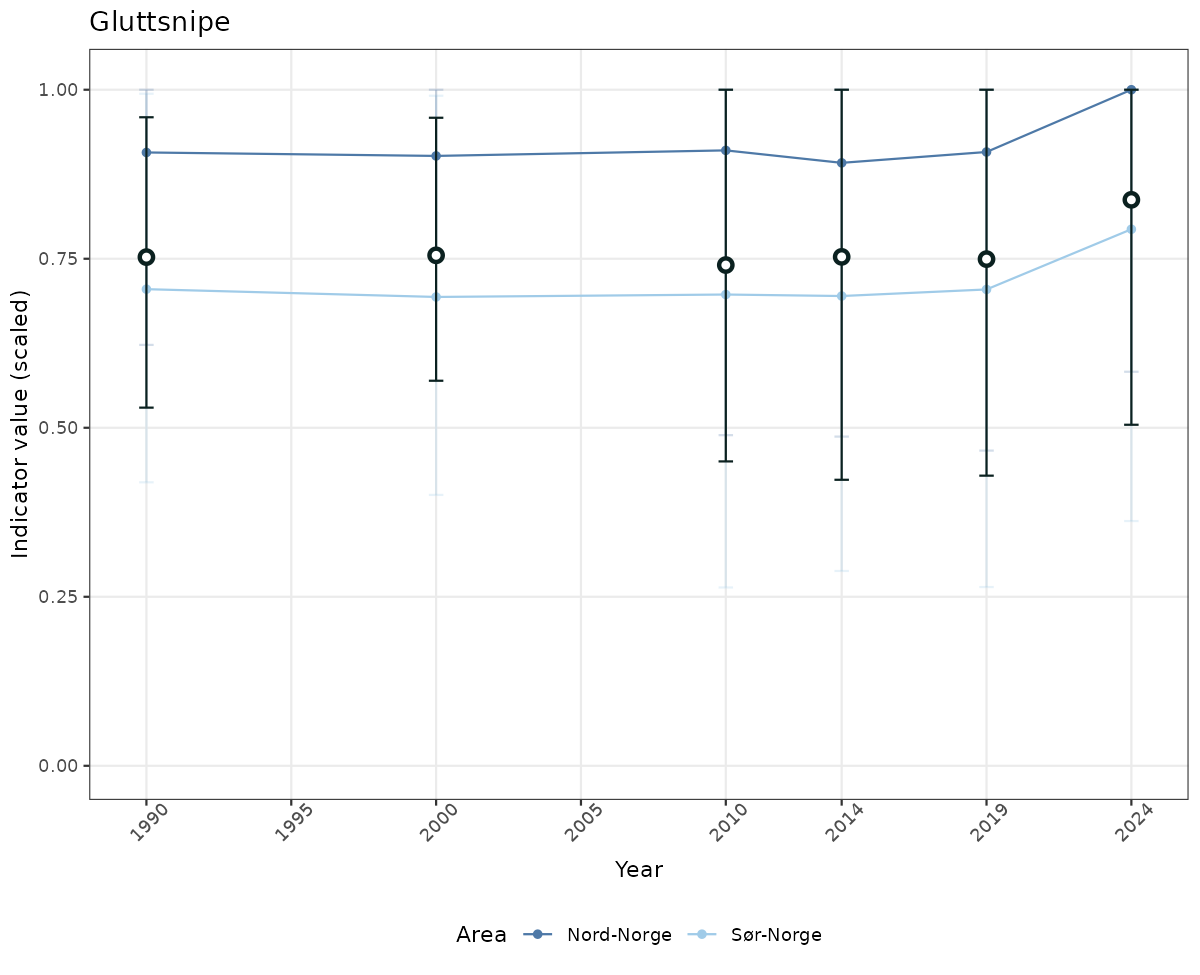
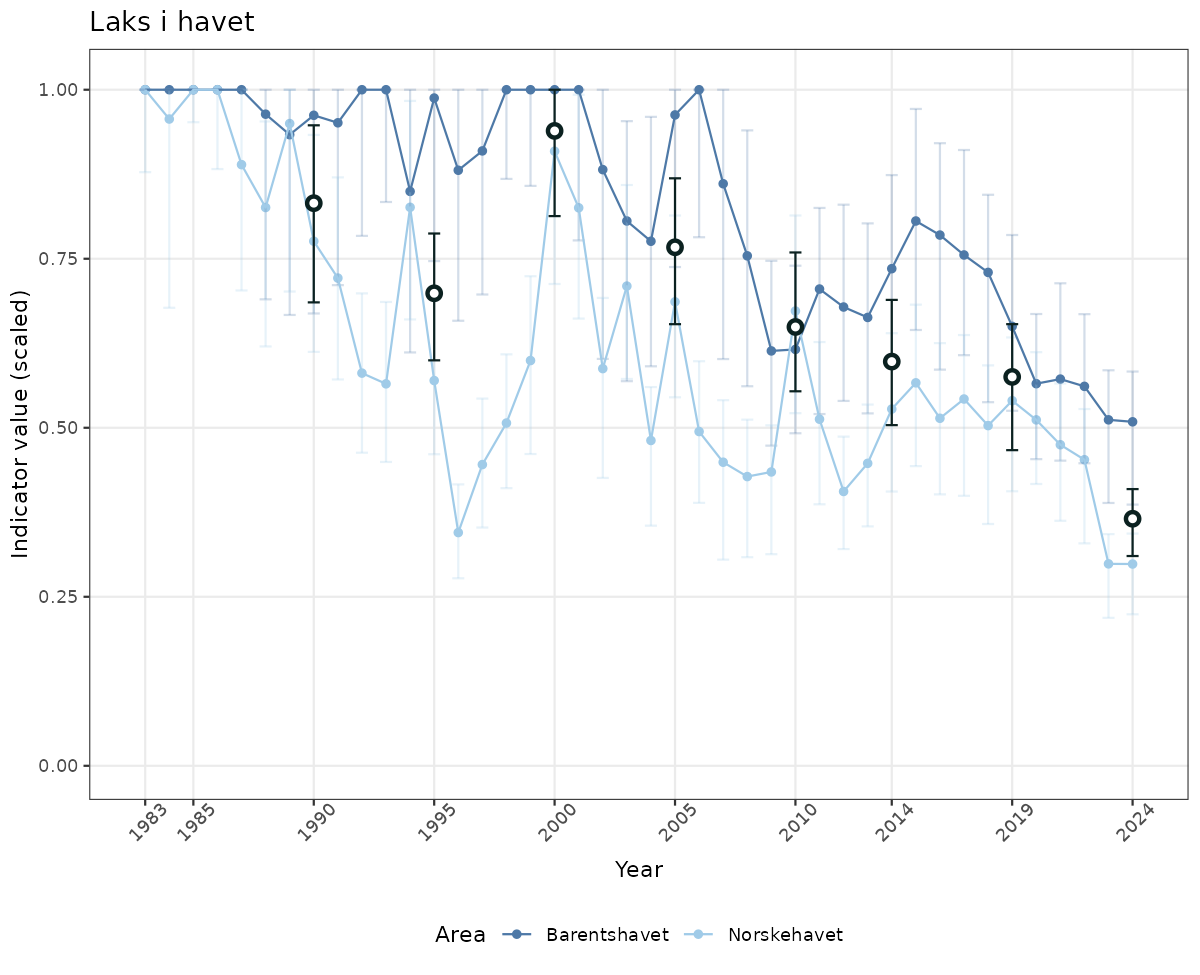
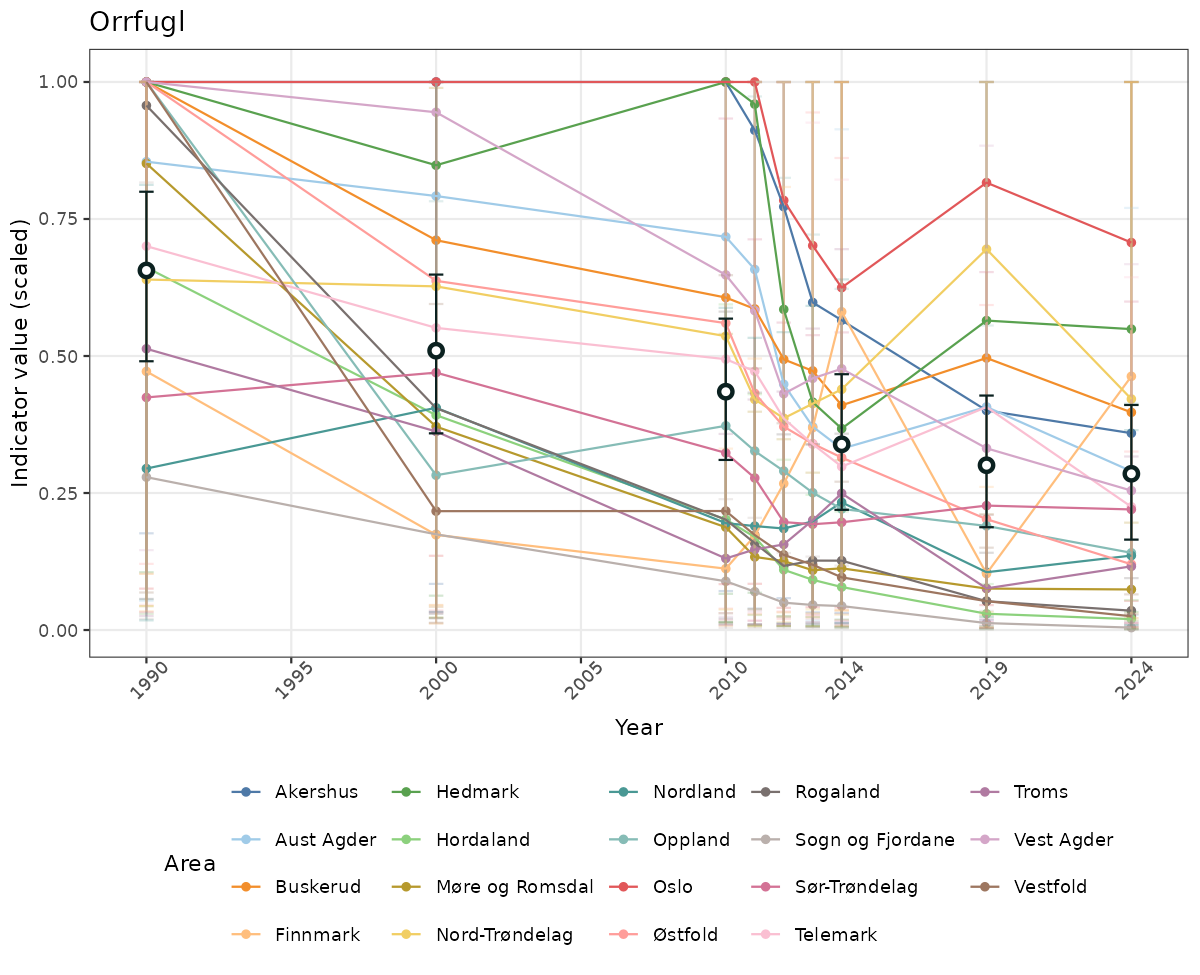
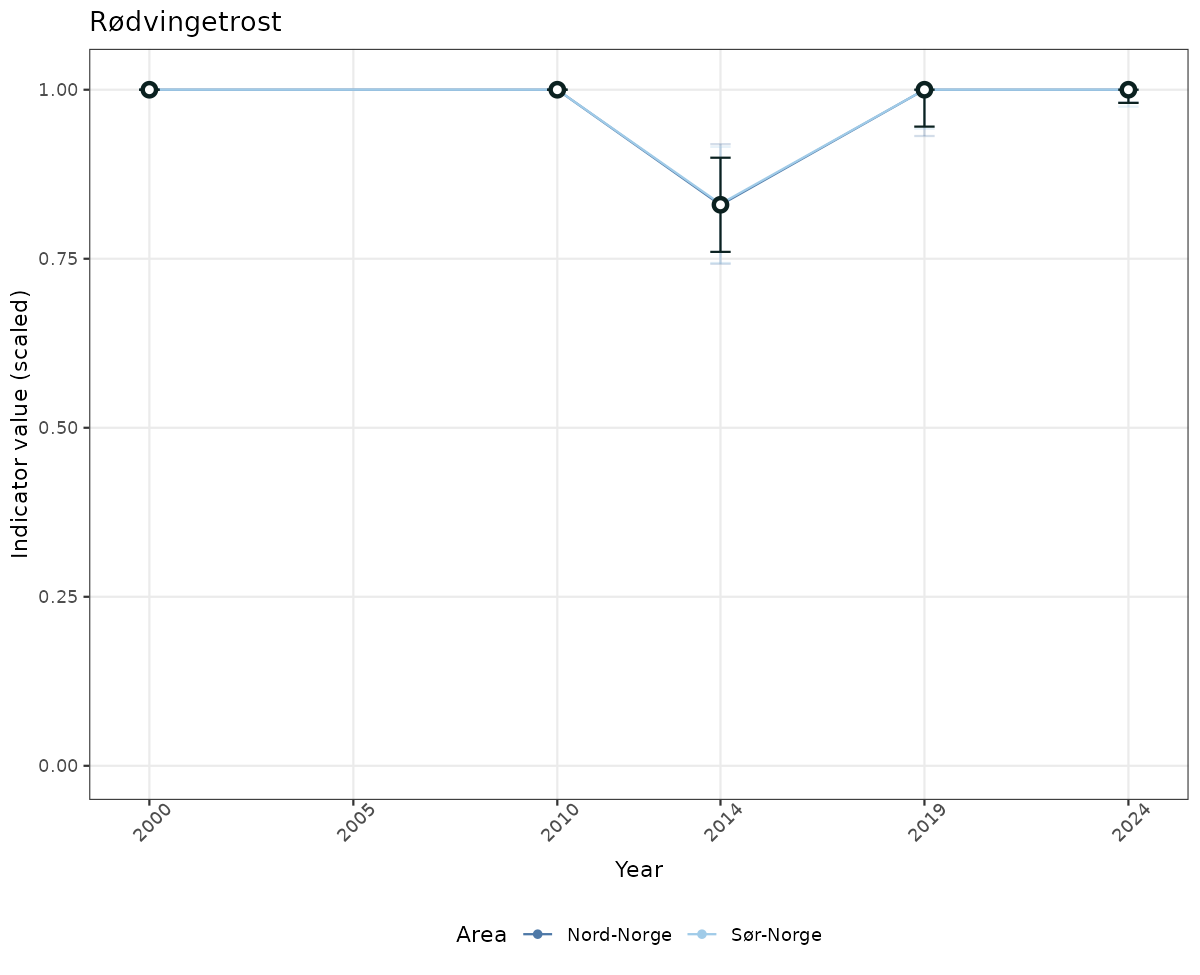
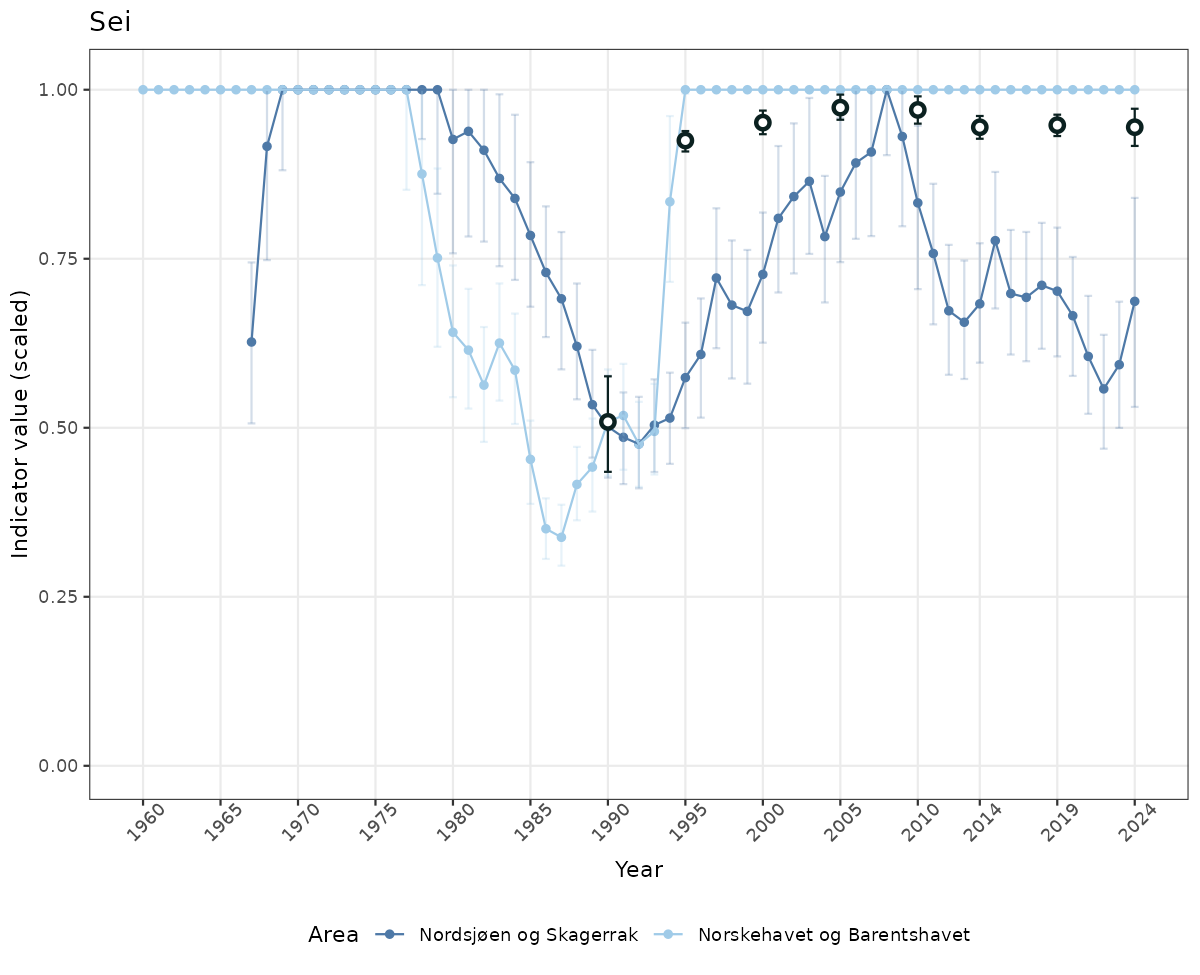
- The defaults for visualization of scaled indicator values are using uncertainty statistics truncated after summarising, and treating the calculations based on fixed references as the baseline. This can be changed using the toggles
truncAfterSummaryanduncertainRefValueat the start of this script. - Visualizations will be partially displayed in this document, and also saved as files.
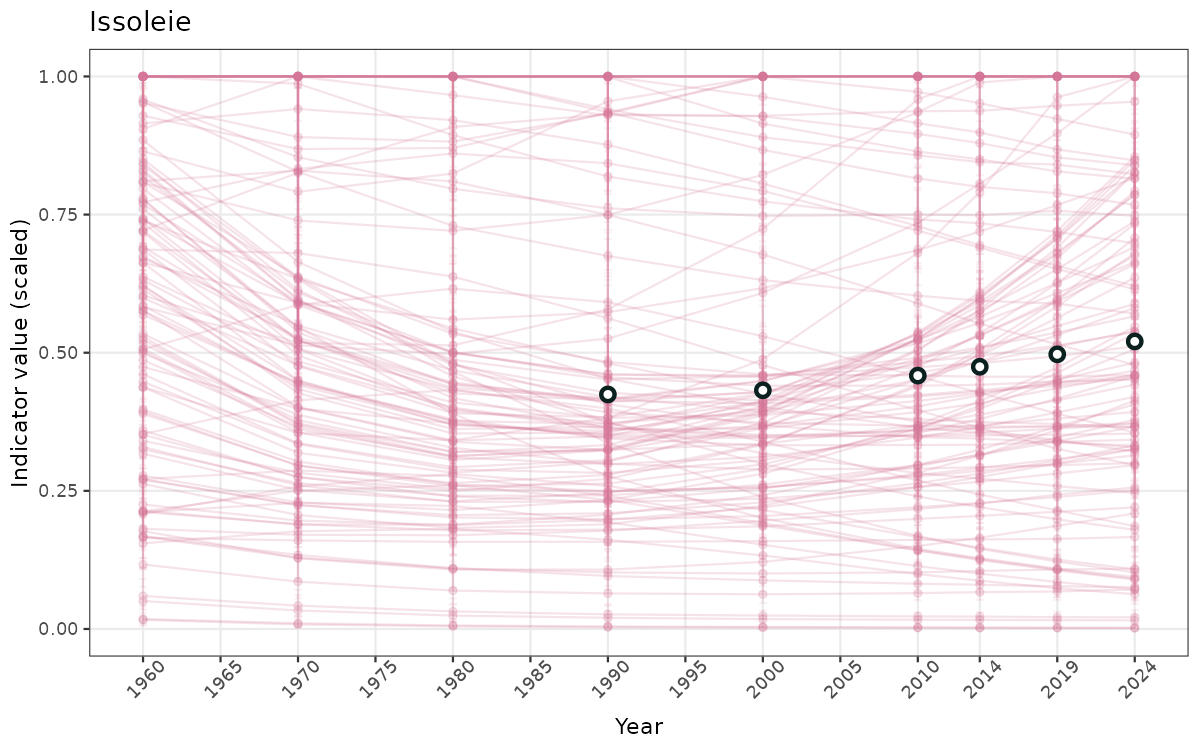
- In this document, we will display 1. featuring all areas. We will not draw and display maps here as these will be part of the presentation of data and results on the new NI webpage.
- For now, the visualizations are static. The goal for the website is to have interactive elements in the figures, including: menus to toggle on and off different time series (and reference levels), “sliders” to change year for maps, simultaneous highlights in median and uncertainty maps, etc.
- Visualizations of the entire value distributions (see under 5.1. here: https://ninanor.github.io/NIviz/other-figures.html) will not be done for individual indicators (only for the aggregated NI for the different ecosystems).
First, we set up directories, determine which scaled values file to use, define a plot color to use in cases where we have time series for many areas, and set up lists to store the plots in R:
## Set up folder for plots (if not present already)
if(!dir.exists("plots")){
dir.create("plots")
}
## Select correct file for scaled data
filename_scaled <- dplyr::case_when(
!uncertainRefValue & truncAfterSummary ~ "fixedRef",
!uncertainRefValue & !truncAfterSummary ~ "fixedRef_truncPreSum",
uncertainRefValue & truncAfterSummary ~ "sampledRef",
uncertainRefValue & !truncAfterSummary ~ "sampledRef_truncPreSum",
)
## Set fixed plotting colors
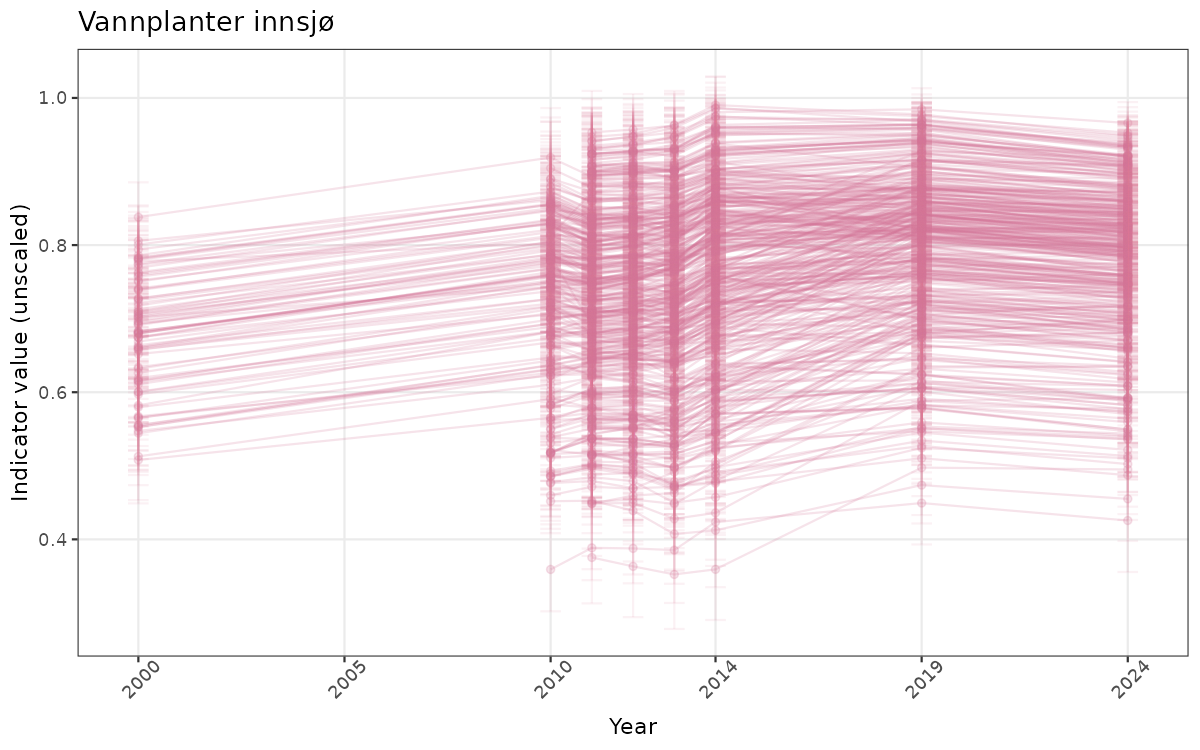
# Time-series plot color for indicators with > 20 areas
plot.col <- "#D37295FF"
# Time-series plot color for indicator indices
plot.col2 <- "#0B2120"
## Make lists to store plots for each indicator
TS_raw <- list()
TS_scaled <- list()
TS_indIdx <- list()
TS_indIdx_std <- list()Then, we proceed to plot time series and save plots to files:
for(i in 1:nrow(indicatorList)){
## Extract indicator name and id
ind_id <- indicatorList$id[i]
ind_name <- indicatorList$name[i]
message(crayon::bold(crayon::green(paste0(ind_name))))
## Make subfolder for species in target directory & set save path
if(!dir.exists(paste0("plots/id_", ind_id, "_", stringr::str_replace_all(ind_name, " ", "_")))){
dir.create(paste0("plots/id_", ind_id, "_", stringr::str_replace_all(ind_name, " ", "_")))
}
savePath_i <- paste0("plots/id_", ind_id, "_", stringr::str_replace_all(ind_name, " ", "_"))
## Load indicator datasets
message("Processing indicator data...")
data_raw <- readRDS(paste0("data/IndicatorData_StatSummaries/statSummary_id_", ind_id, "_raw.rds")) %>%
dplyr::mutate(year_t = as.integer(year))
data_scaled <- readRDS(paste0("data/IndicatorData_StatSummaries/statSummary_id_", ind_id, "_scaled_", filename_scaled, ".rds")) %>%
dplyr::mutate(year_t = as.integer(year))
output_indIdx <- readRDS(paste0("results_Indicators/indIndex_id_", ind_id, ".rds"))
data_indIdx <- output_indIdx$summary %>%
dplyr::filter(indexArea == "wholeArea", year_t %in% ind_focalYears & !data_singleArea) %>%
dplyr::mutate(year_t = as.integer(year_t))
## Truncate scaled values (if necessary)
data_rows <- c("mean", "median", "q025", "q05", "q25", "q75", "q95", "q975")
data_scaled <- data_scaled %>%
dplyr::mutate(dplyr::across(.cols = all_of(data_rows), ~ ifelse(. > 1, 1, .)))
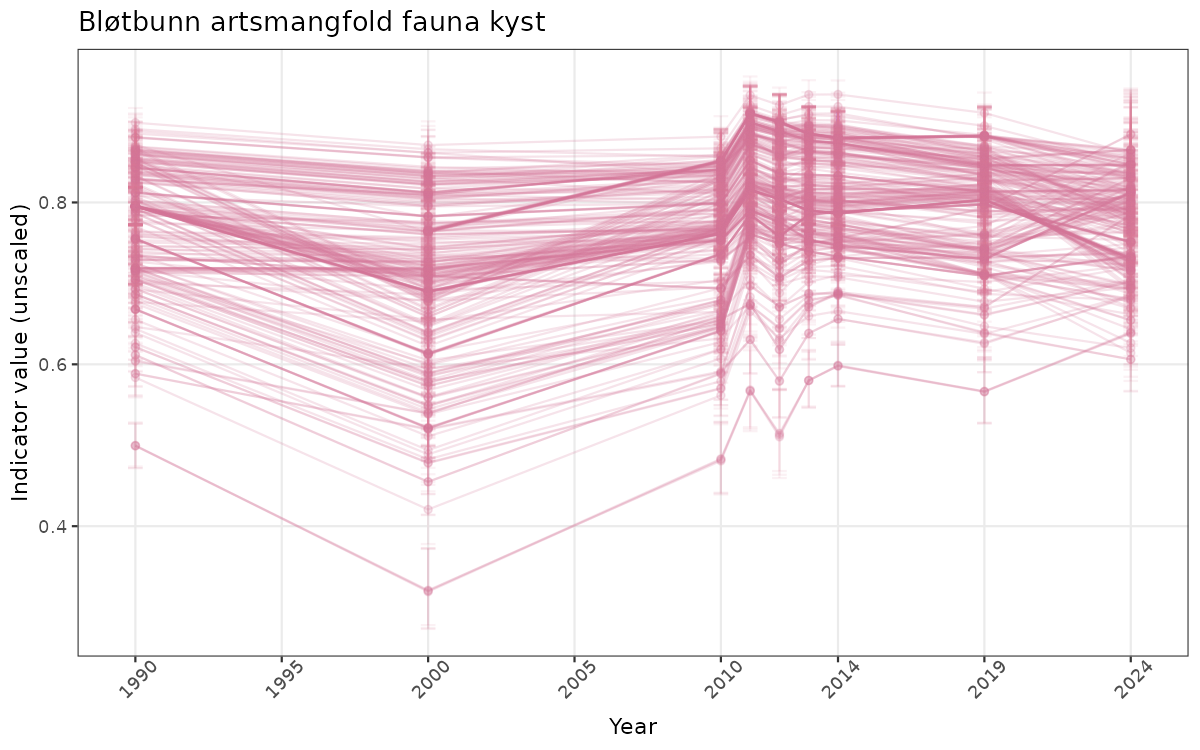
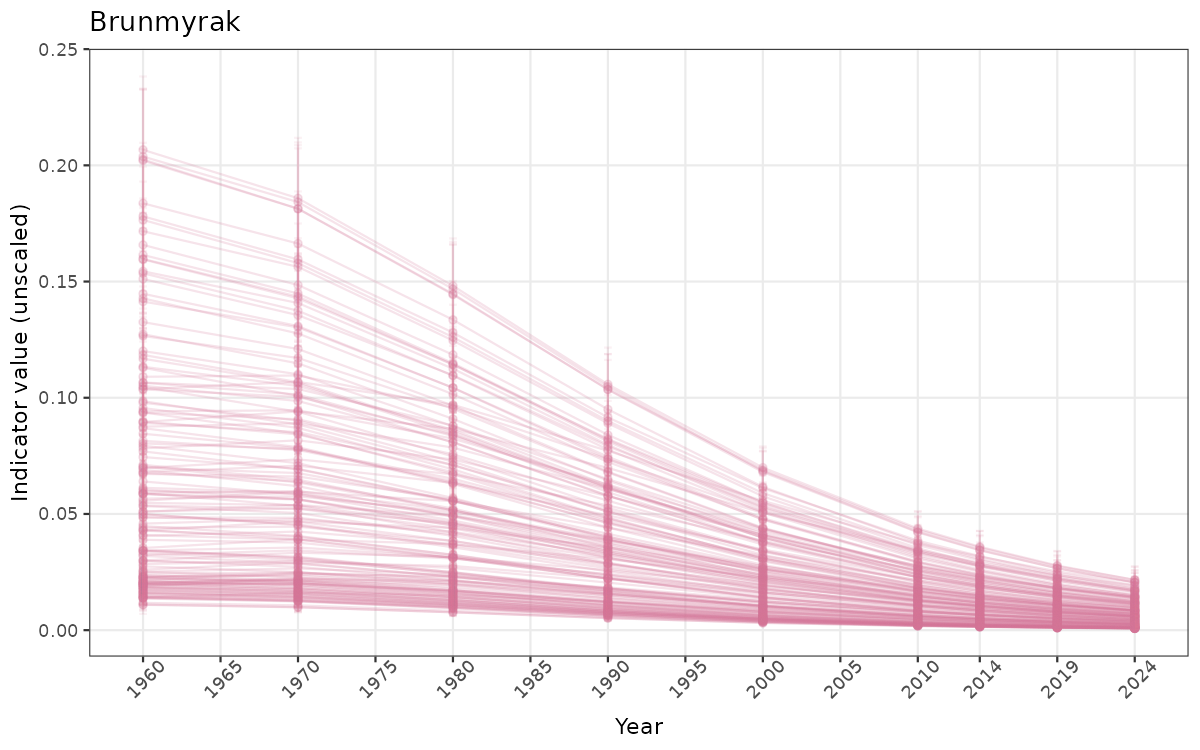
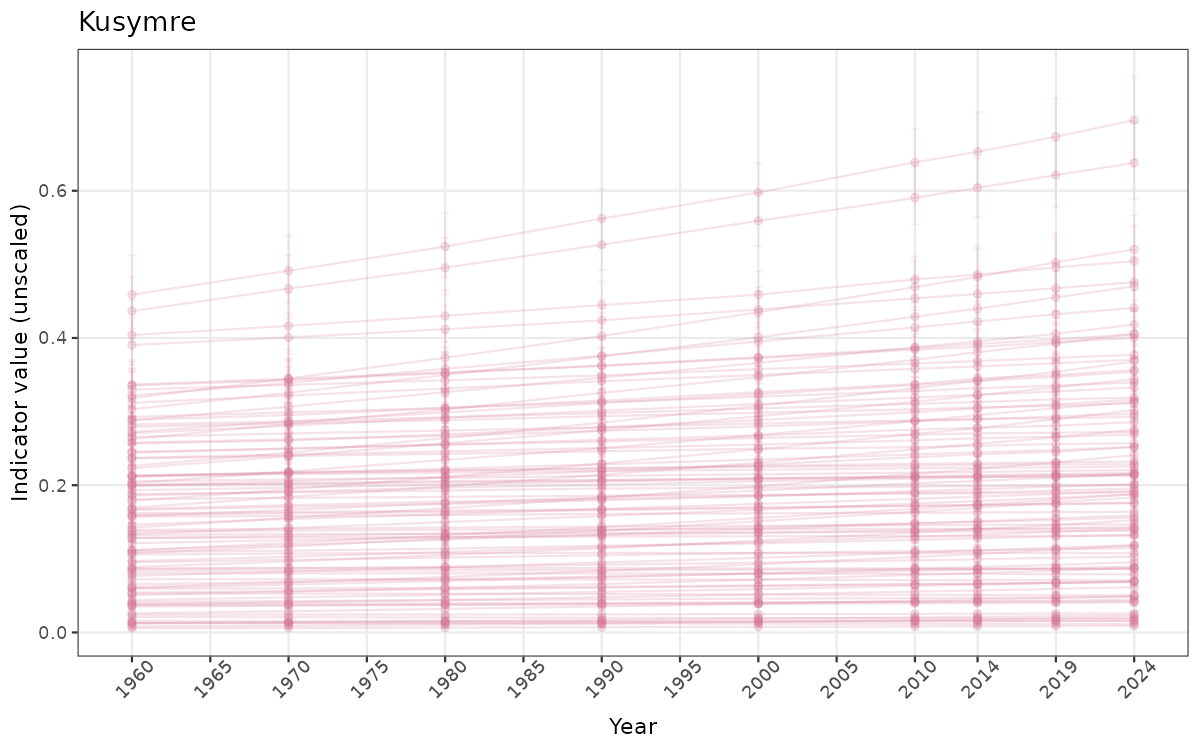
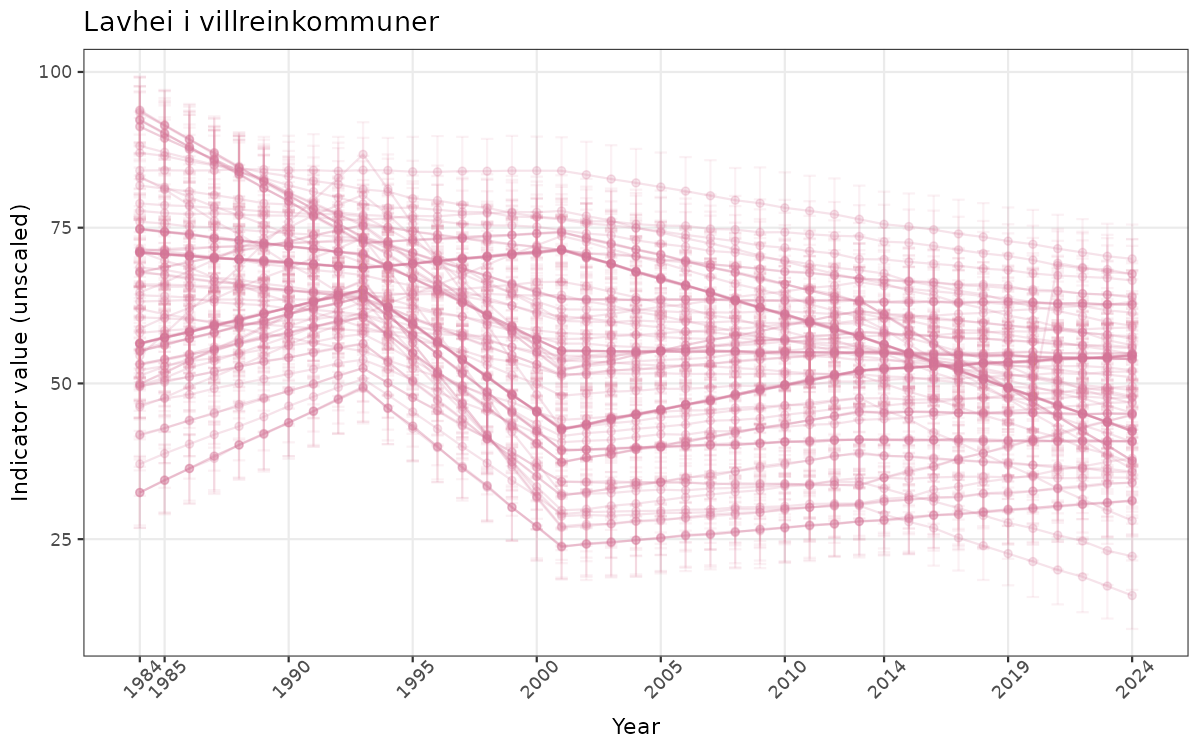
## Count areas for picking time series display
manyAreas <- ifelse(length(unique(data_raw$ICunitName)) > 20, TRUE, FALSE)
## Set years to highlight
minYear <- min(data_raw$year_t)
maxYear <- max(data_raw$year_t)
if(minYear >= ind_focalYears[1]){
plot.yearH <- ind_focalYears[which(ind_focalYears >= minYear)]
}else{
plot.yearH <- unique(c(minYear, rev(seq(from = ind_focalYears[1], to = minYear, by = -5)), ind_focalYears))
}
## Prepare time series plots for each indicator
message("Drawing time-series plots...")
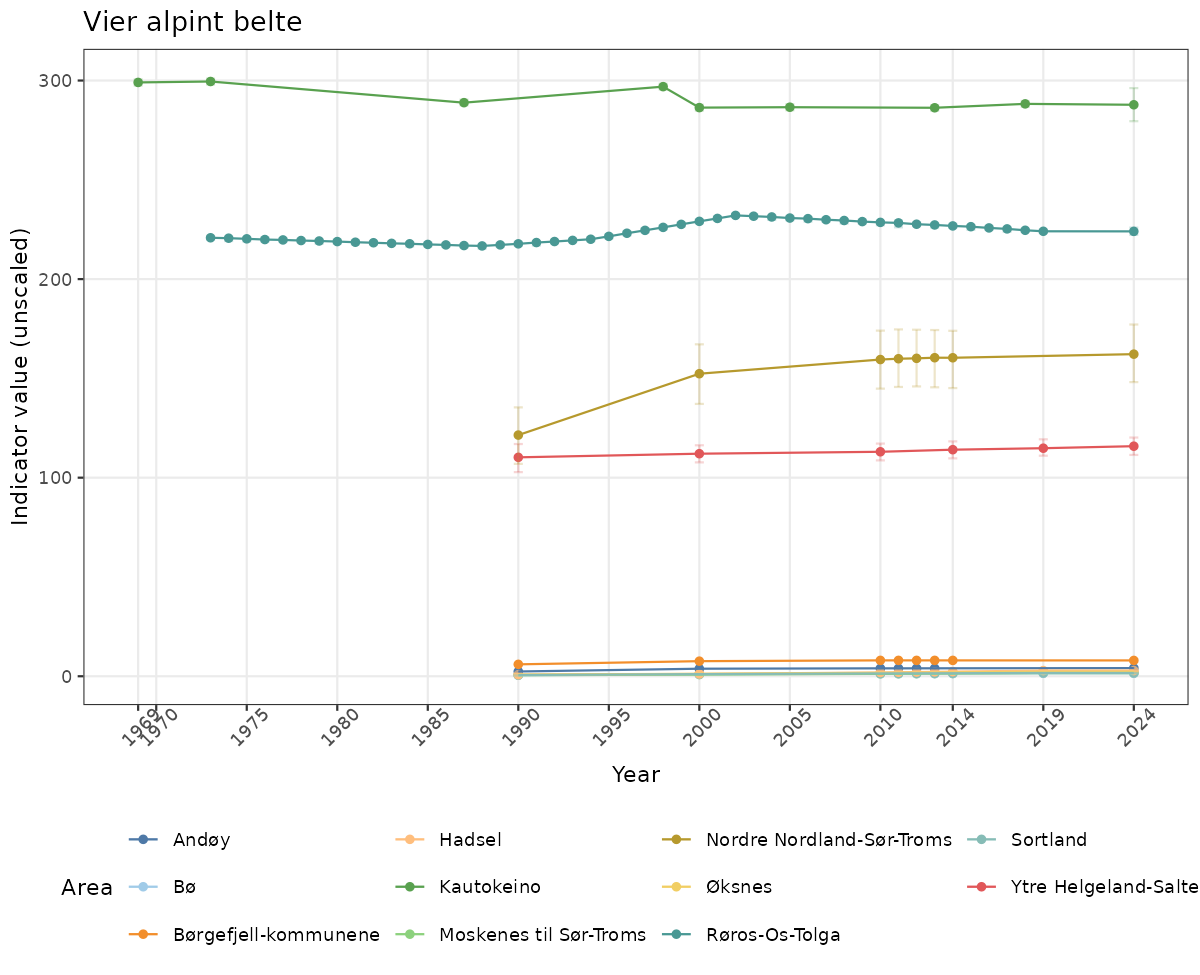
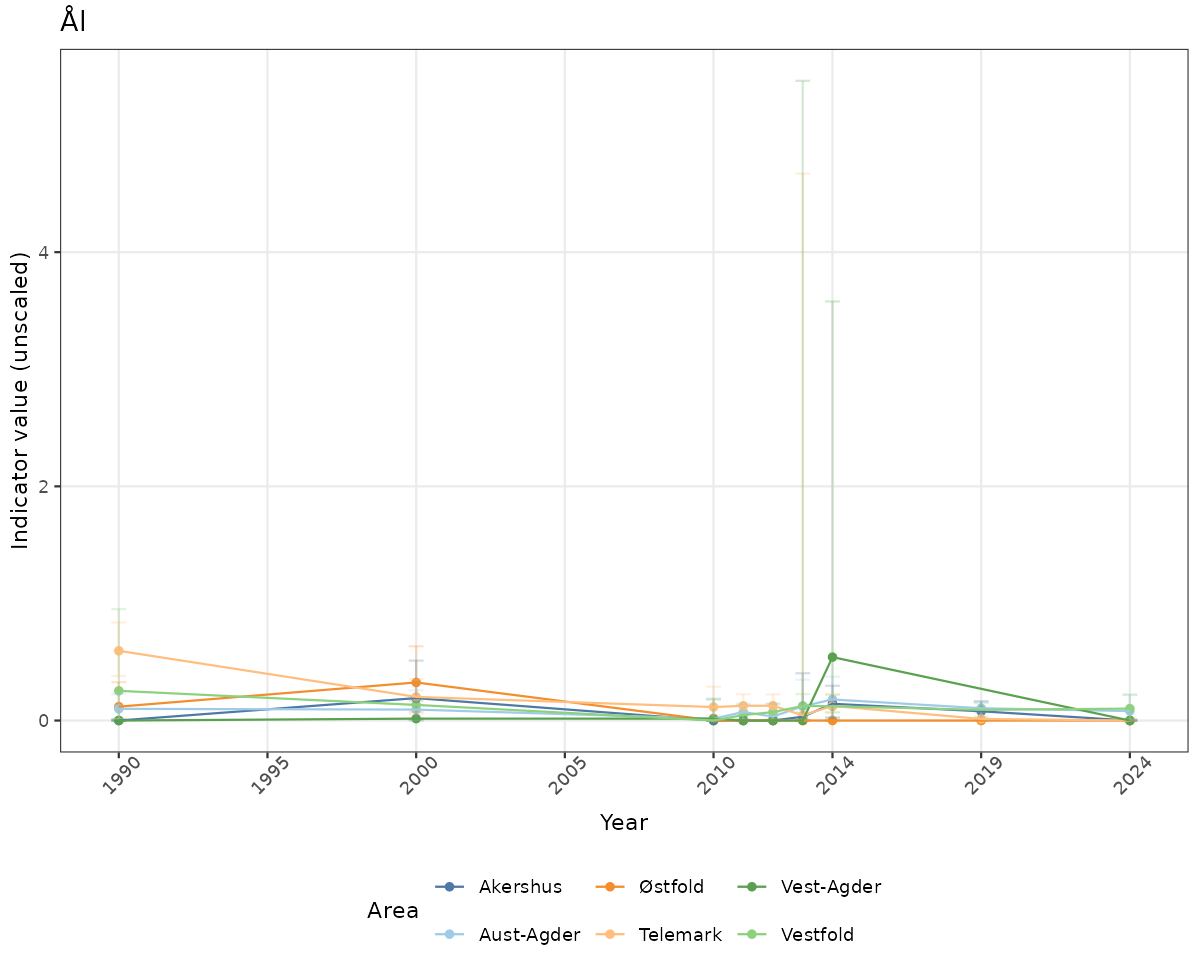
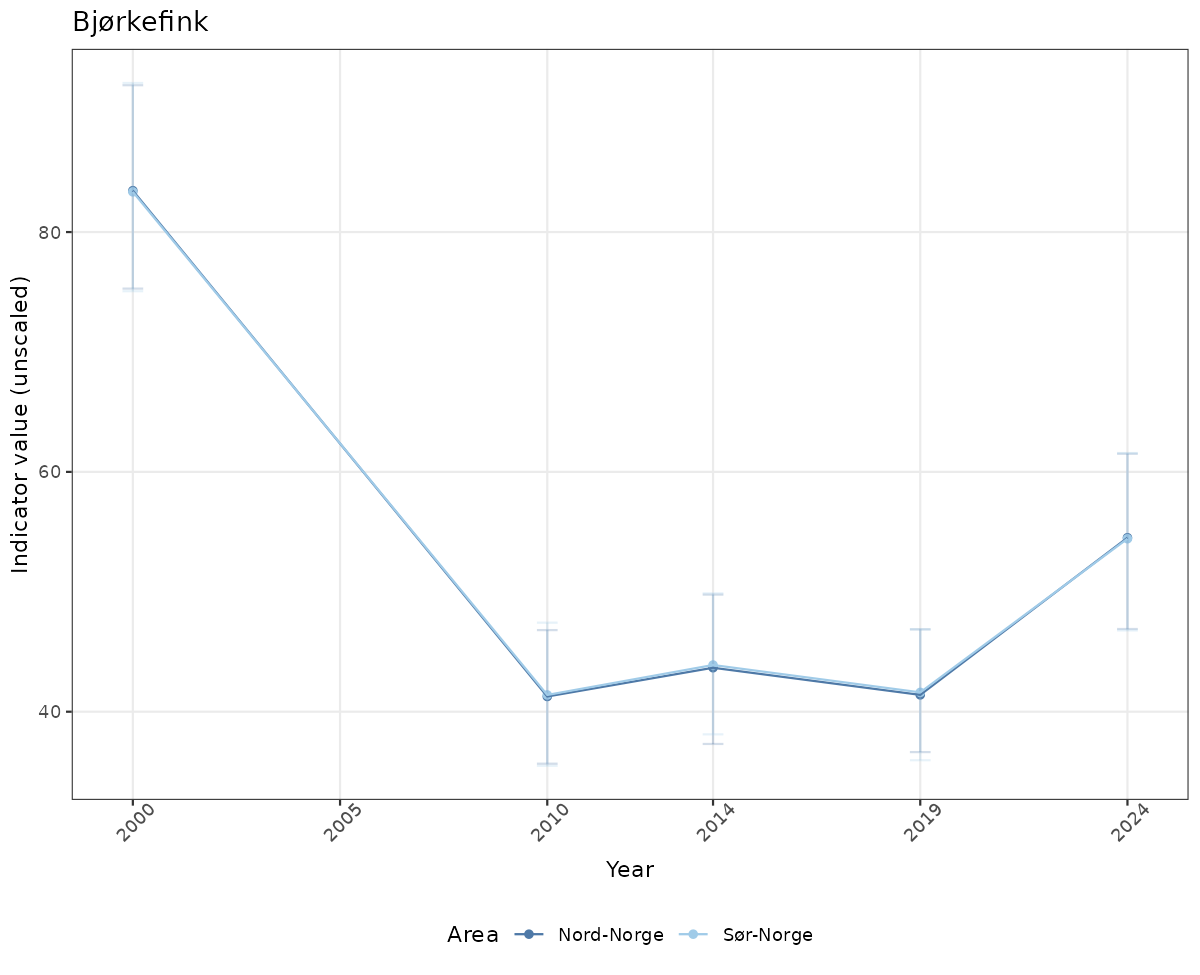
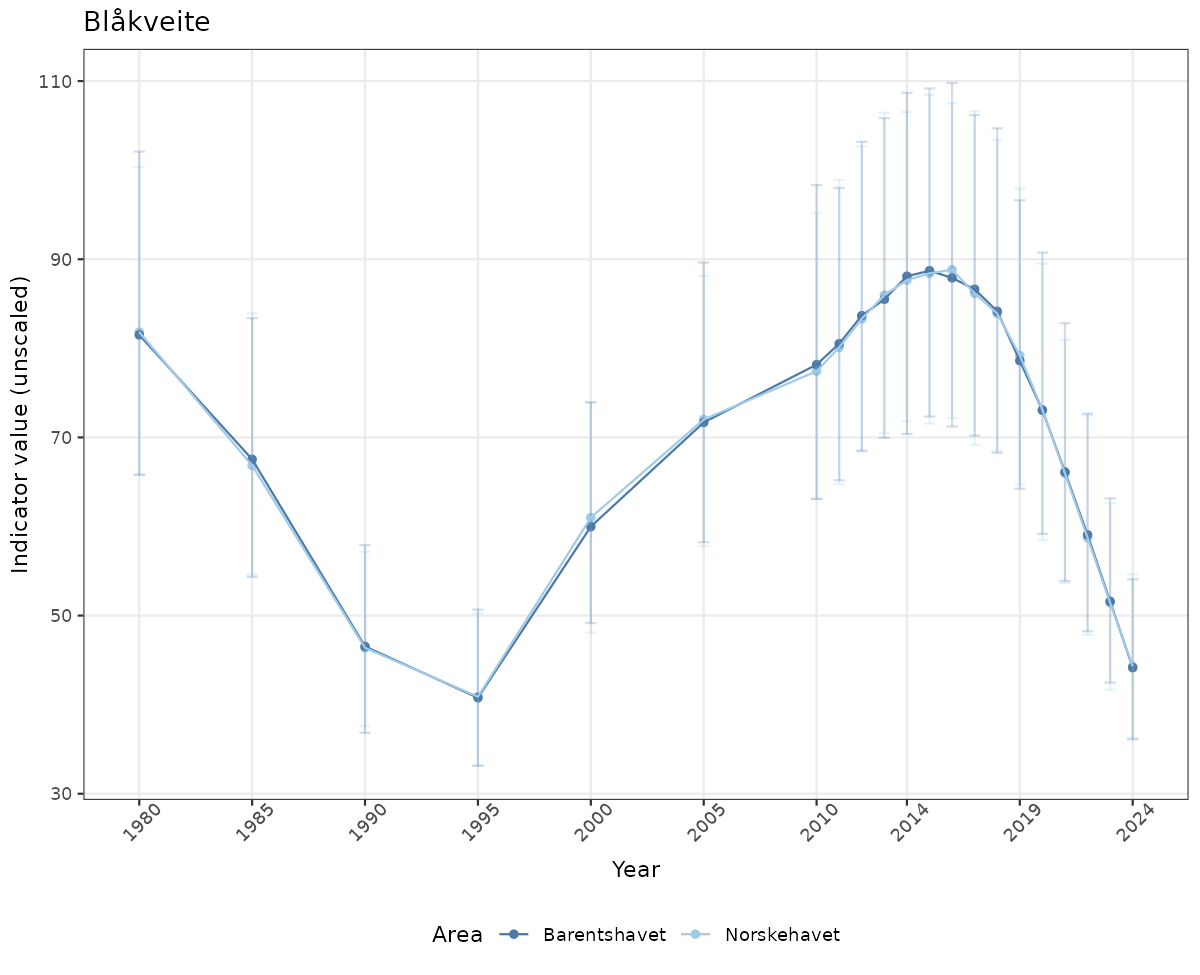
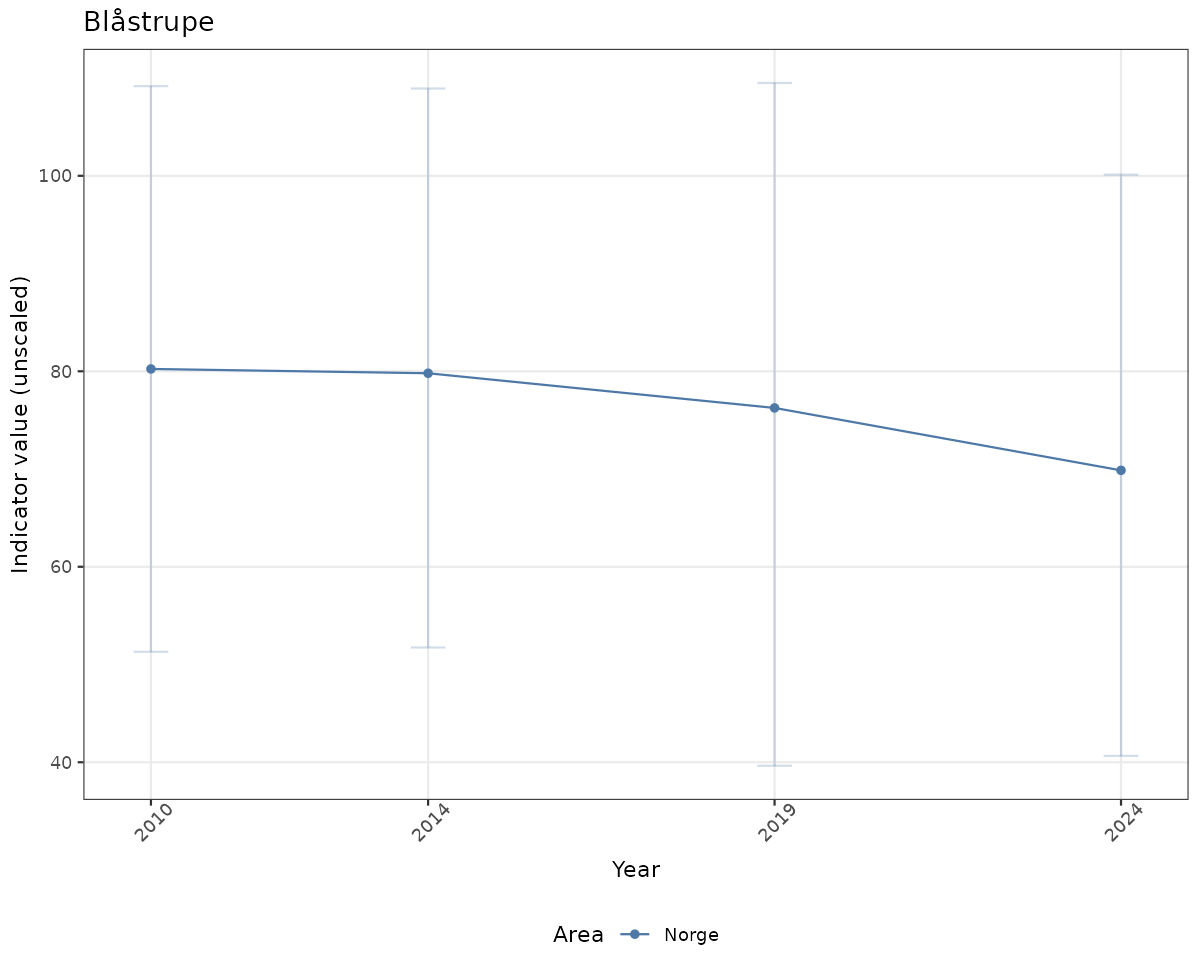
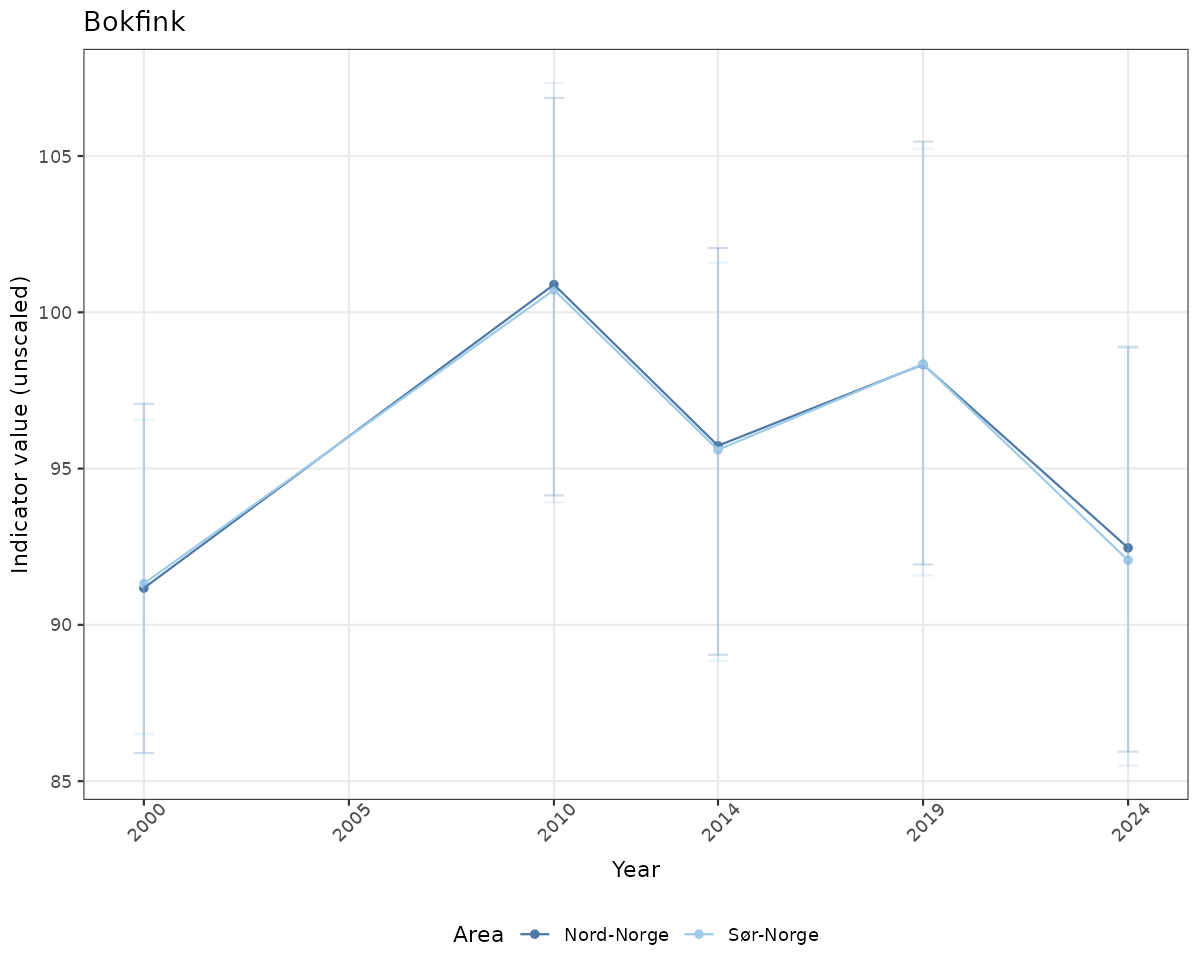
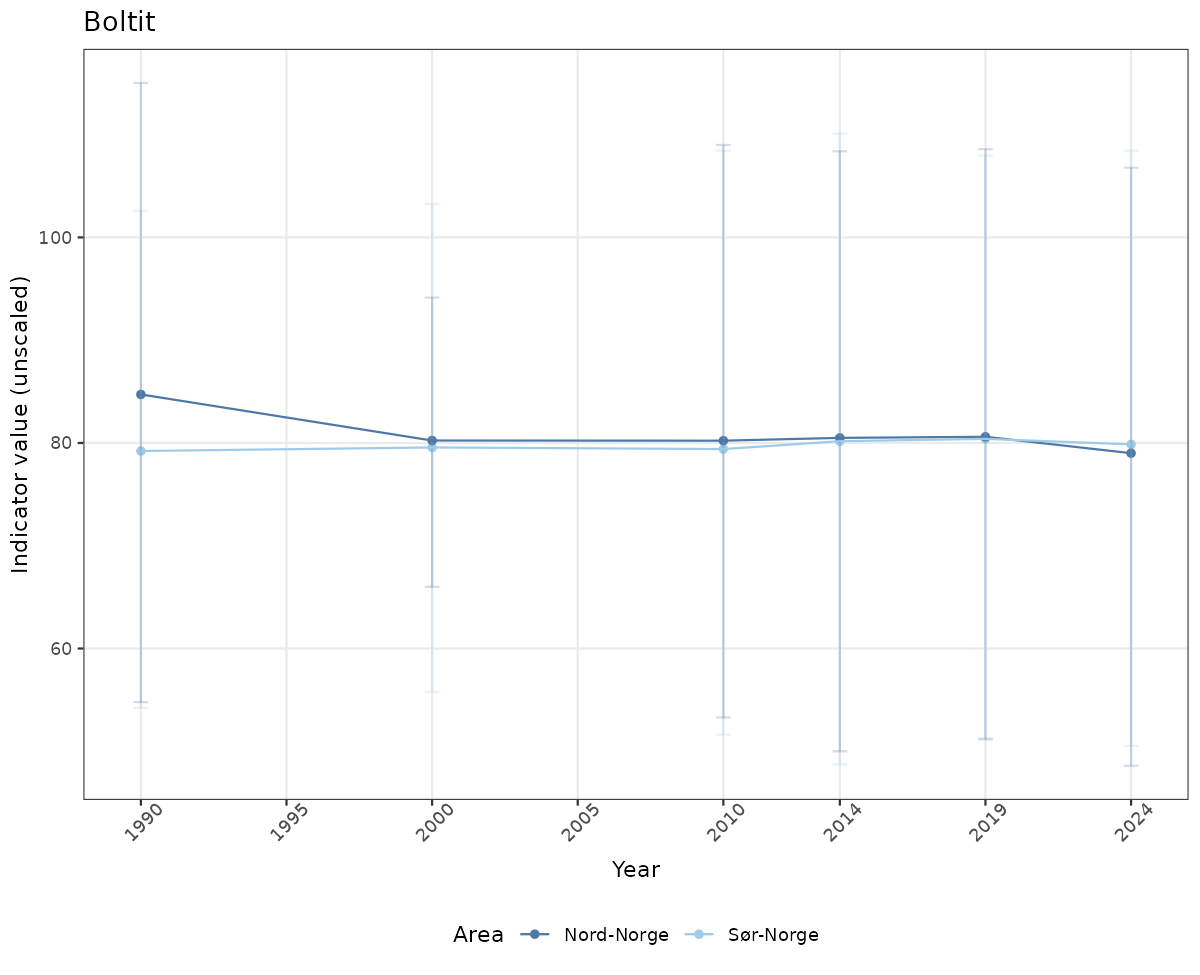
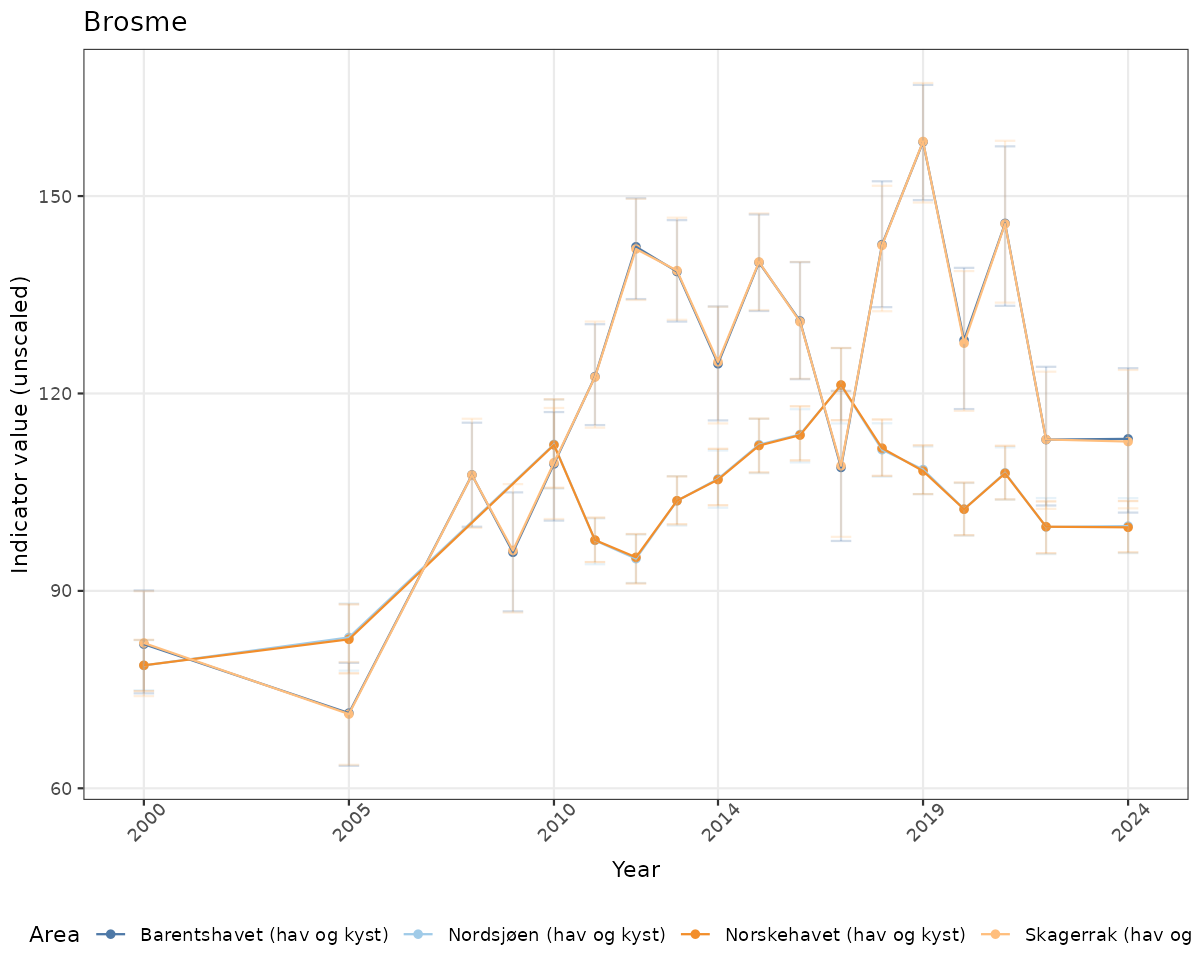
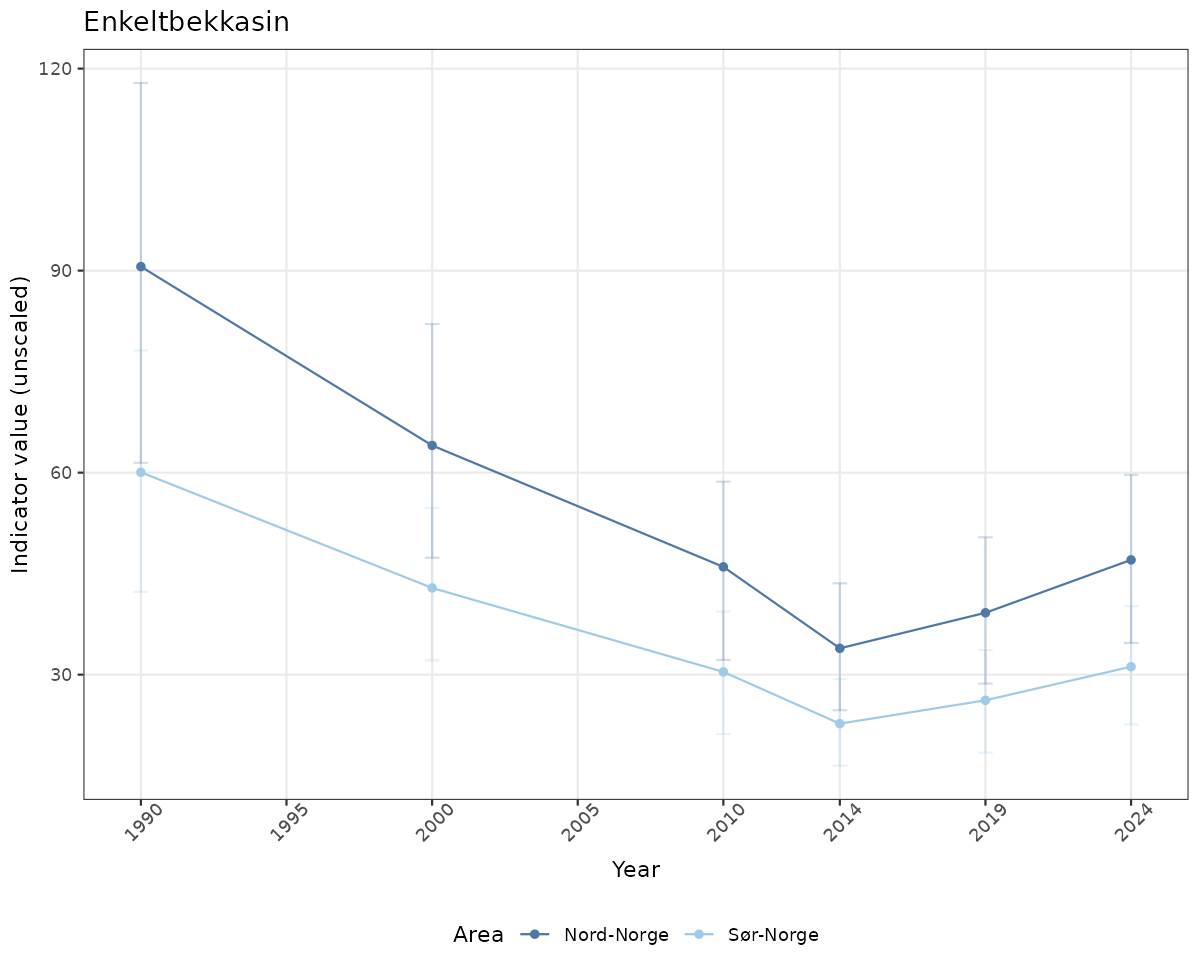
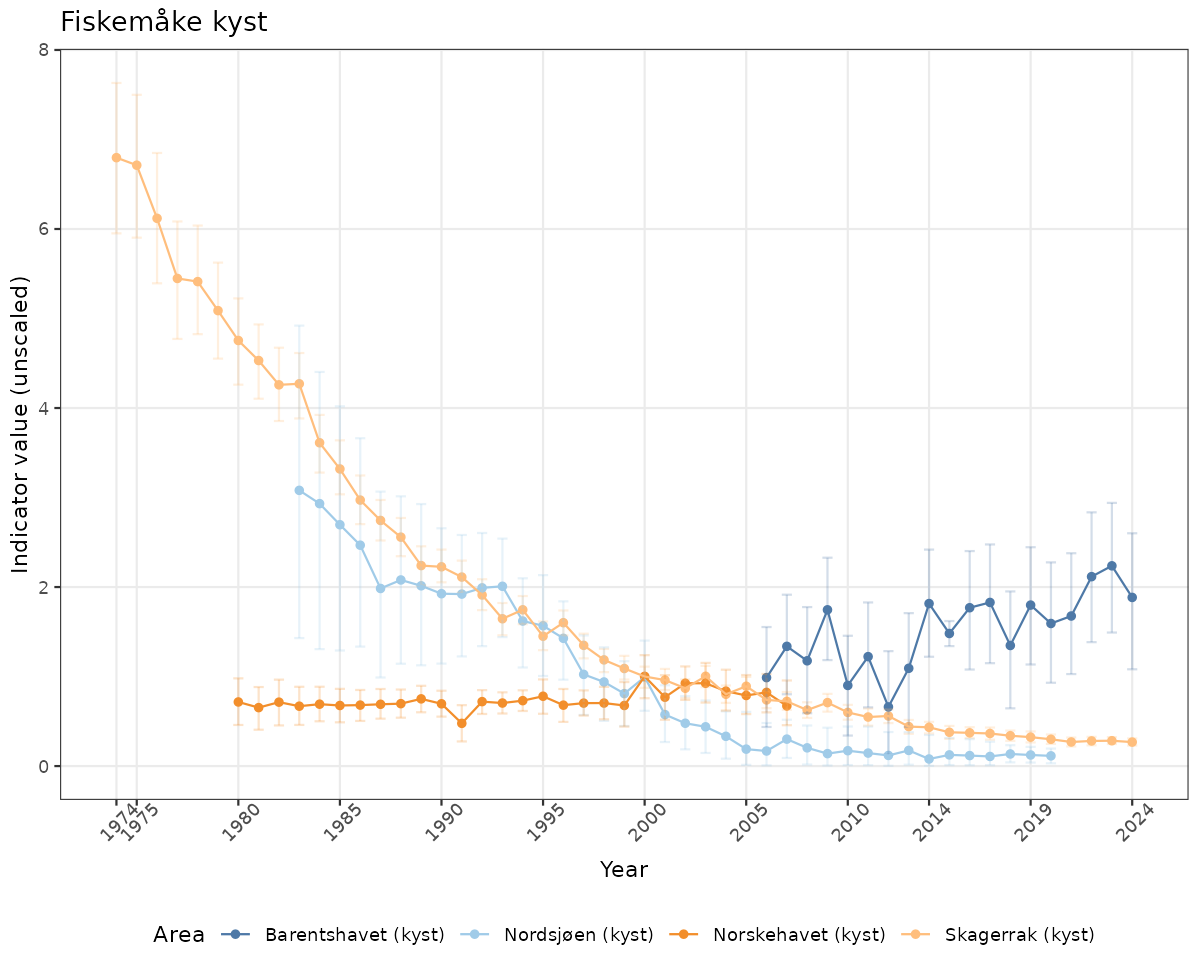
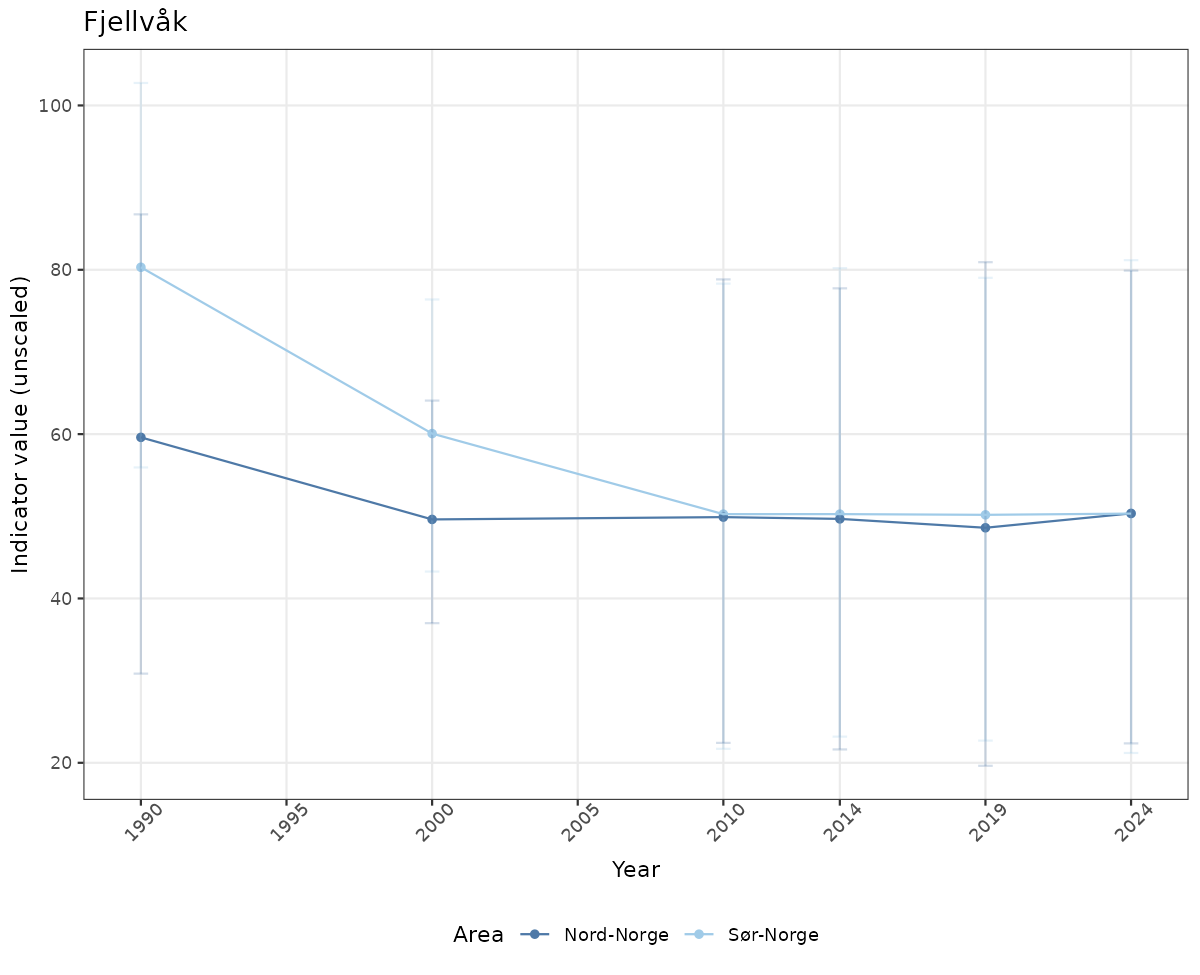
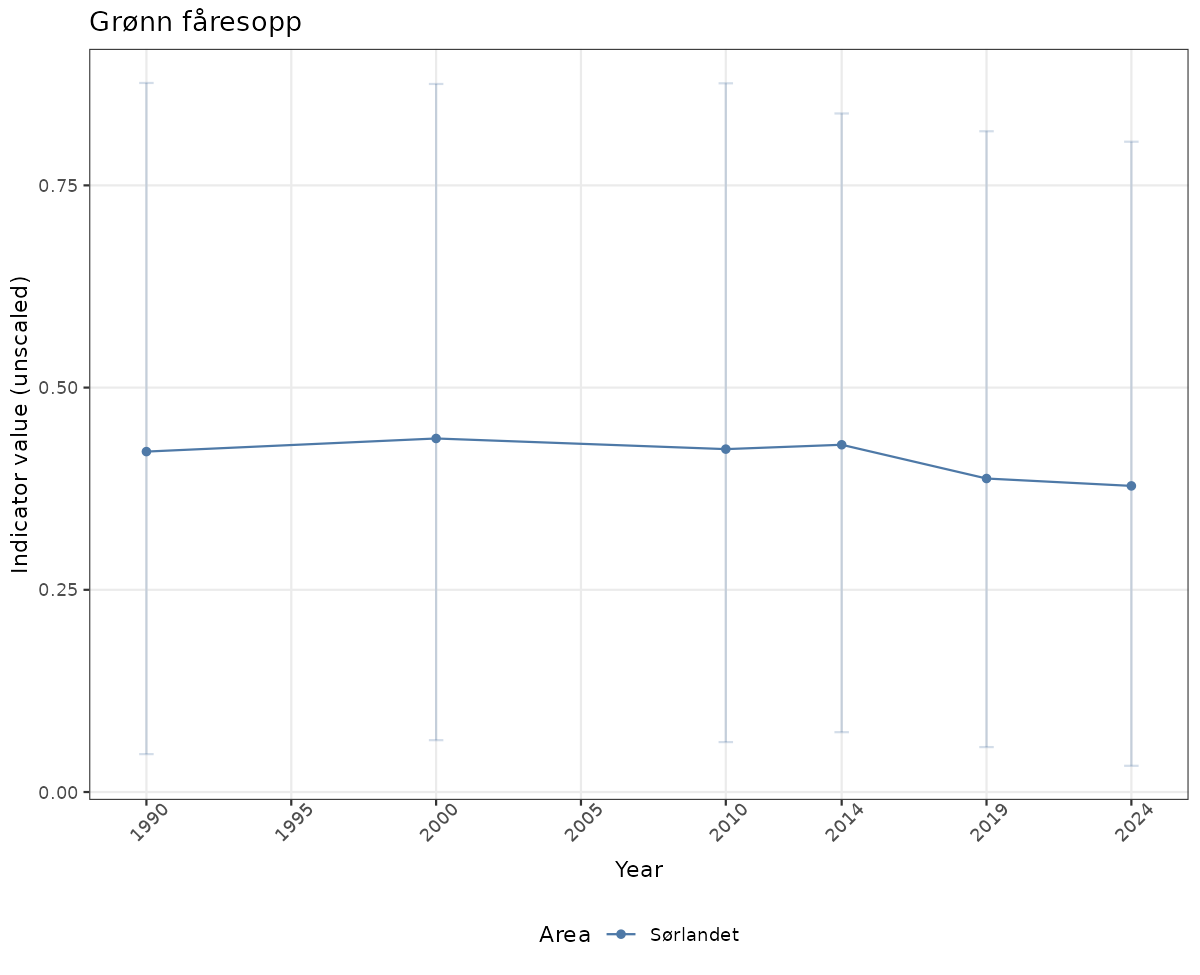
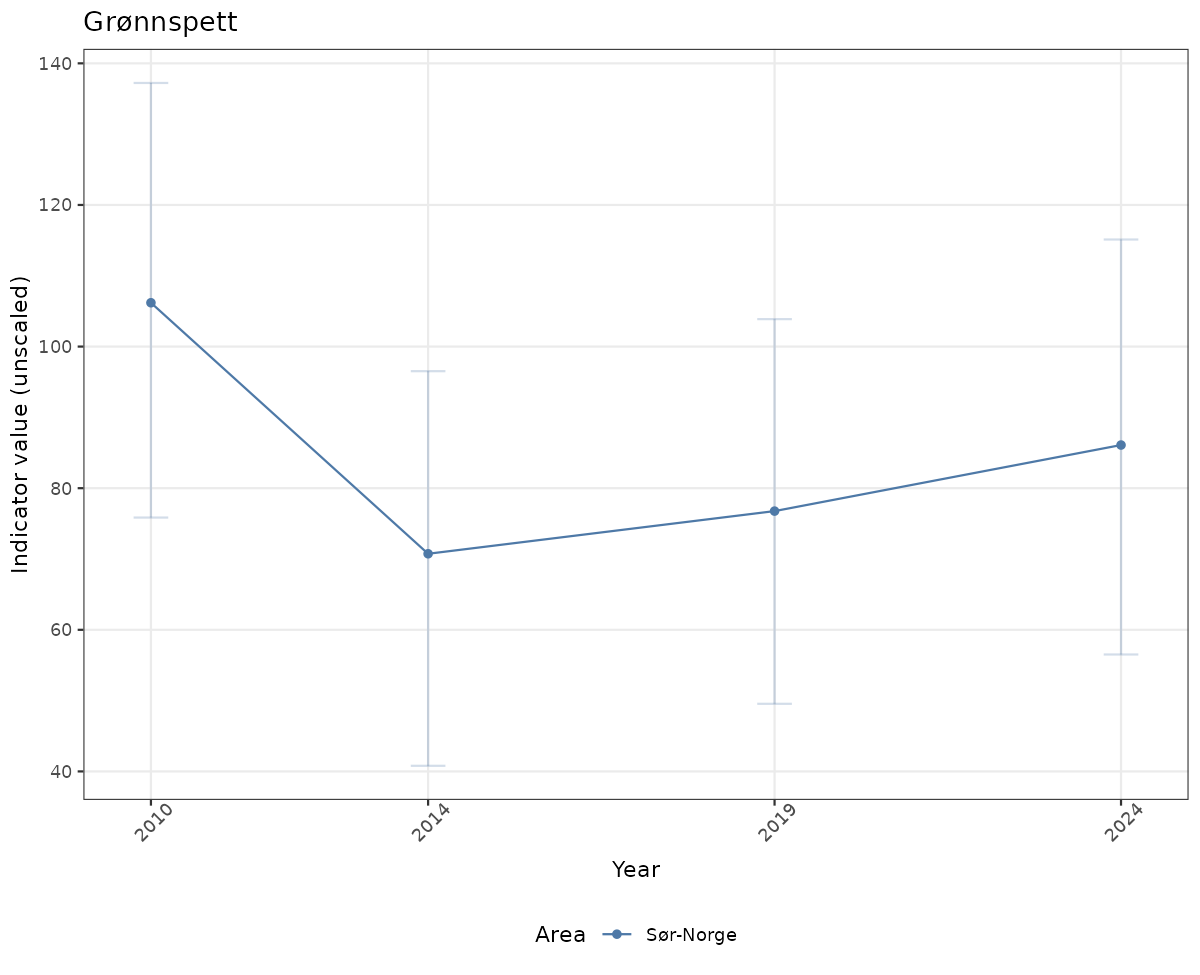
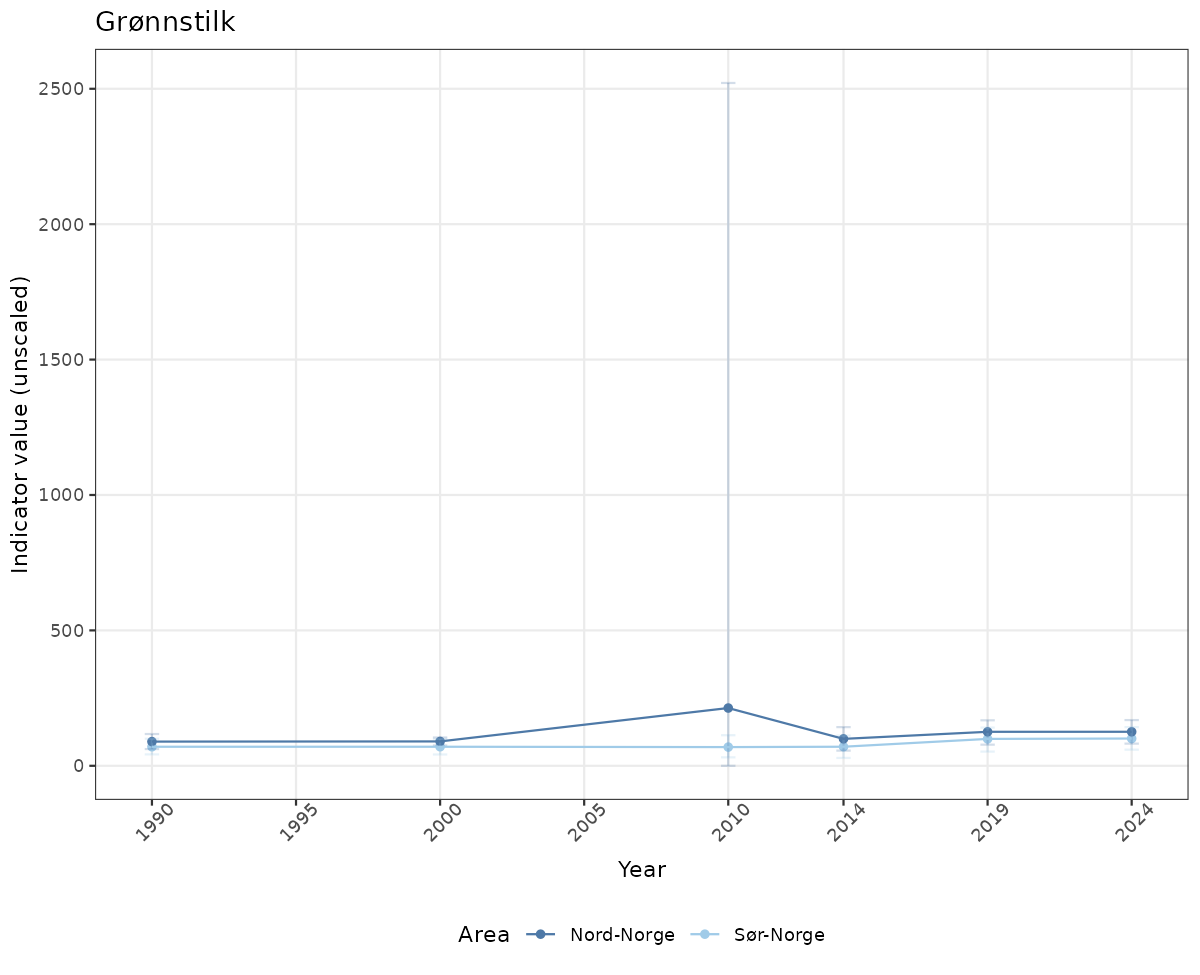
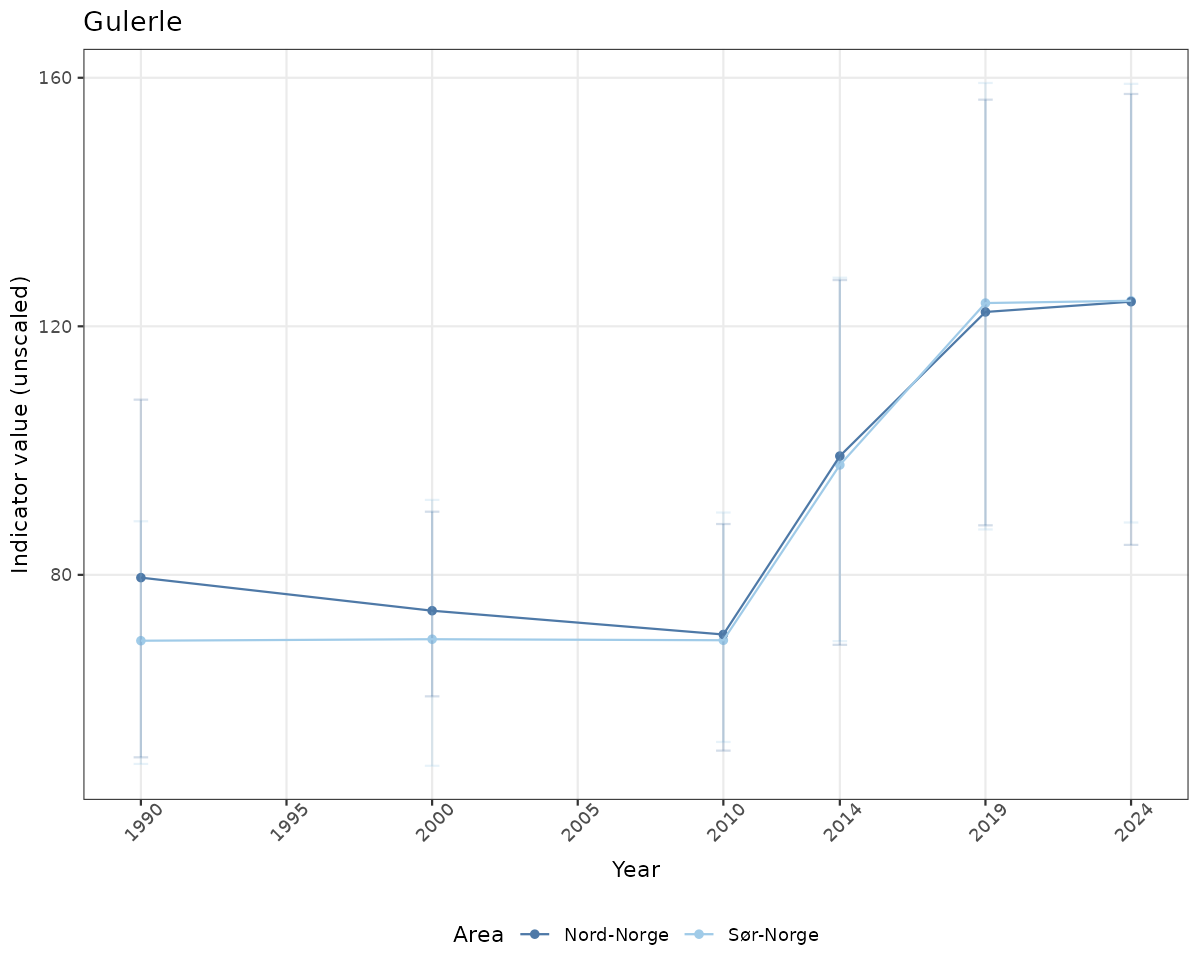
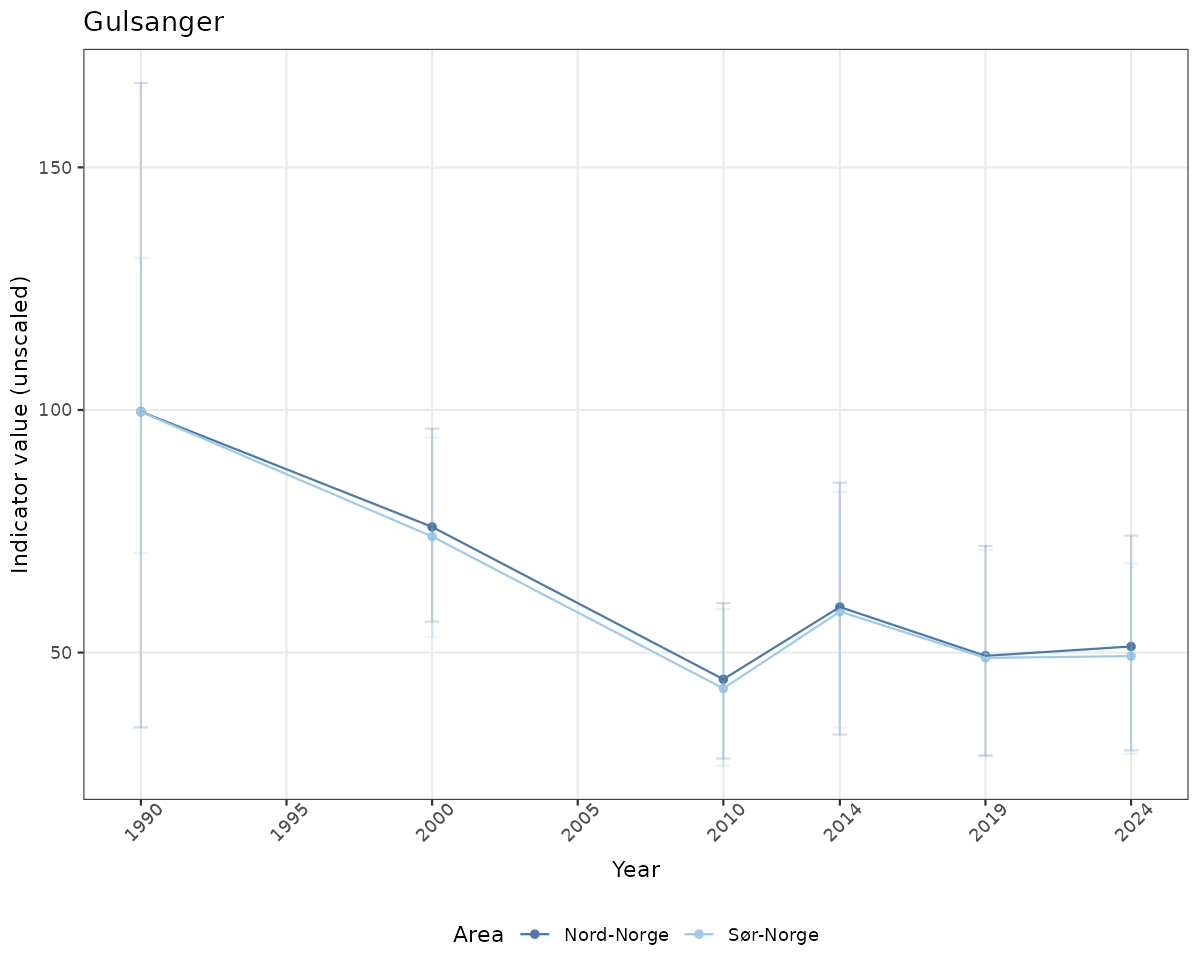
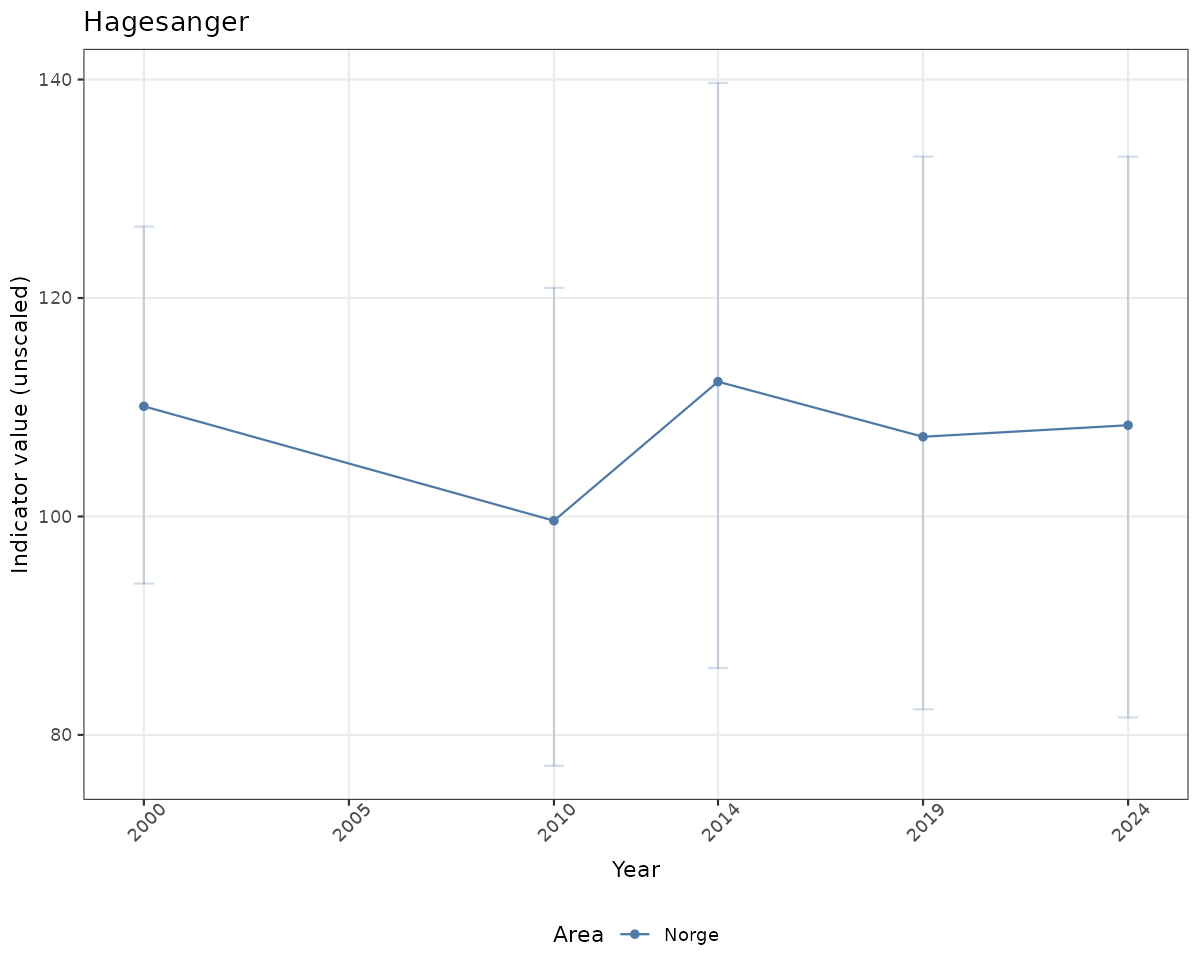
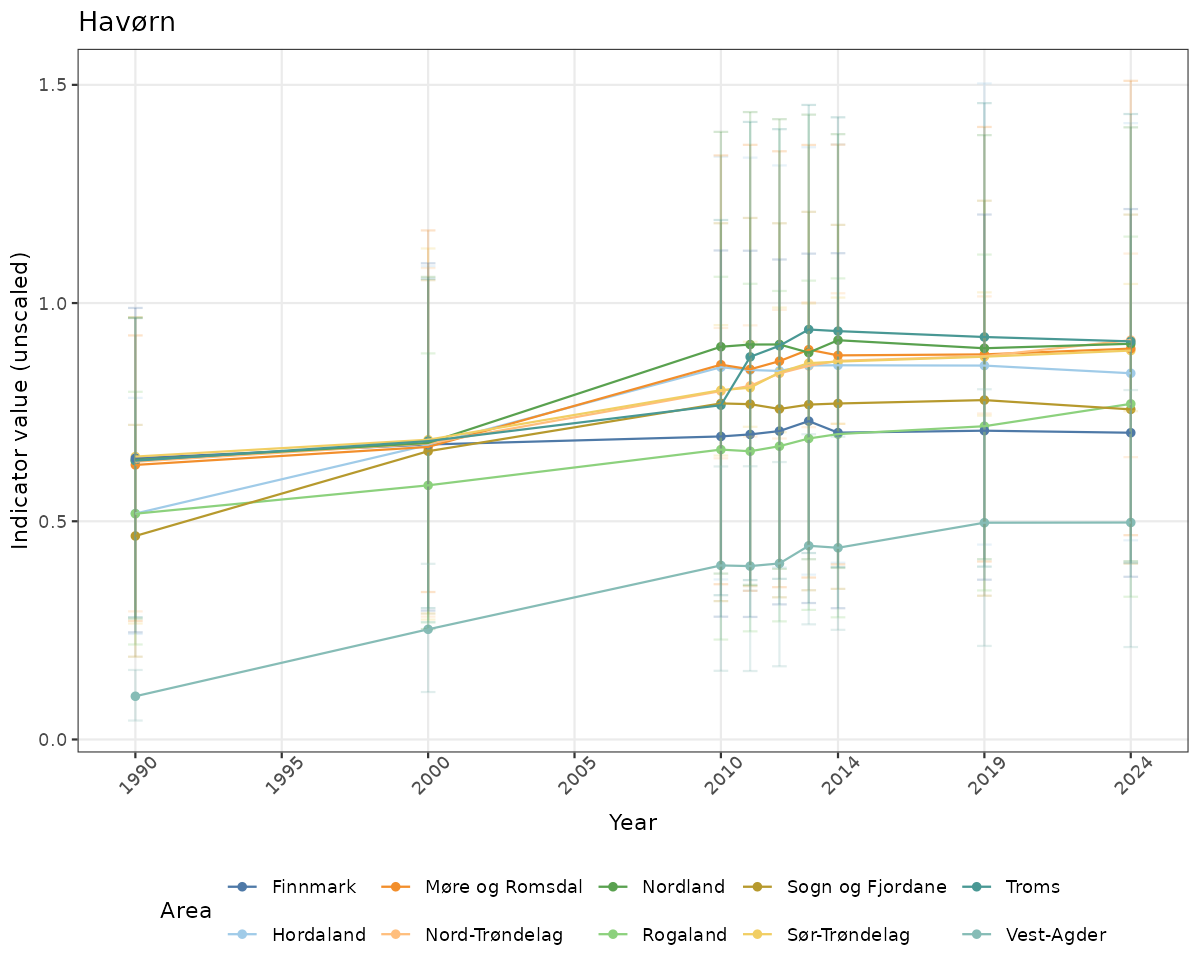
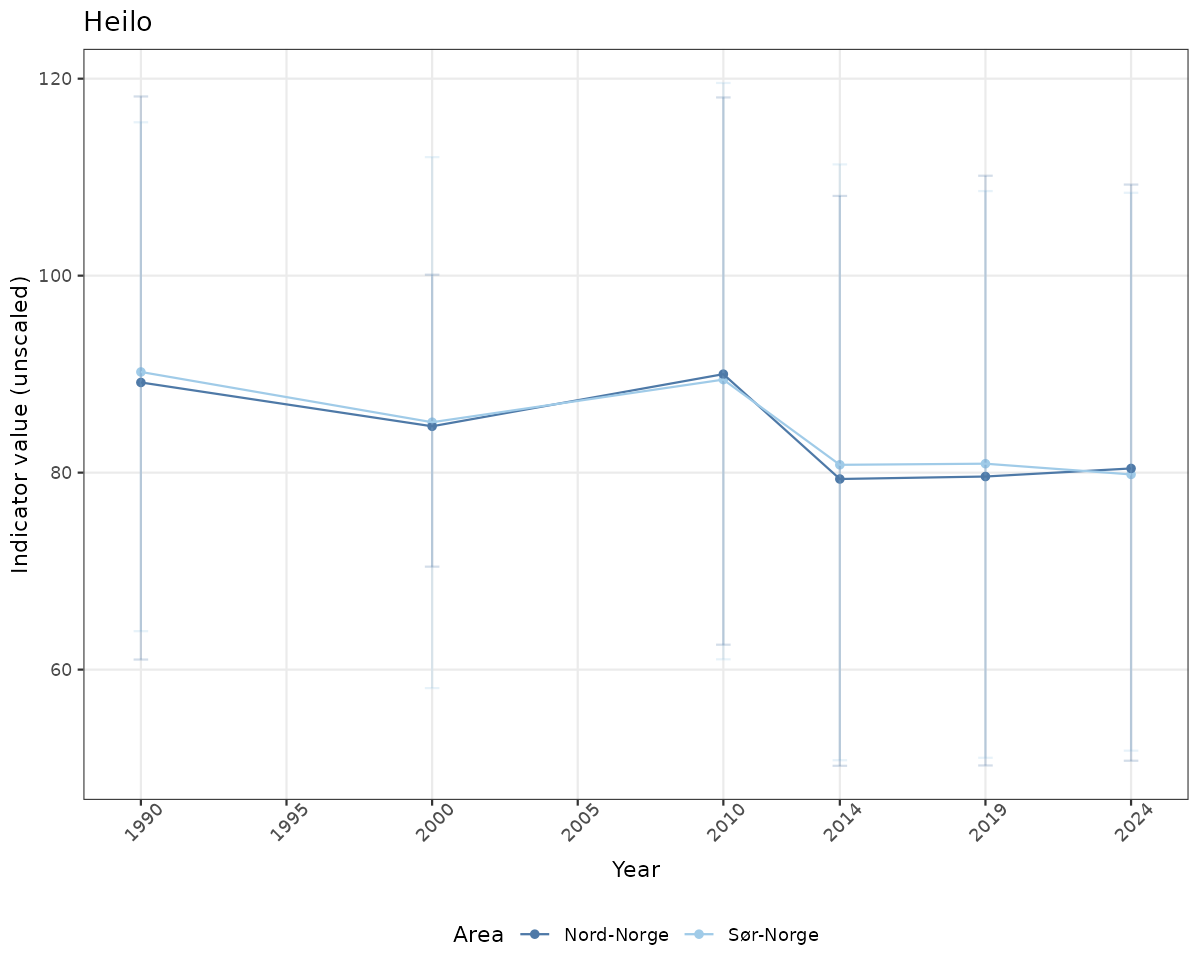
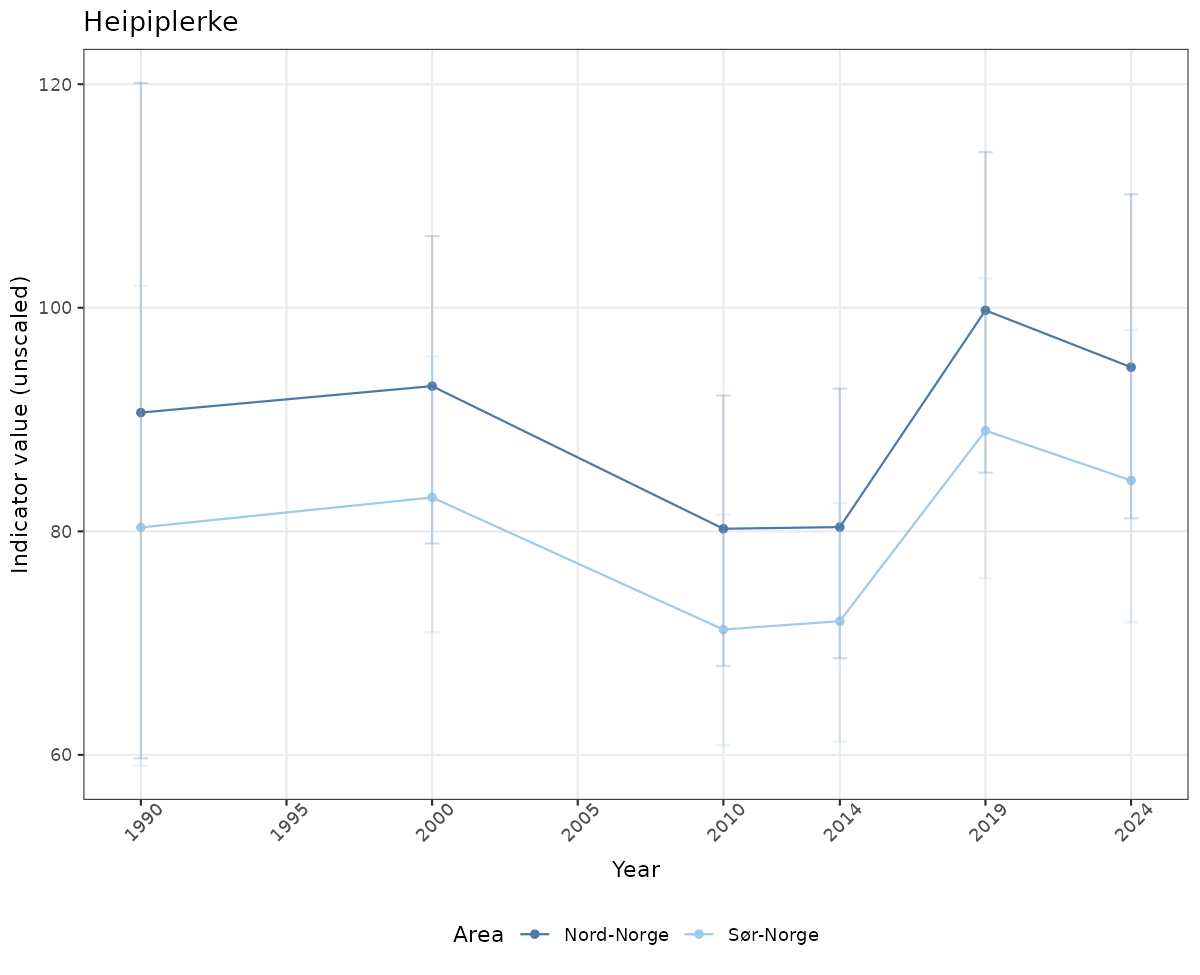
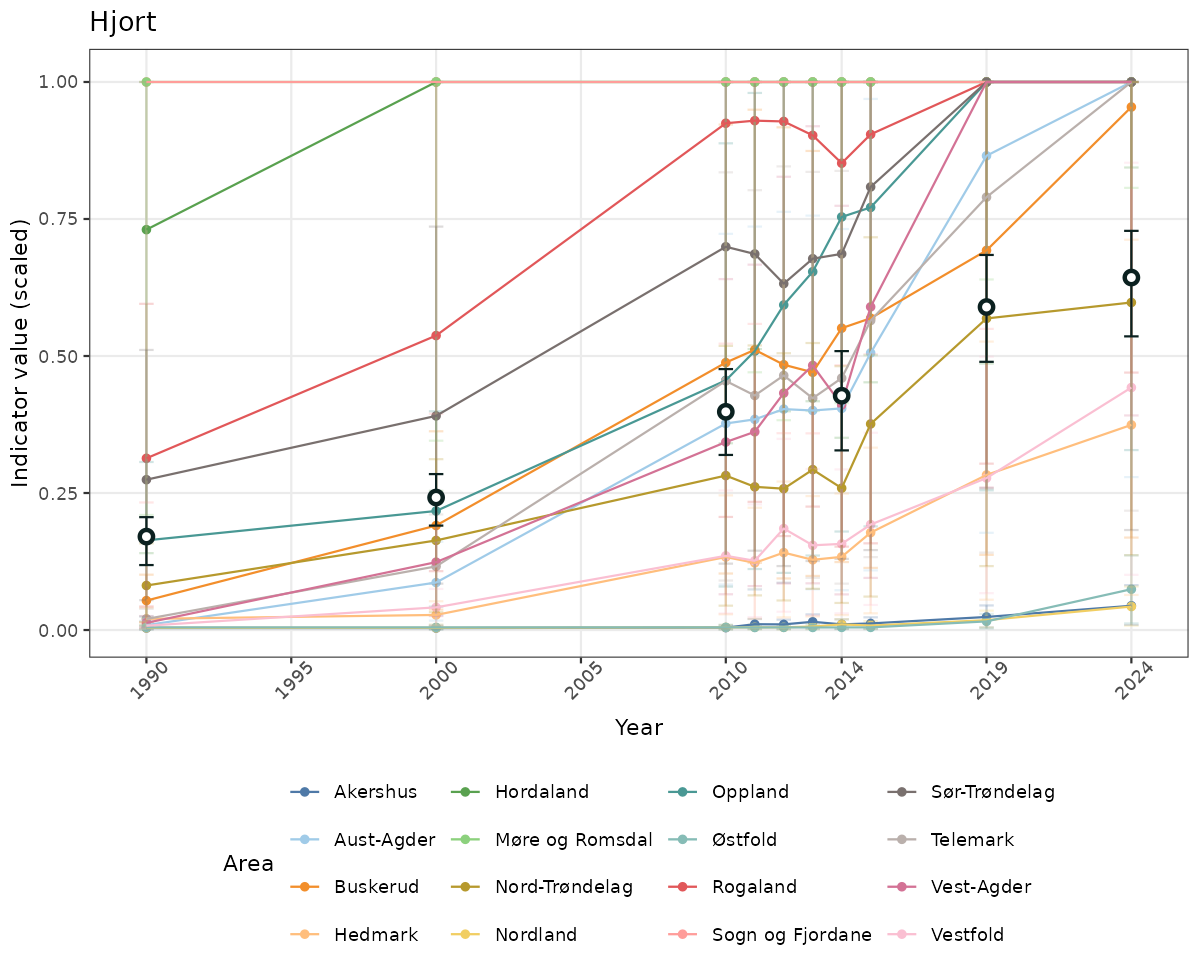
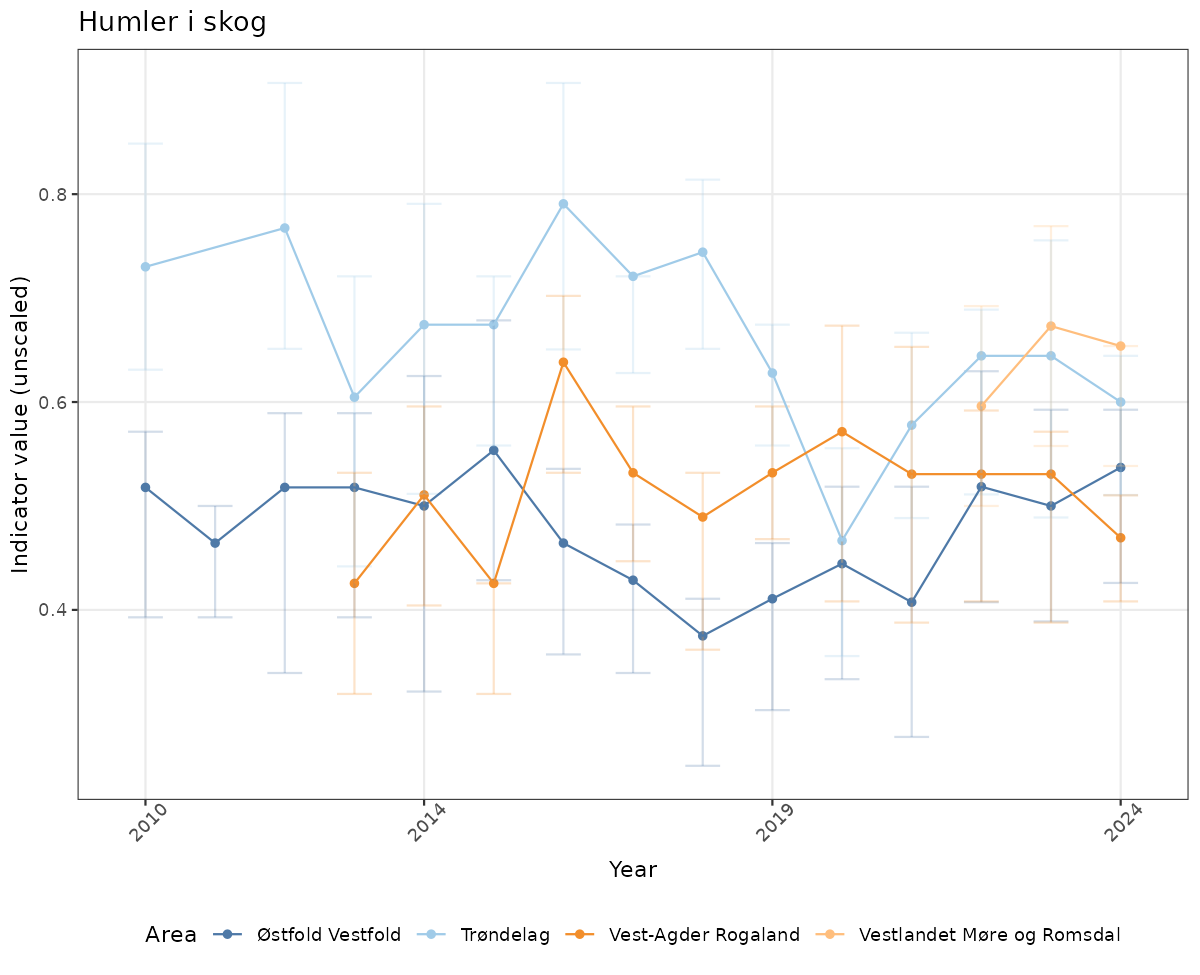
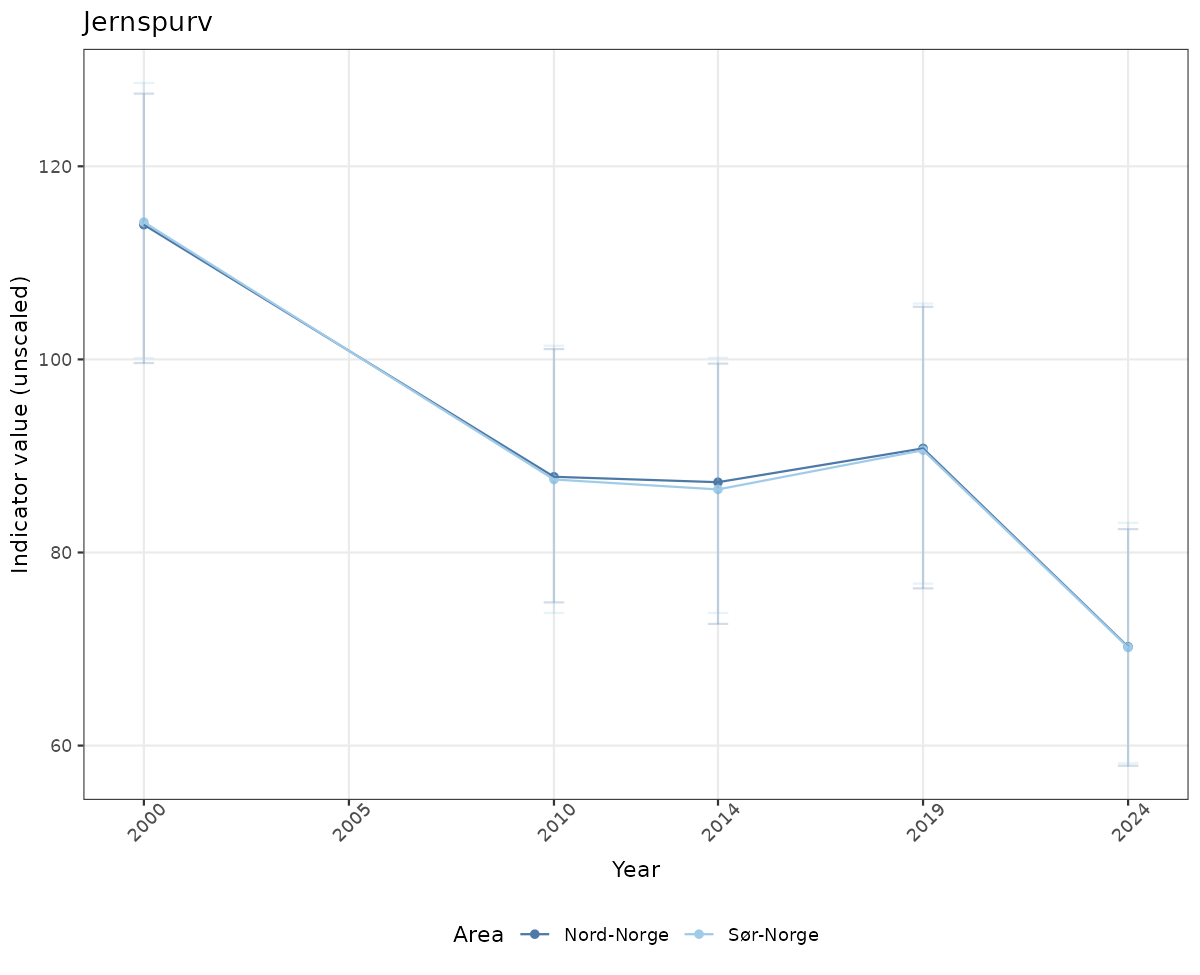
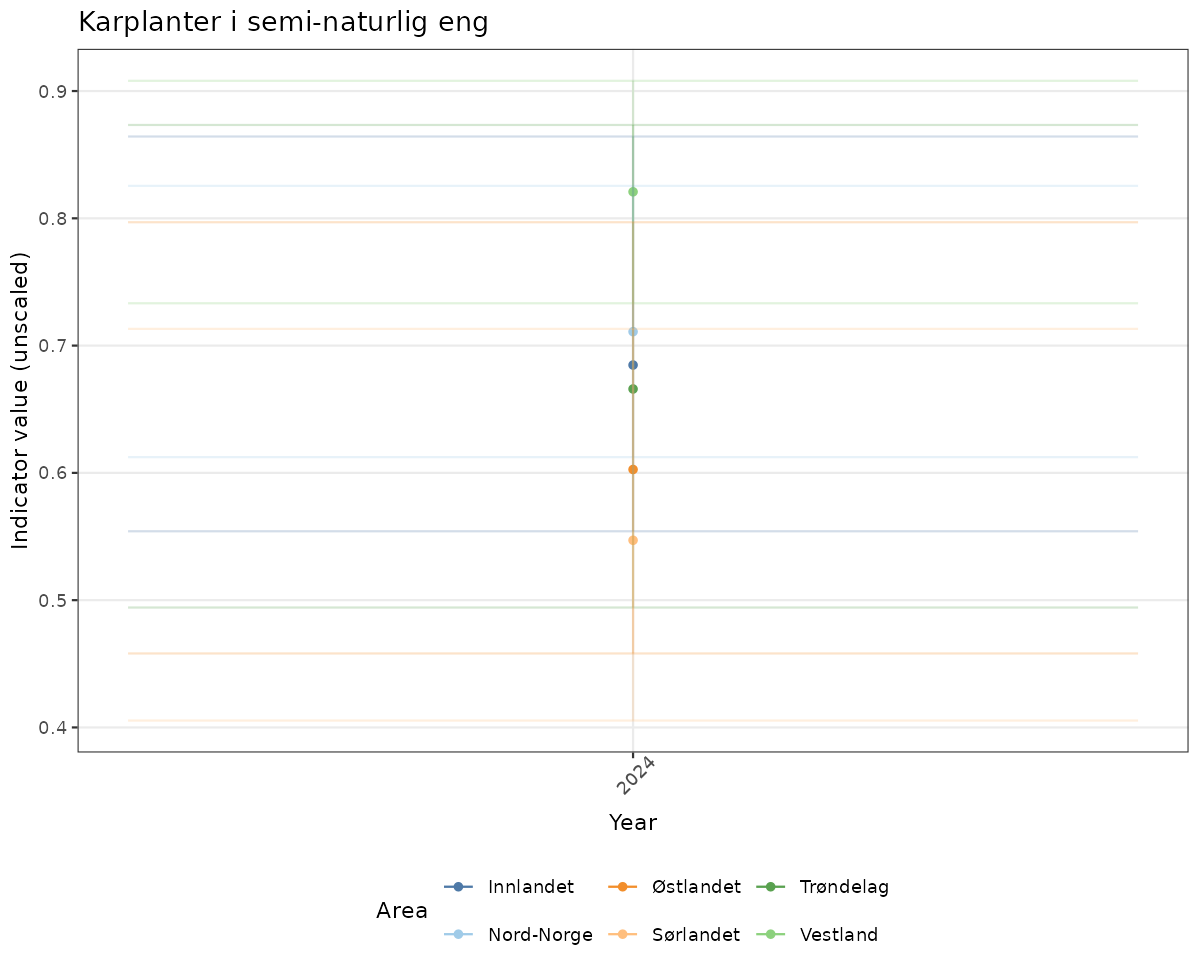
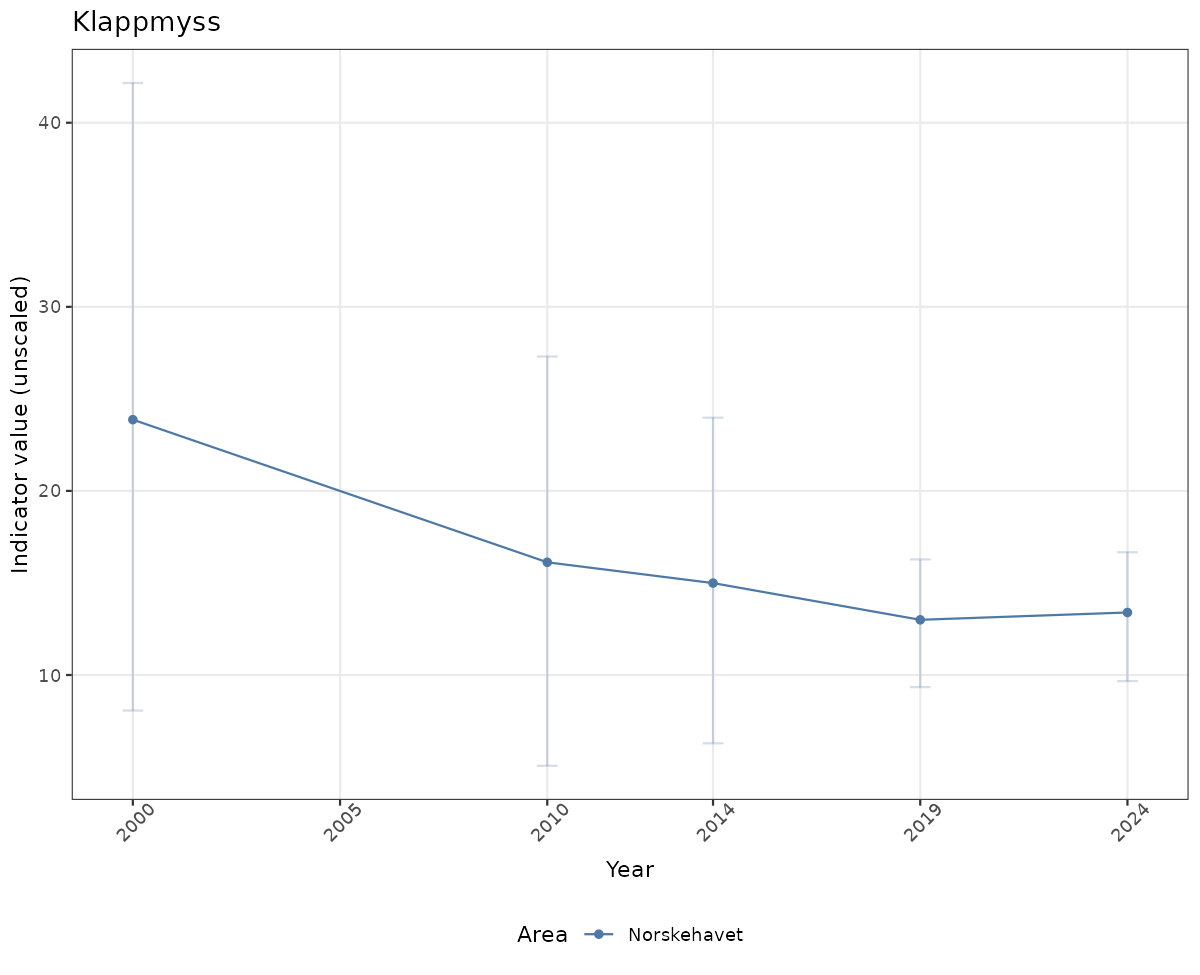
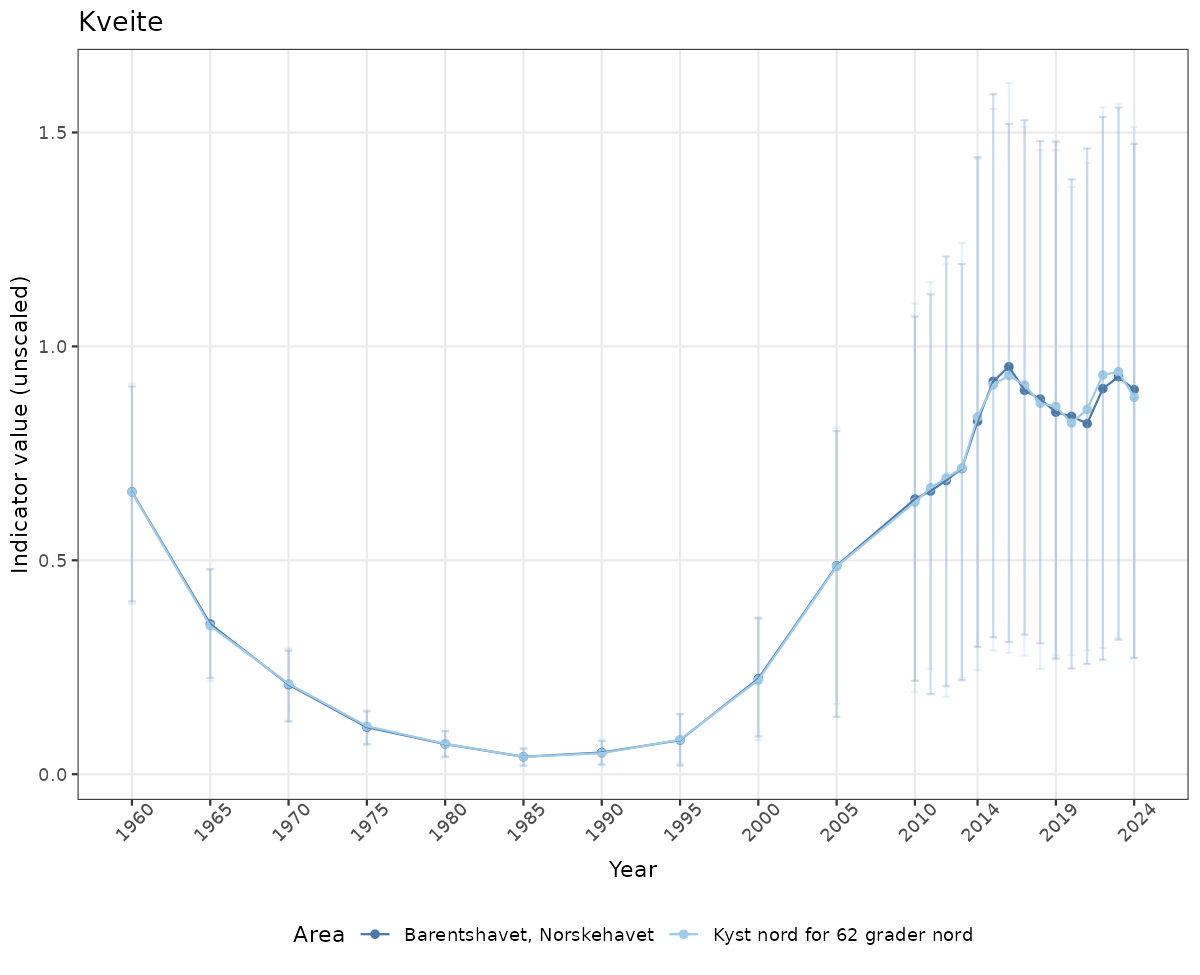
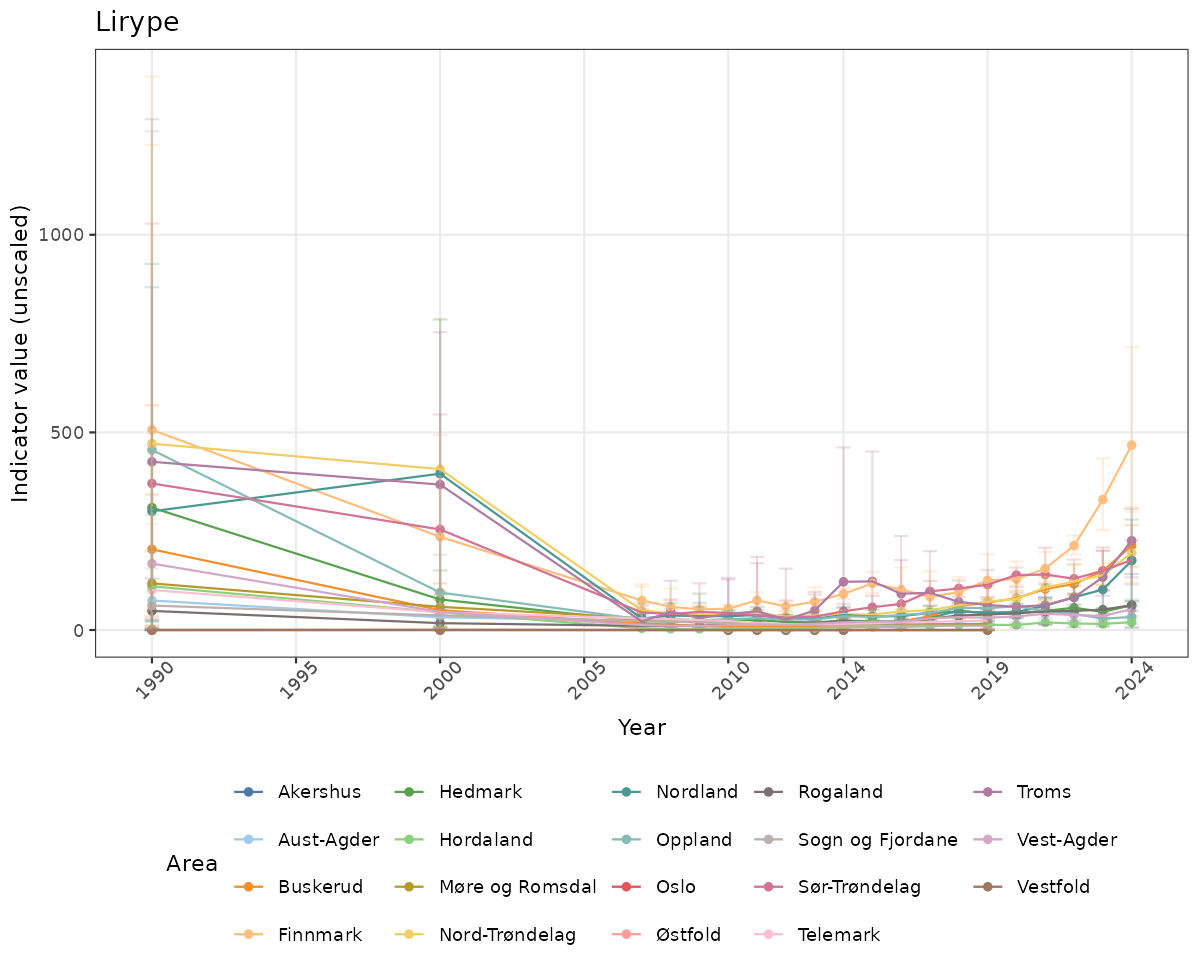
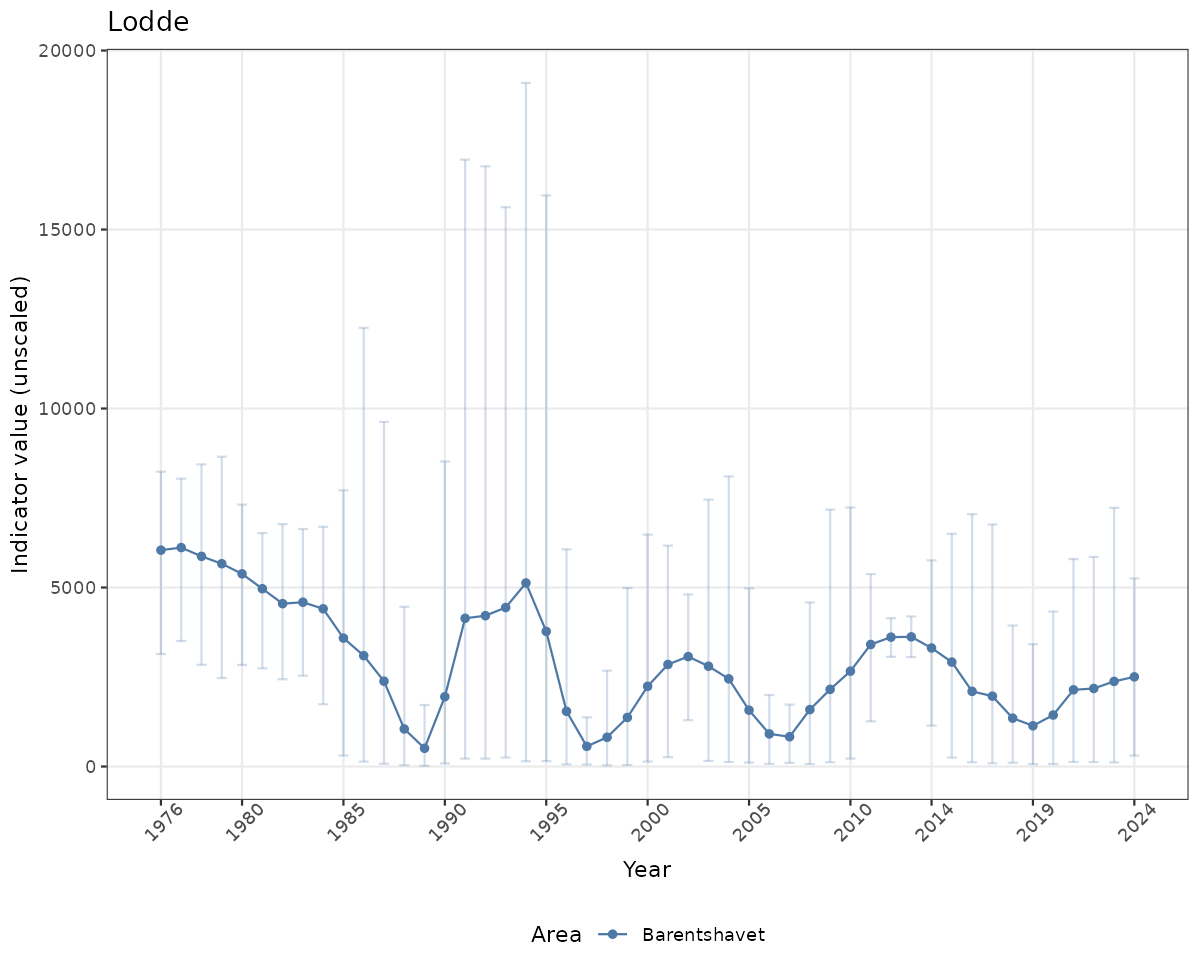
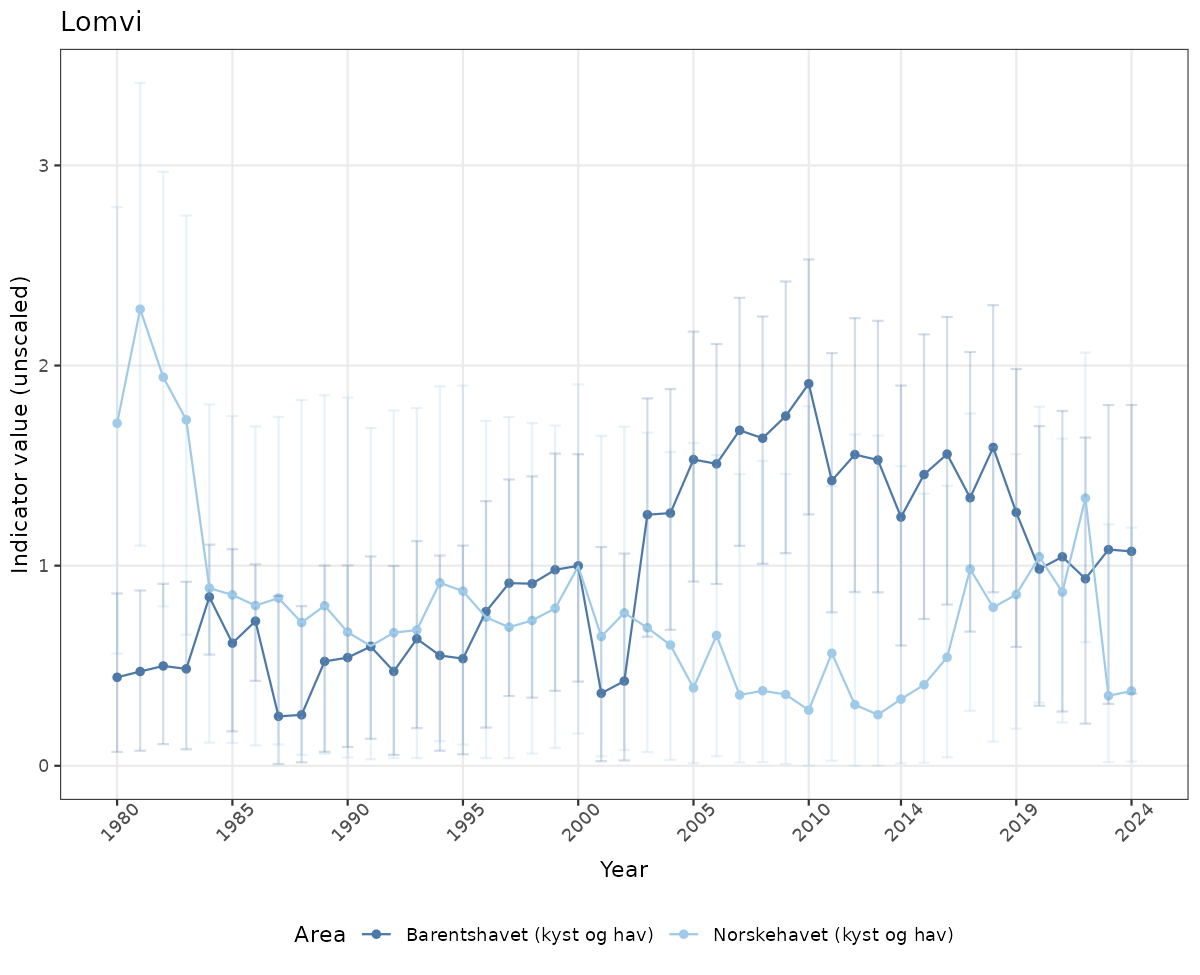
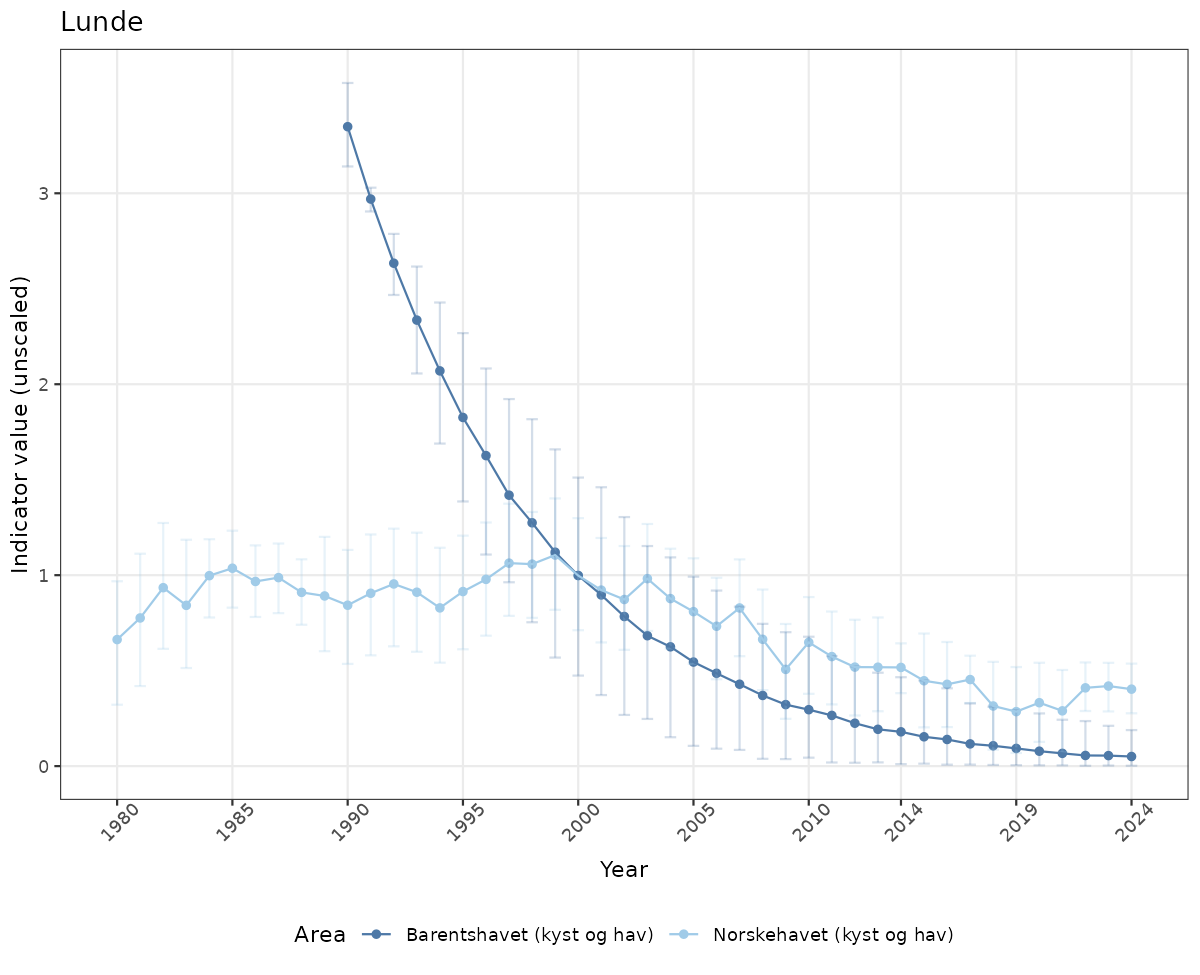
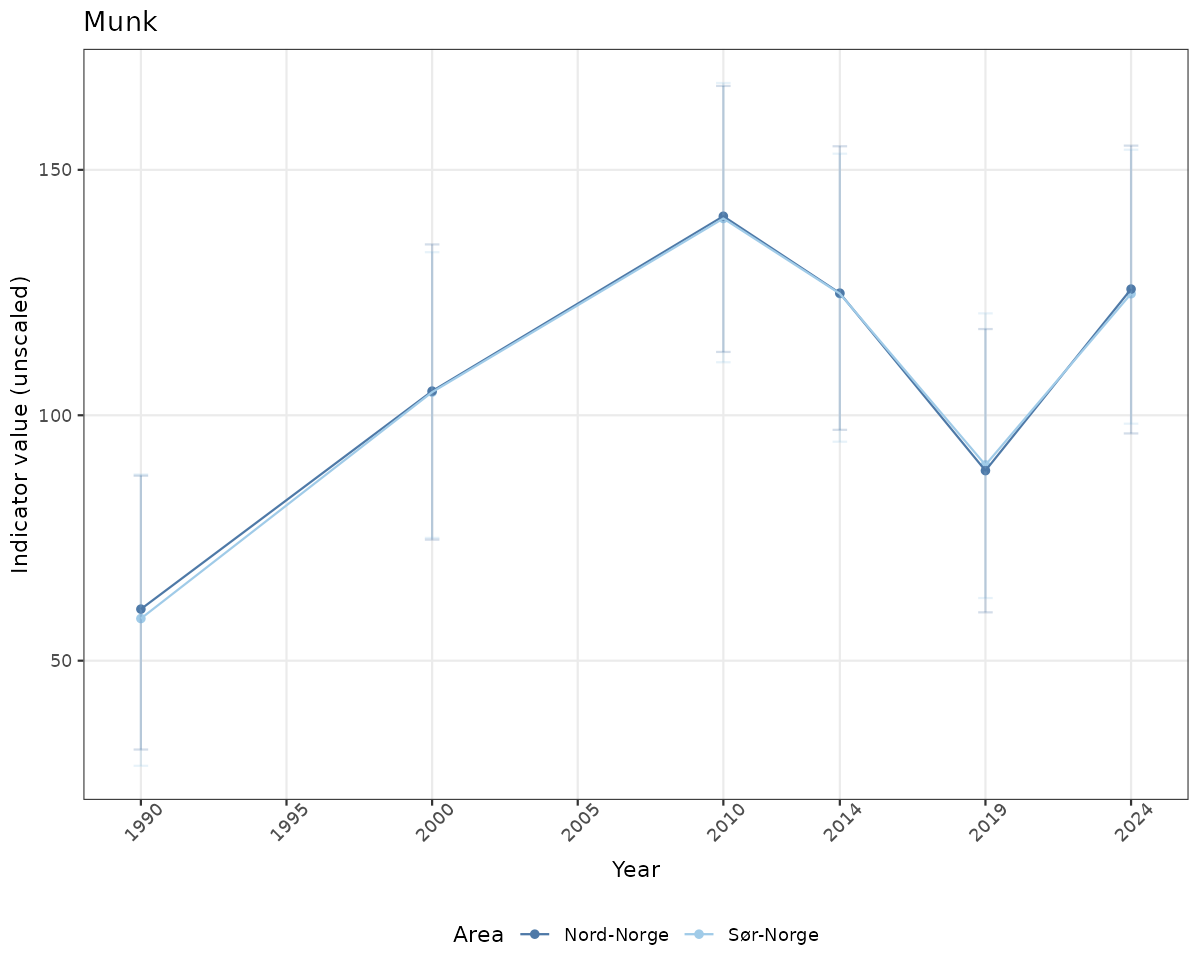
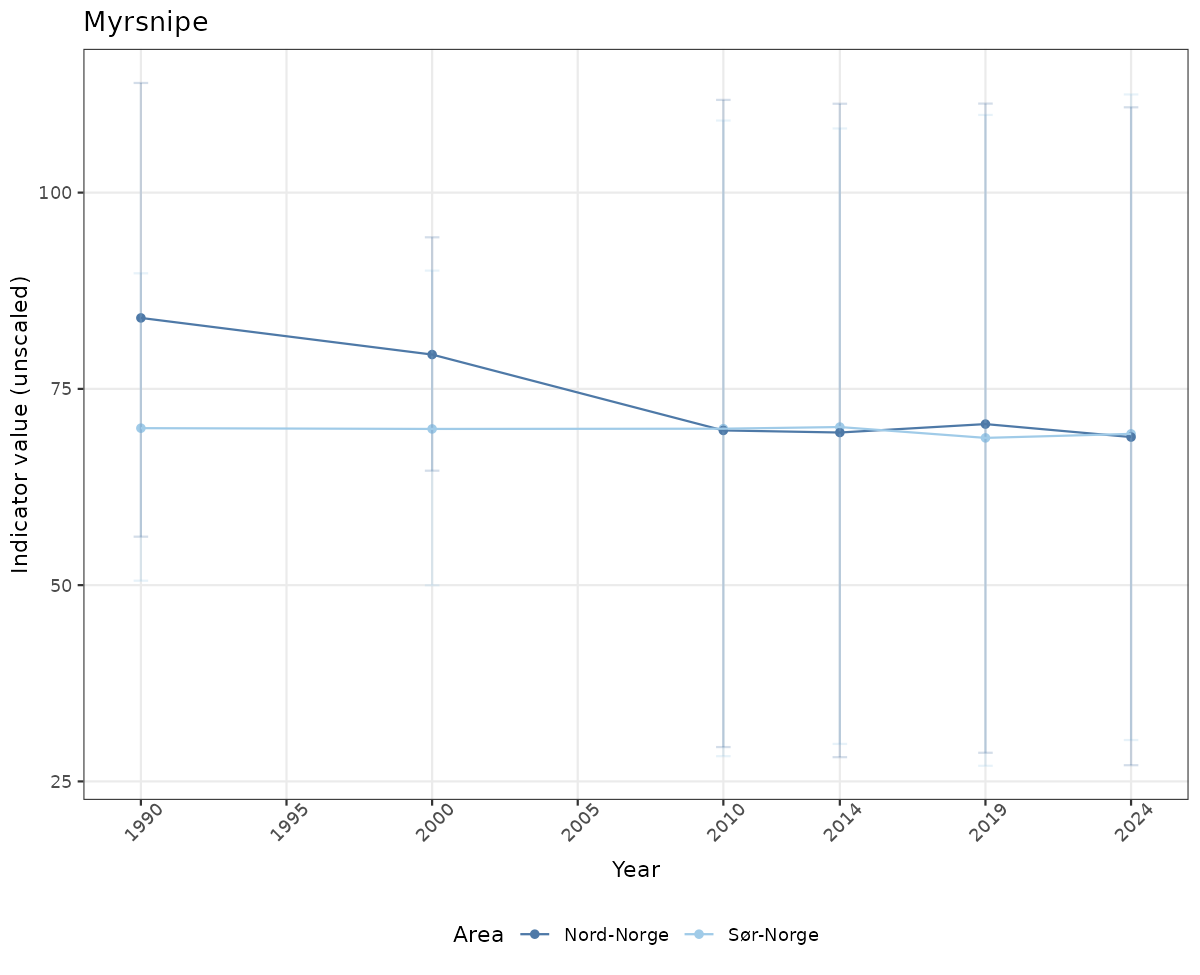
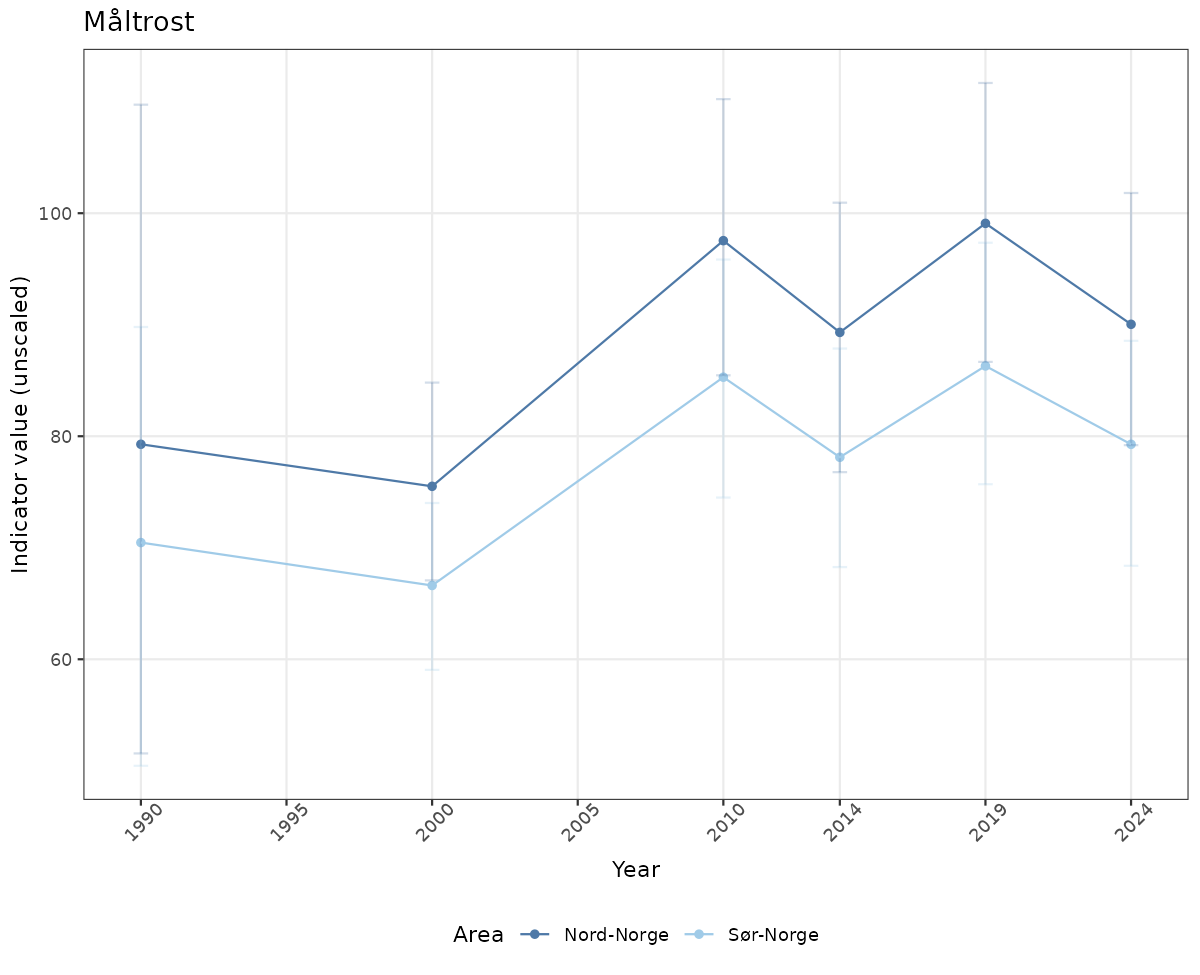
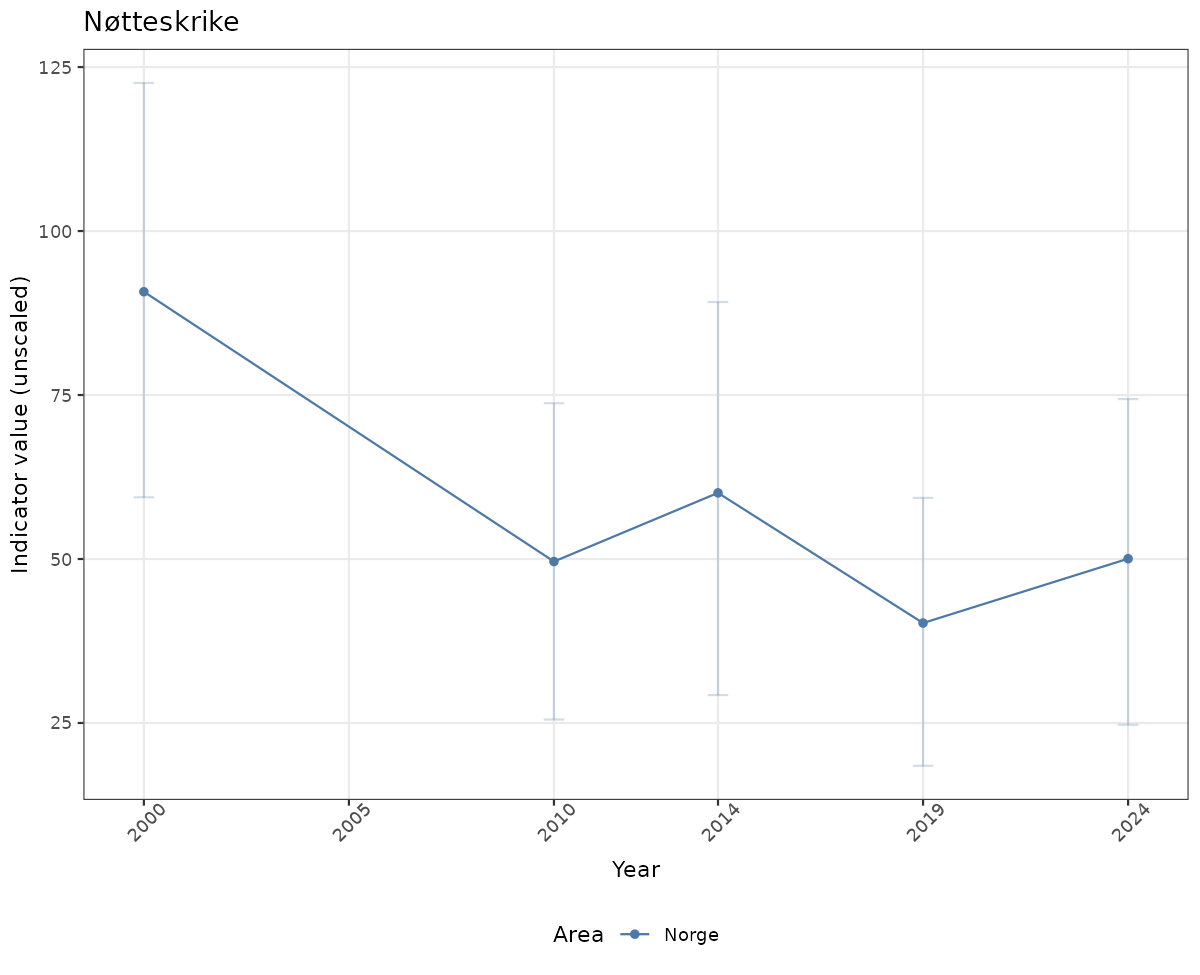
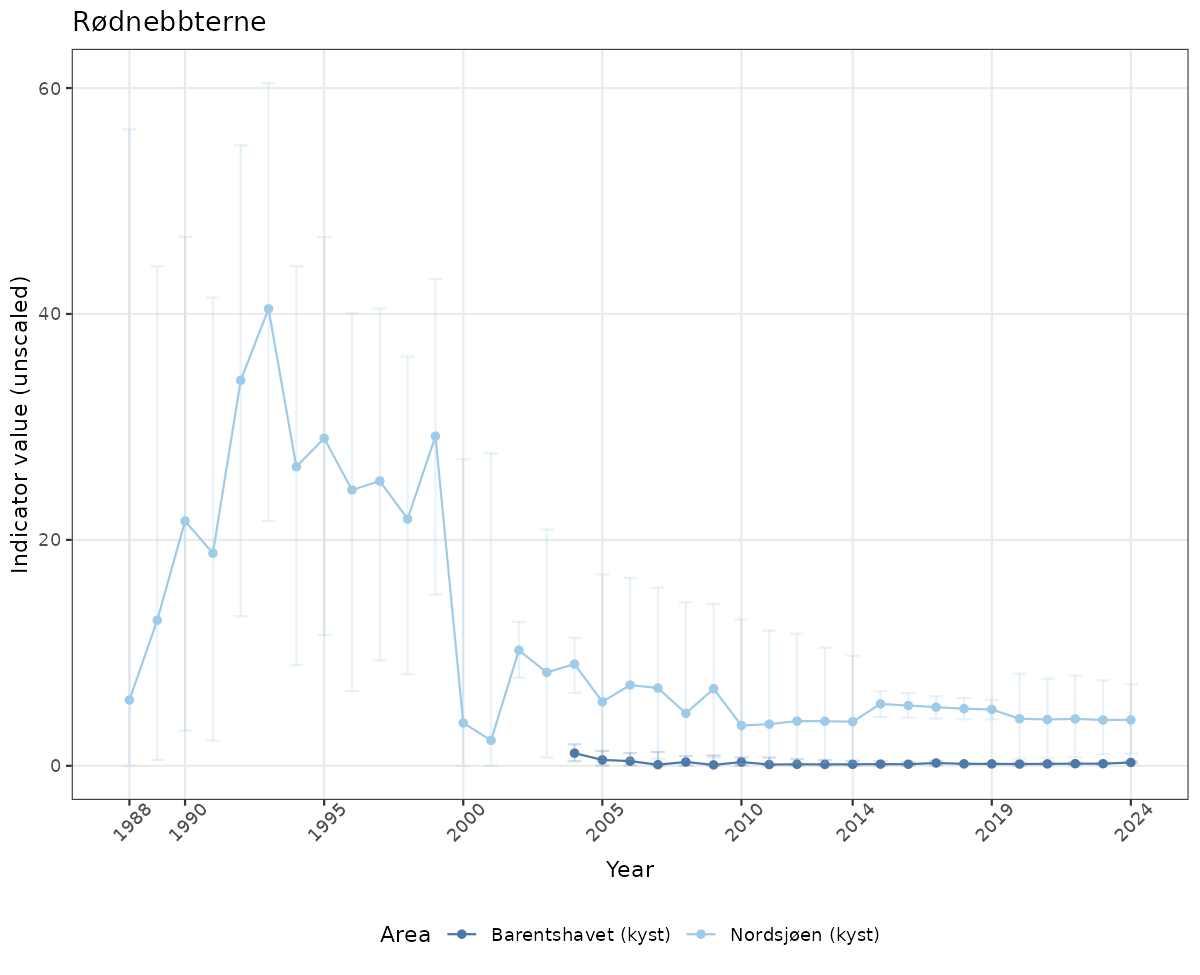
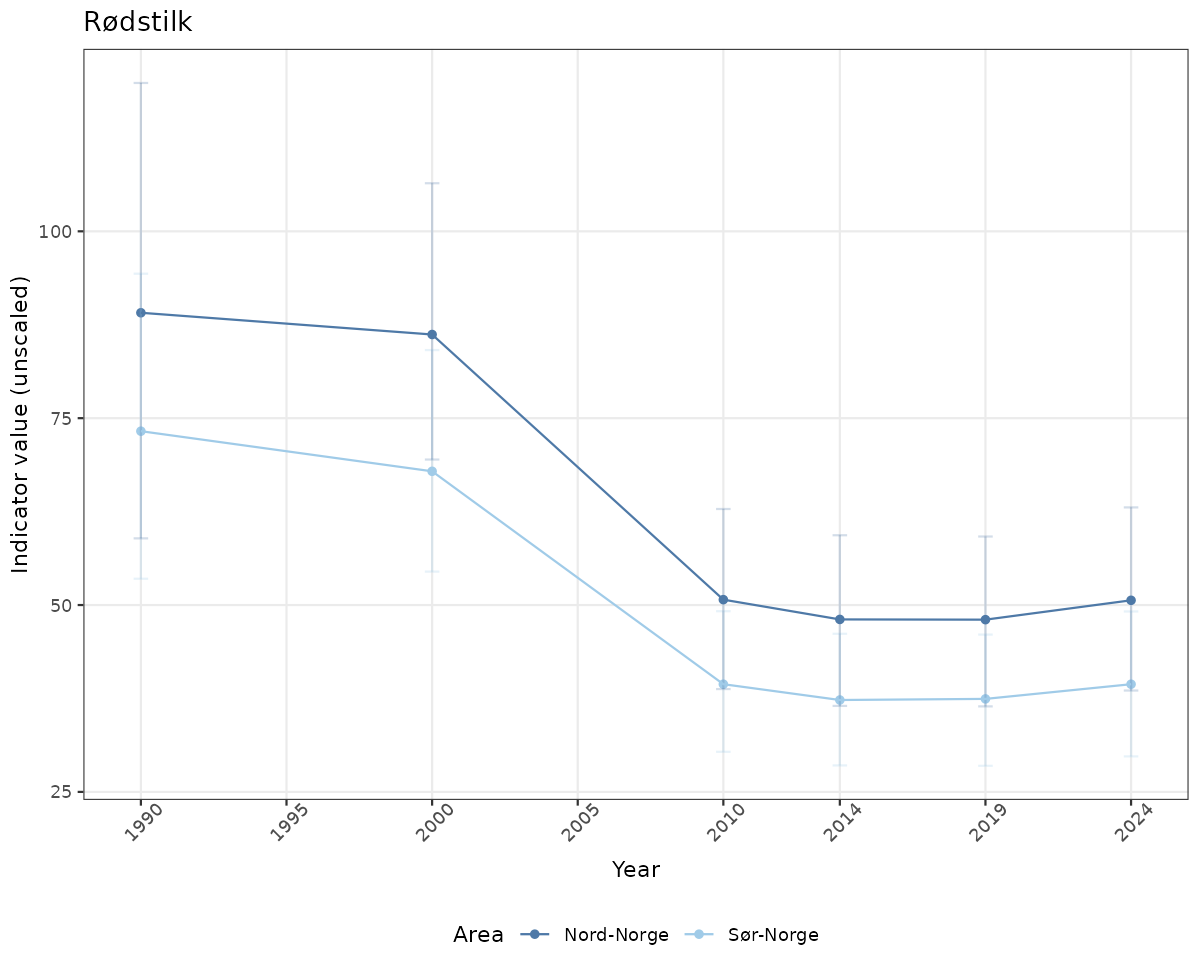
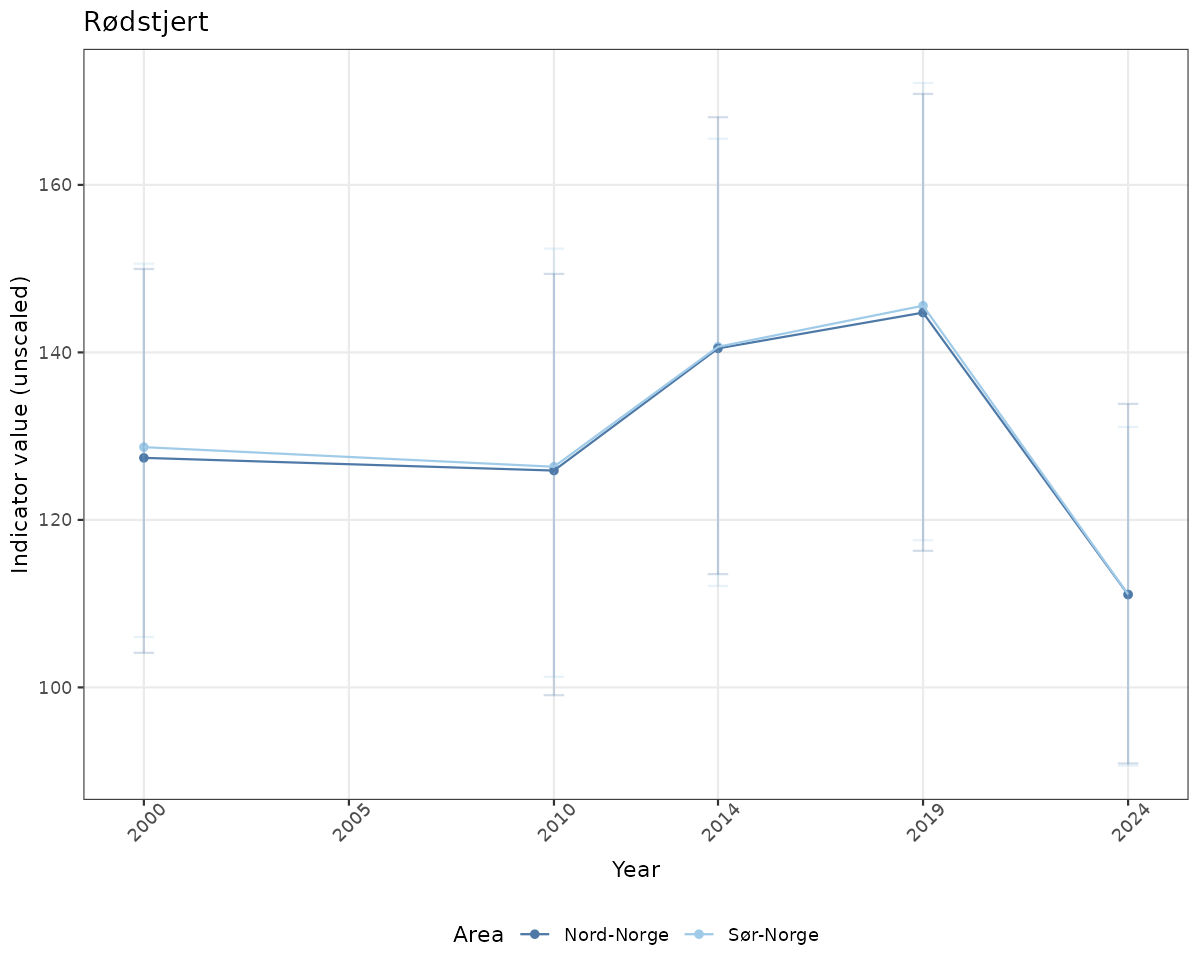
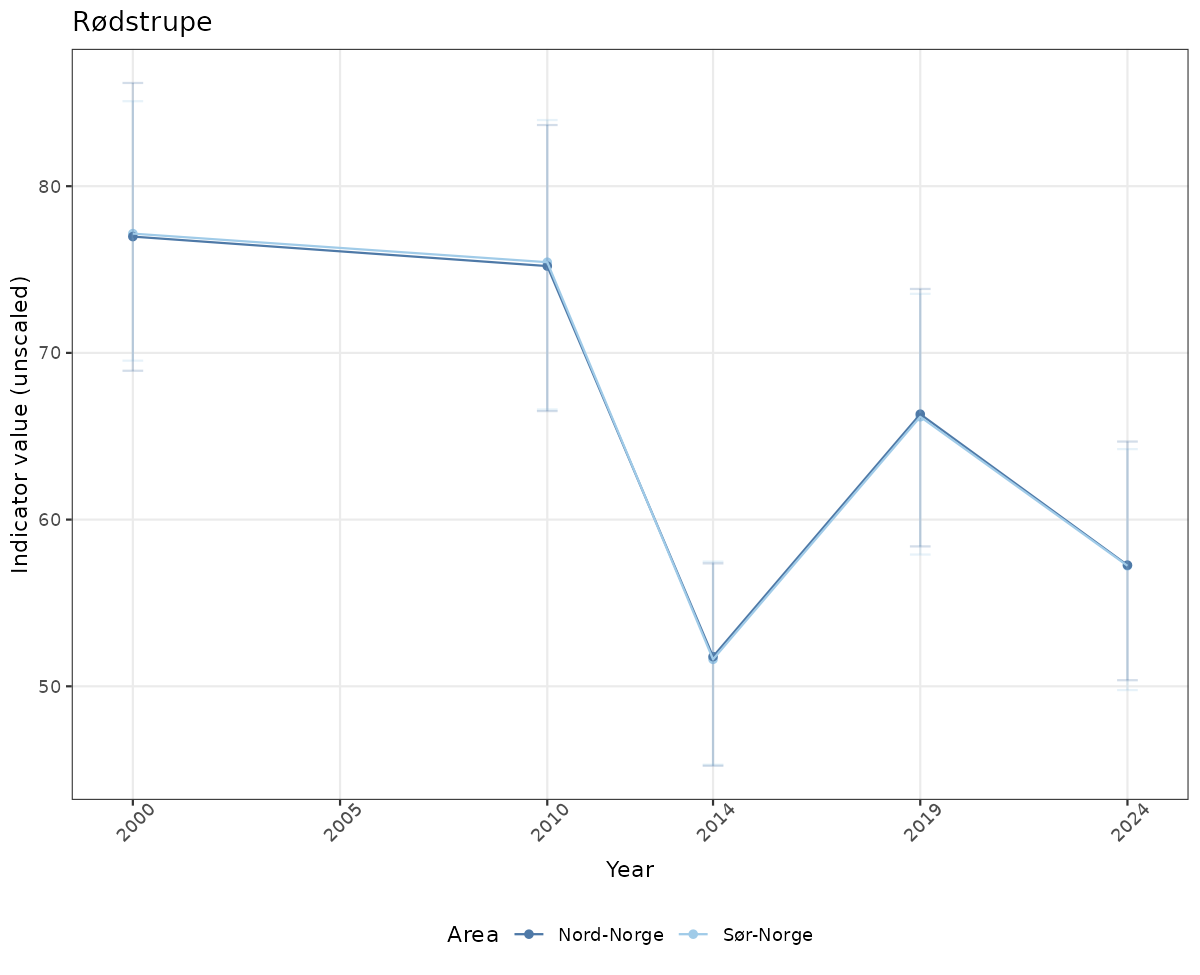
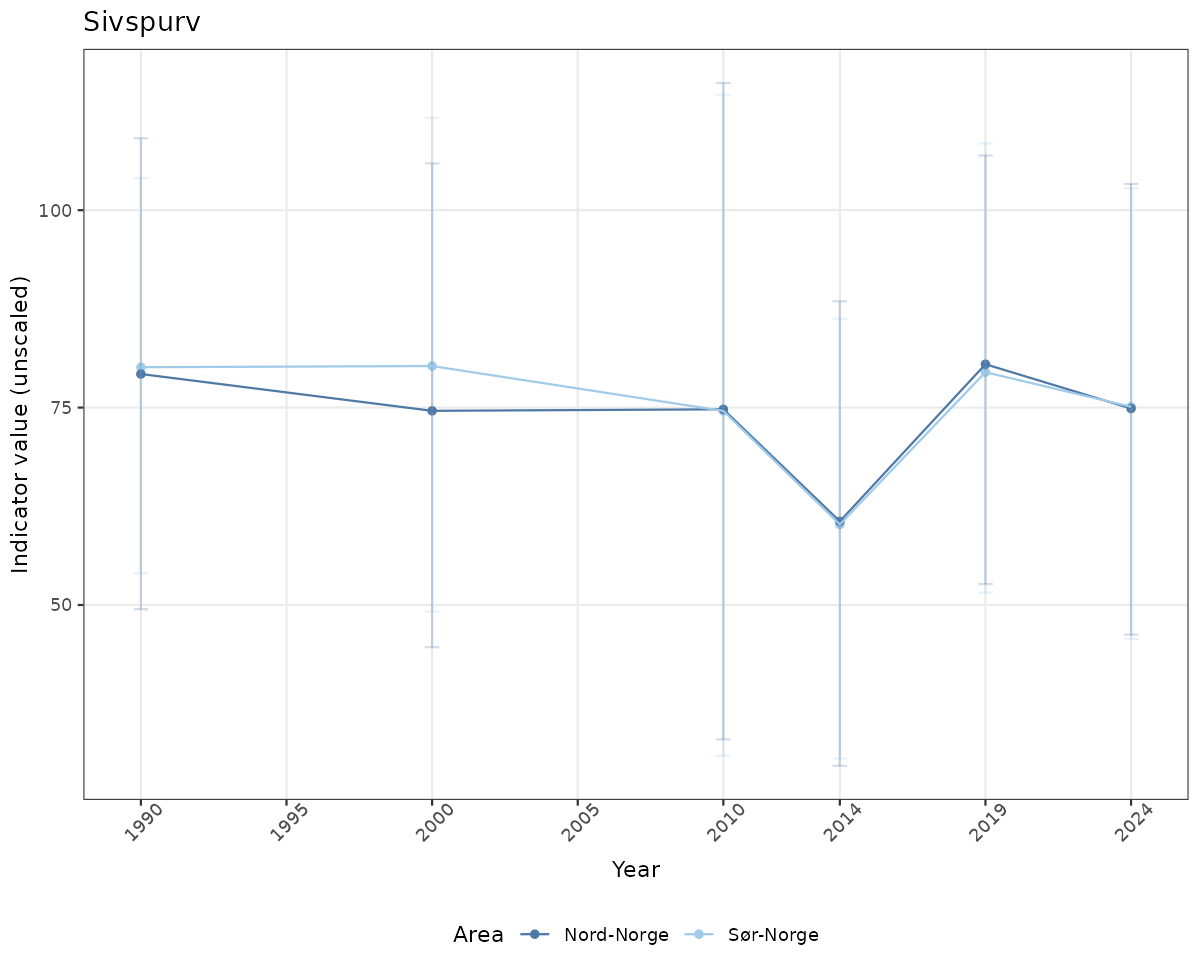
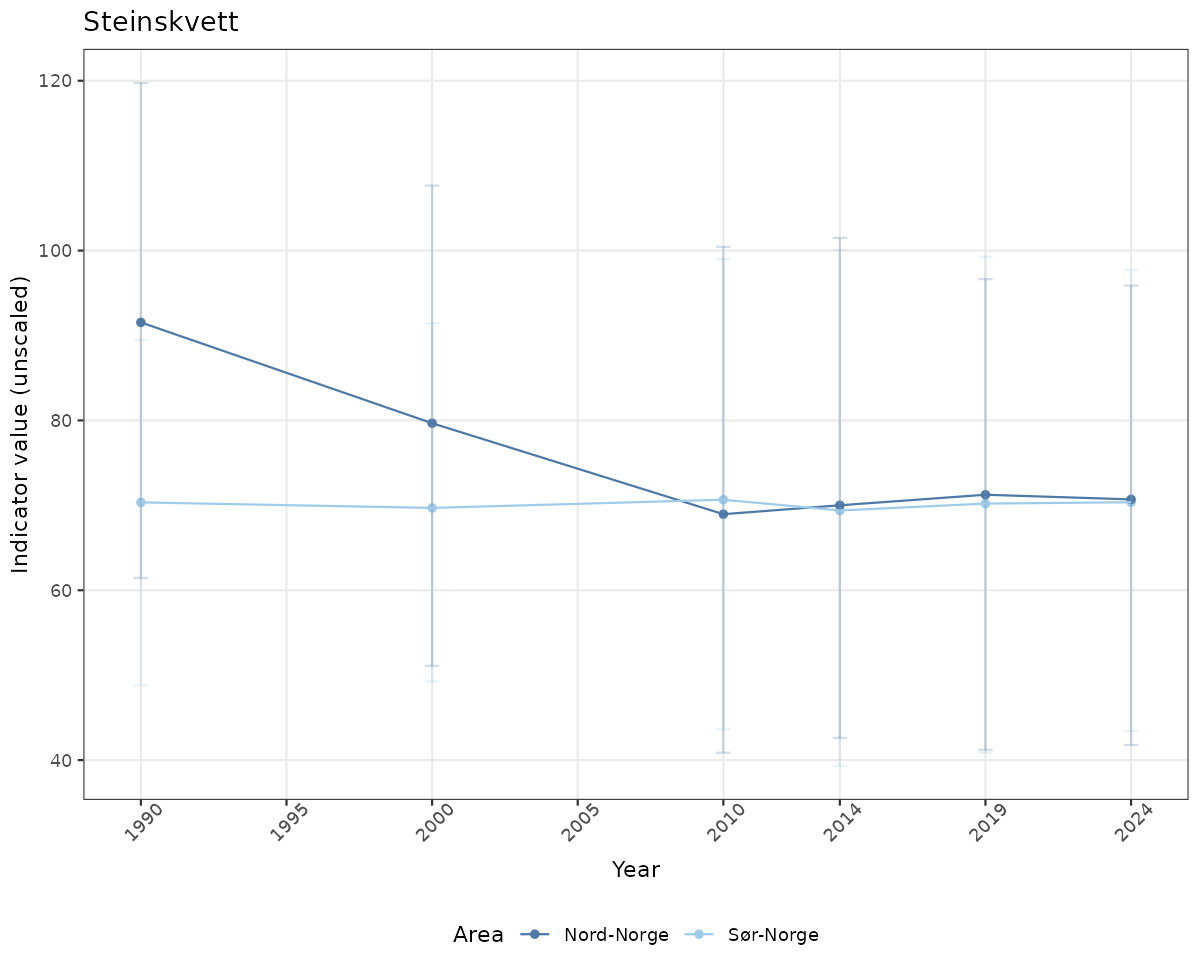
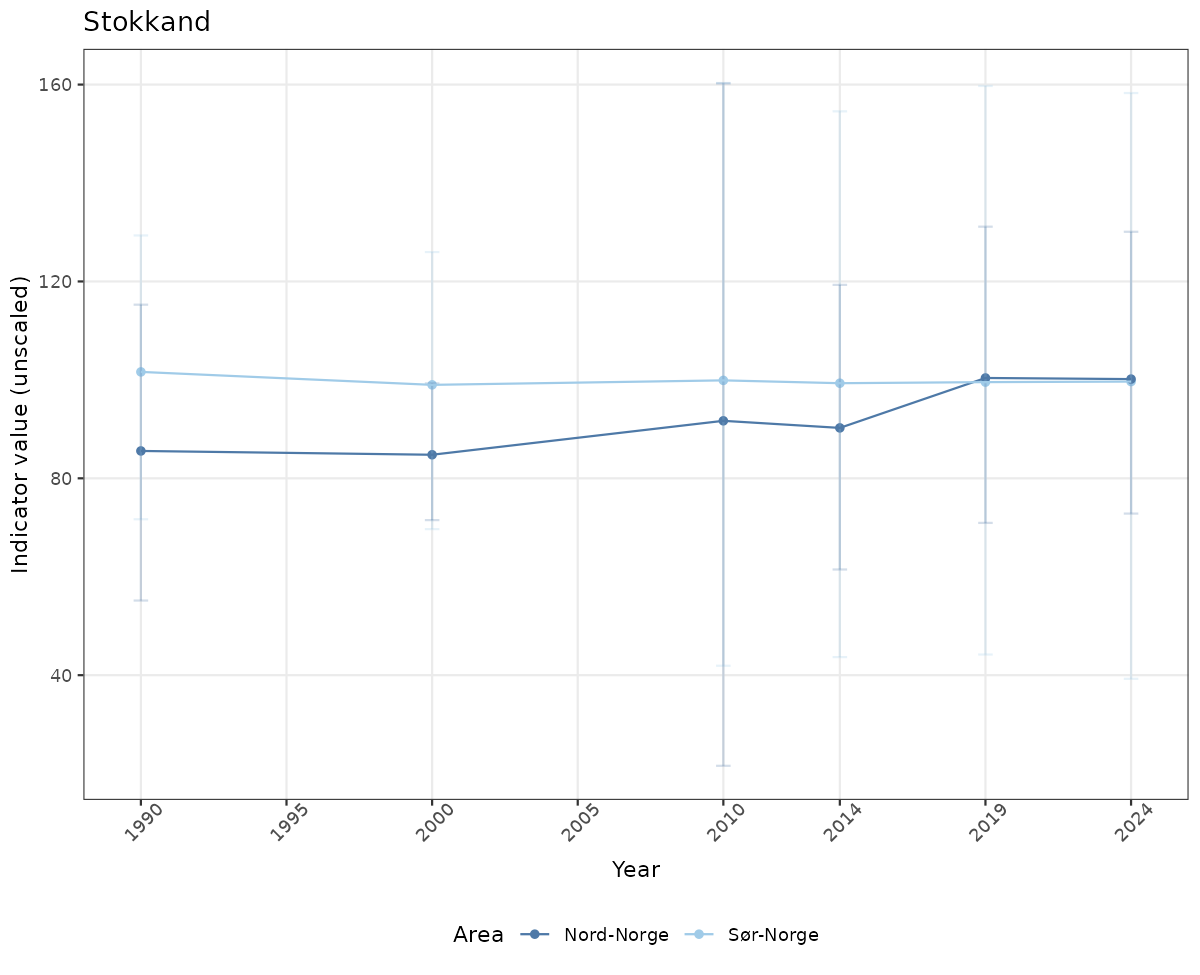
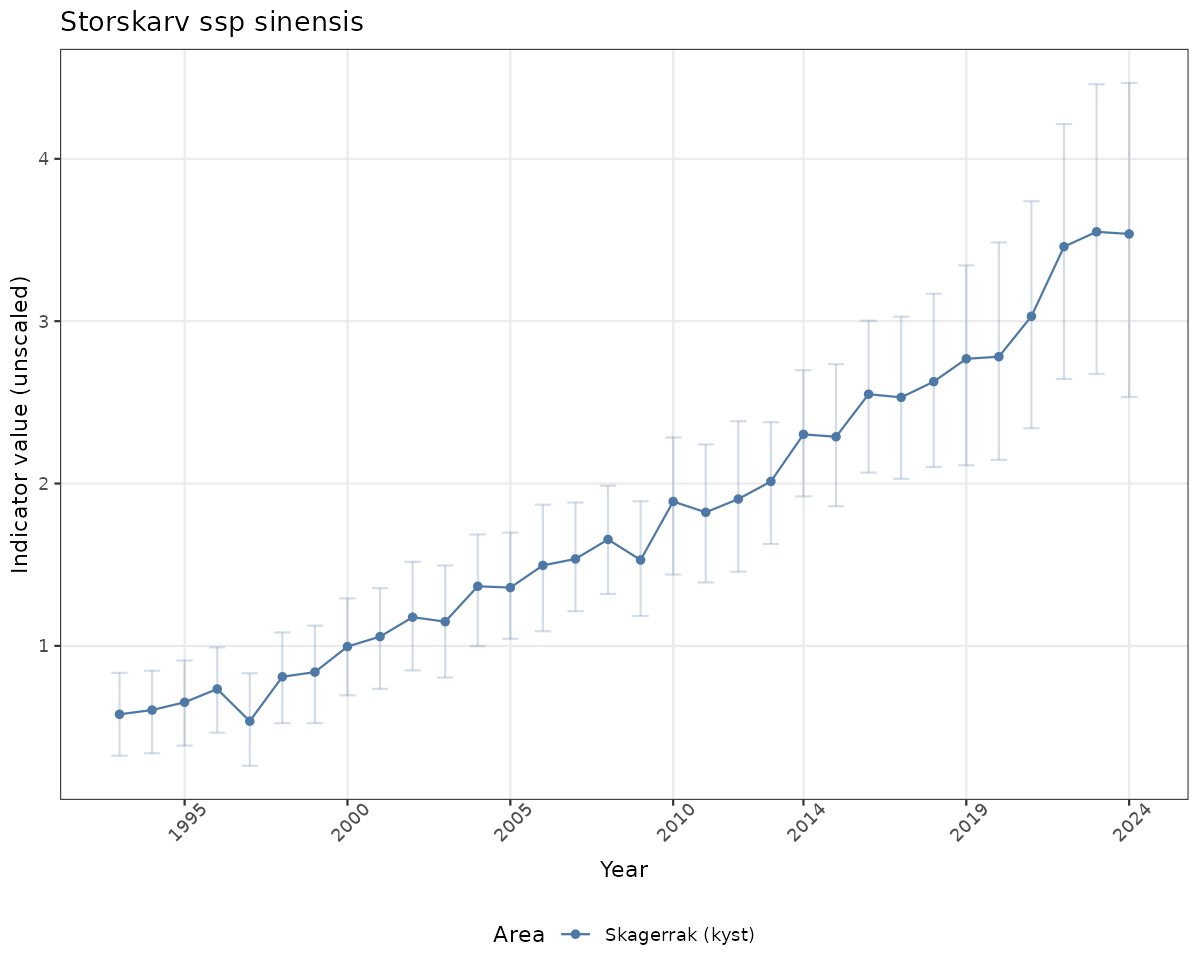
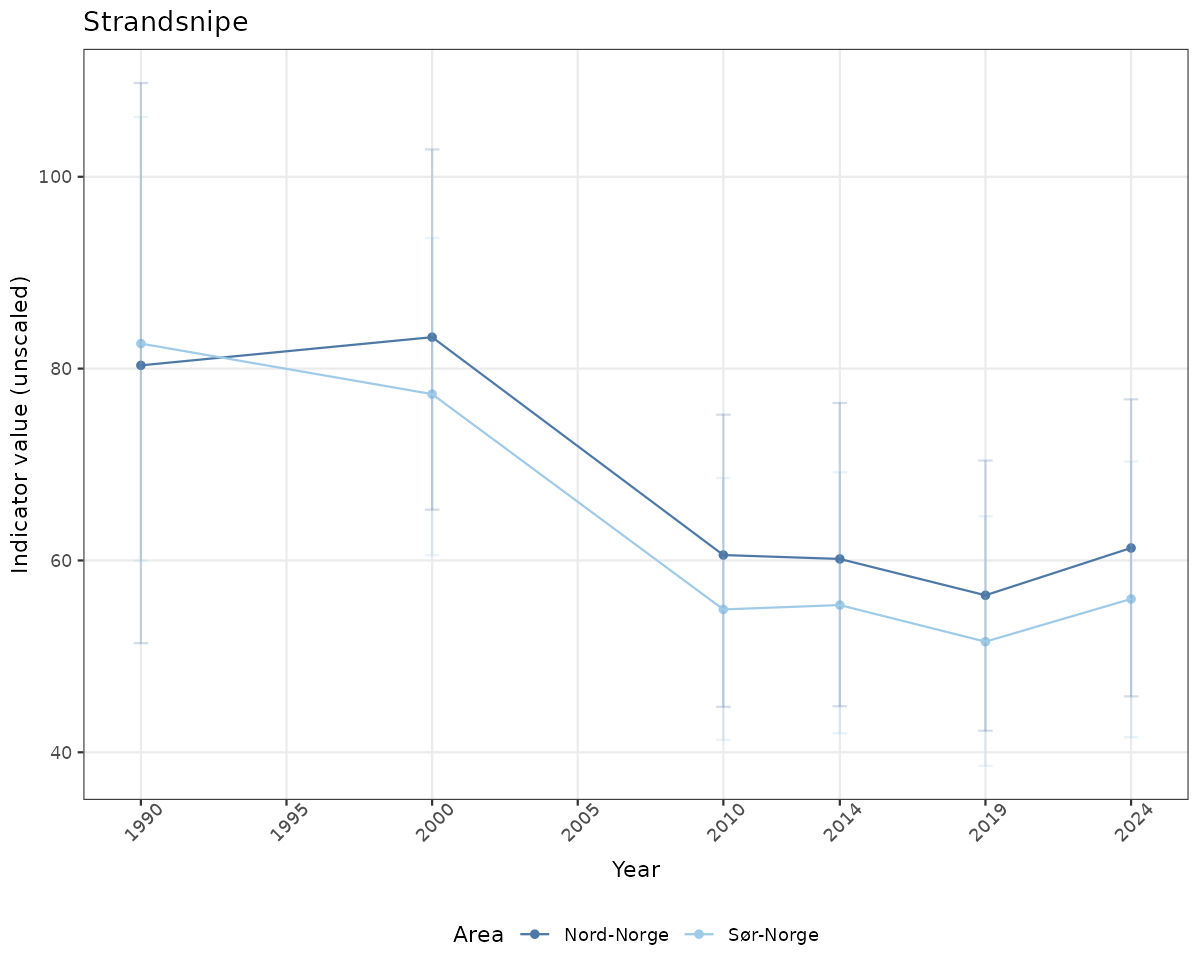
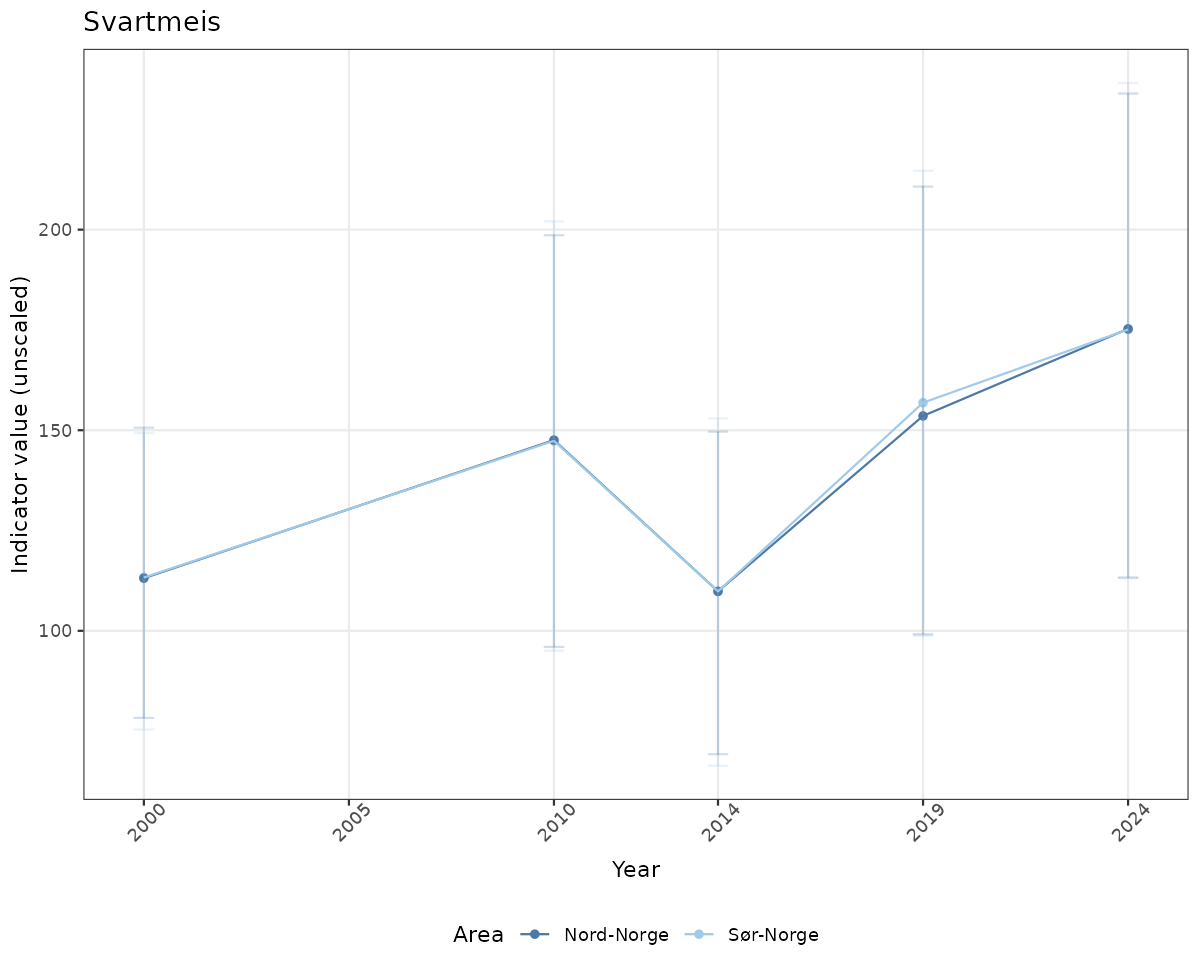
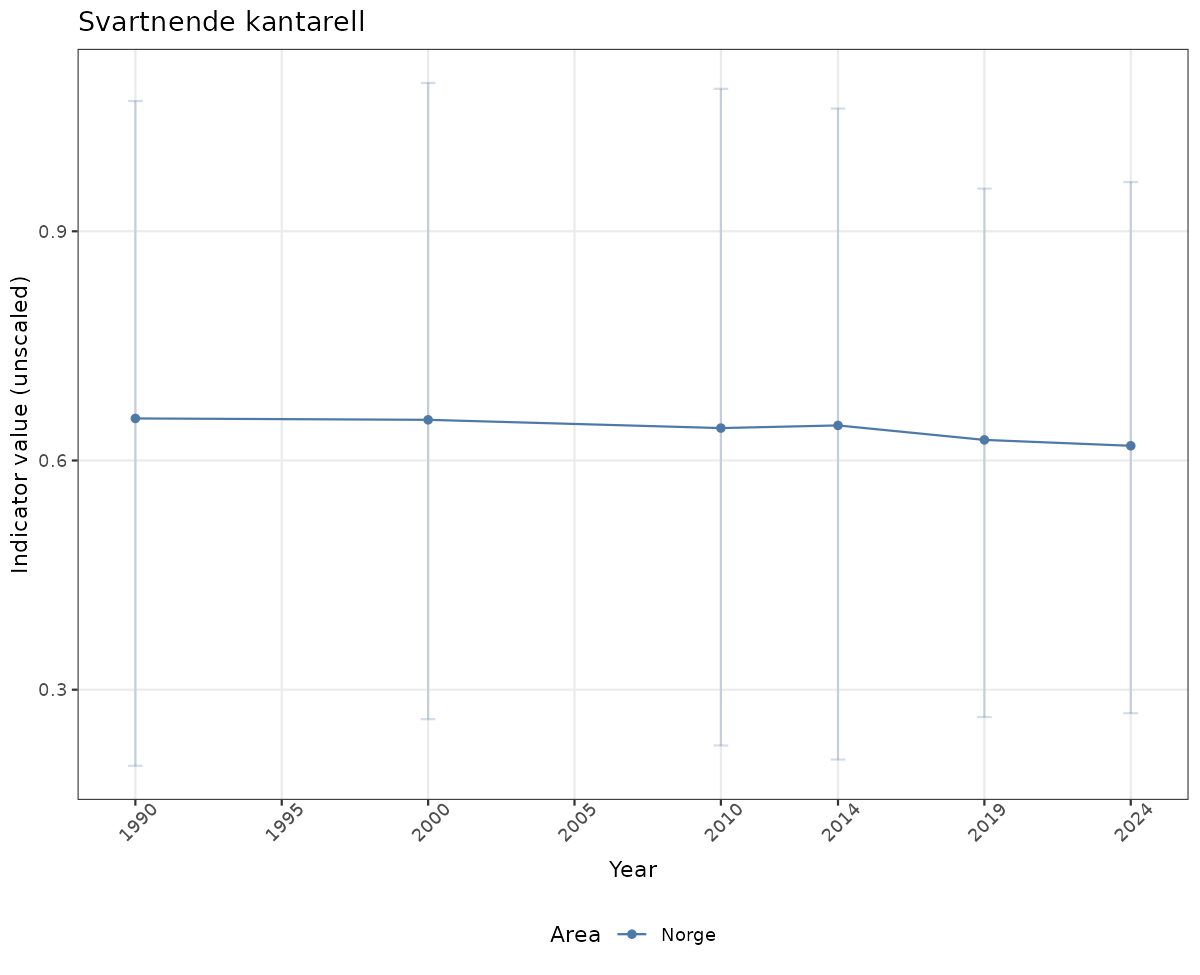
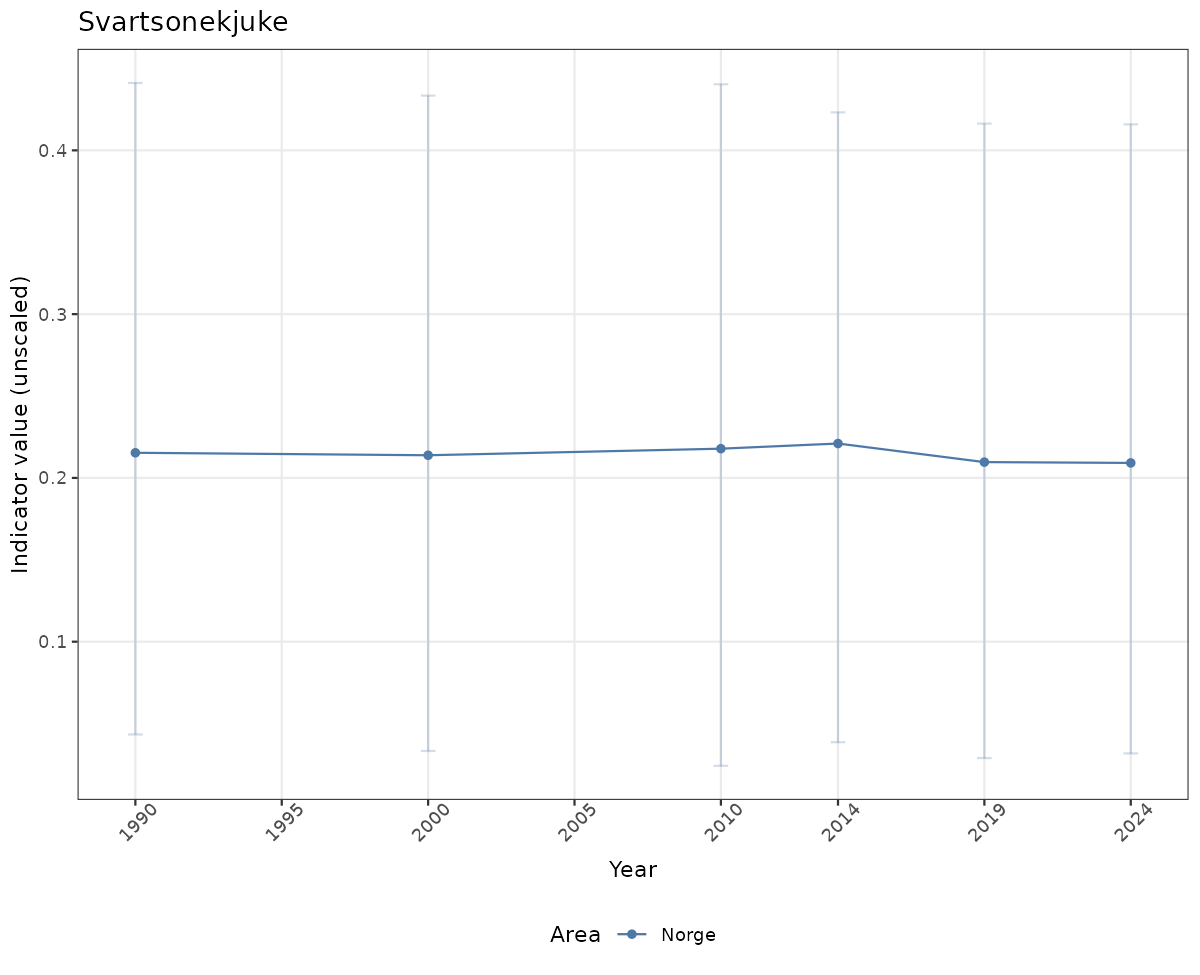
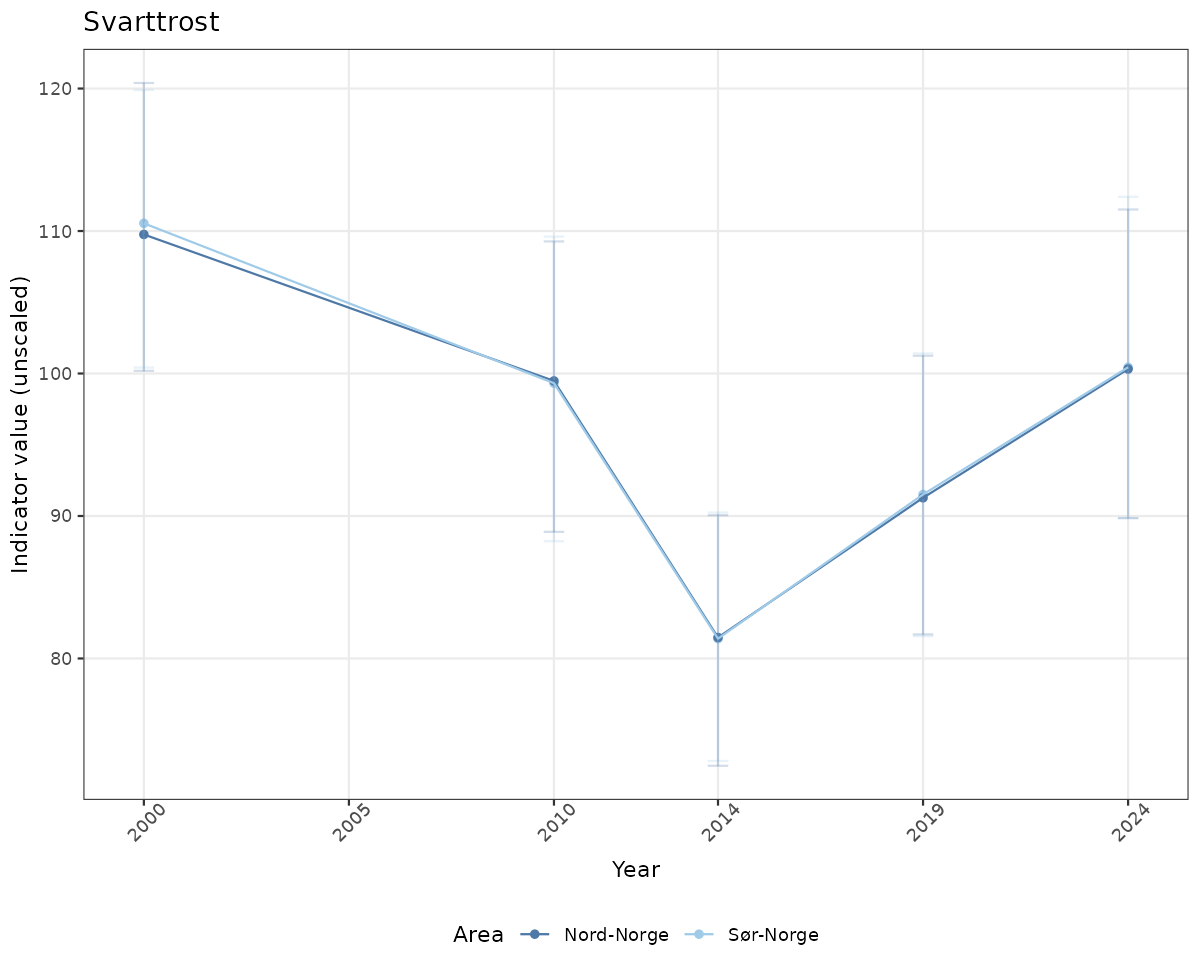
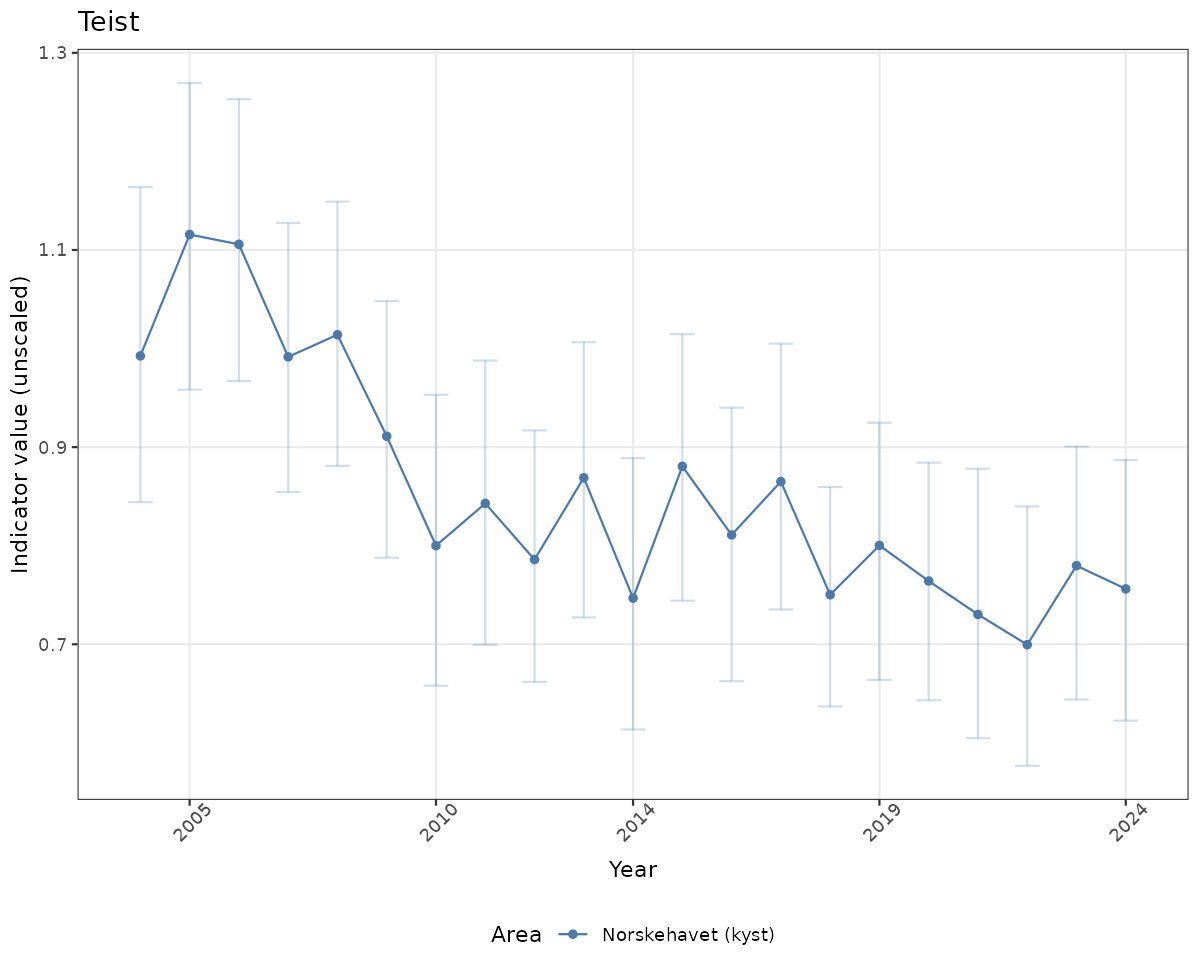
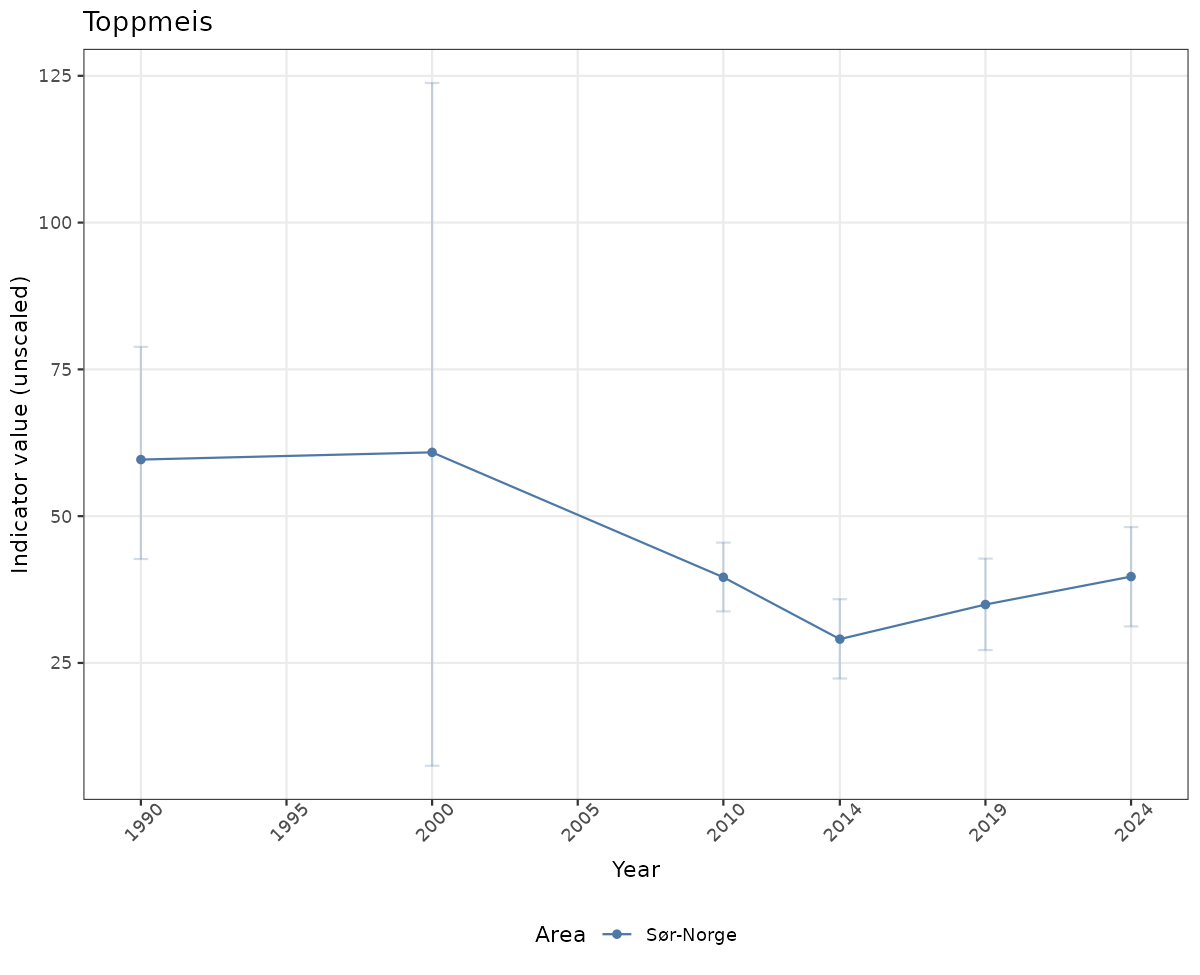
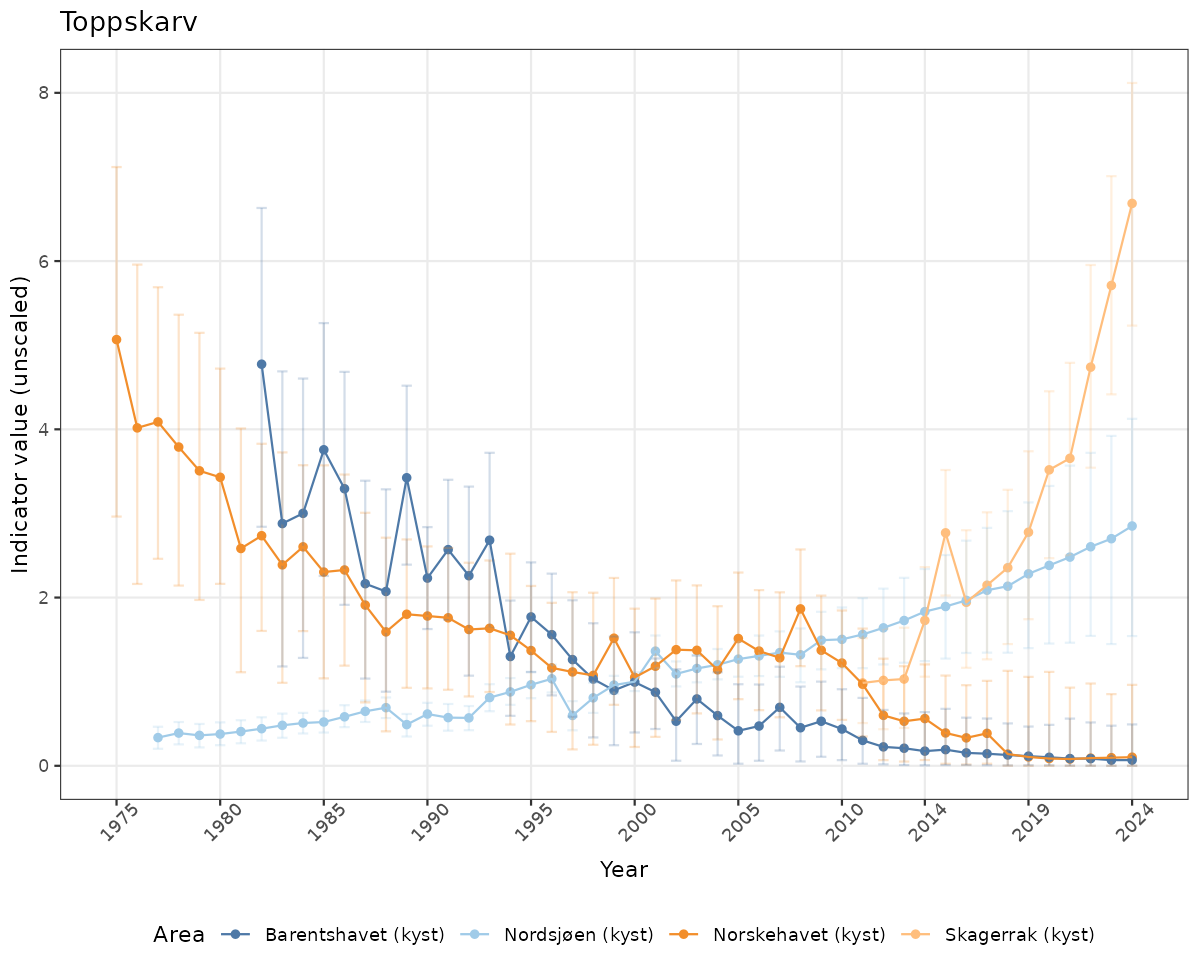
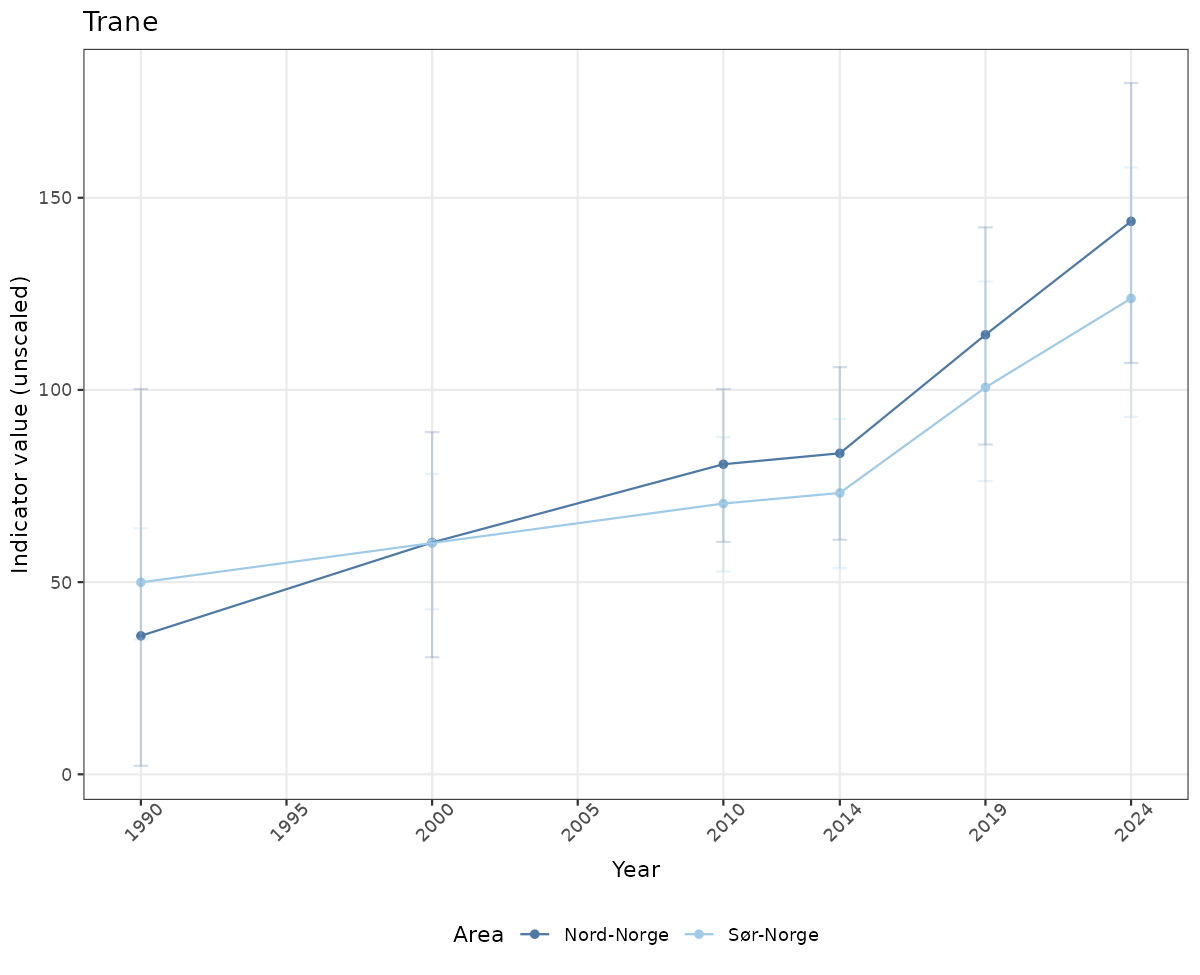
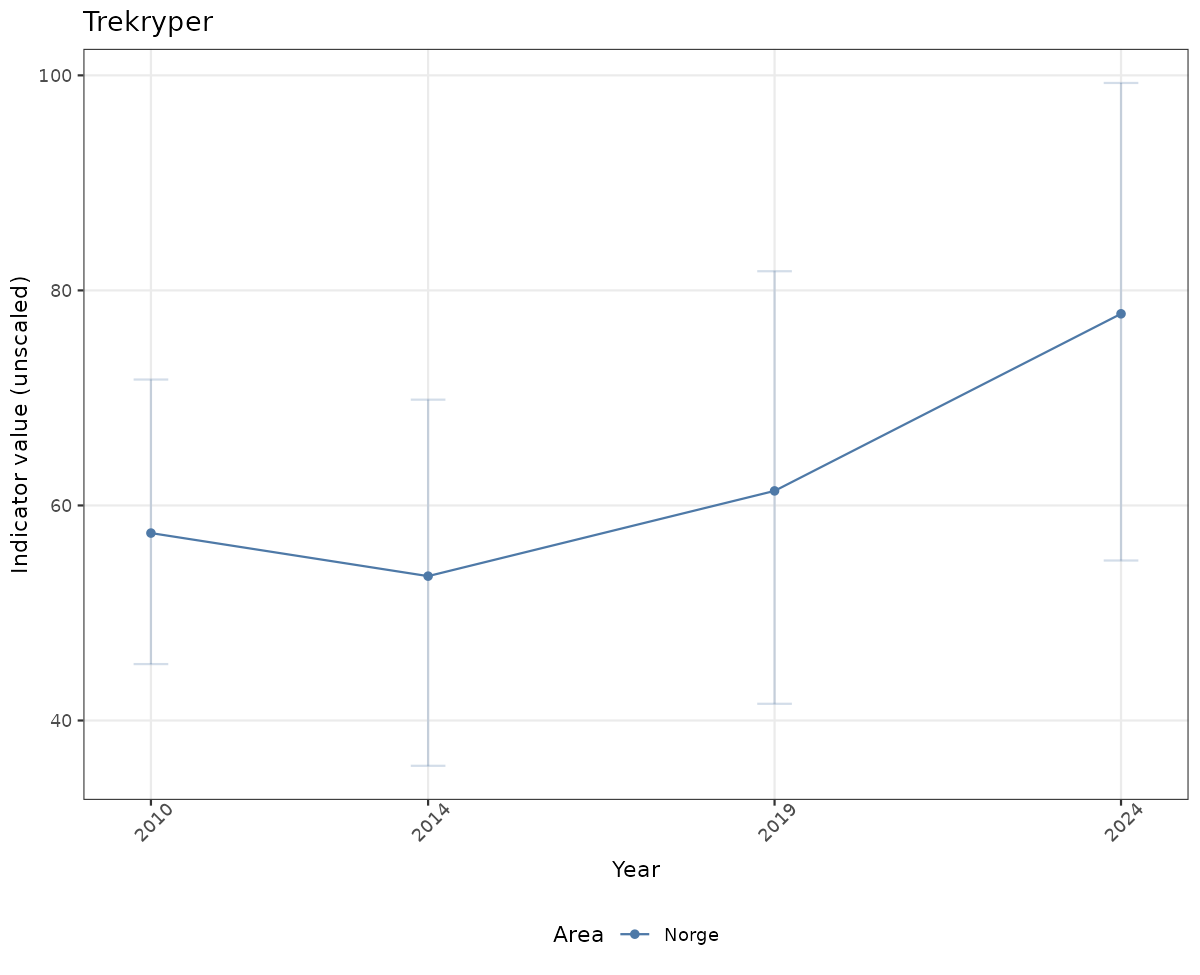
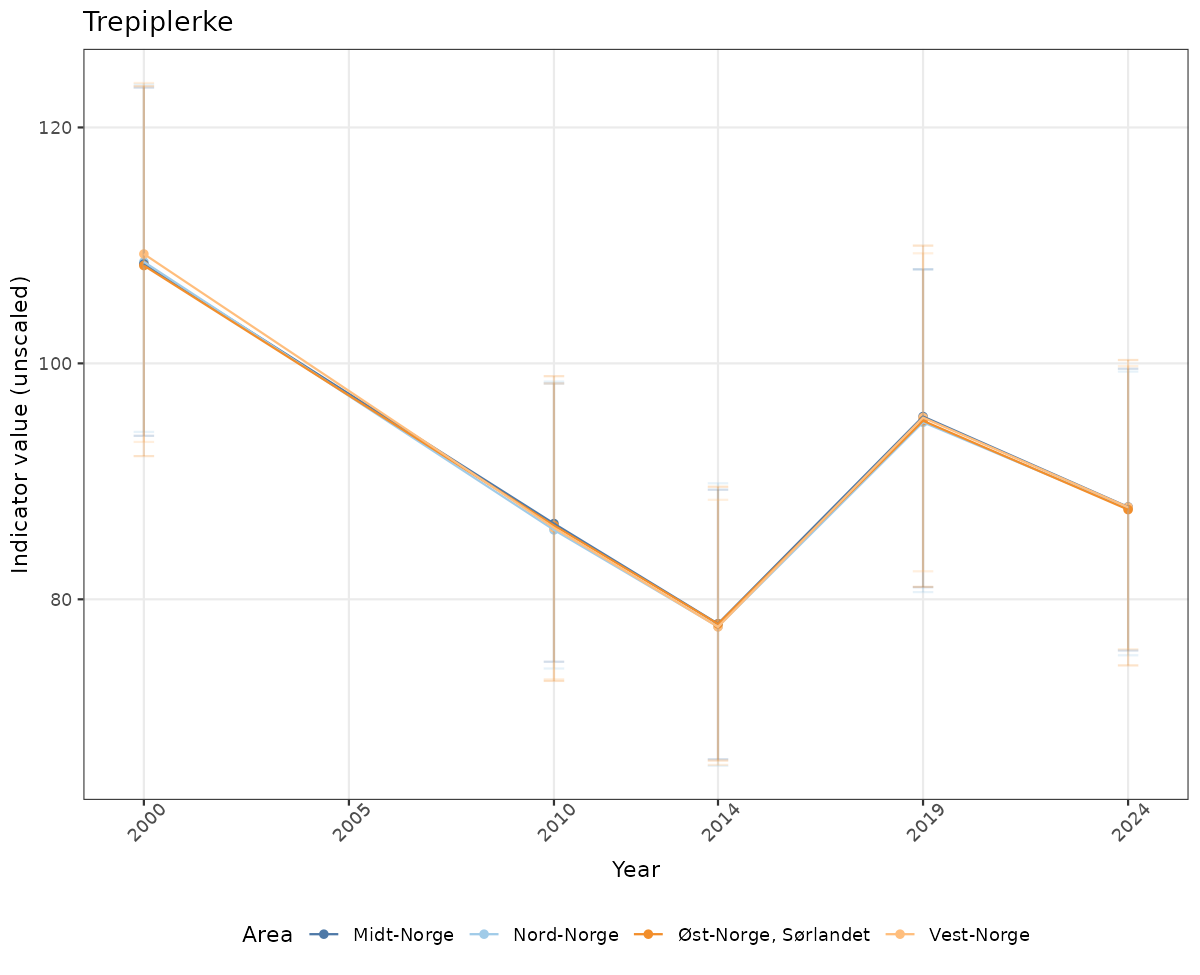
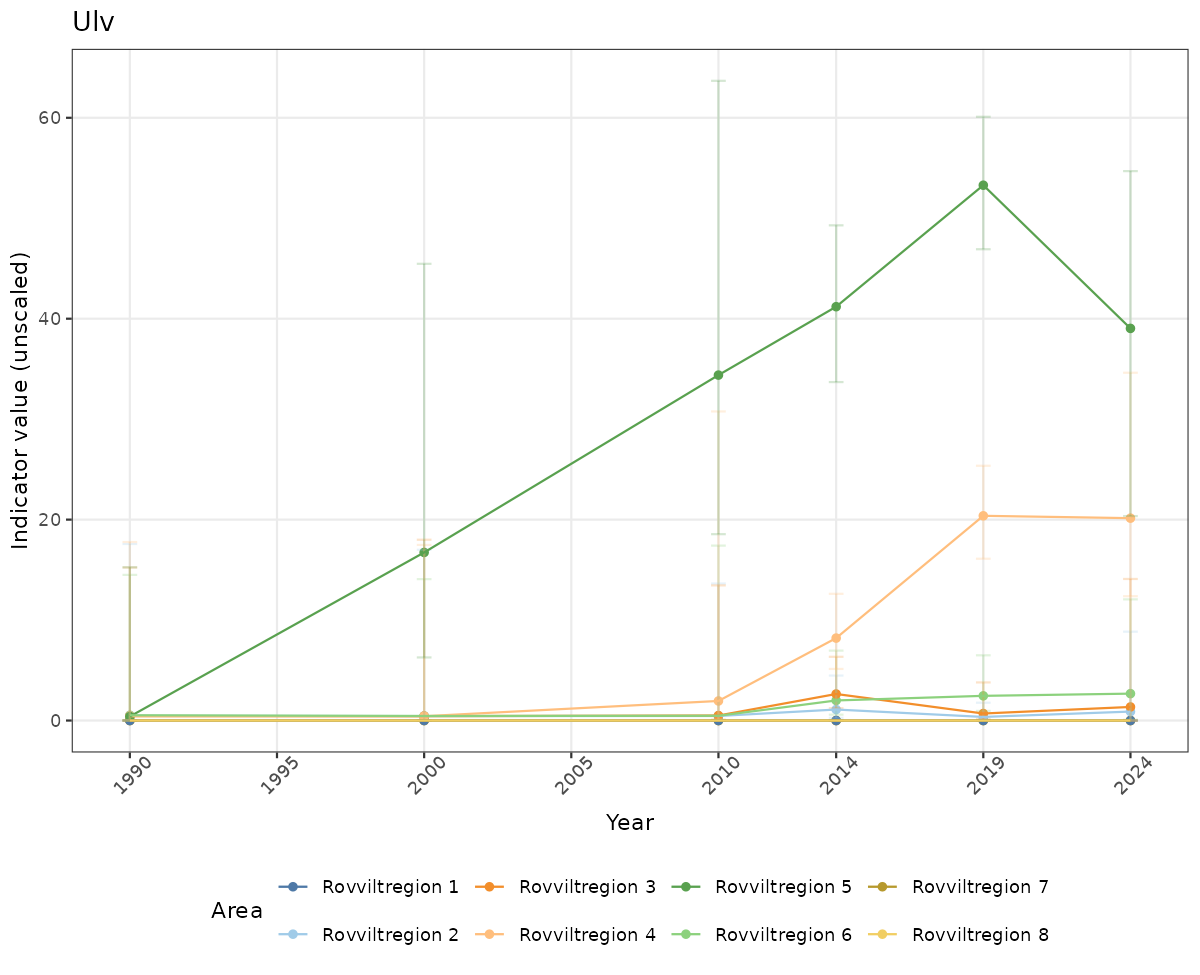
# Raw data
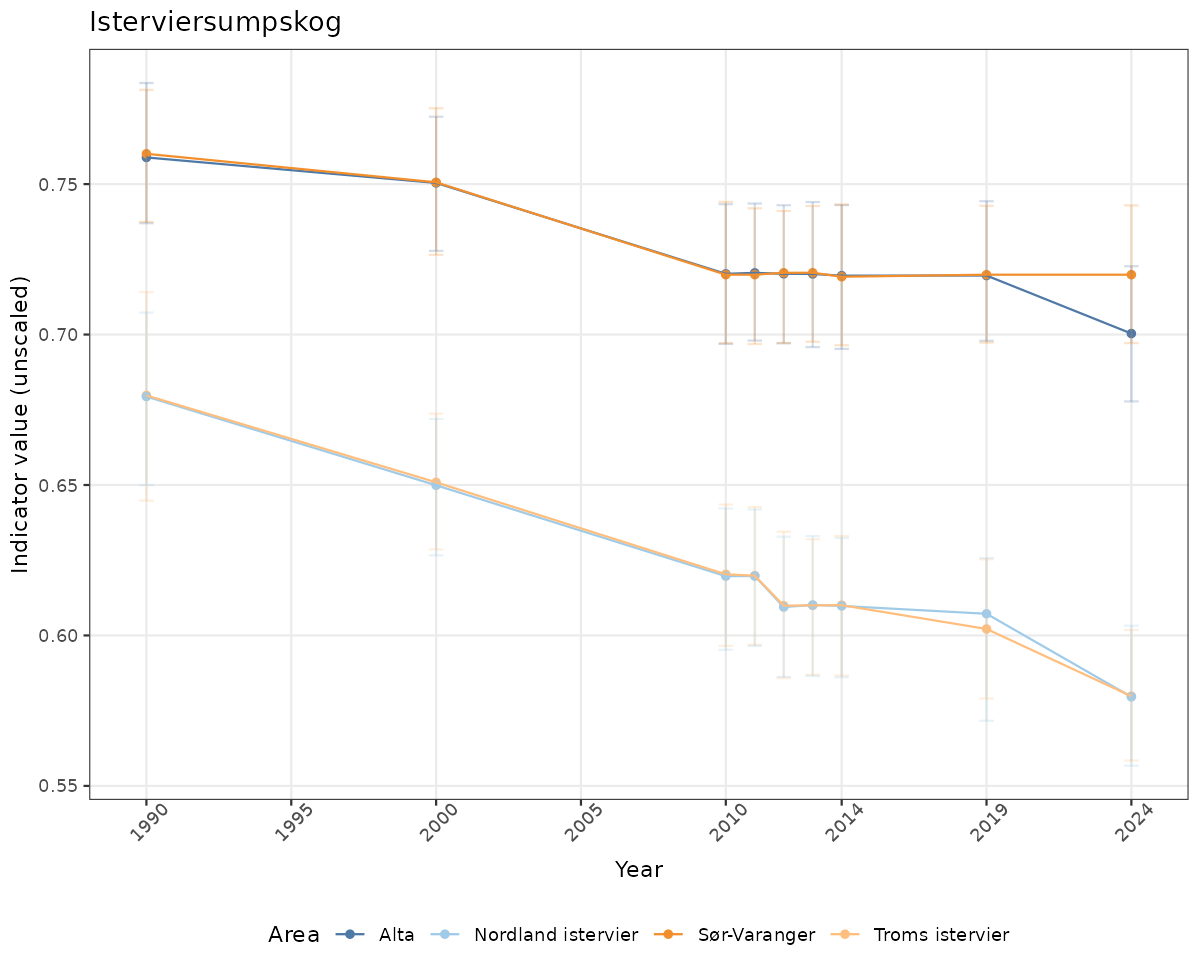
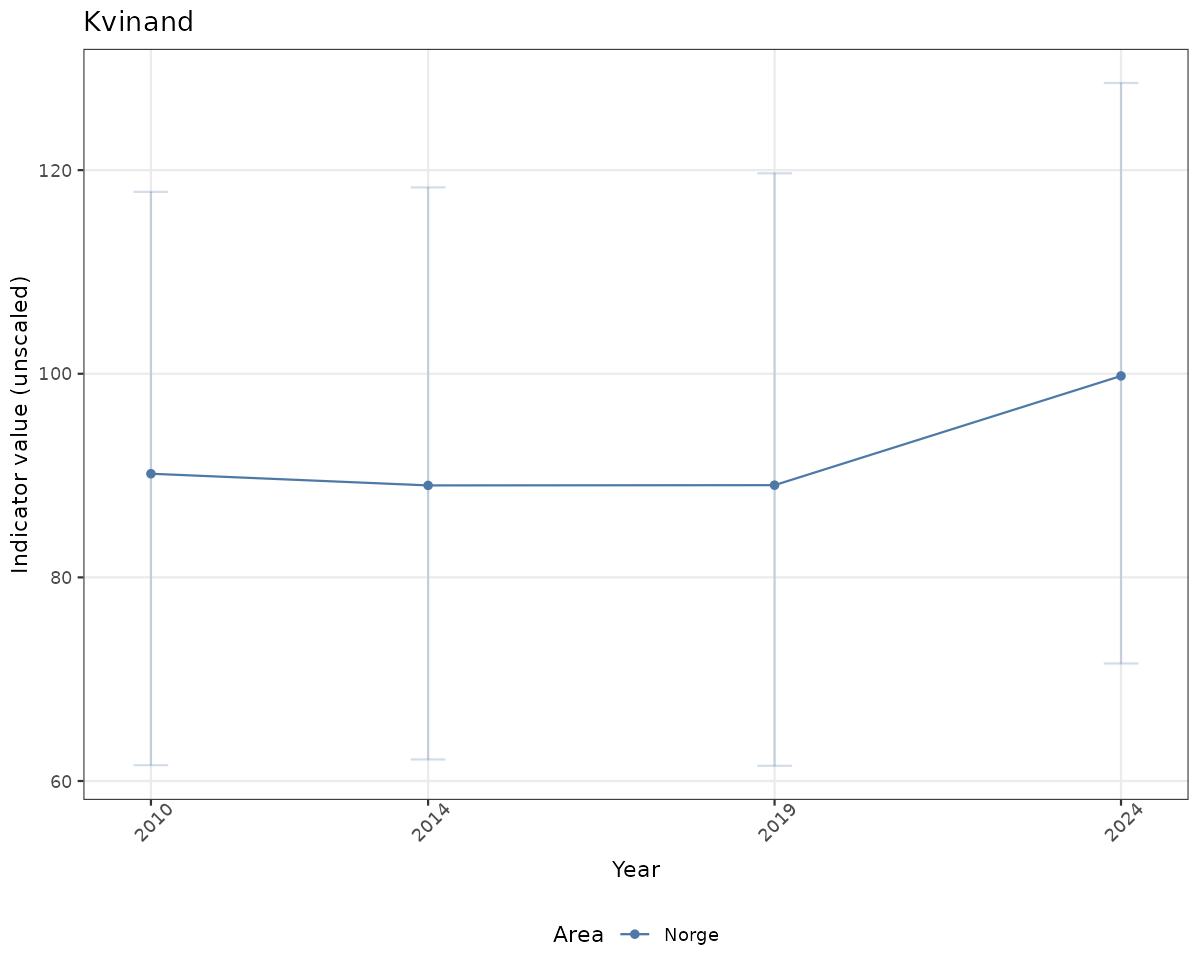
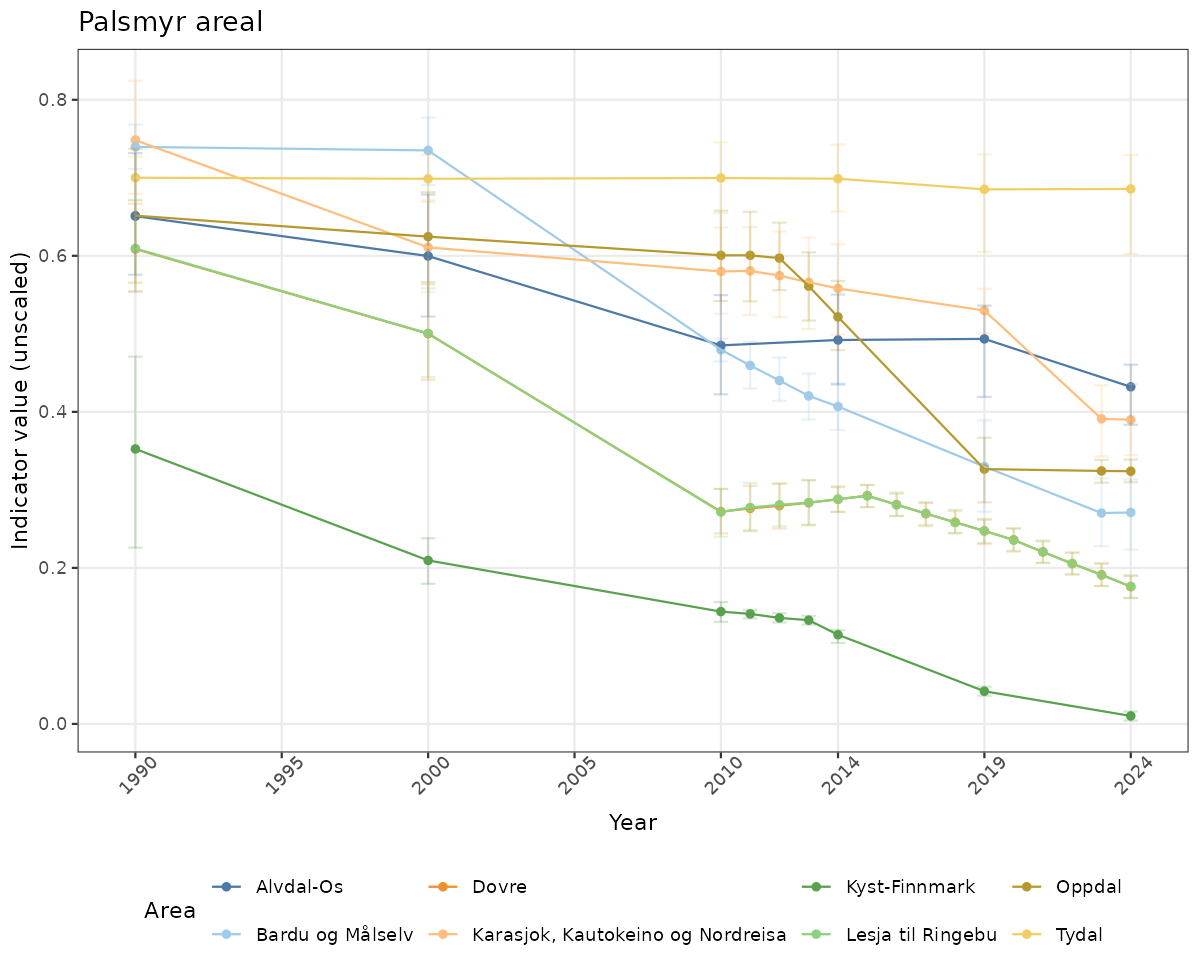
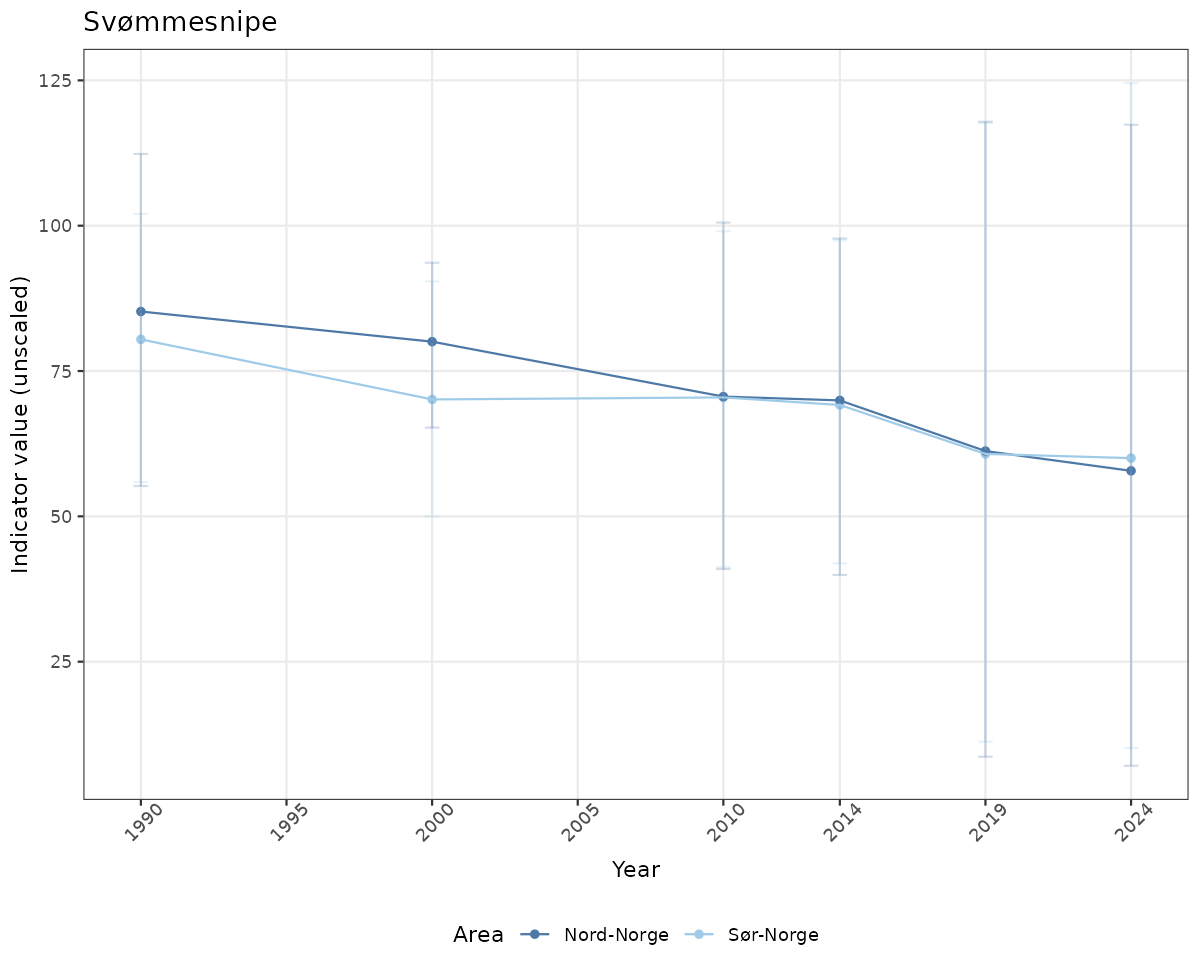
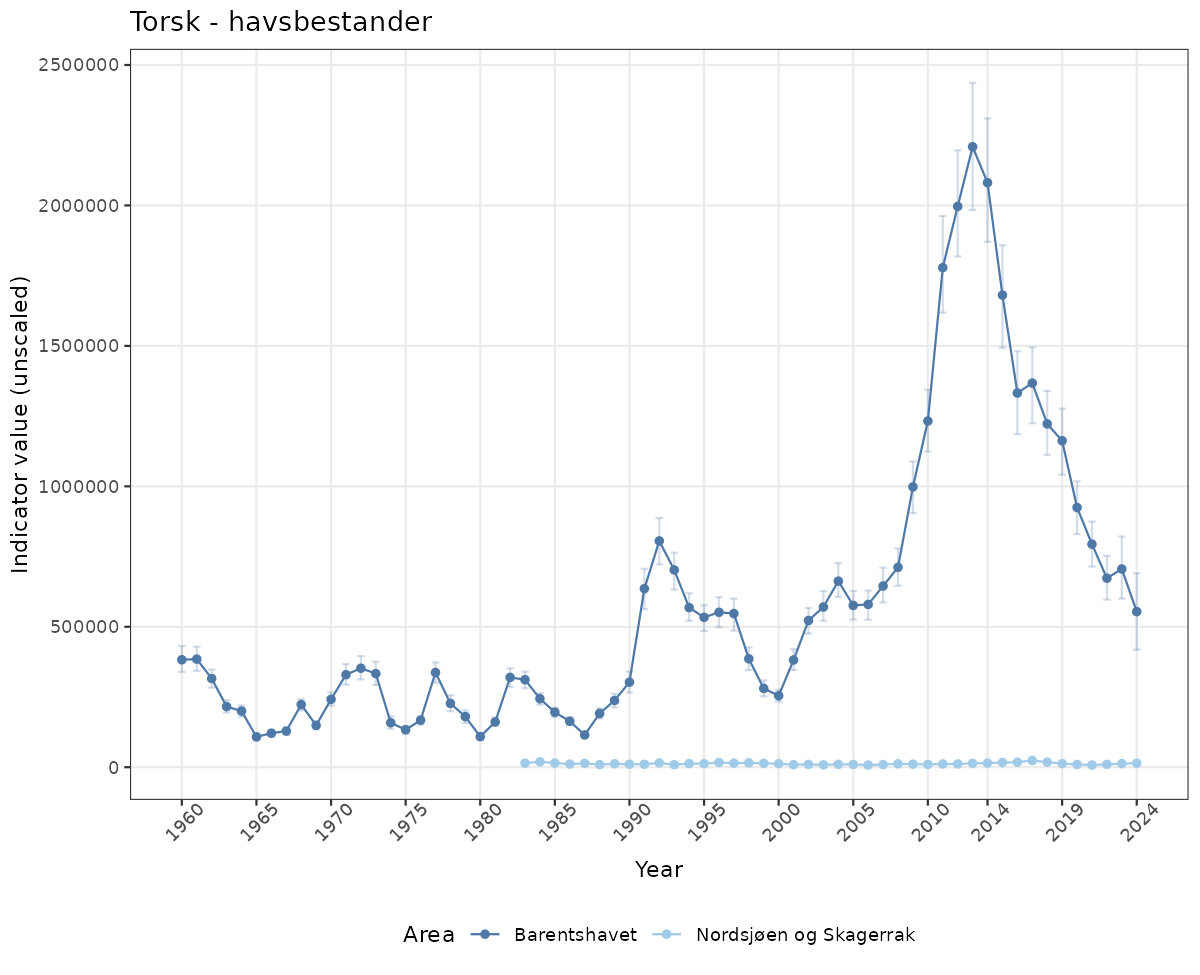
if(!manyAreas){
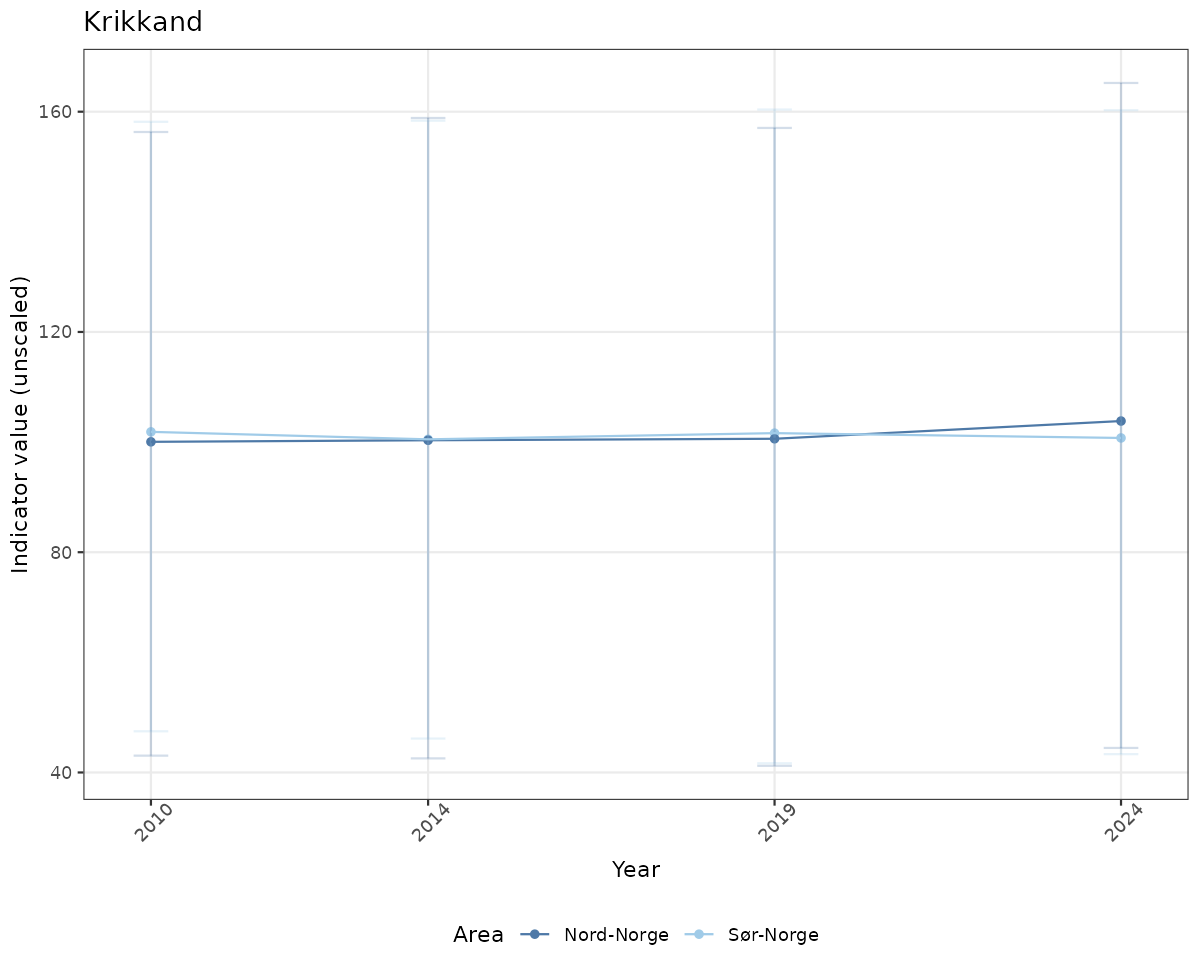
ts_raw <- ggplot(data_raw, aes(x = year_t)) +
geom_point(aes(y = median, color = ICunitName)) +
geom_errorbar(aes(ymin = q025, ymax = q975, color = ICunitName), width = 0.5, alpha = 0.25) +
geom_line(aes(y = median, color = ICunitName)) +
xlab("Year") + ylab("Indicator value (unscaled)") +
ggtitle(ind_name) +
paletteer::scale_color_paletteer_d(palette = "ggthemes::Tableau_20", name = "Area") +
scale_x_continuous(breaks = plot.yearH) +
theme_bw() +
theme(legend.position = "bottom",
panel.grid.minor = element_blank(),
axis.text.x = element_text(angle = 45, vjust = 0.8))
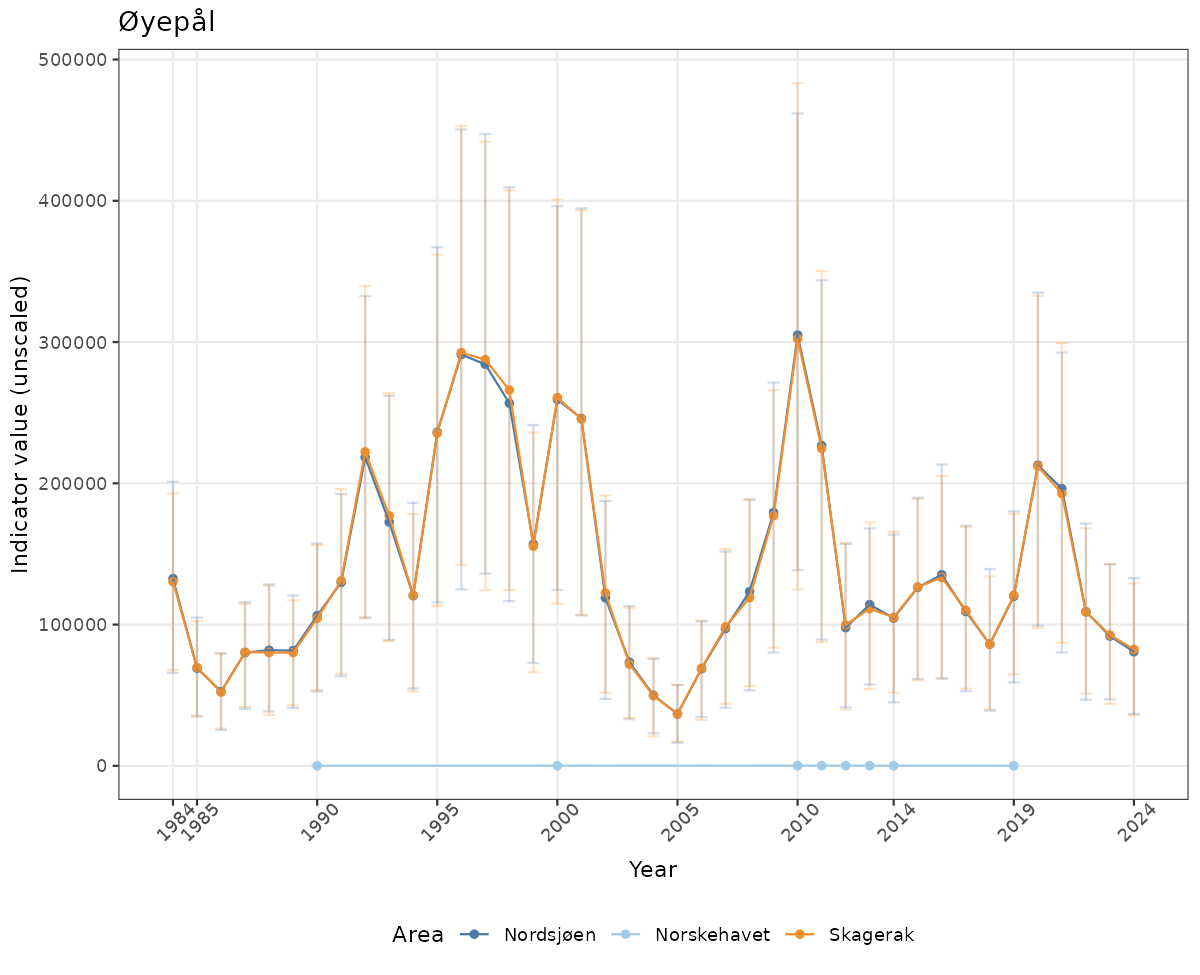
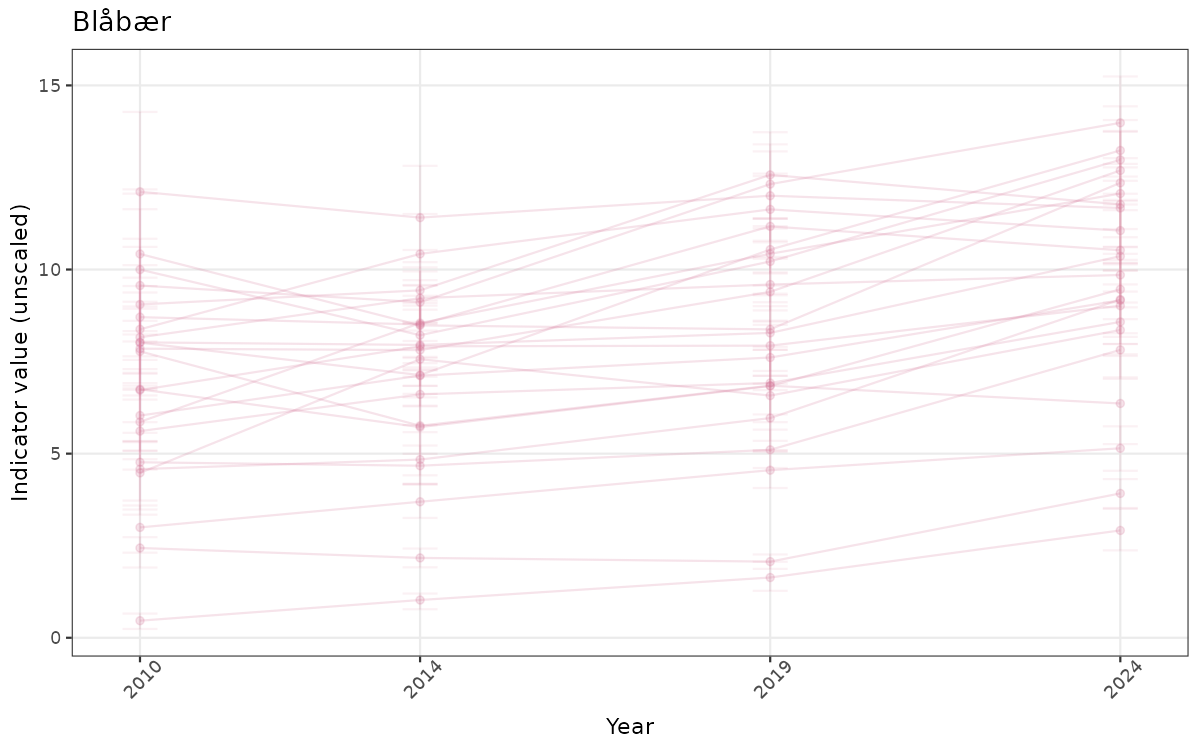
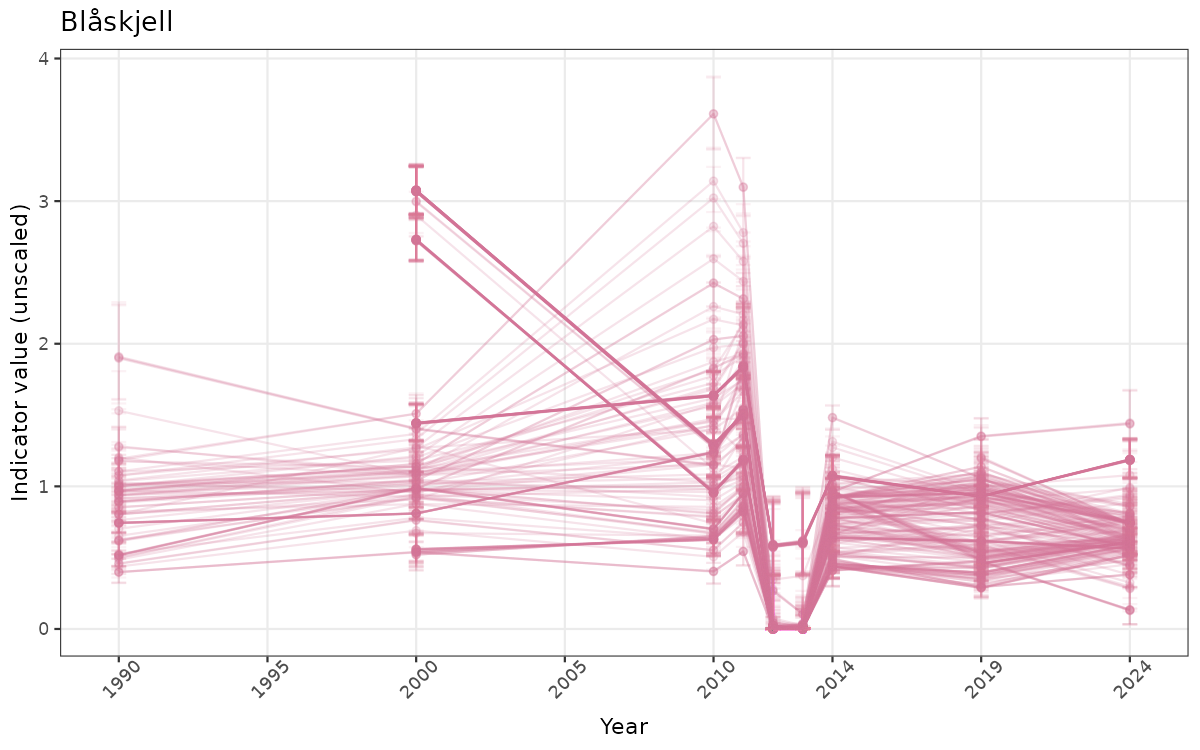
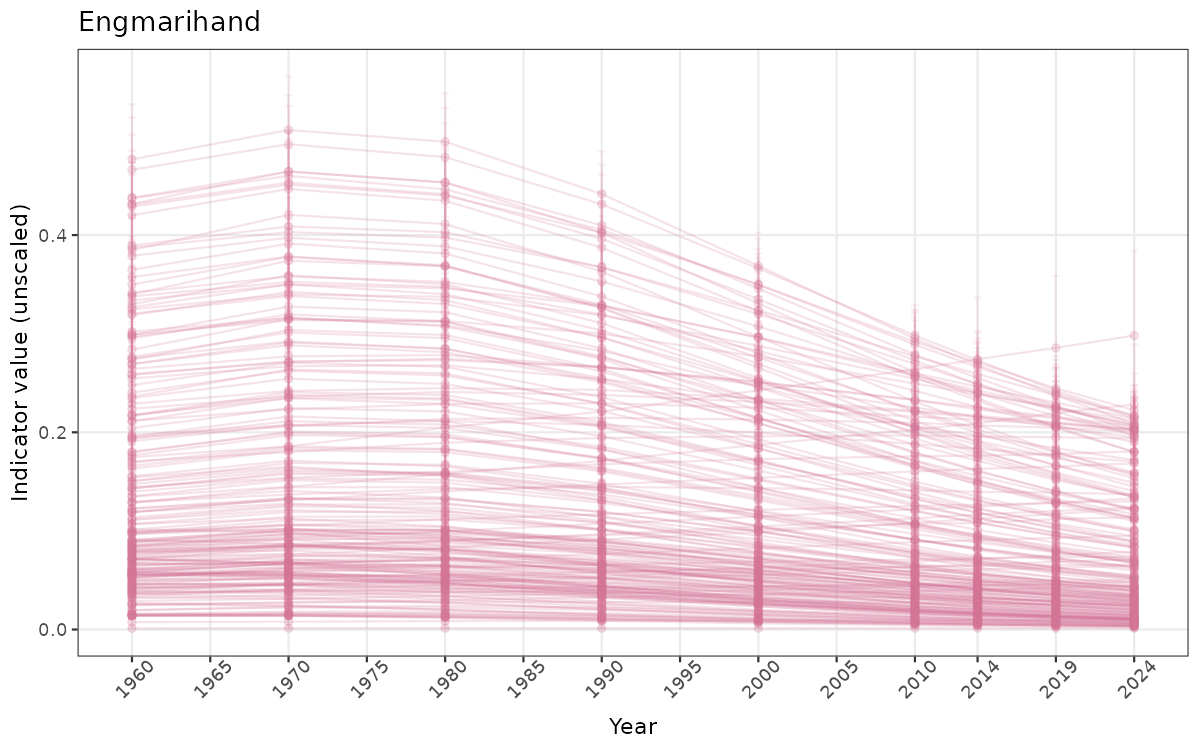
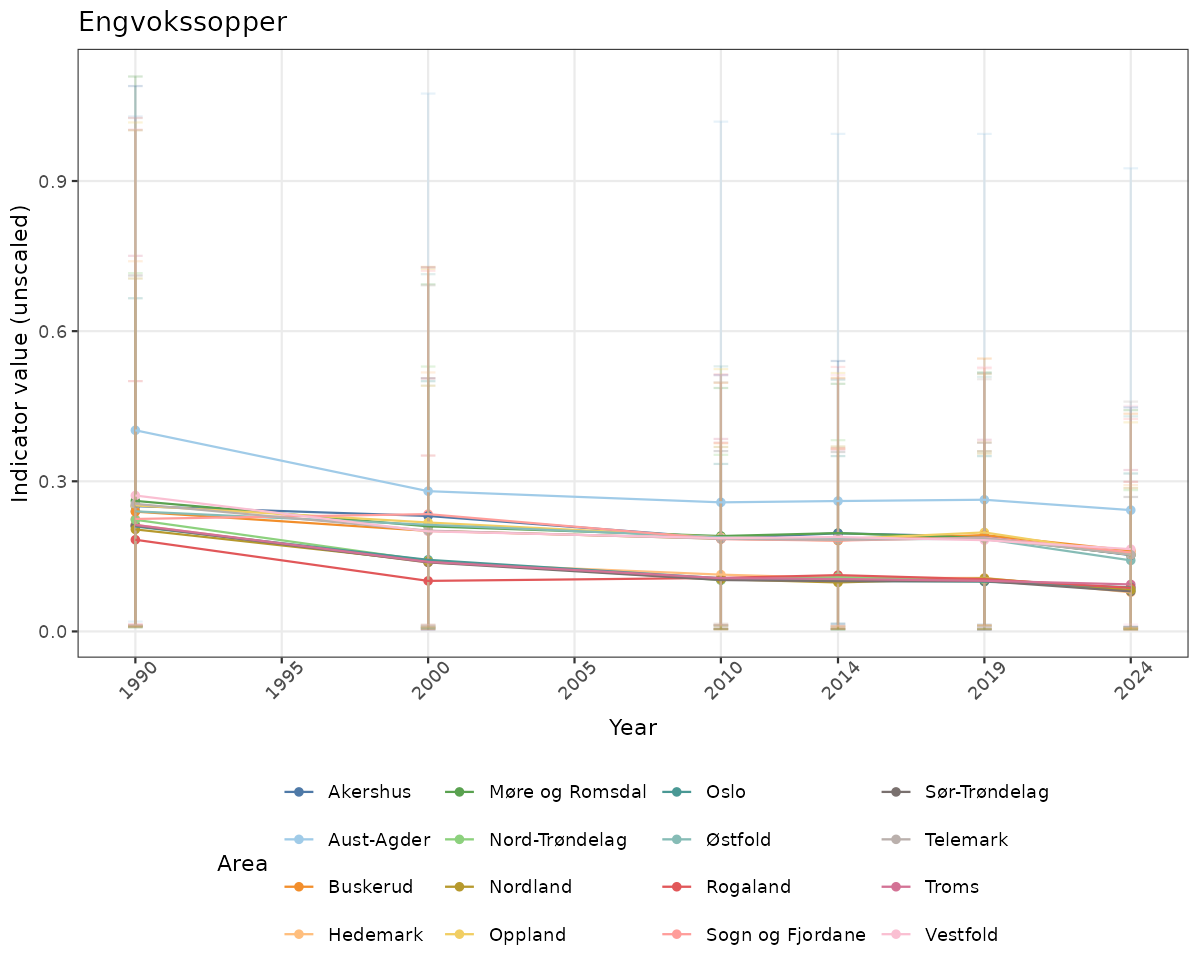
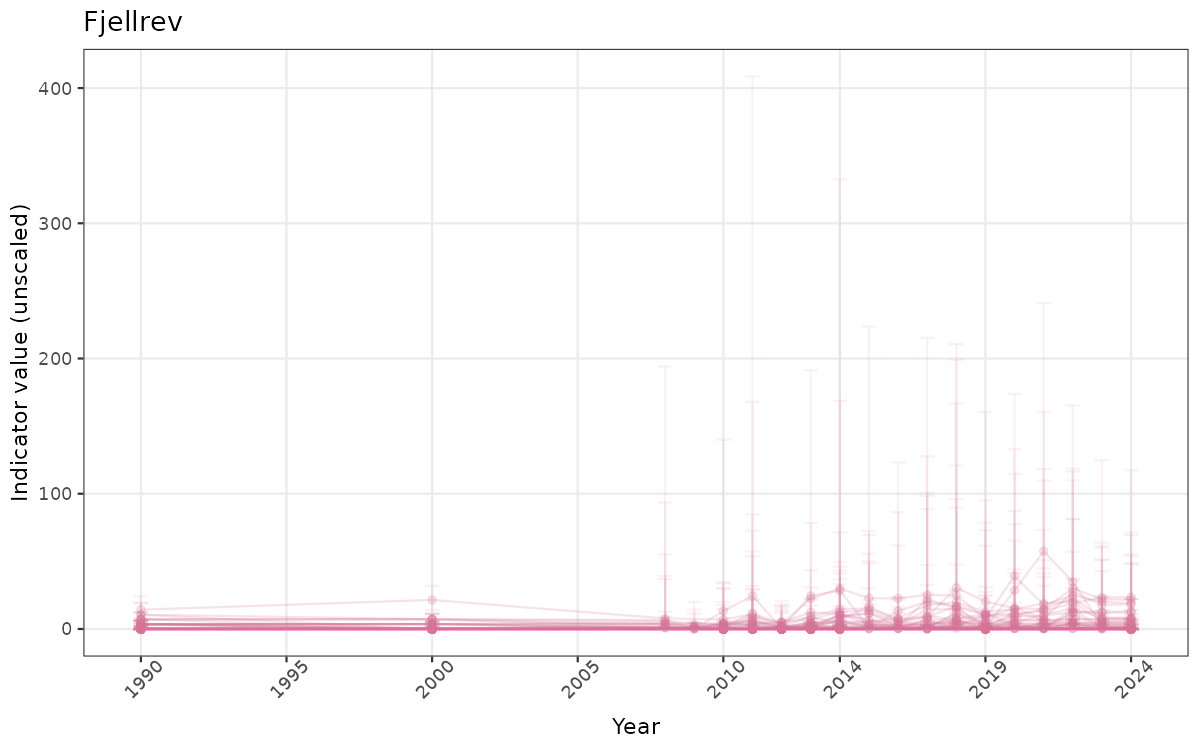
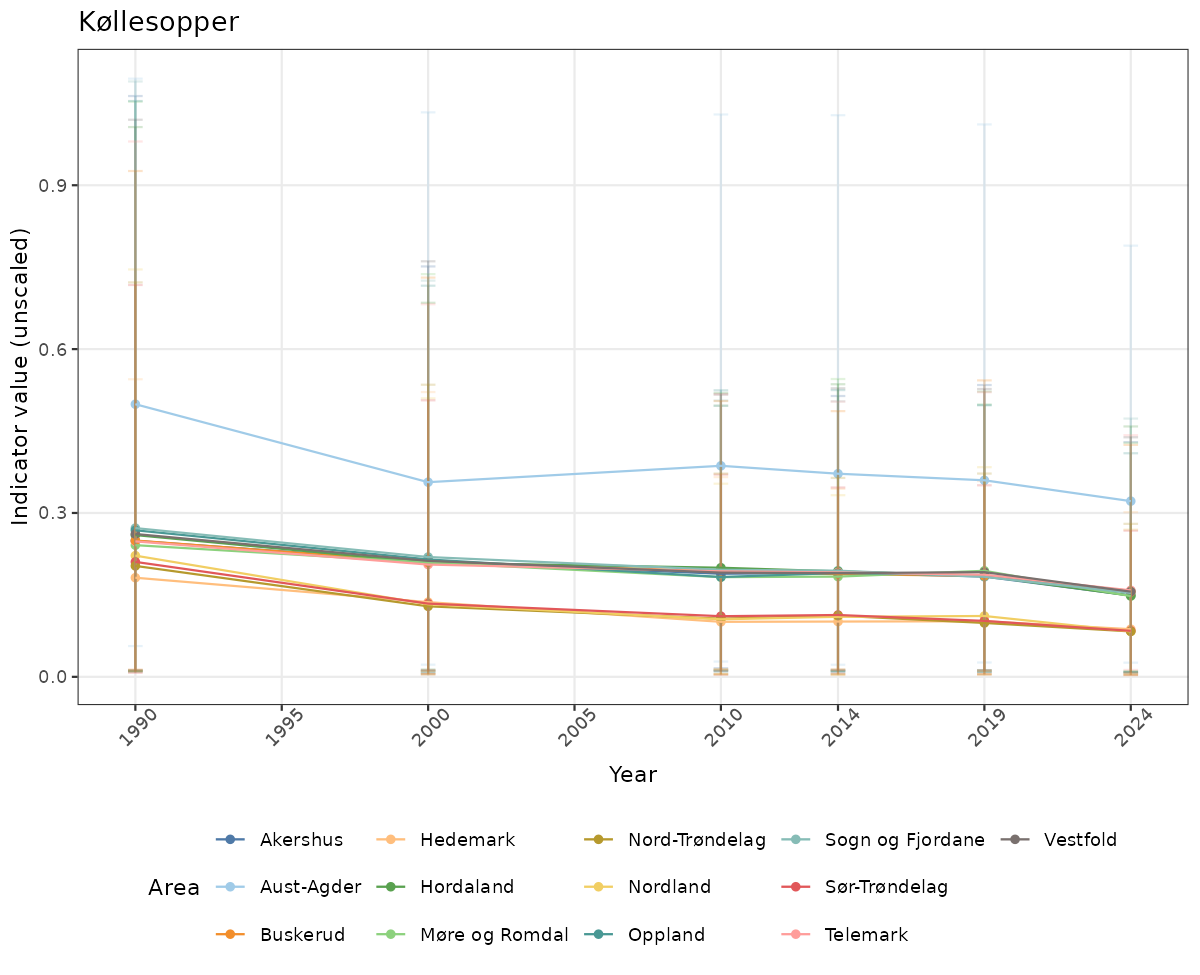
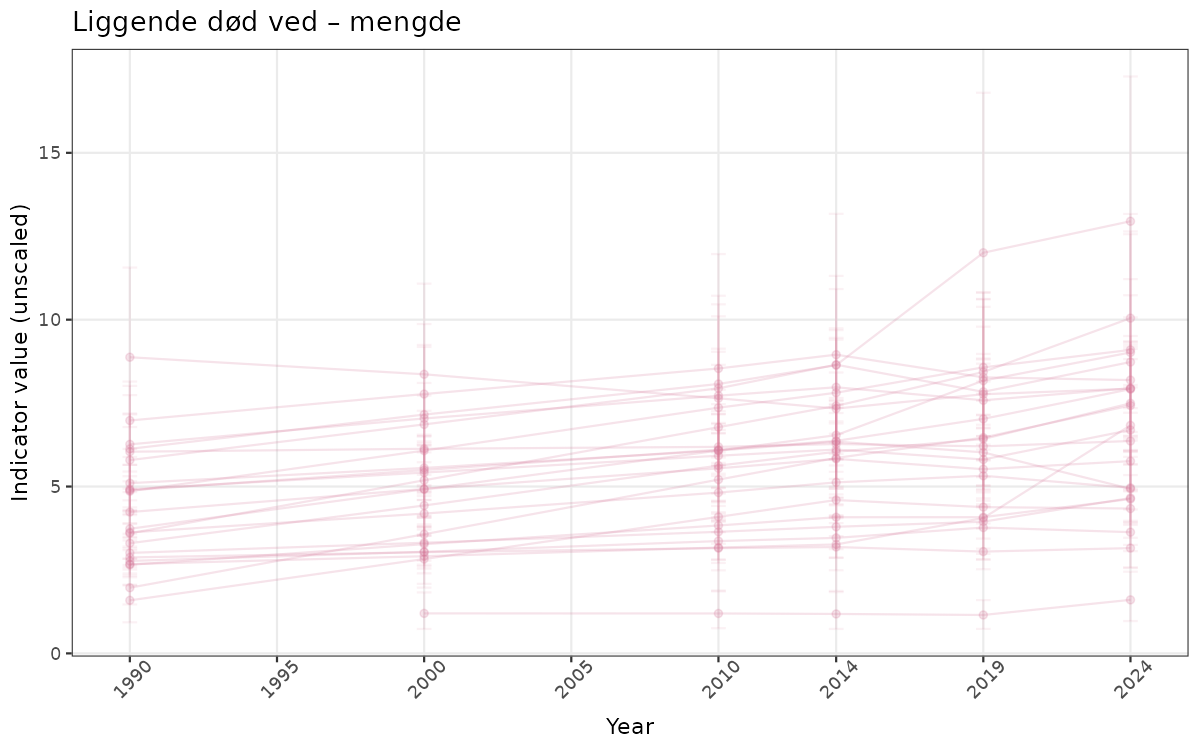
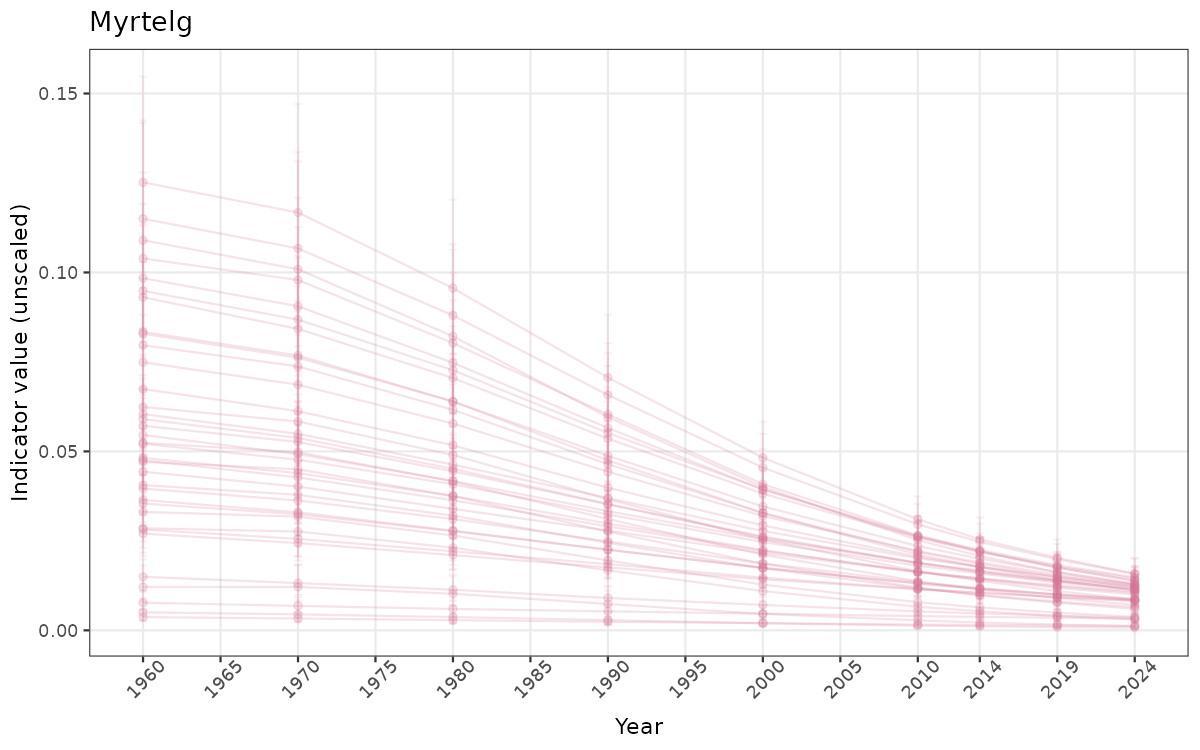
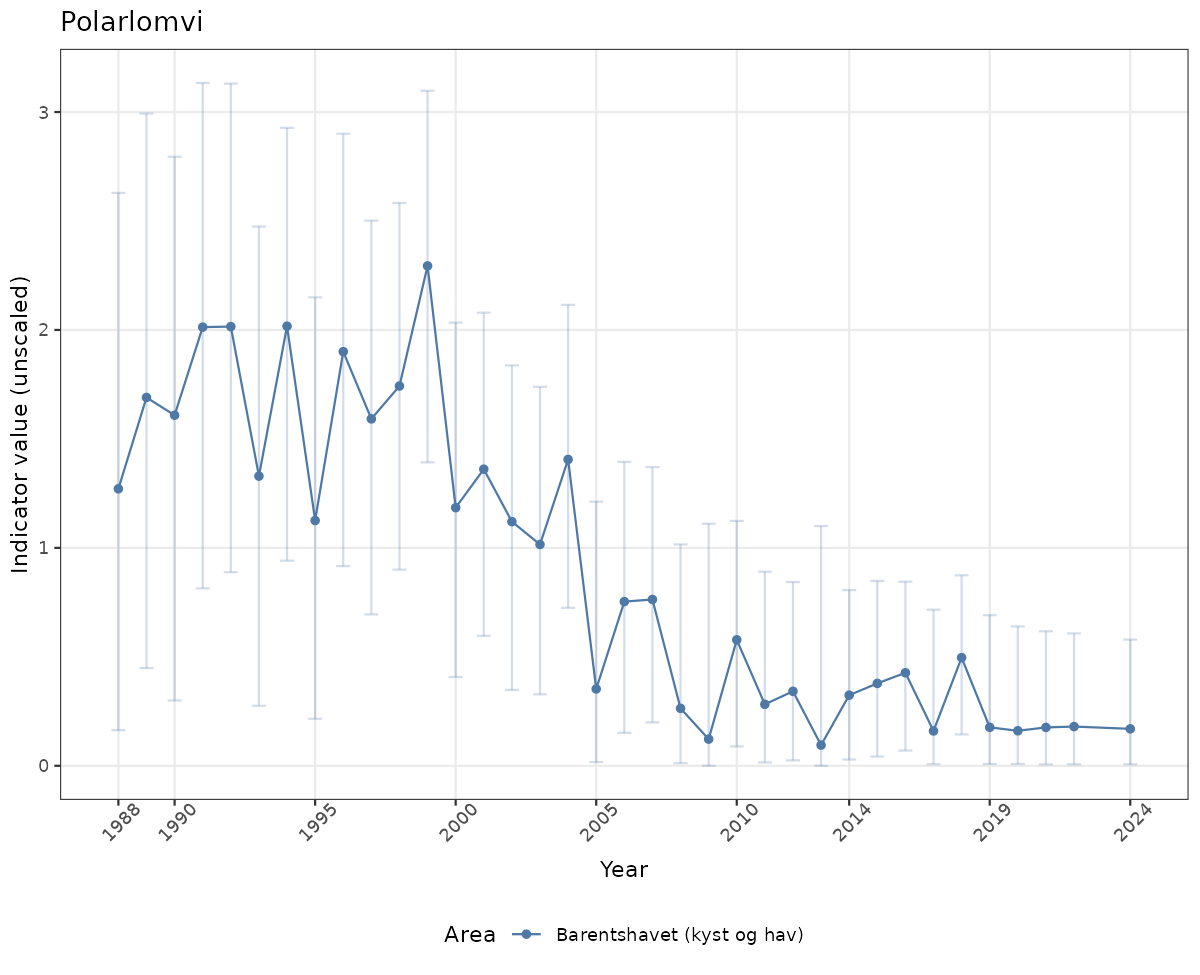
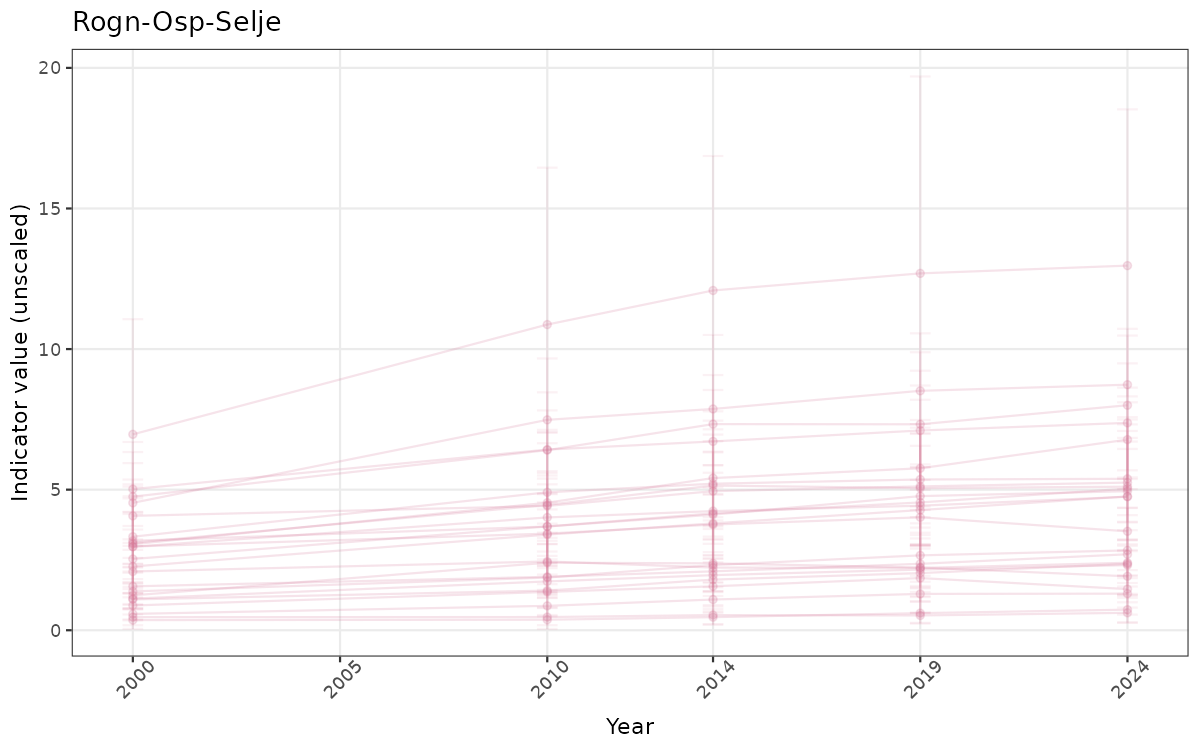
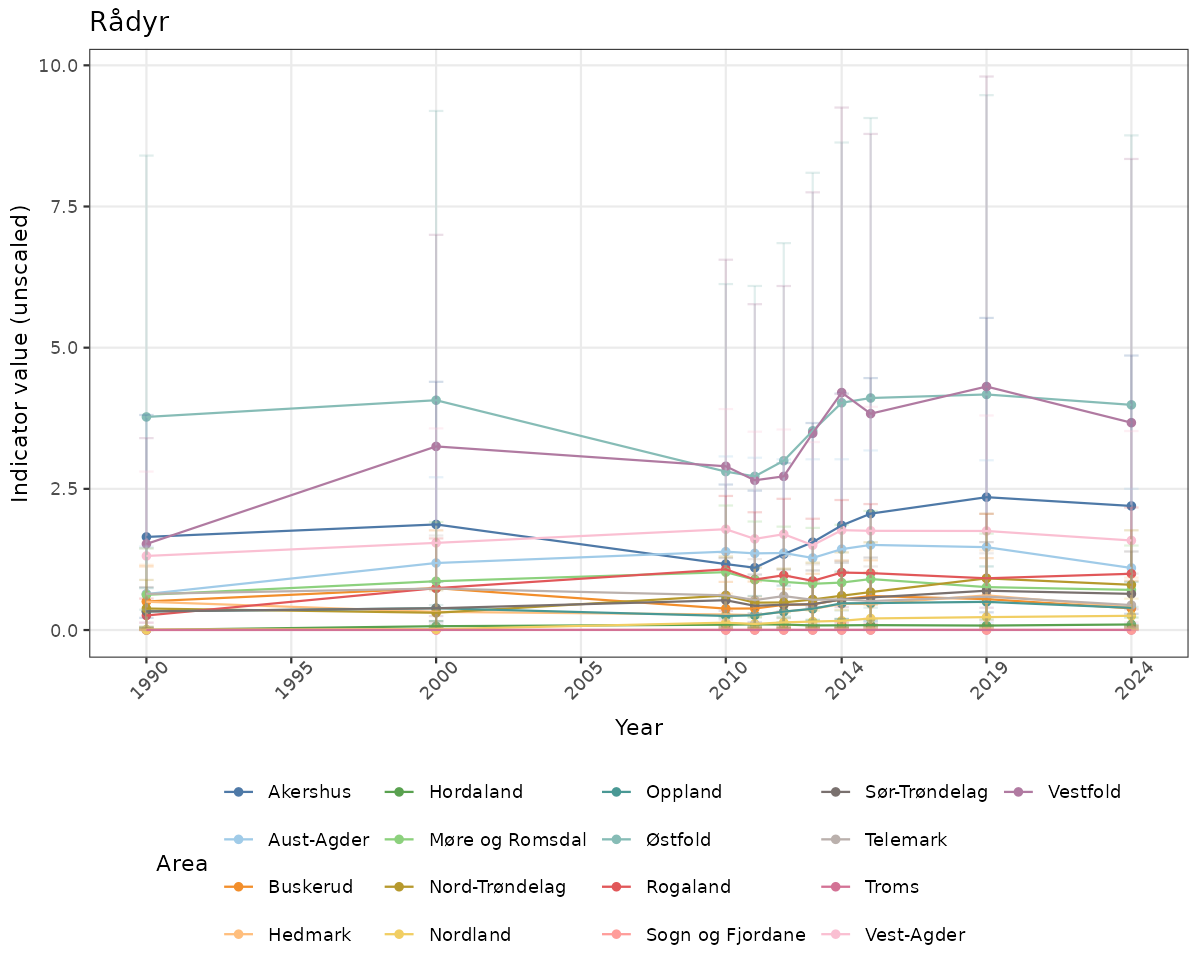
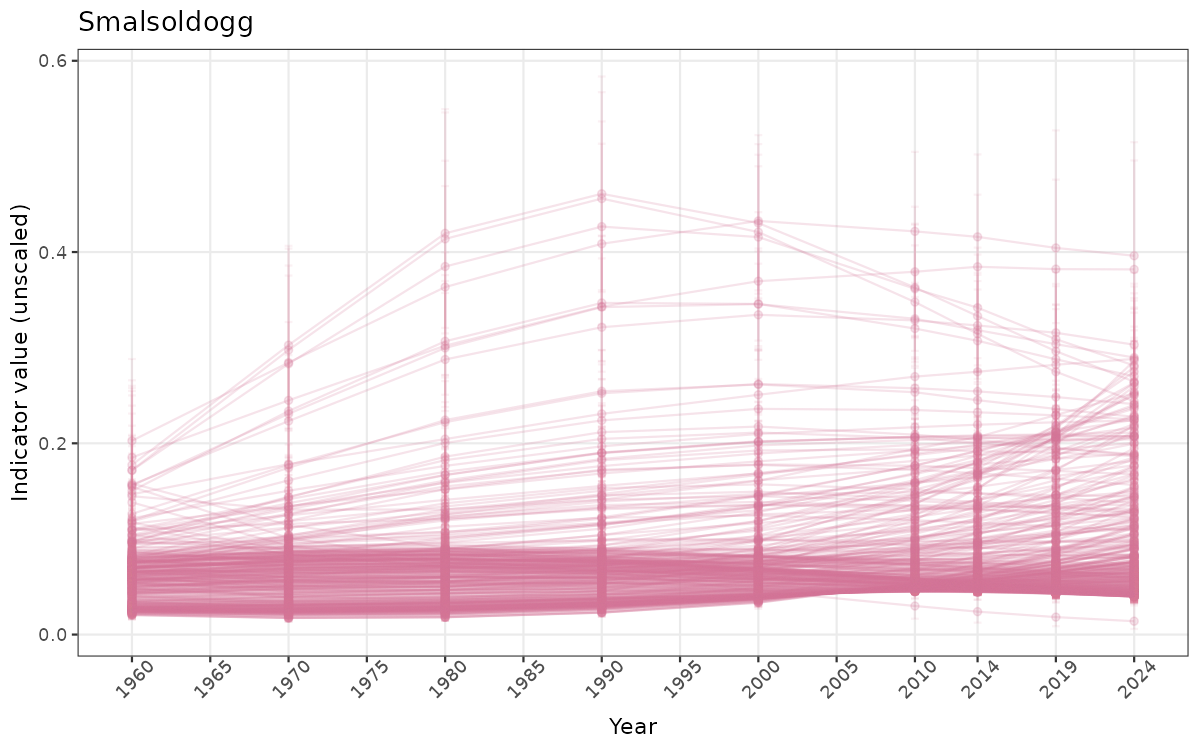
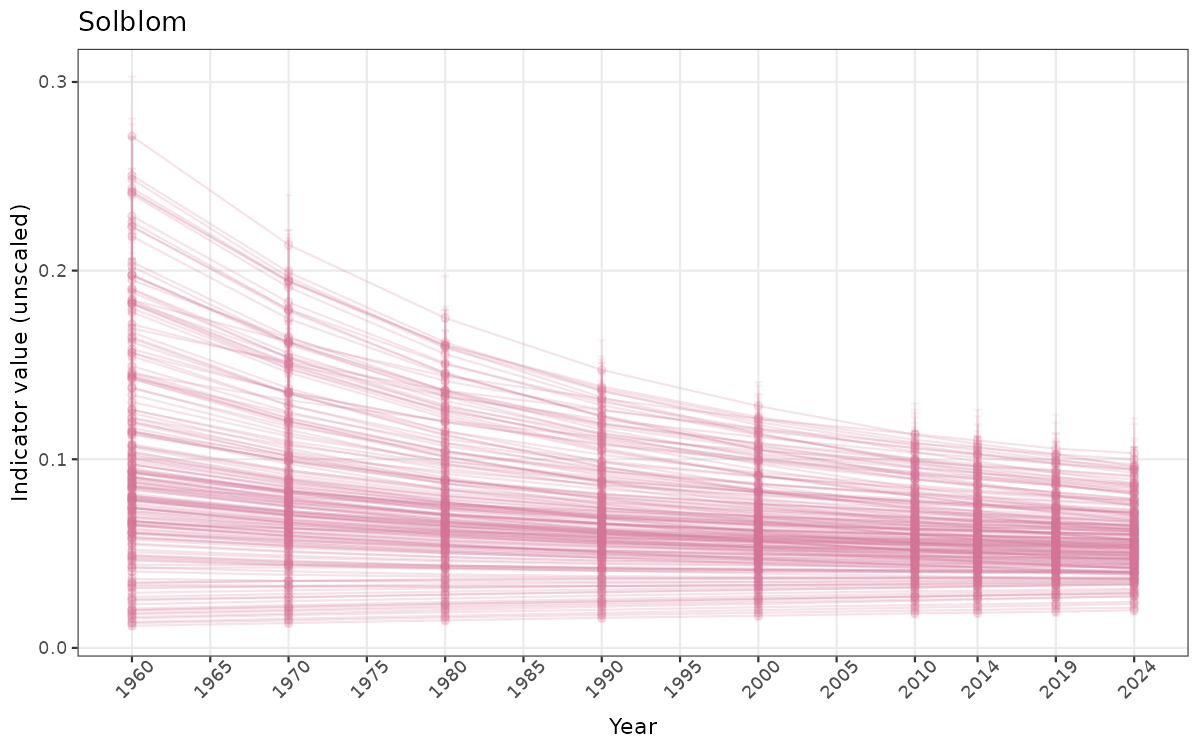
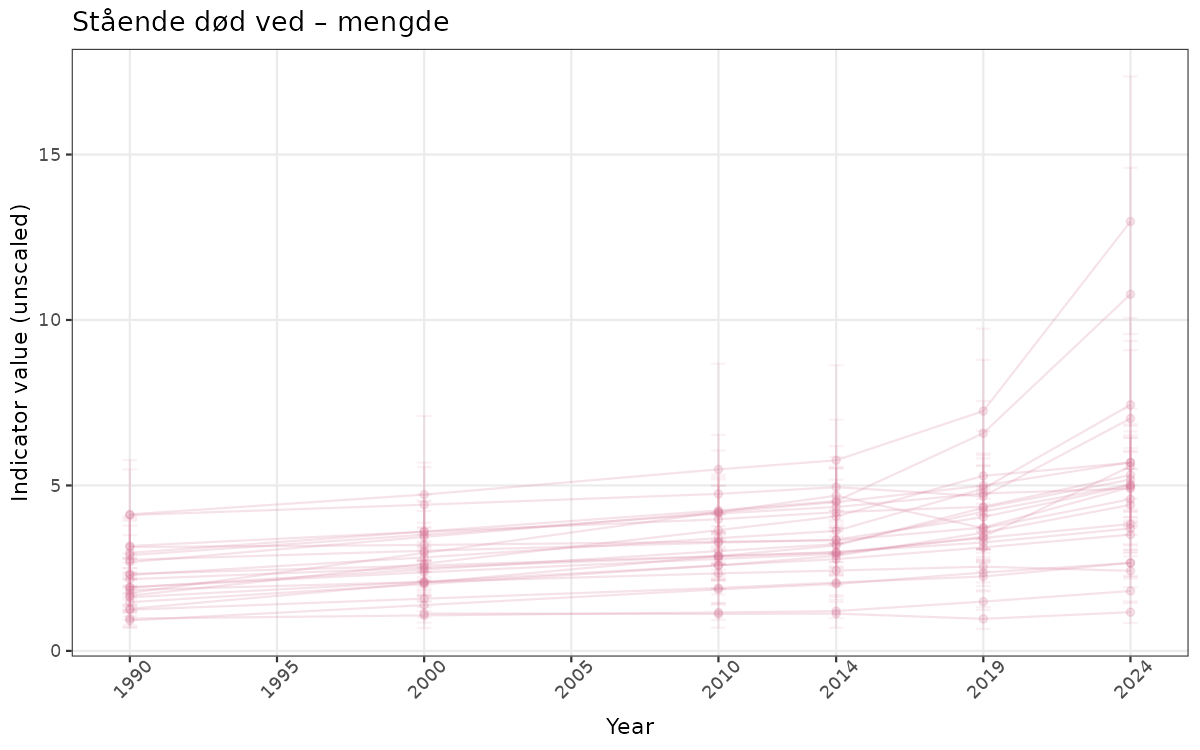
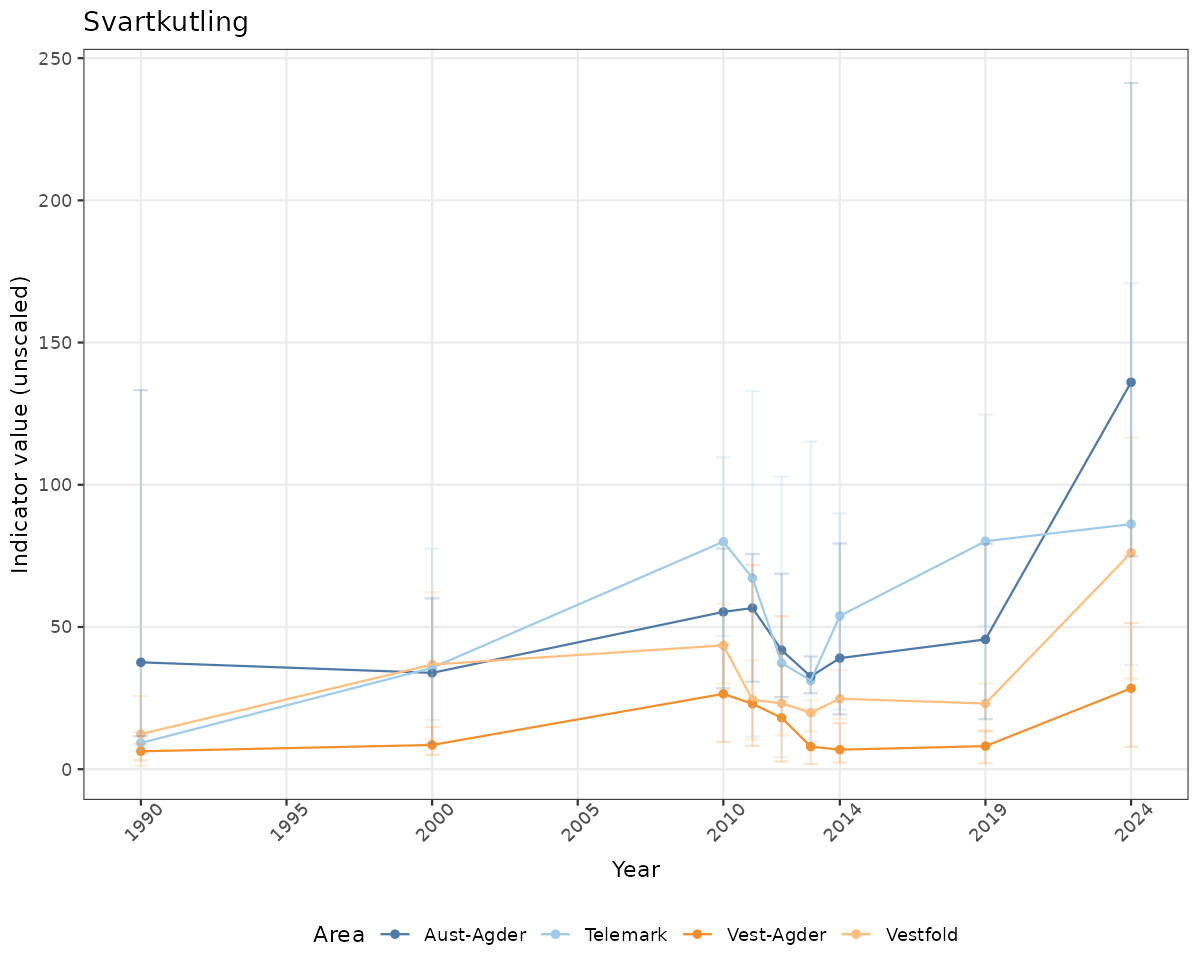
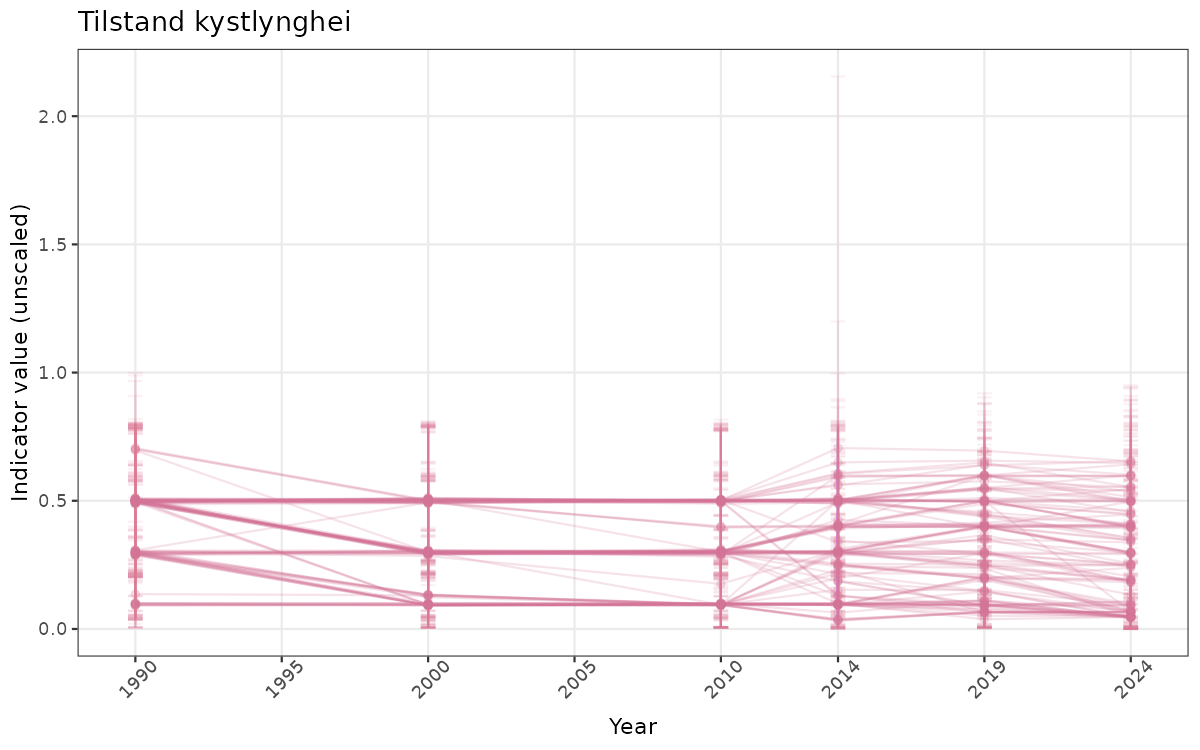
}else{
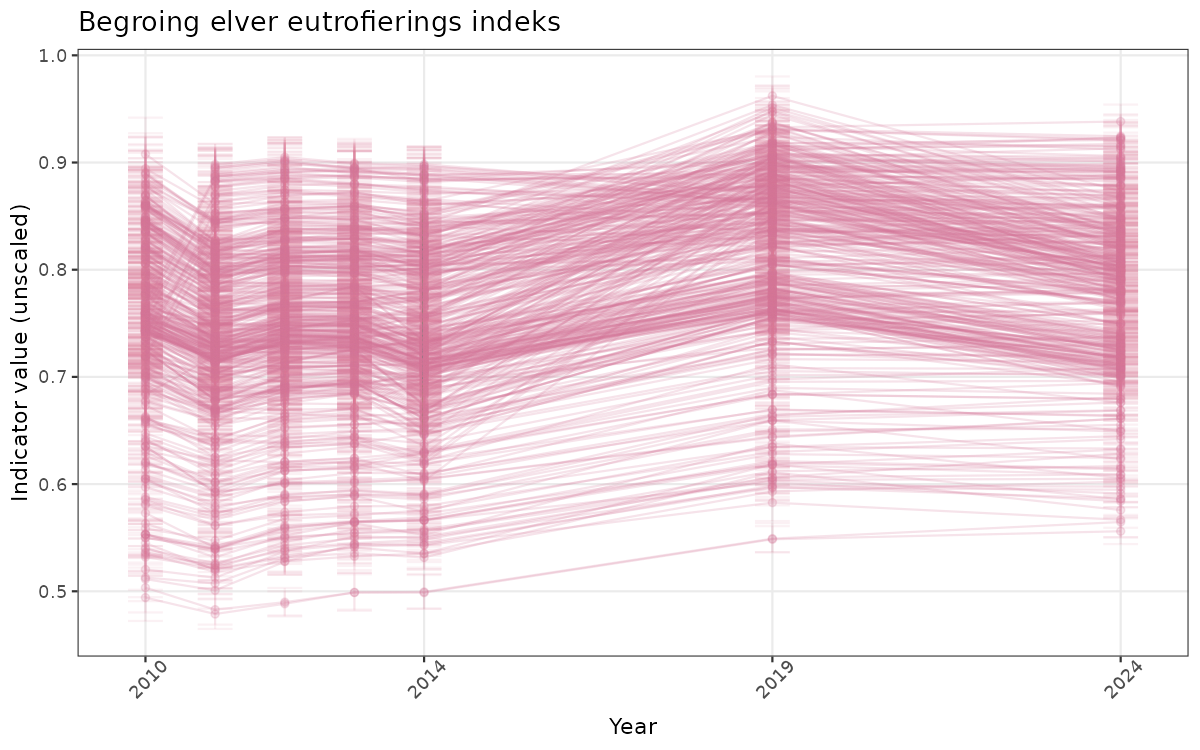
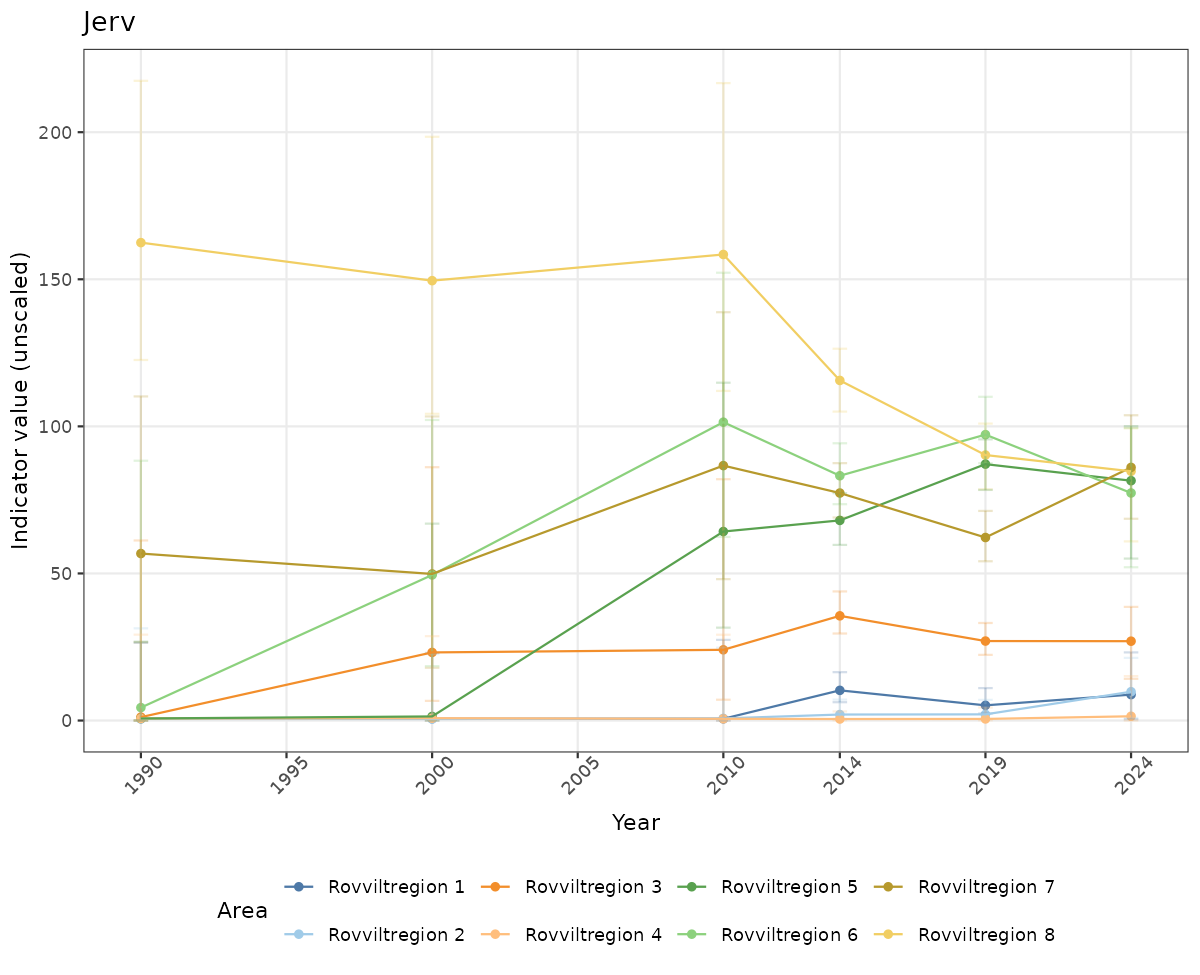
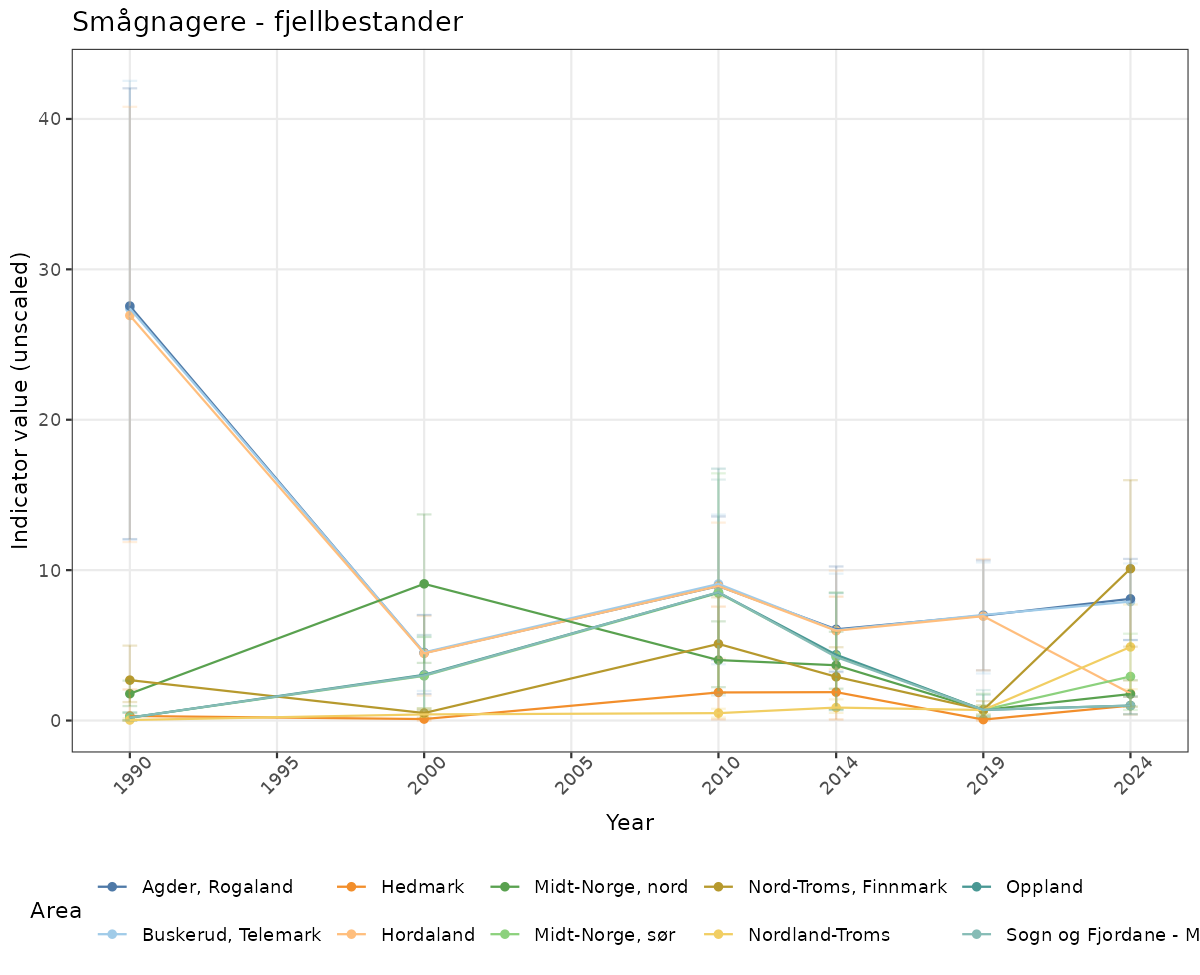
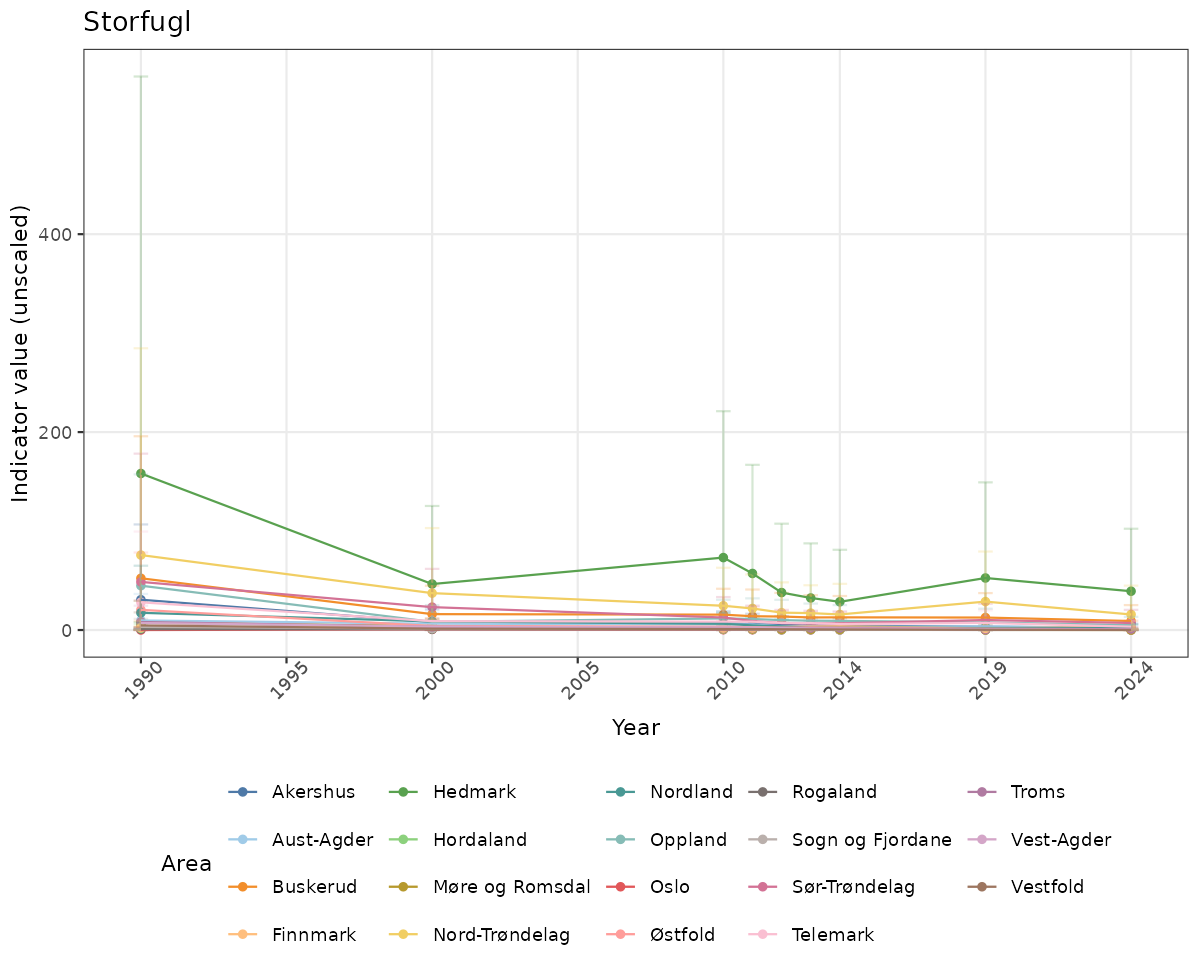
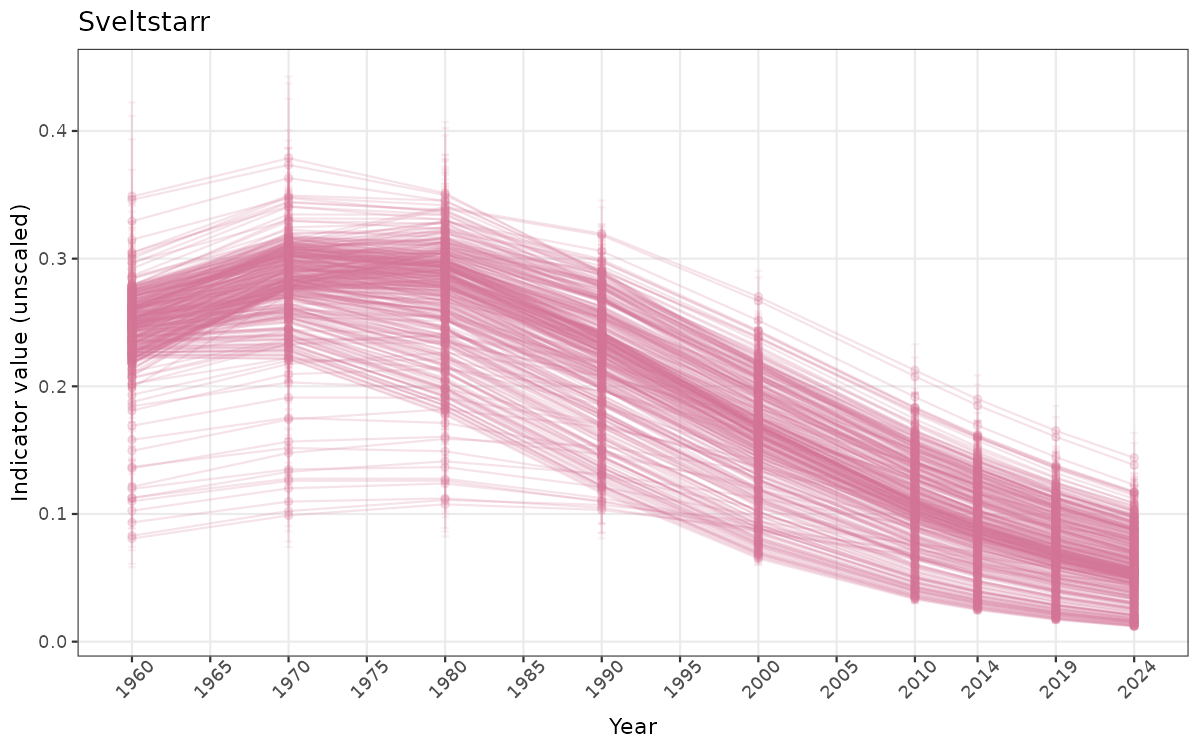
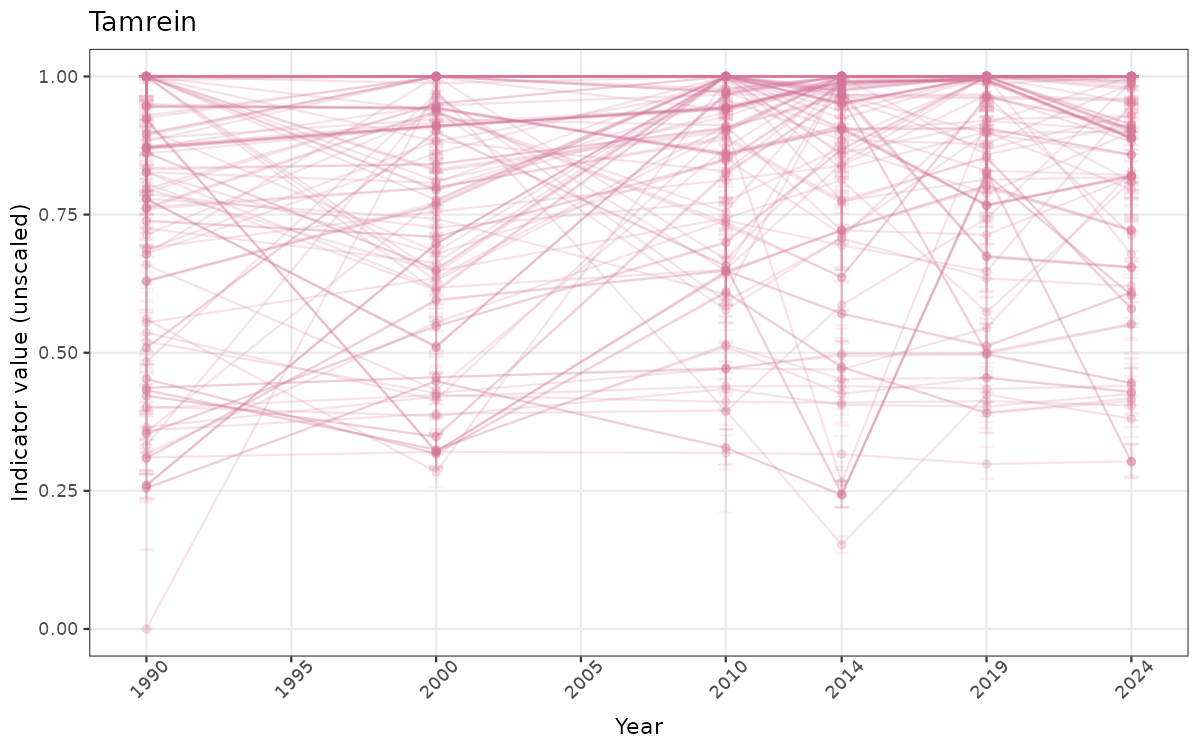
ts_raw <- ggplot(data_raw, aes(x = year_t)) +
geom_point(aes(y = median, group = ICunitName), color = plot.col, alpha = 0.2) +
geom_errorbar(aes(ymin = q025, ymax = q975, group = ICunitName), width = 0.5, color = plot.col, alpha = 0.1) +
geom_line(aes(y = median, group = ICunitName), color = plot.col, alpha = 0.2) +
xlab("Year") + ylab("Indicator value (unscaled)") +
ggtitle(ind_name) +
scale_x_continuous(breaks = plot.yearH) +
theme_bw() +
theme(legend.position = "bottom",
panel.grid.minor = element_blank(),
axis.text.x = element_text(angle = 45, vjust = 0.8))
}
TS_raw[[i]] <- ts_raw
ggsave(file = paste0(savePath_i, "/TimeSeries_raw.svg"), plot = ts_raw, width = 8, height = ifelse(manyAreas, 5, 6.5))
ggsave(file = paste0(savePath_i, "/TimeSeries_raw.pdf"), plot = ts_raw, width = 8, height = ifelse(manyAreas, 5, 6.5))
ggsave(file = paste0(savePath_i, "/TimeSeries_raw.png"), plot = ts_raw,width = 8, height = ifelse(manyAreas, 5, 6.5), dpi = 150)
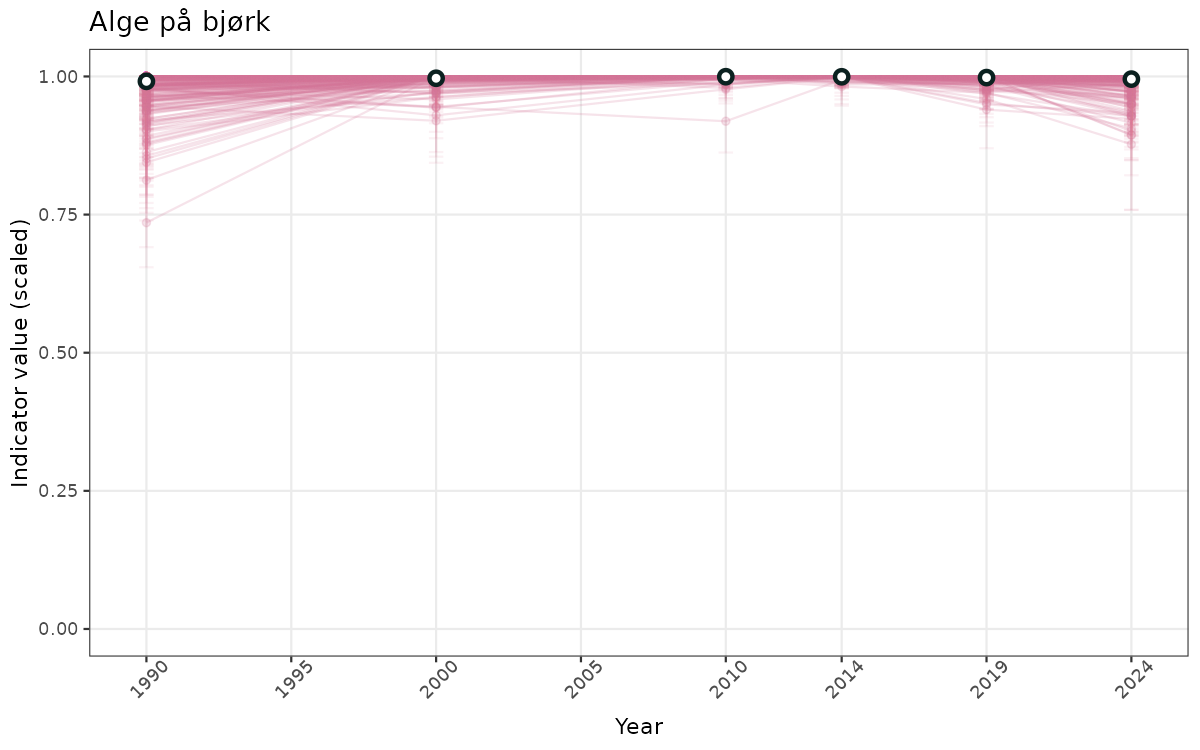
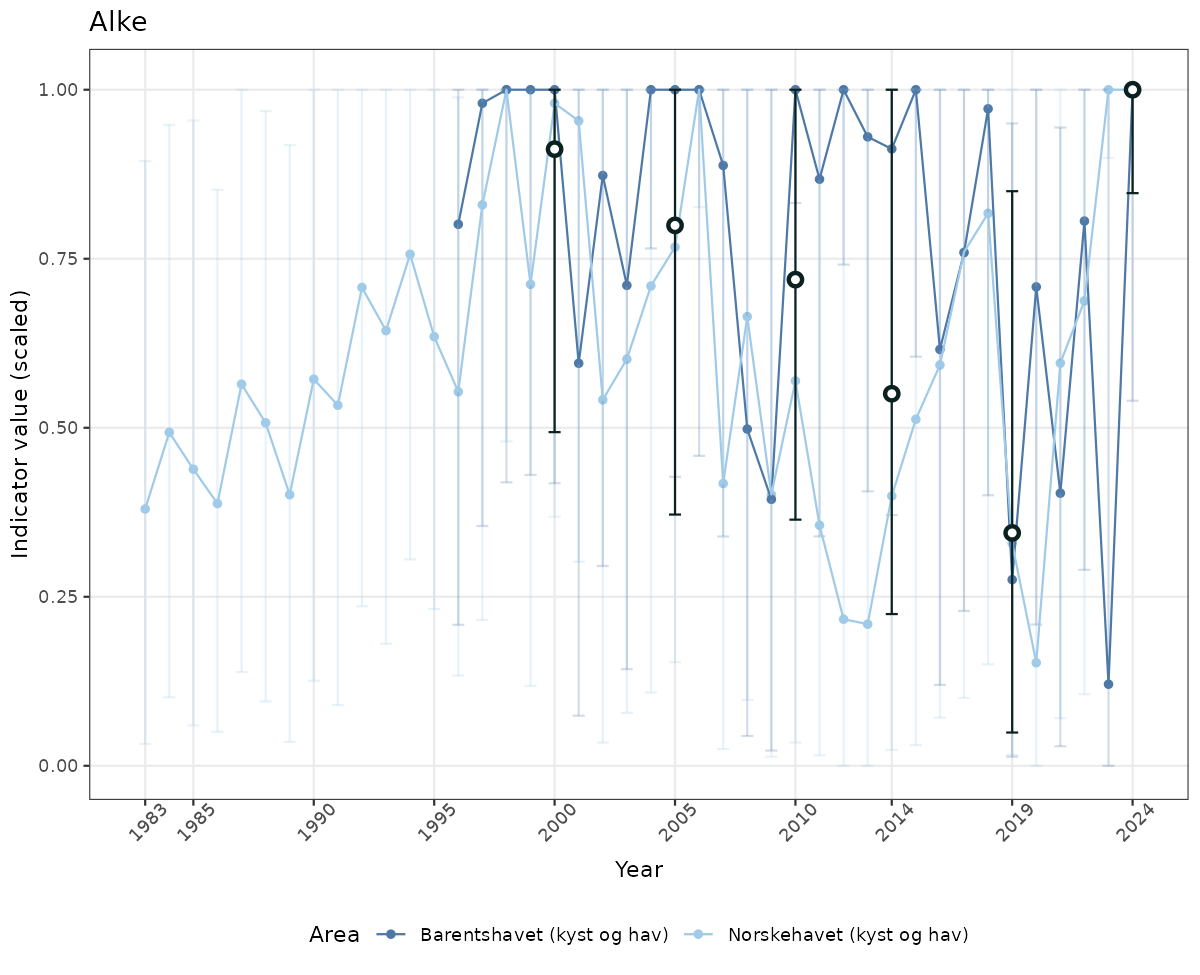
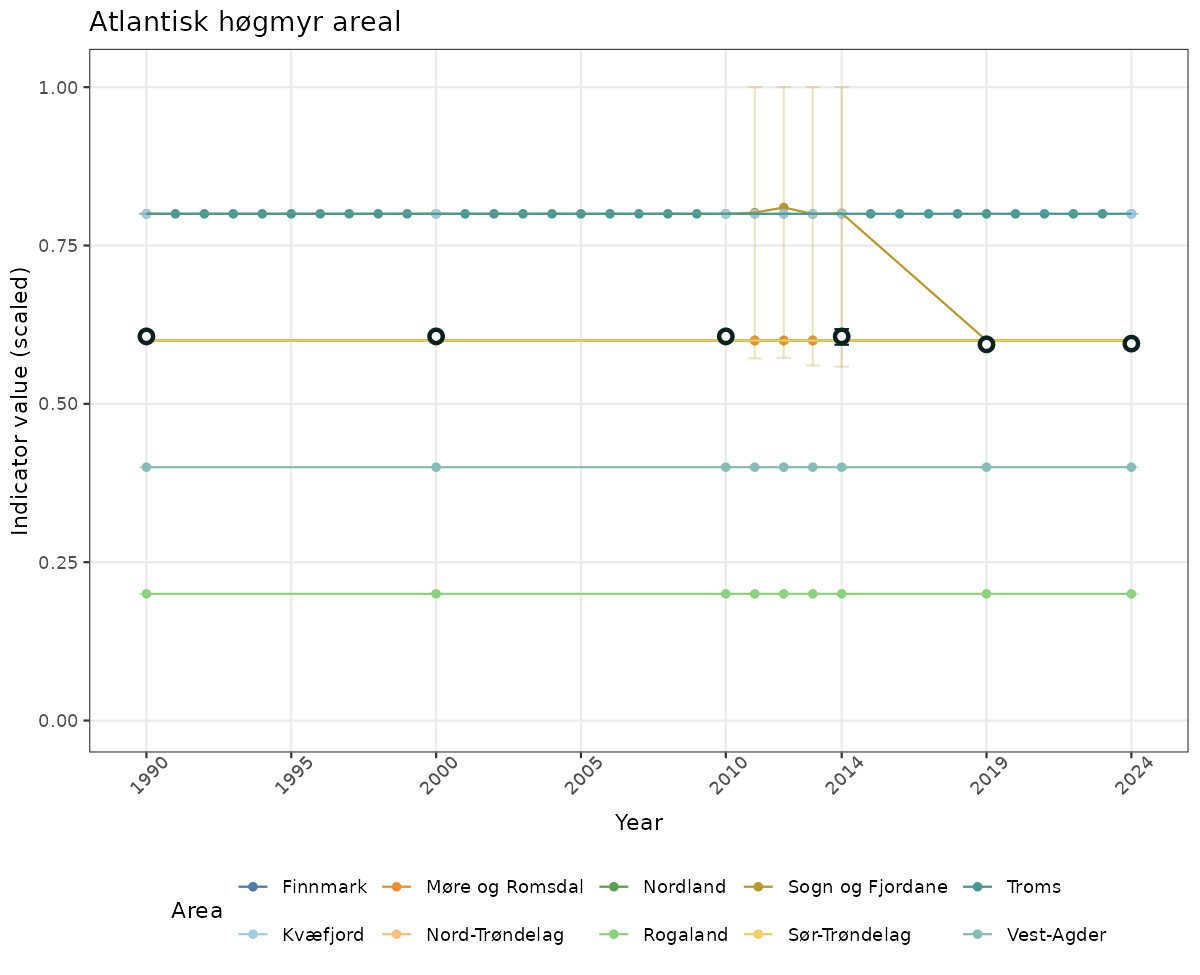
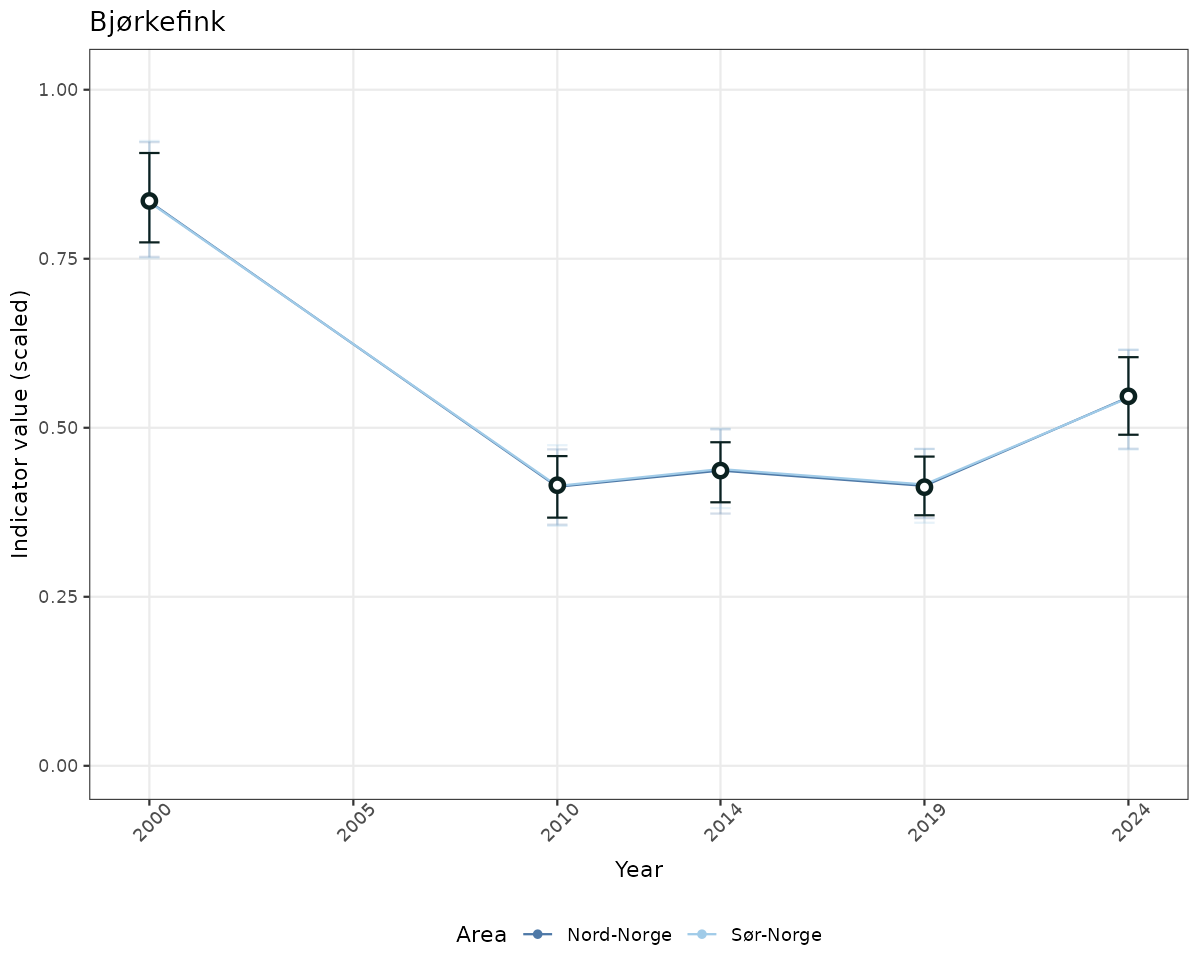
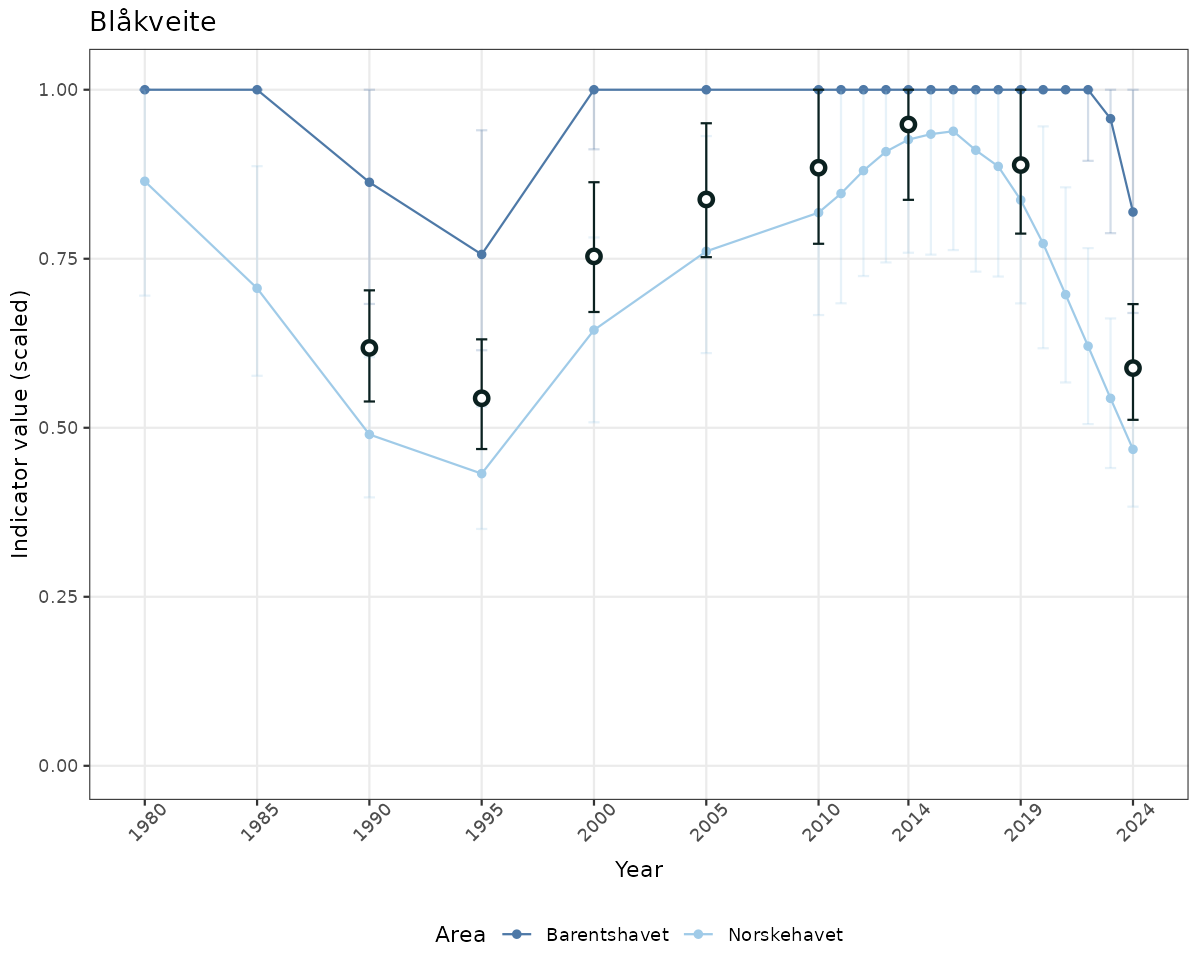
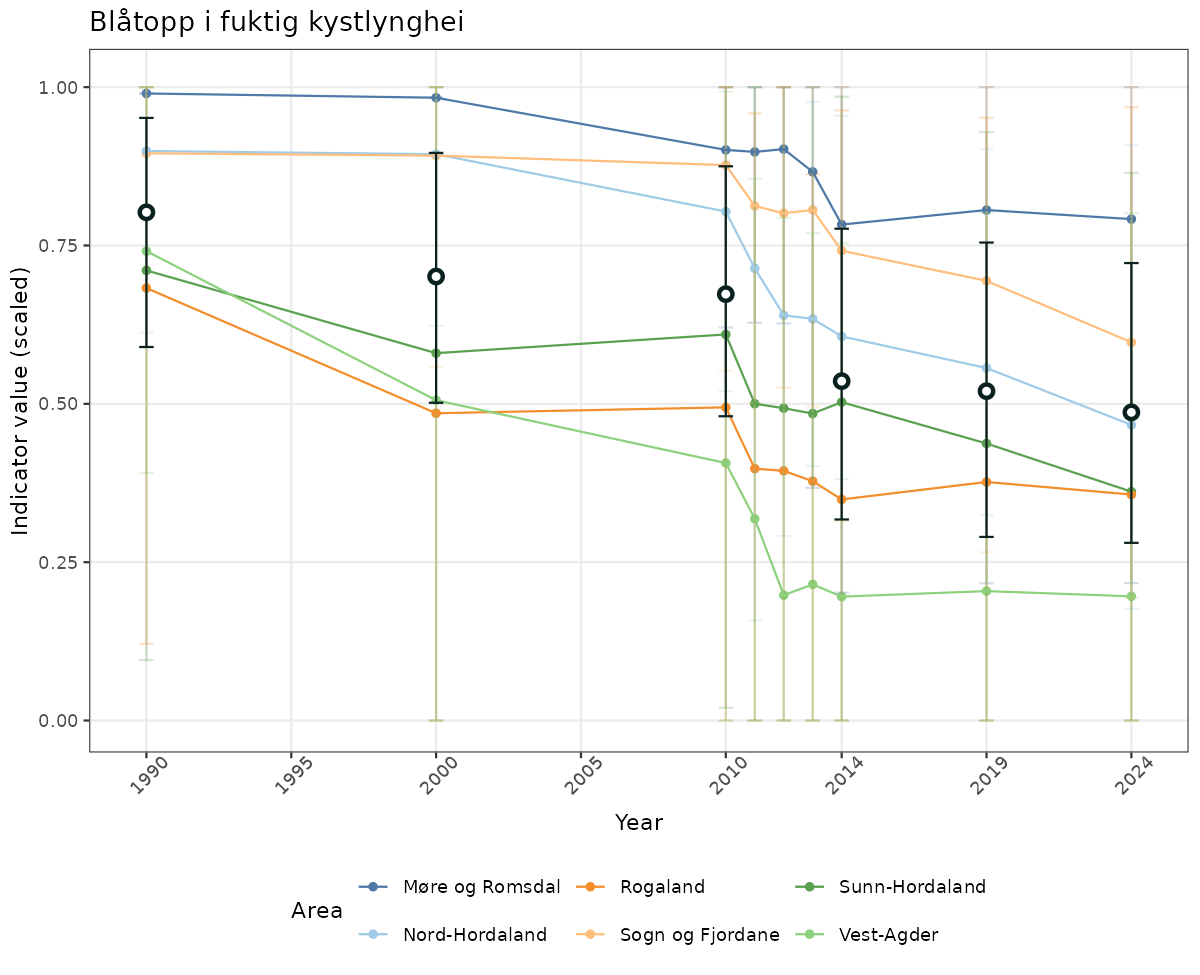
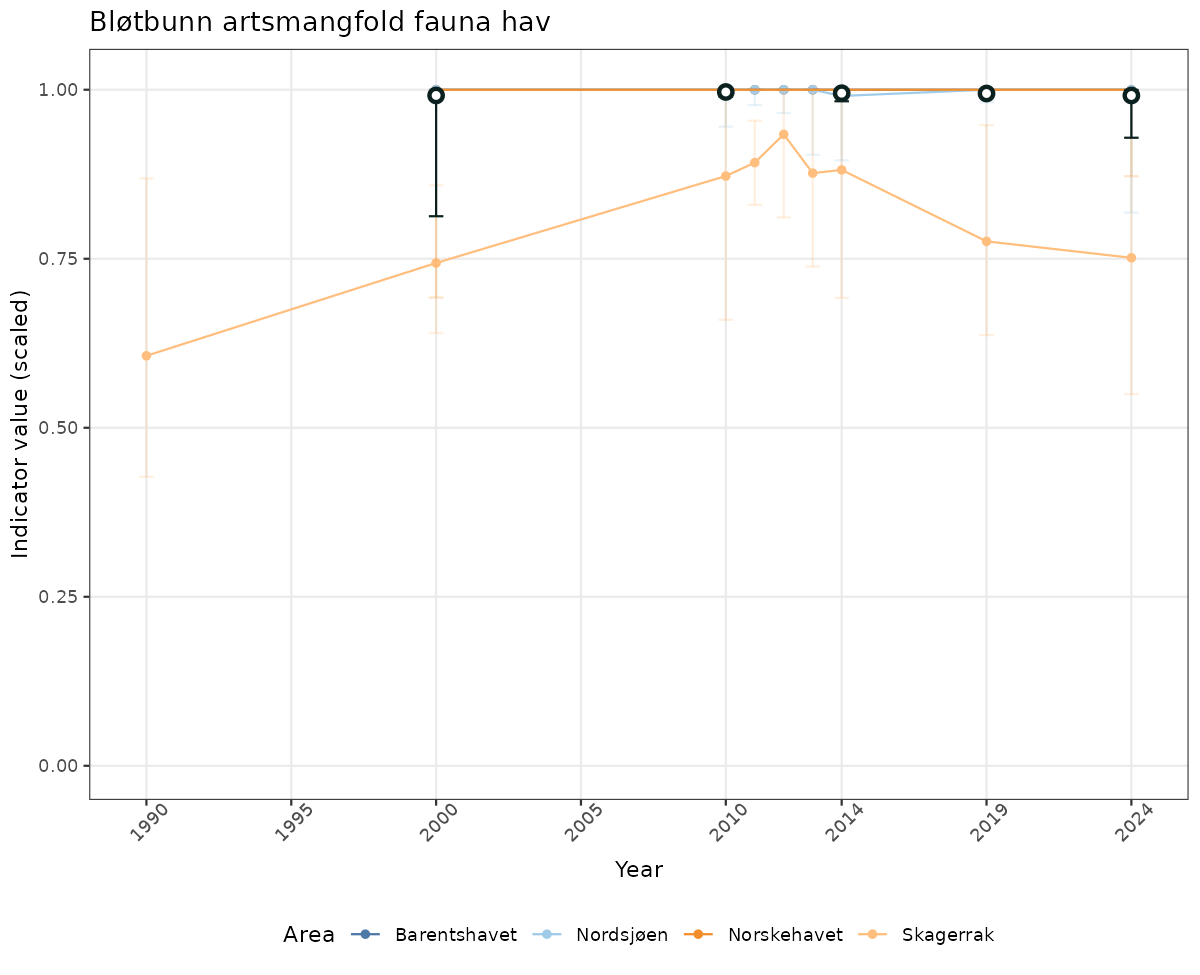
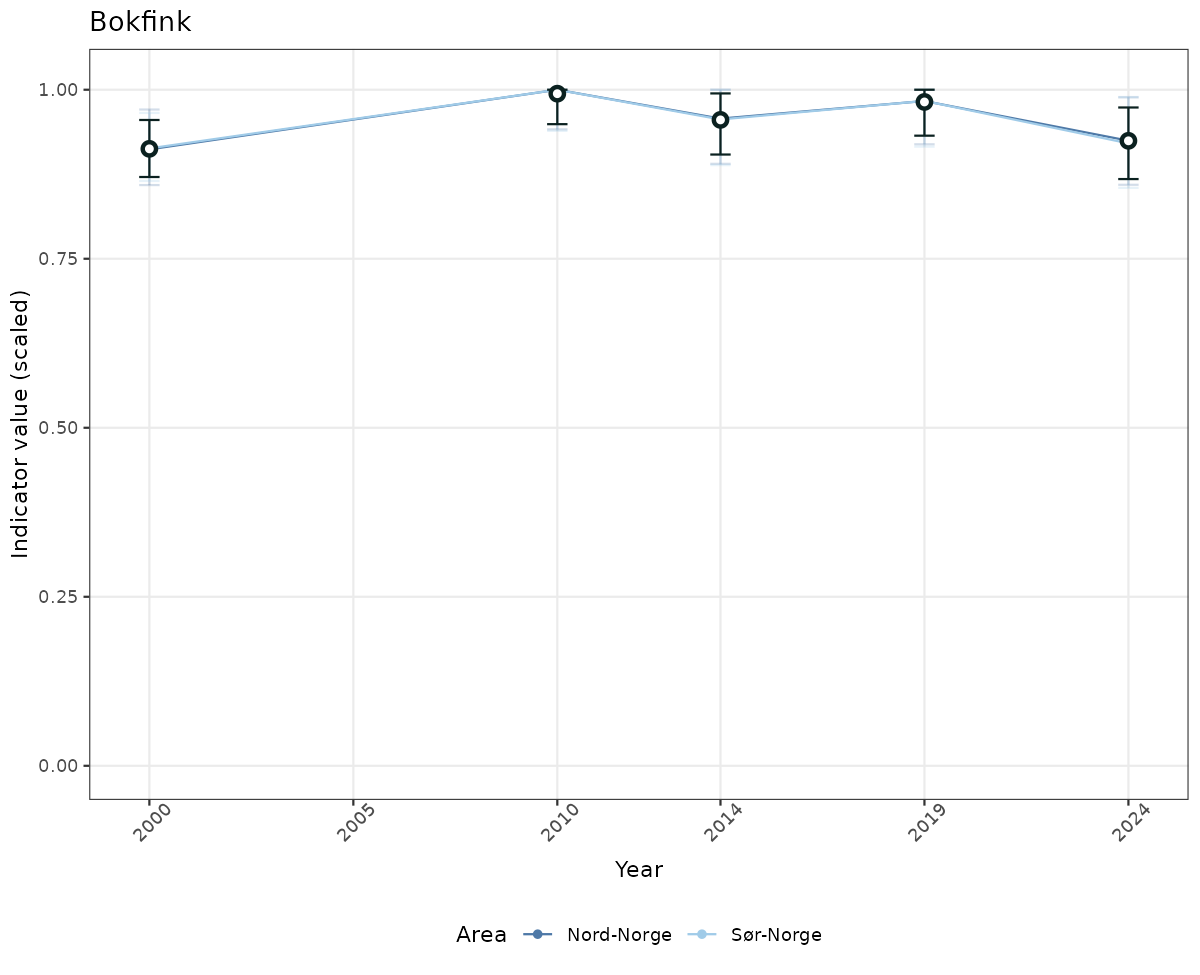
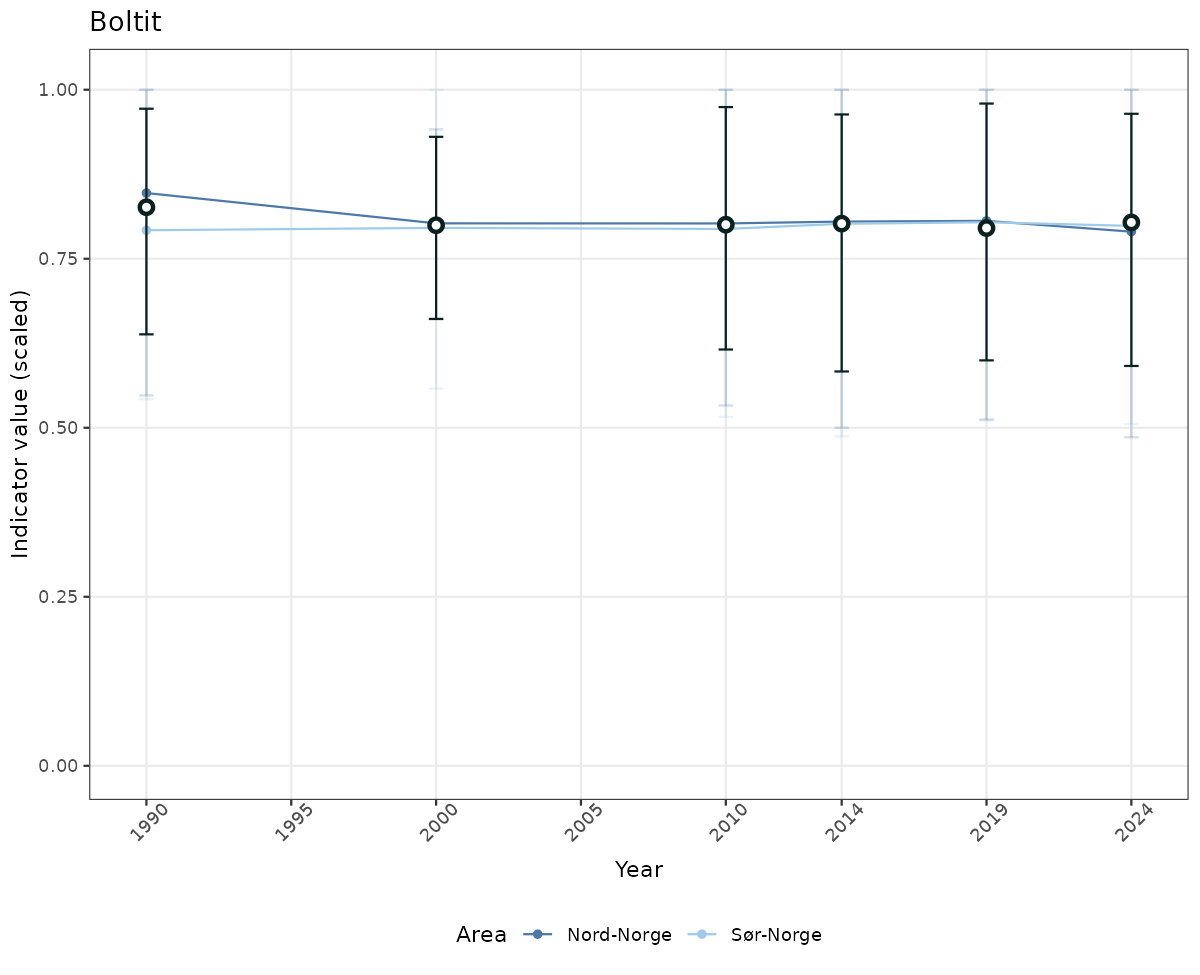
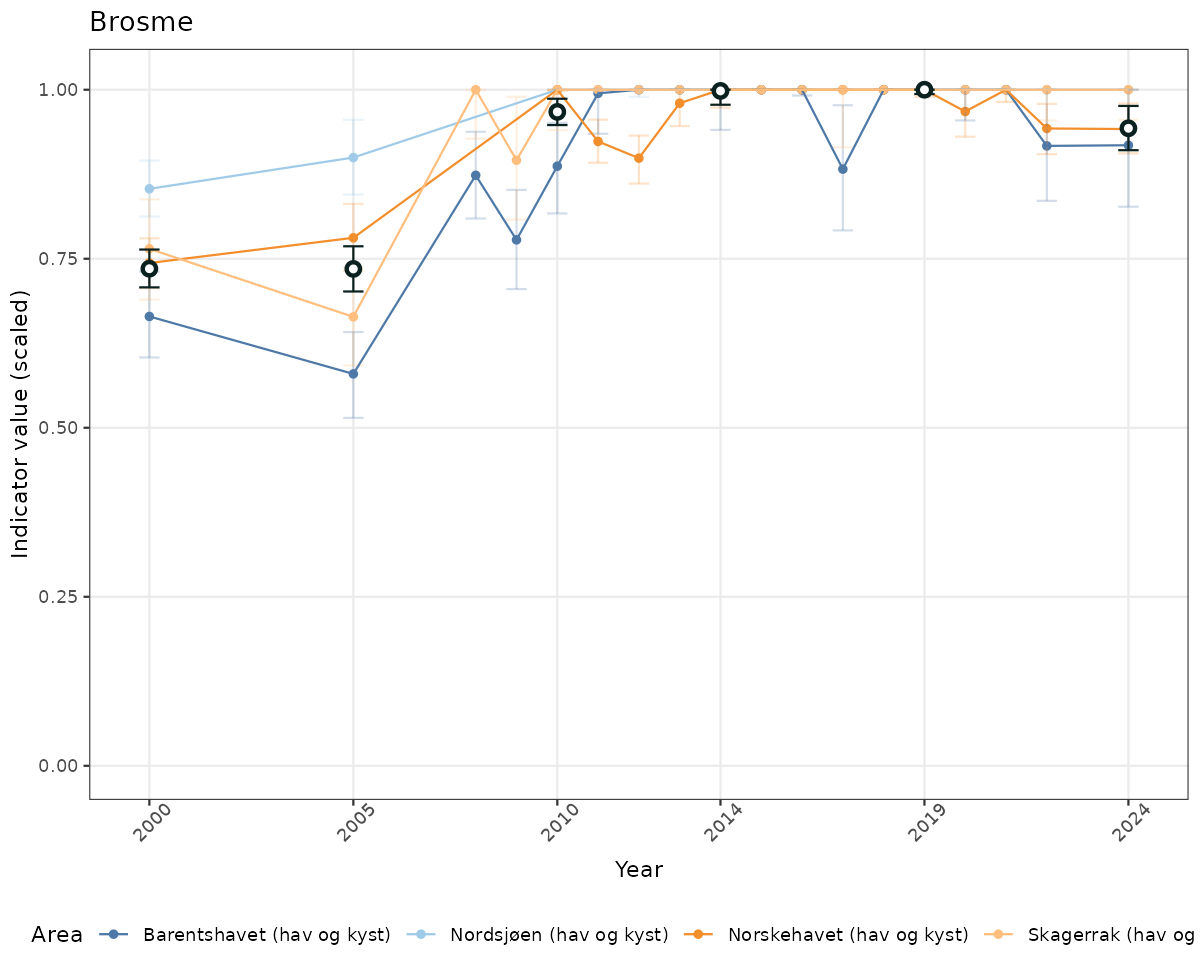
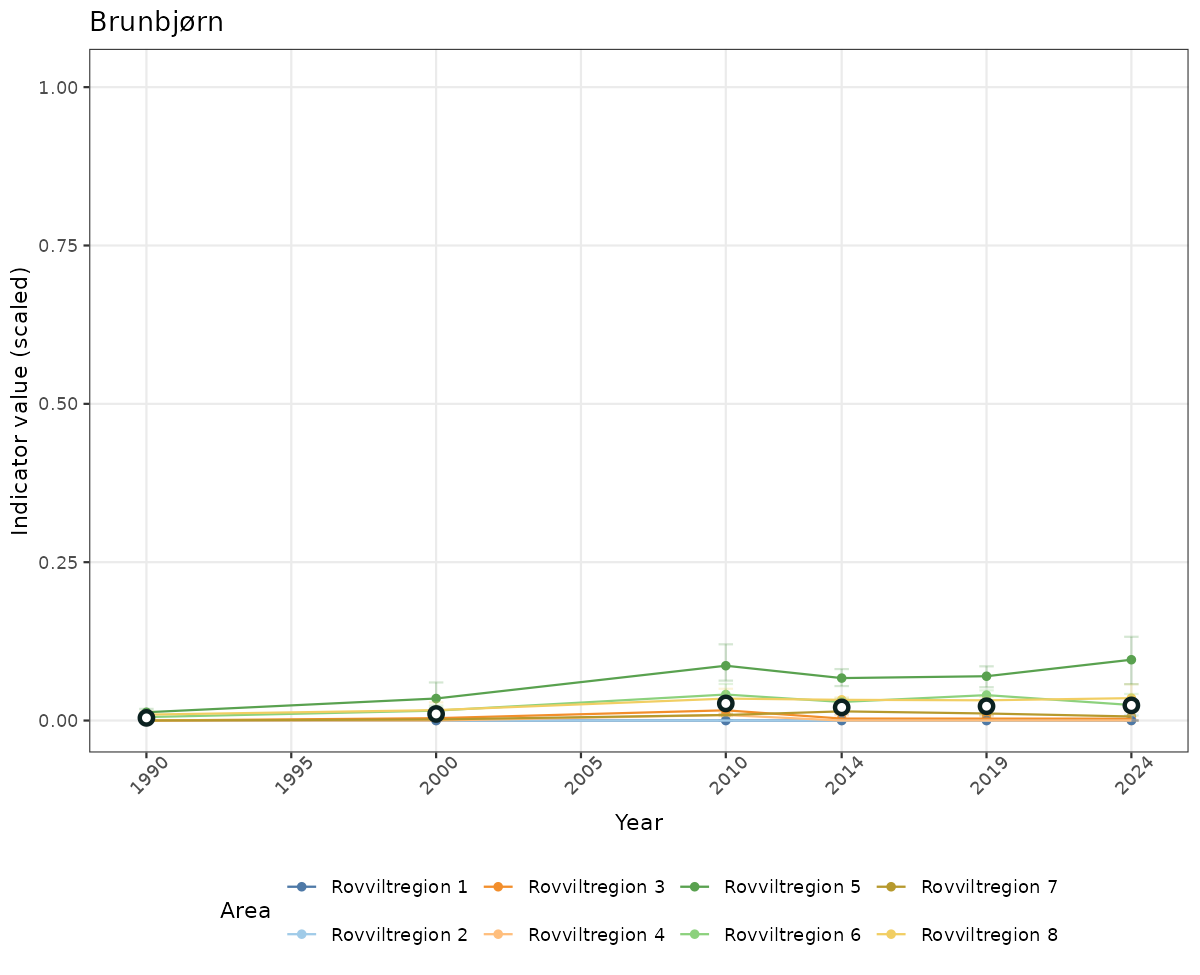
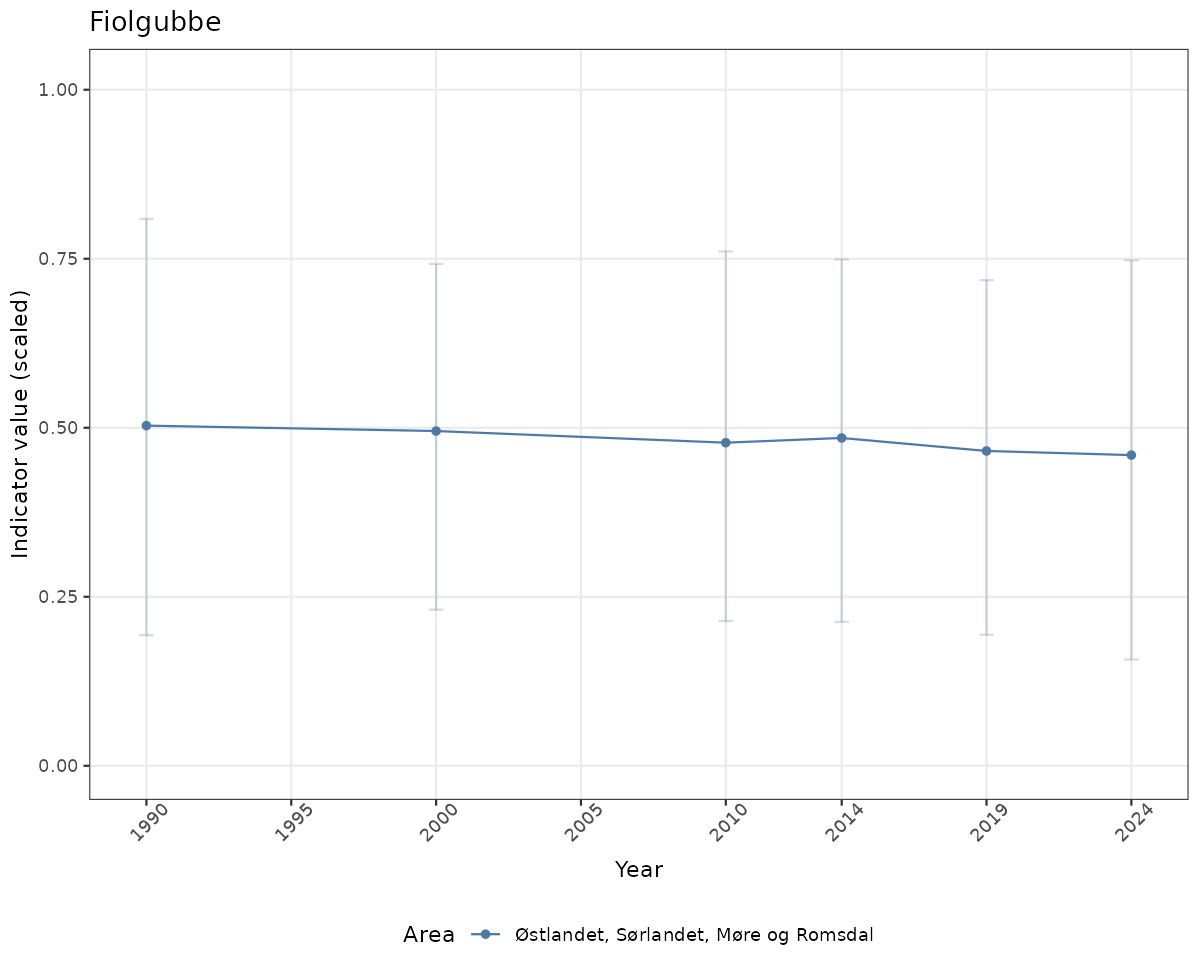
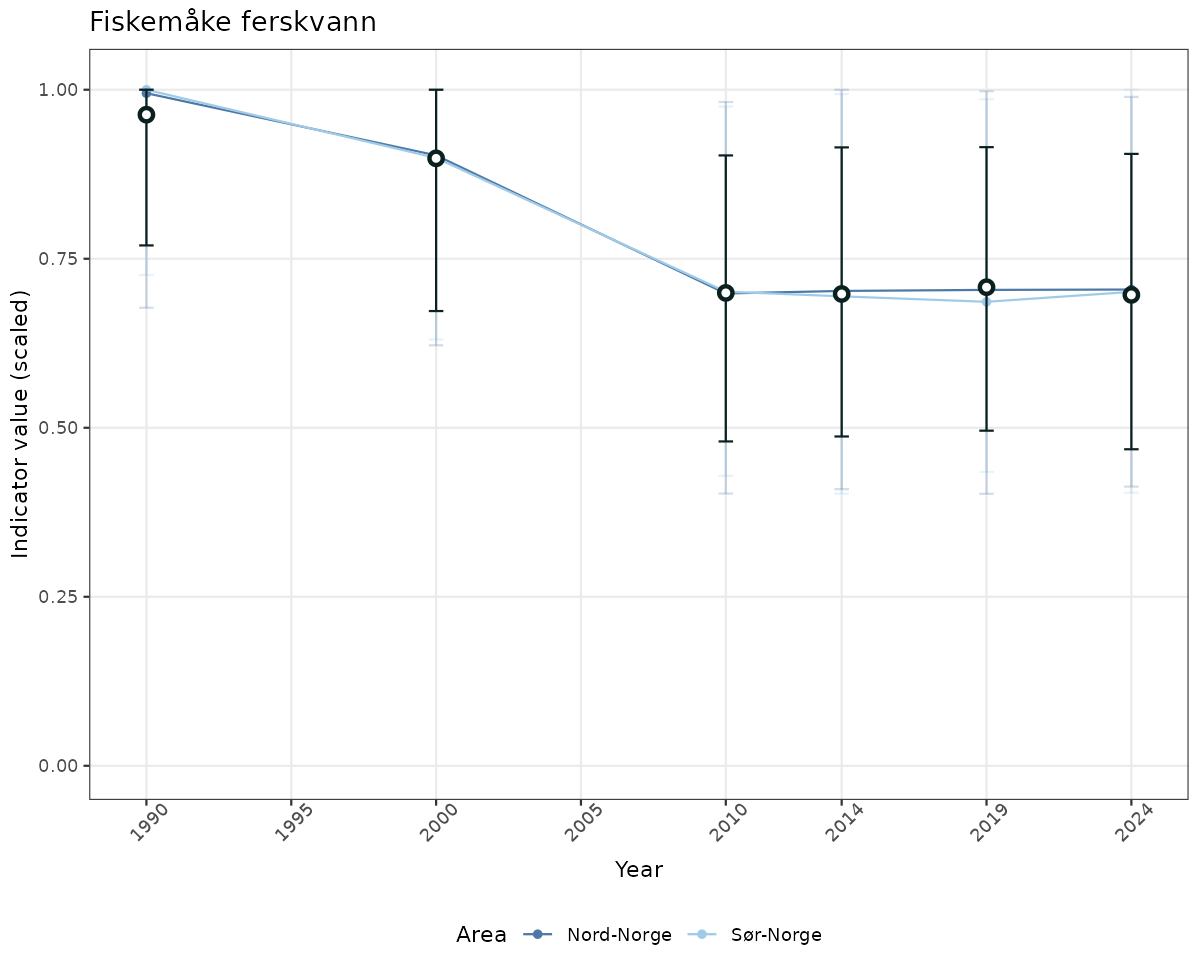
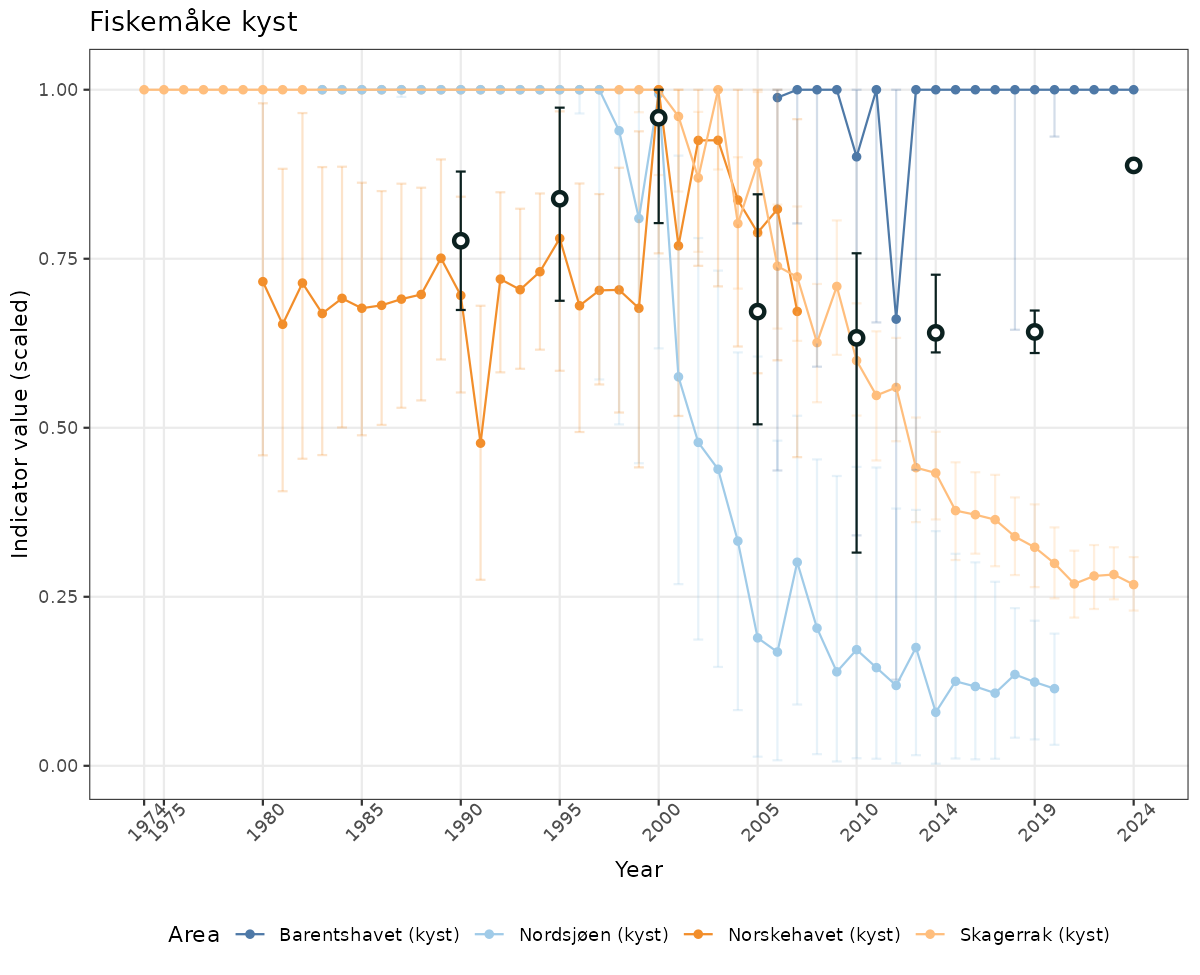
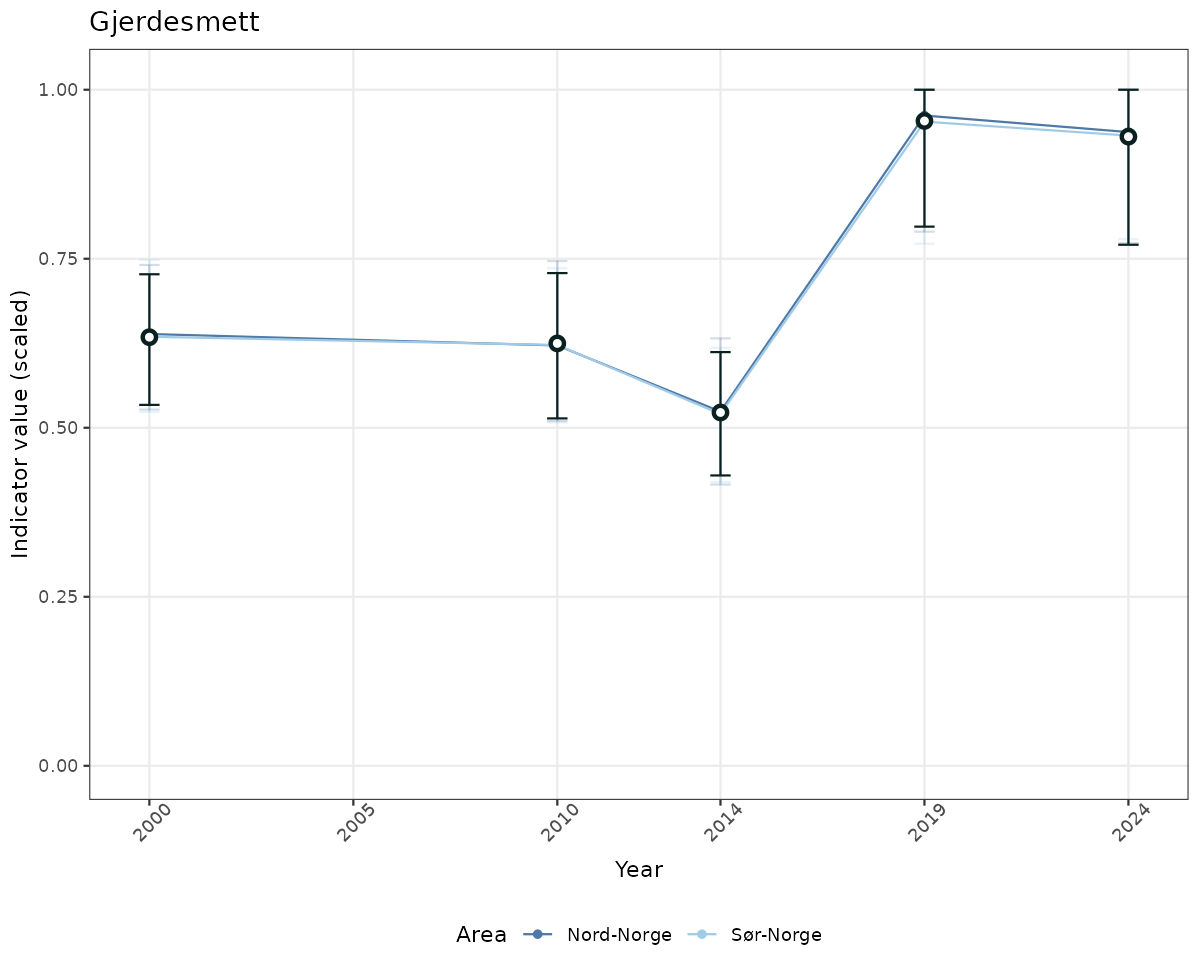
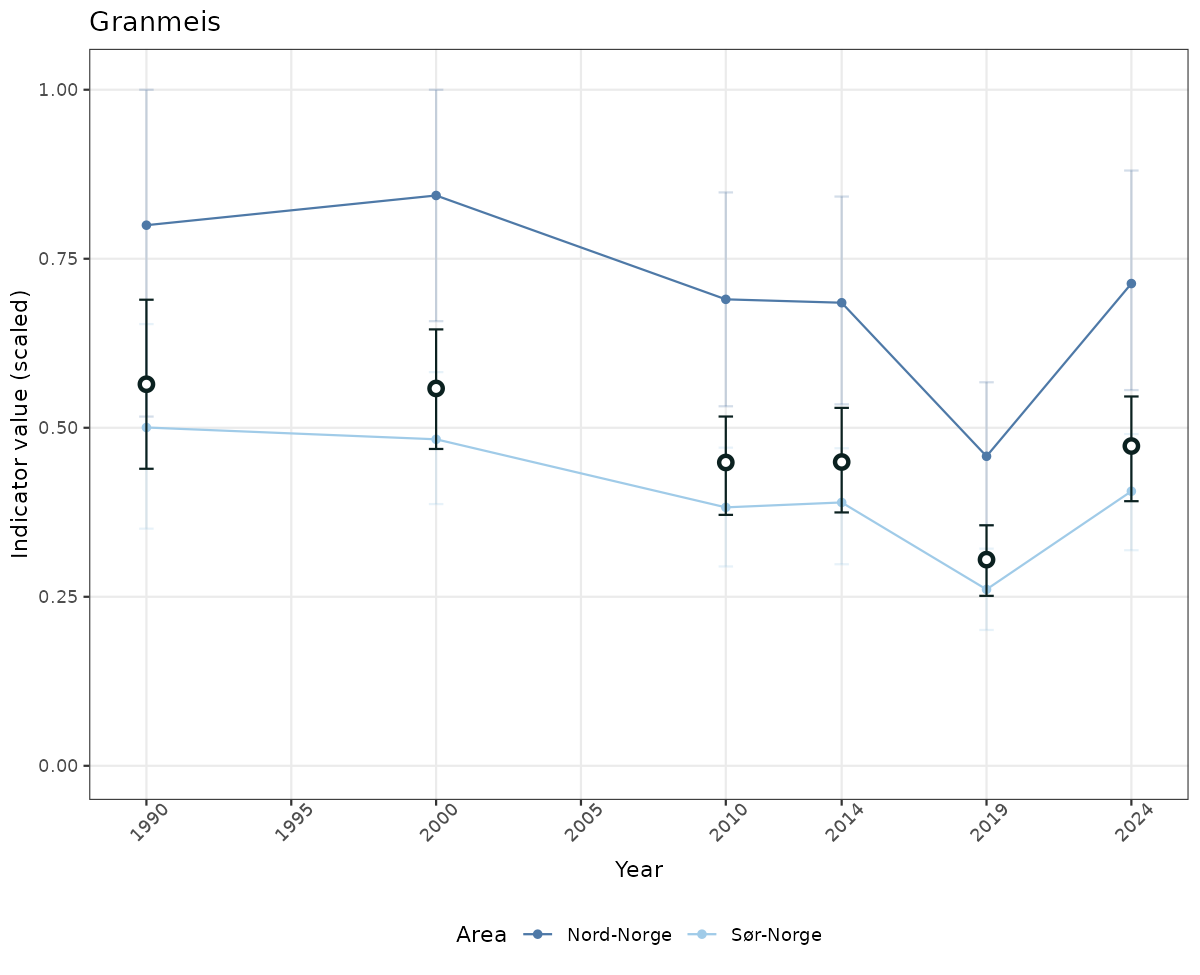
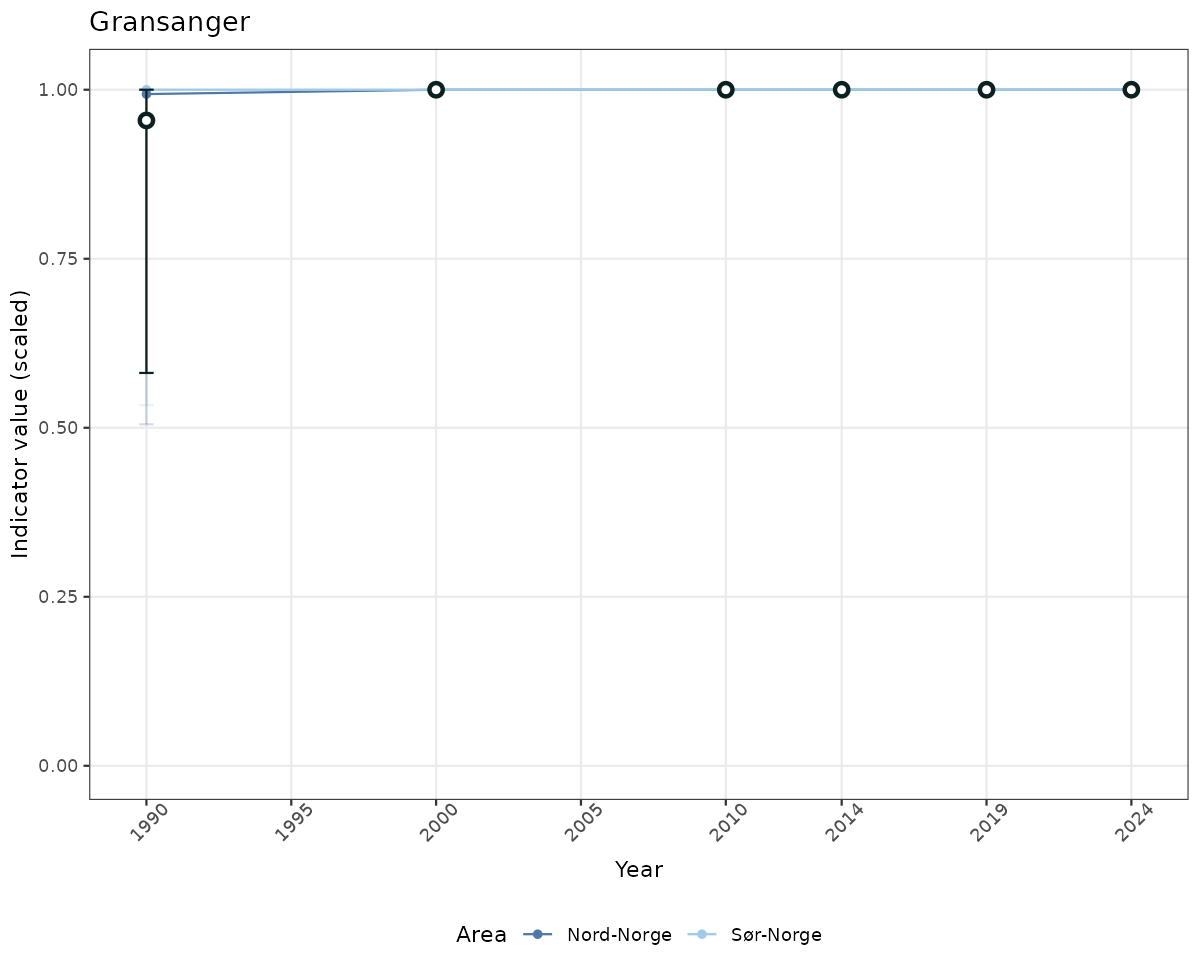
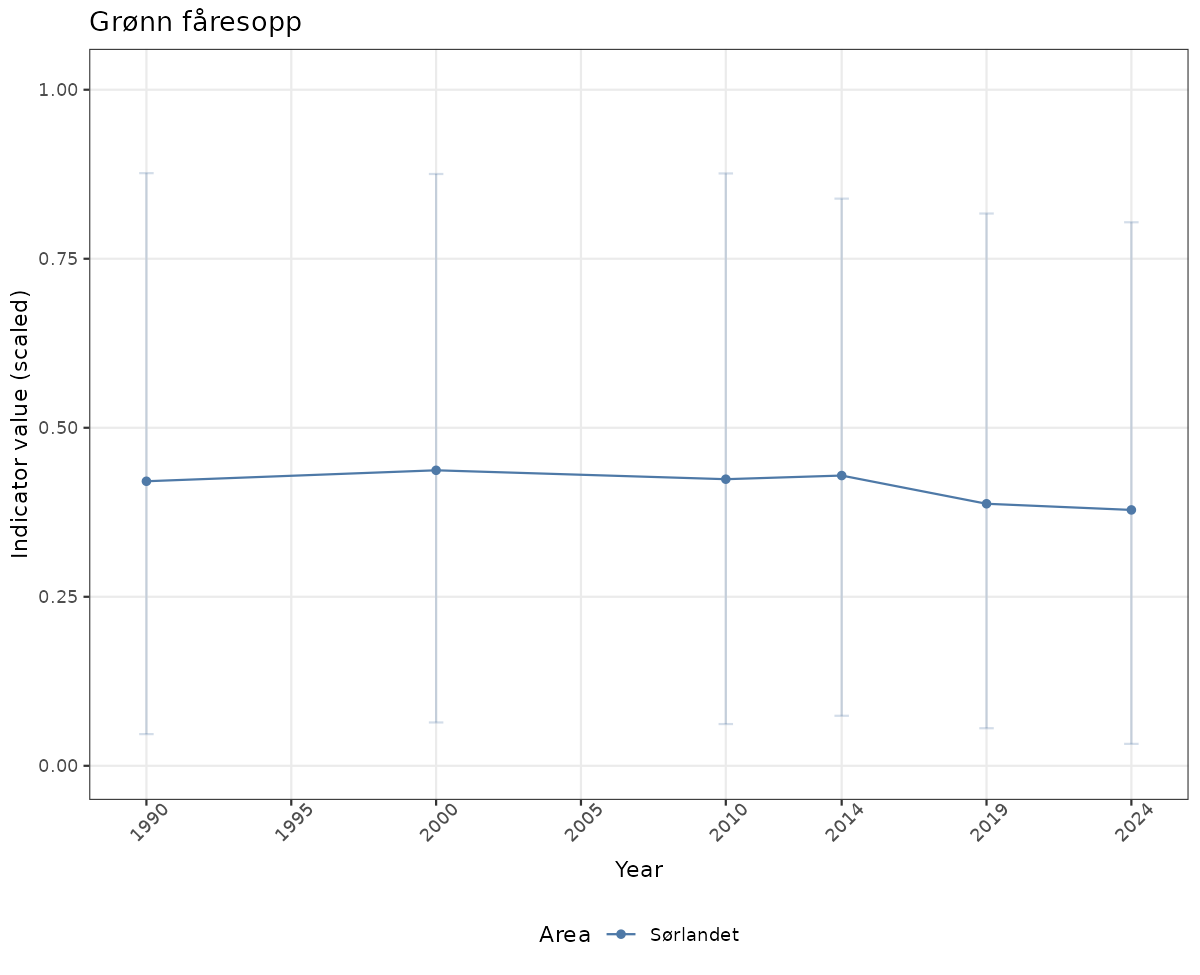
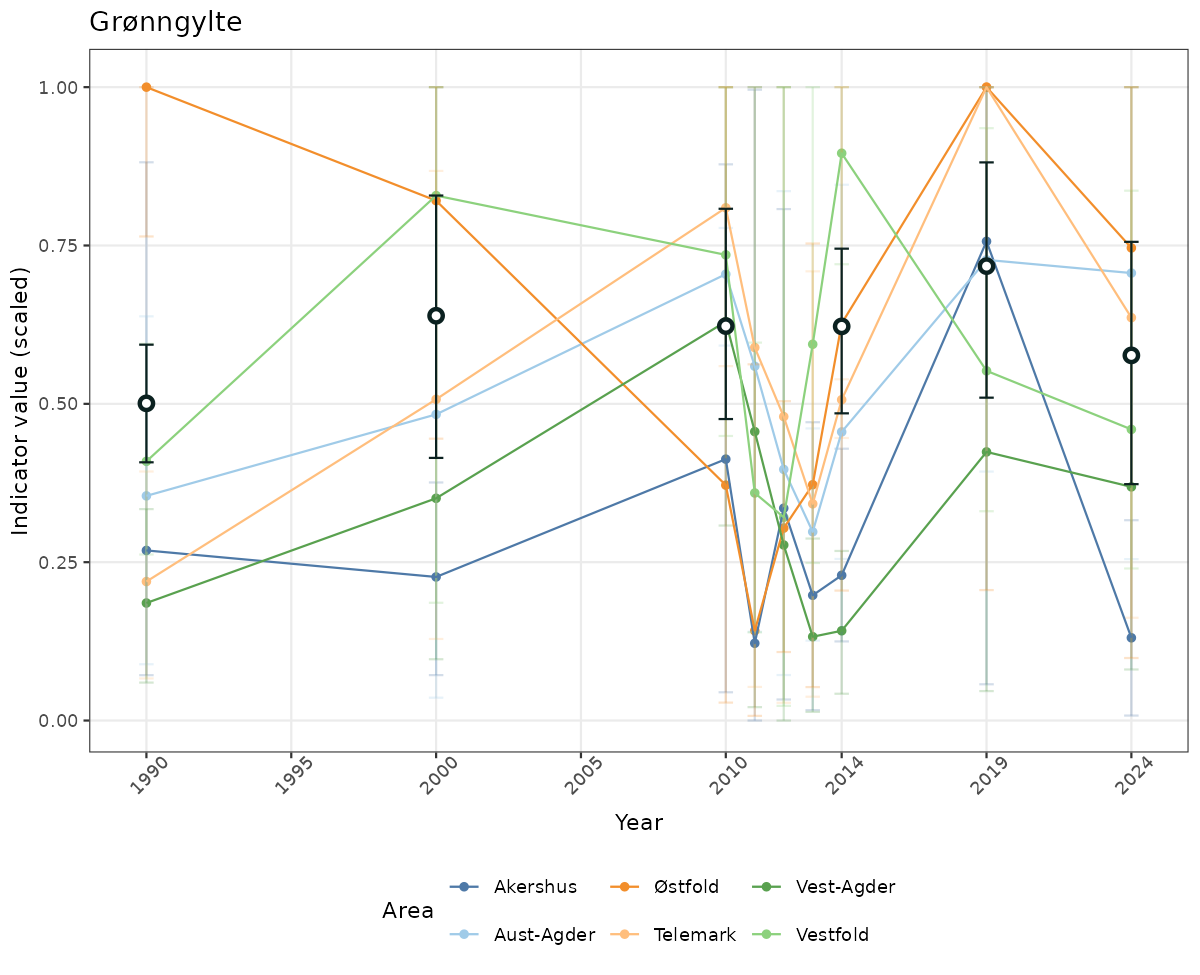
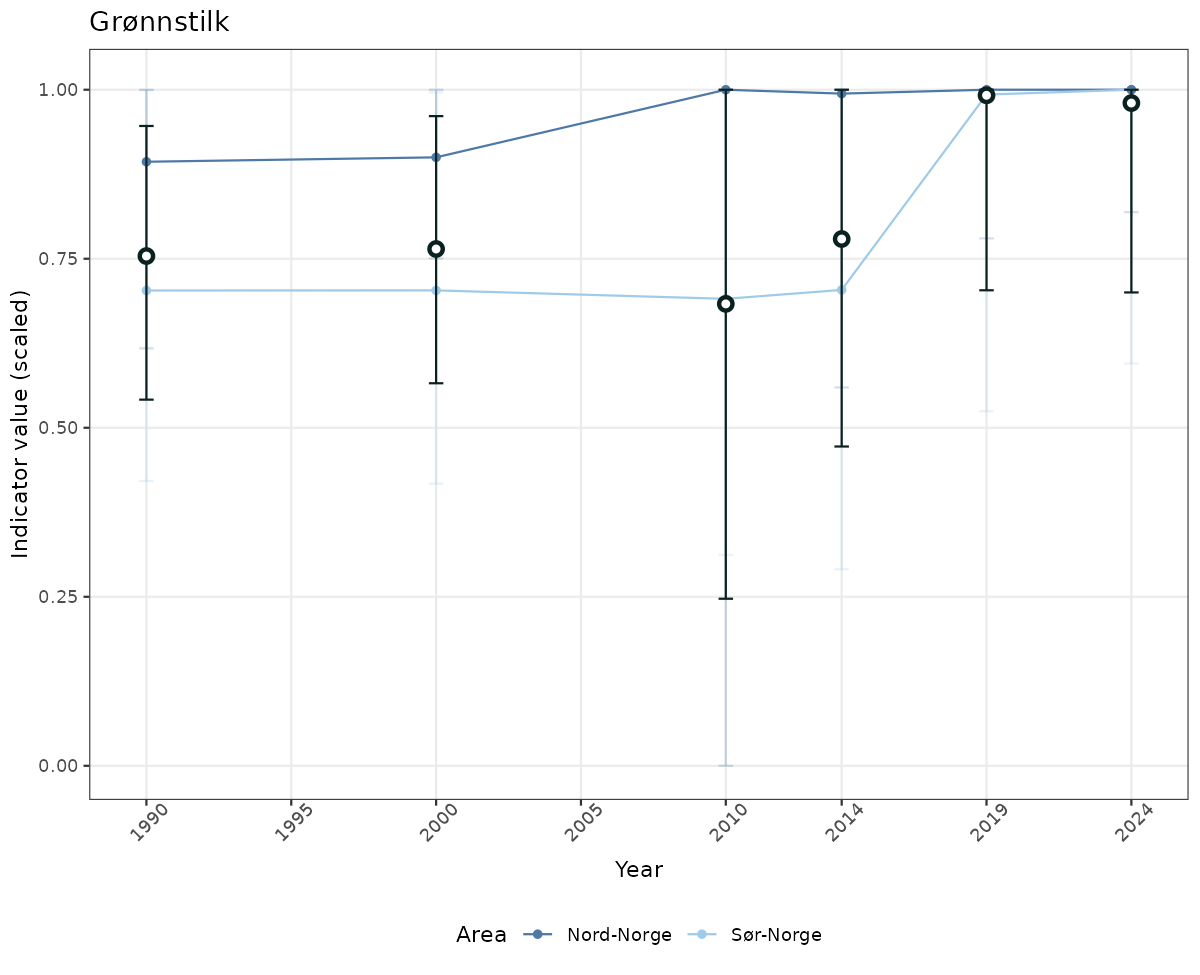
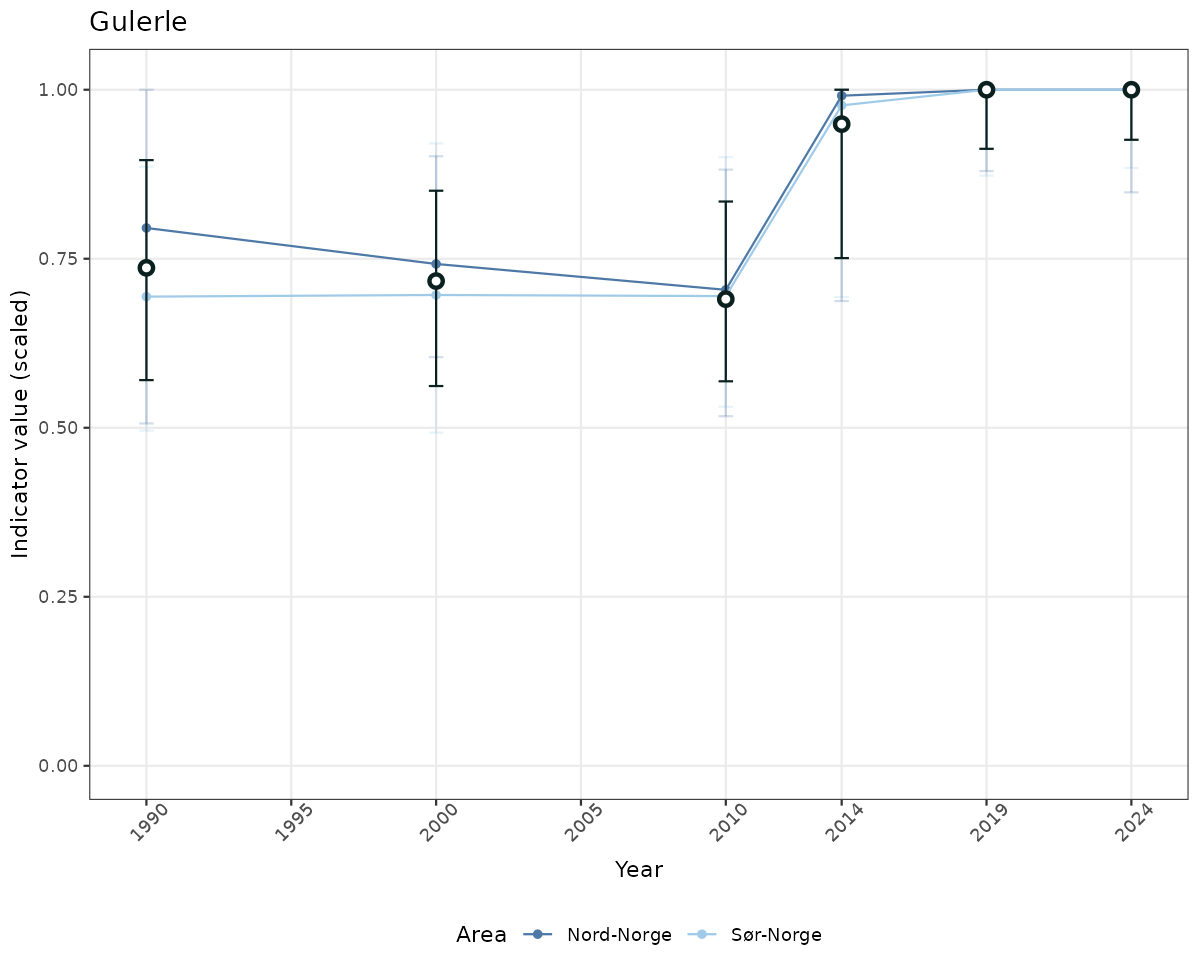
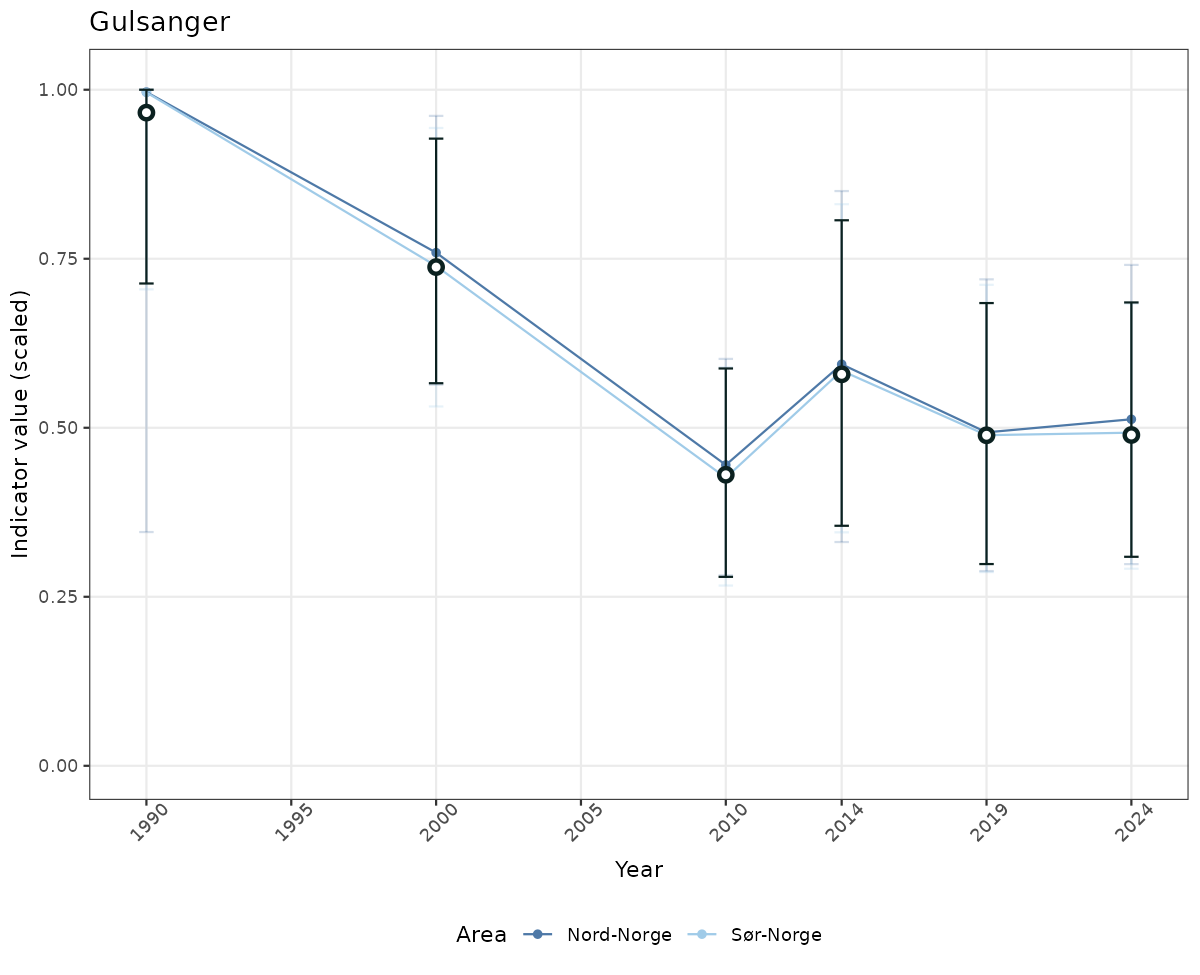
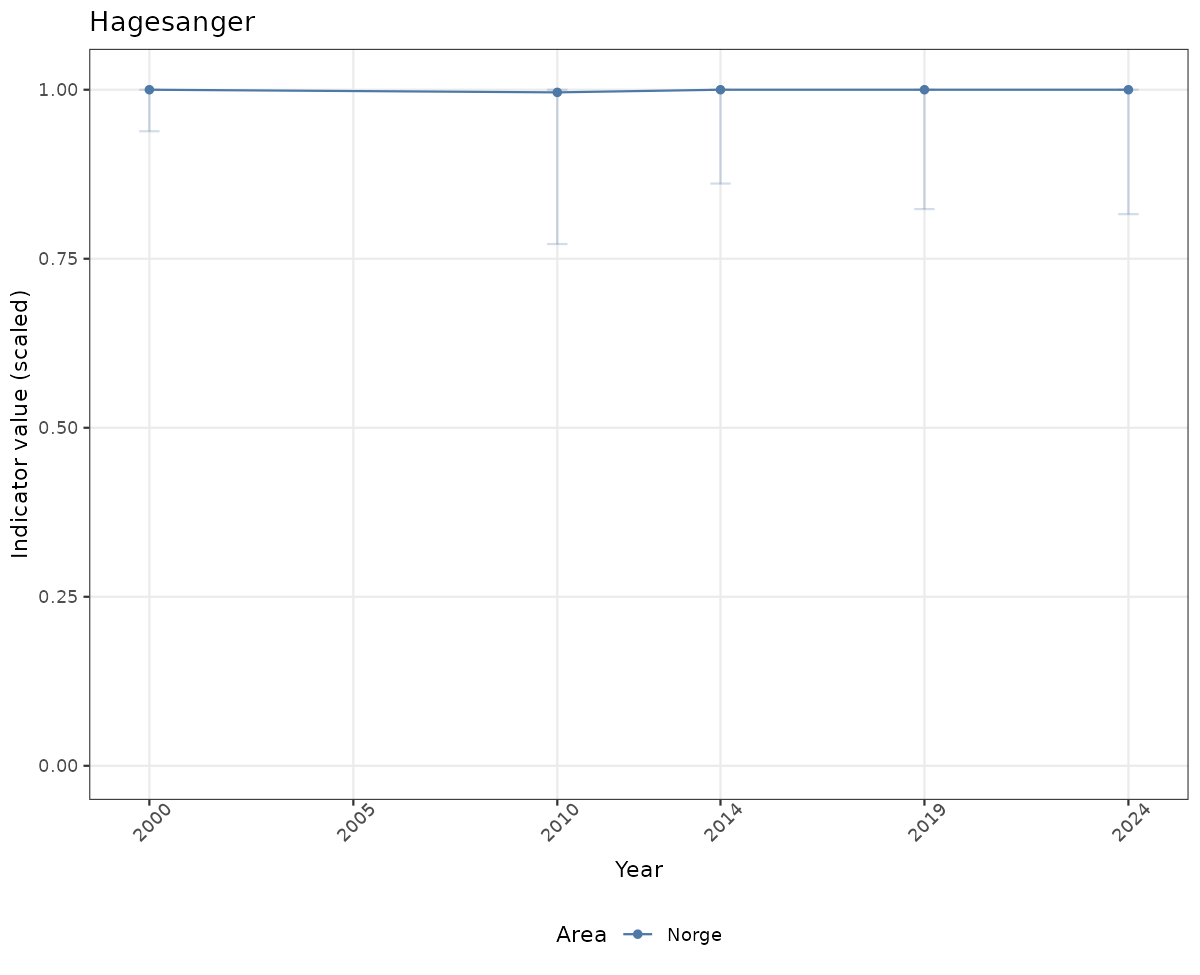
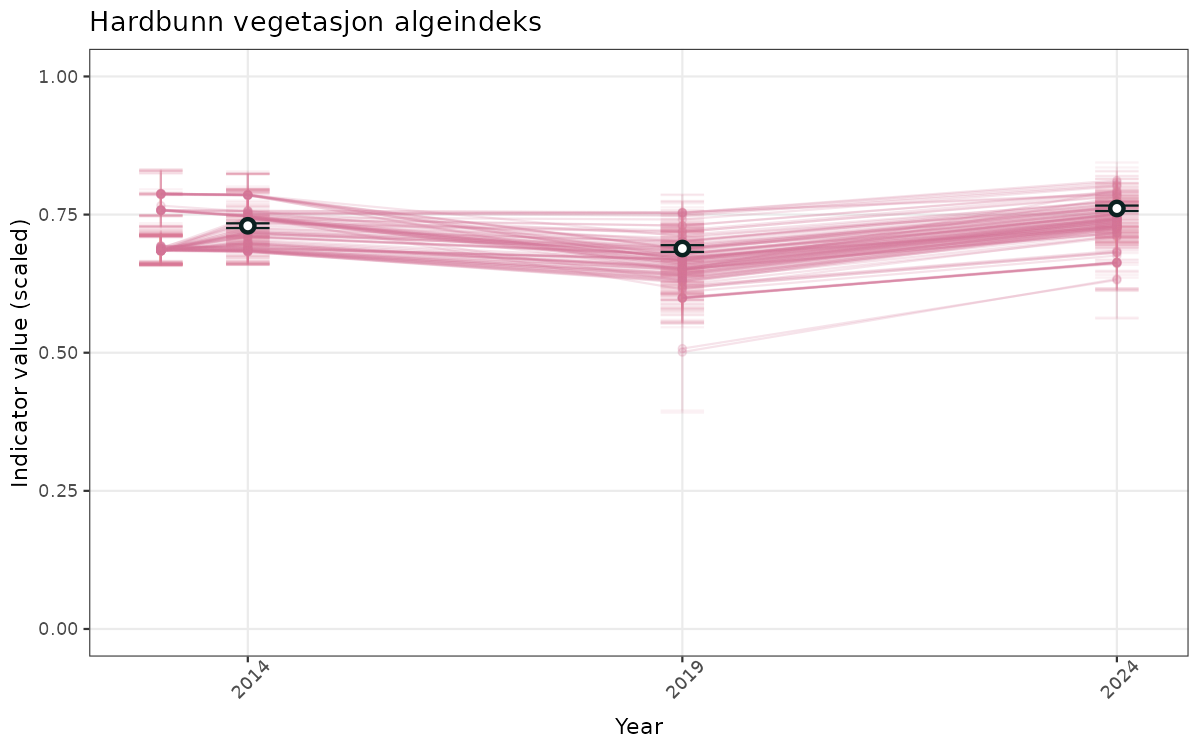
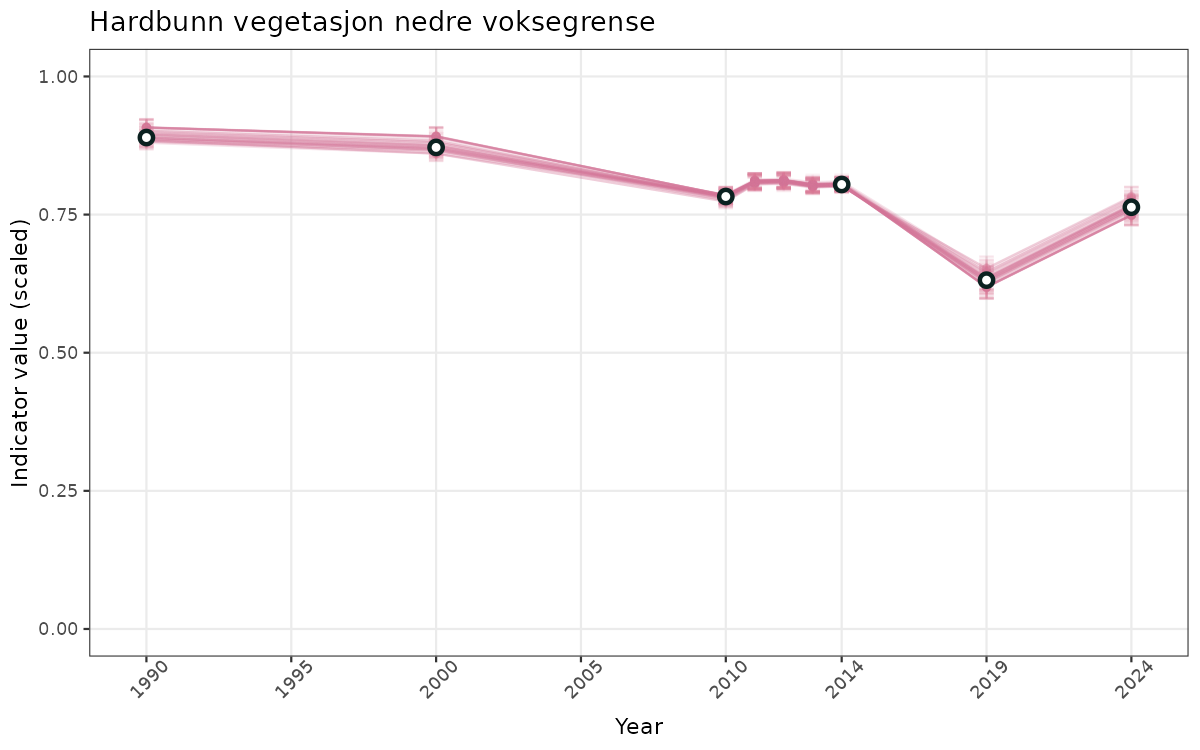
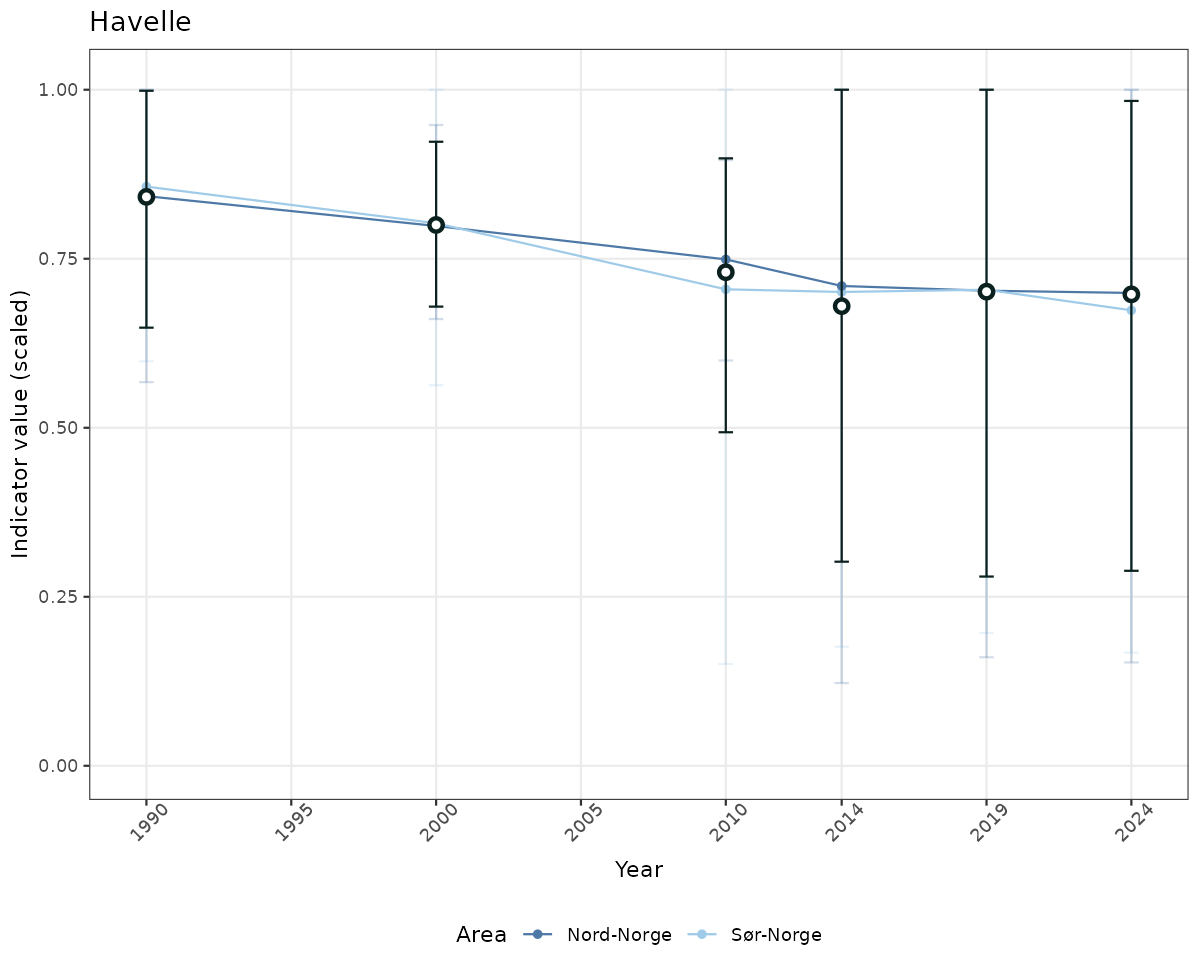
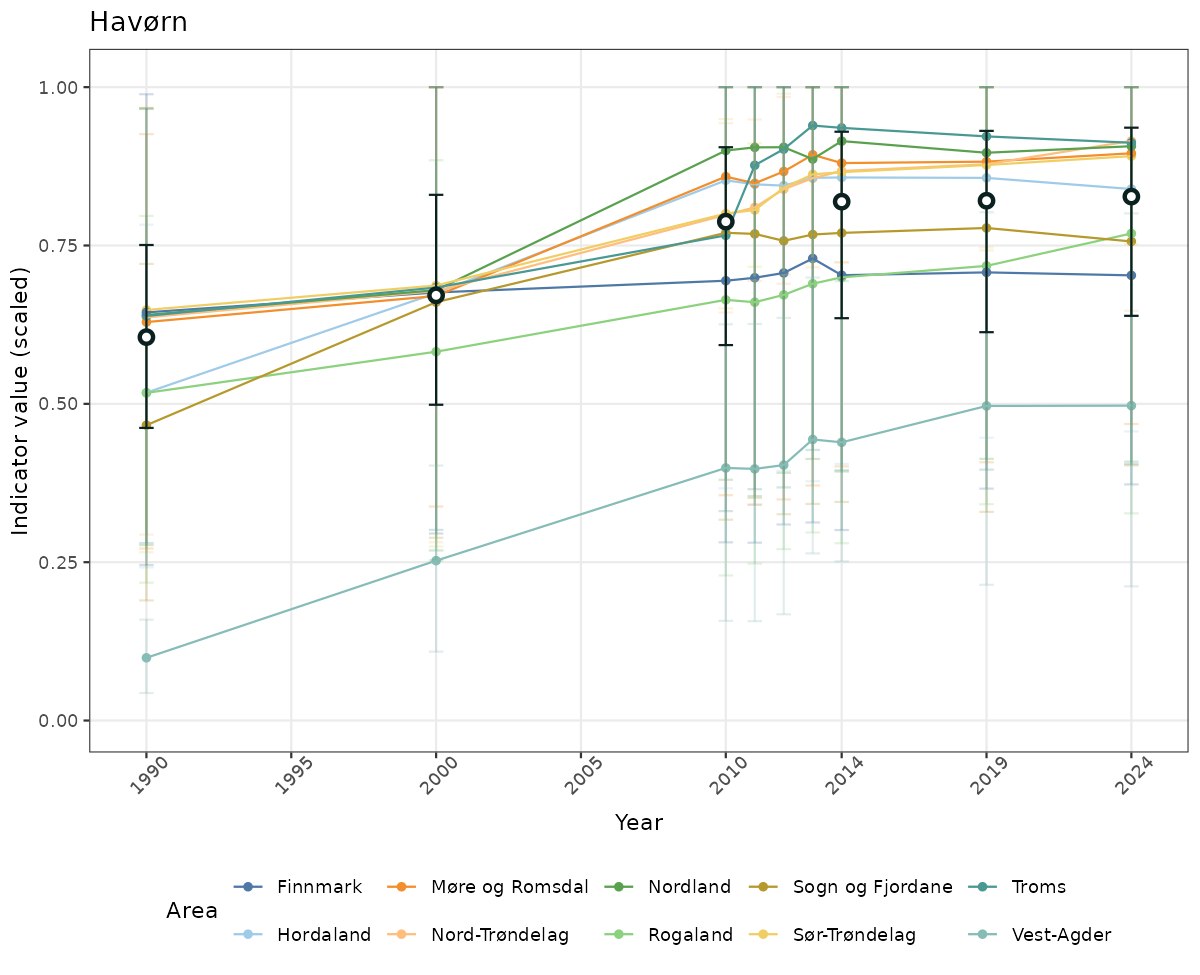
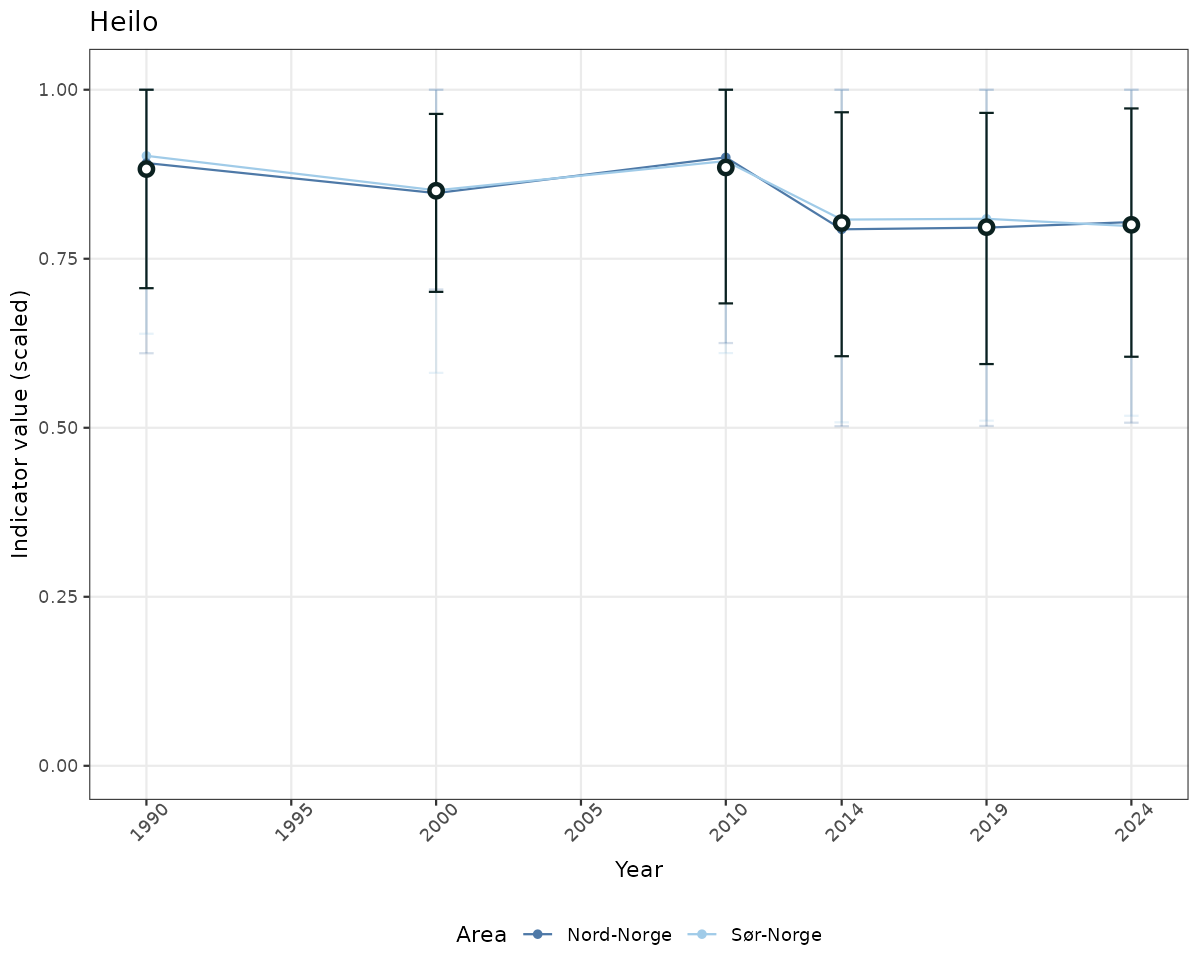
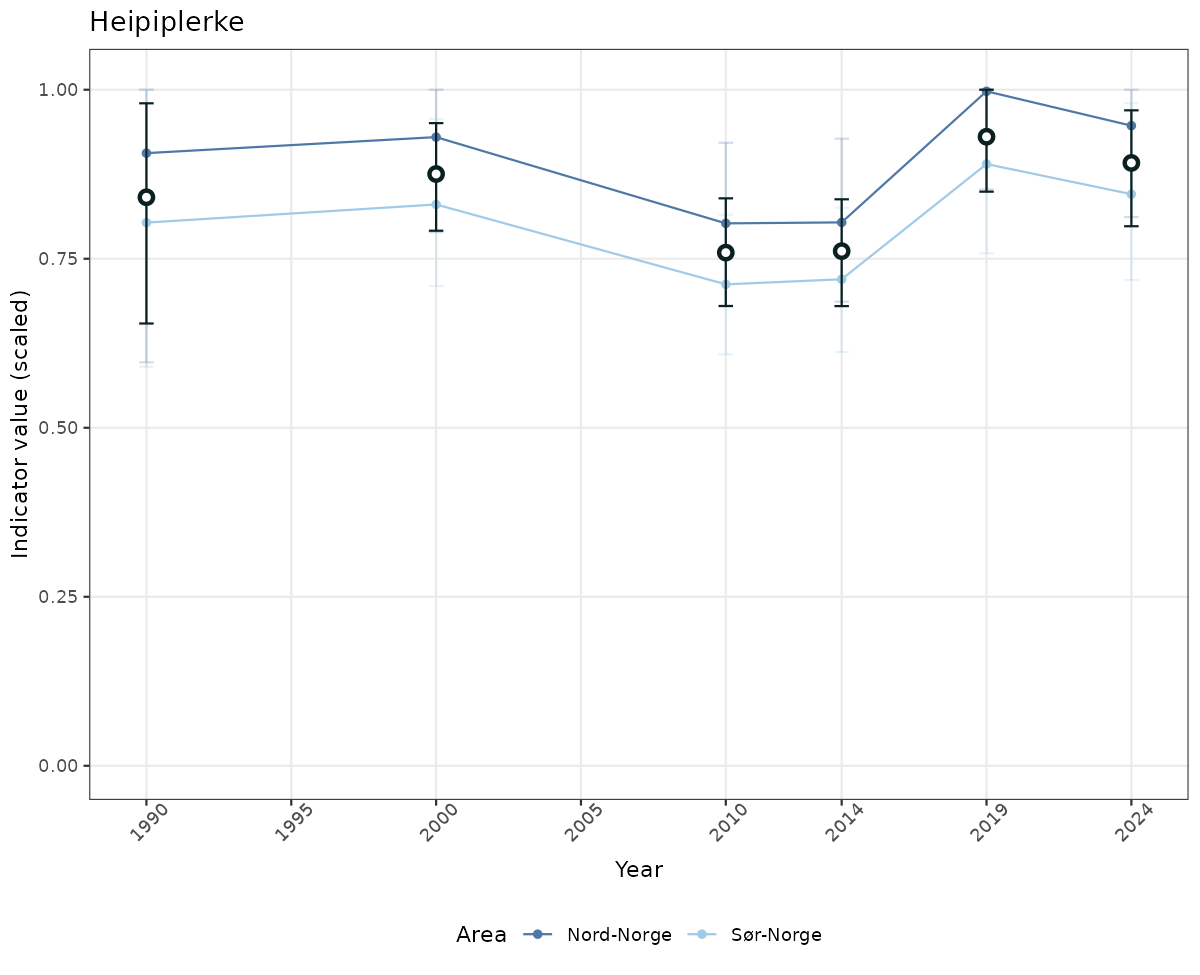
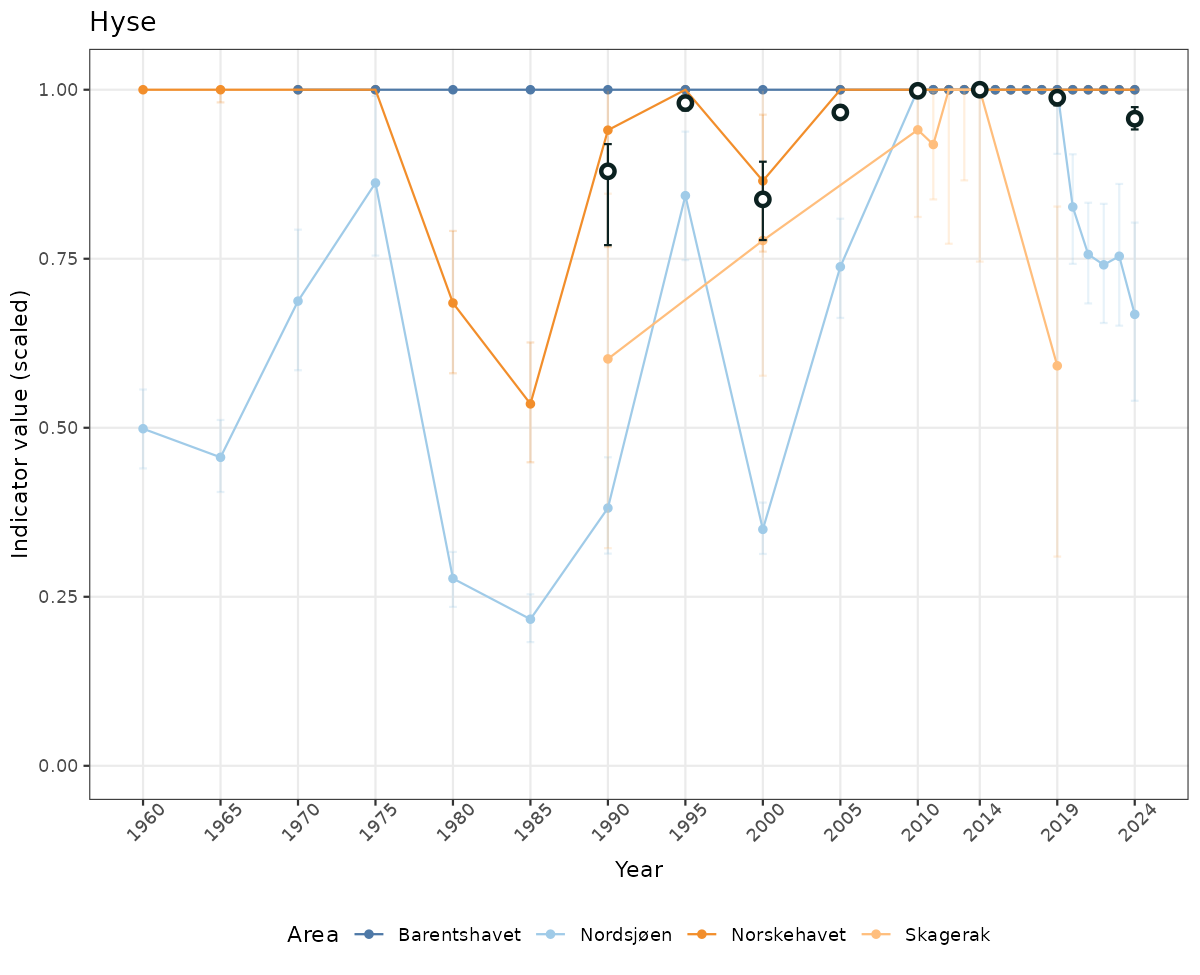
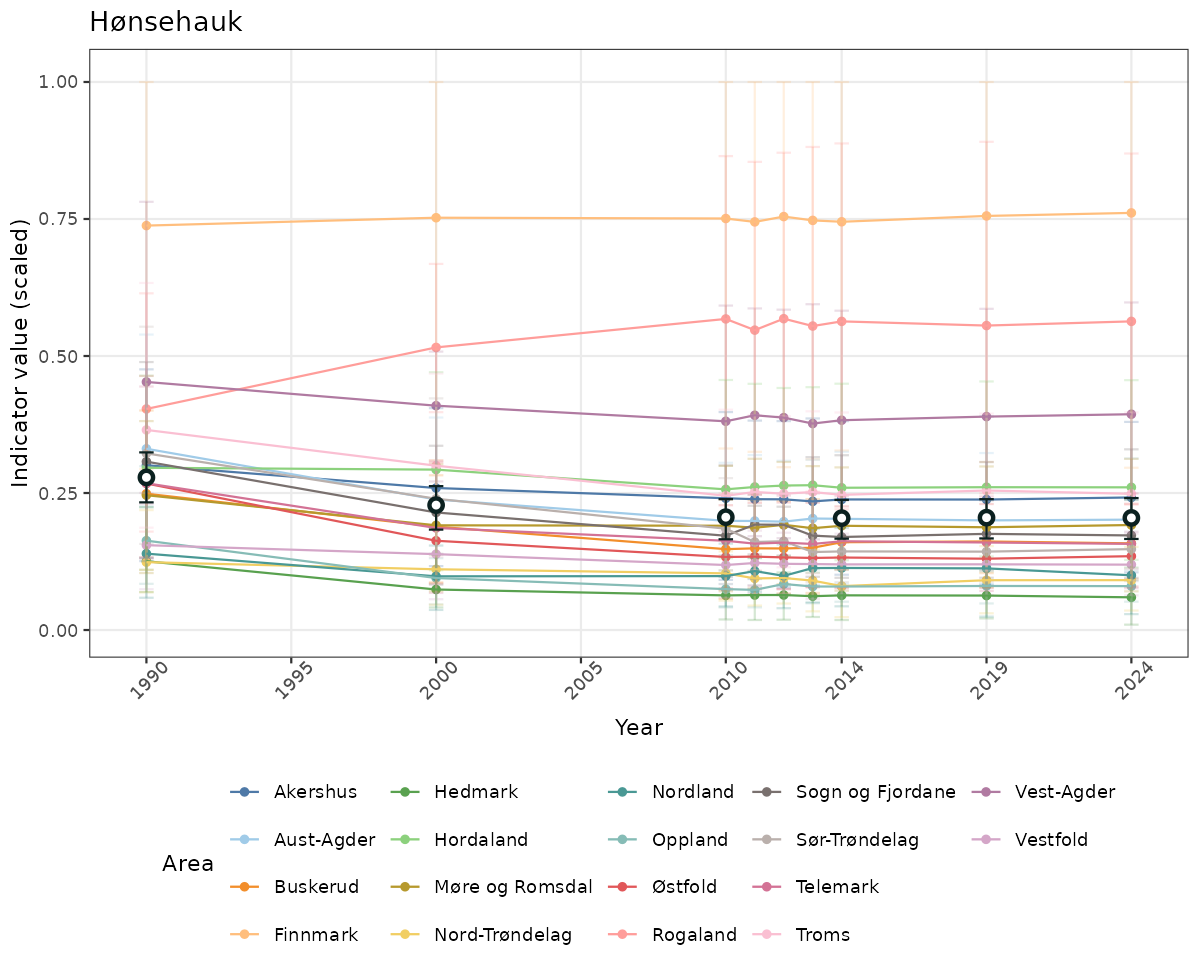
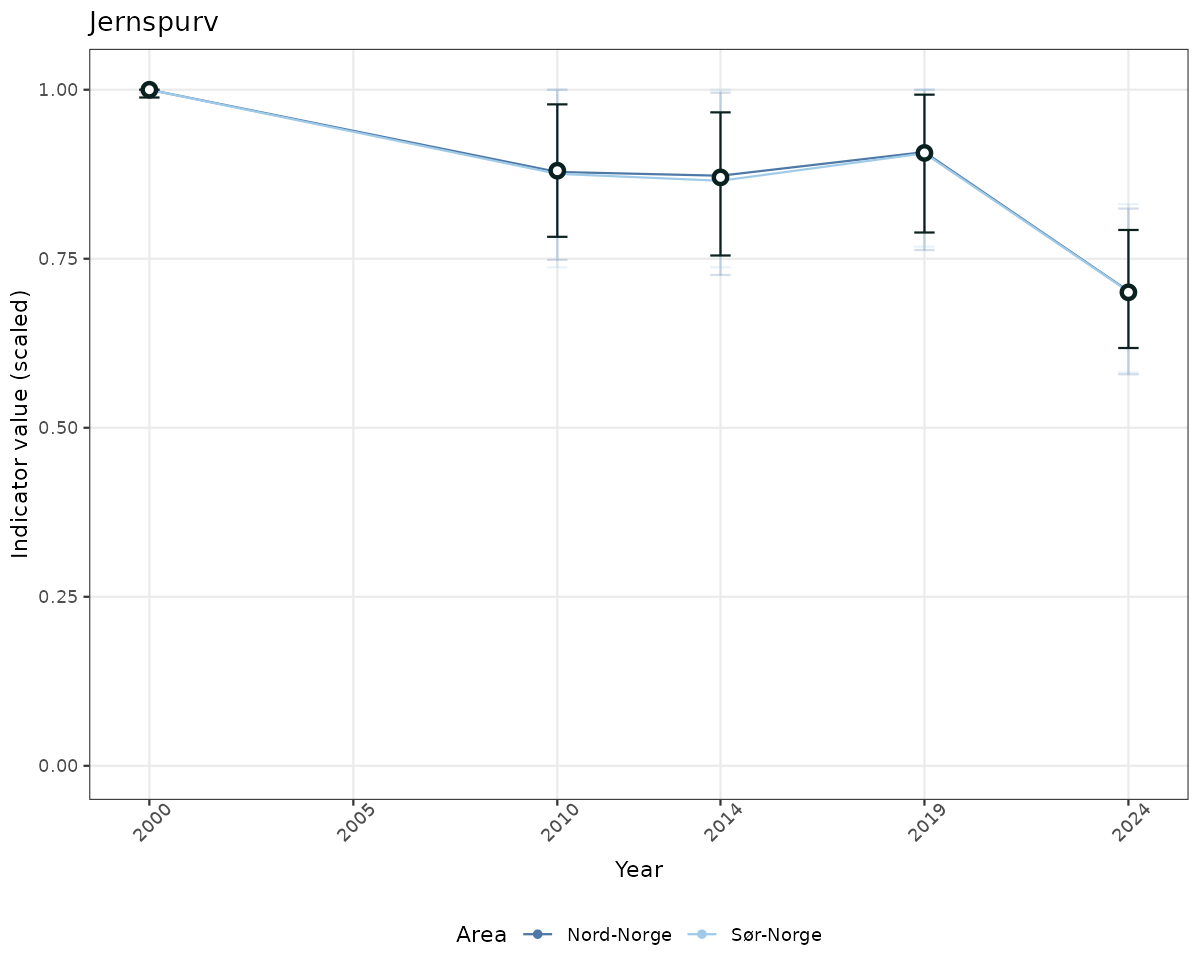
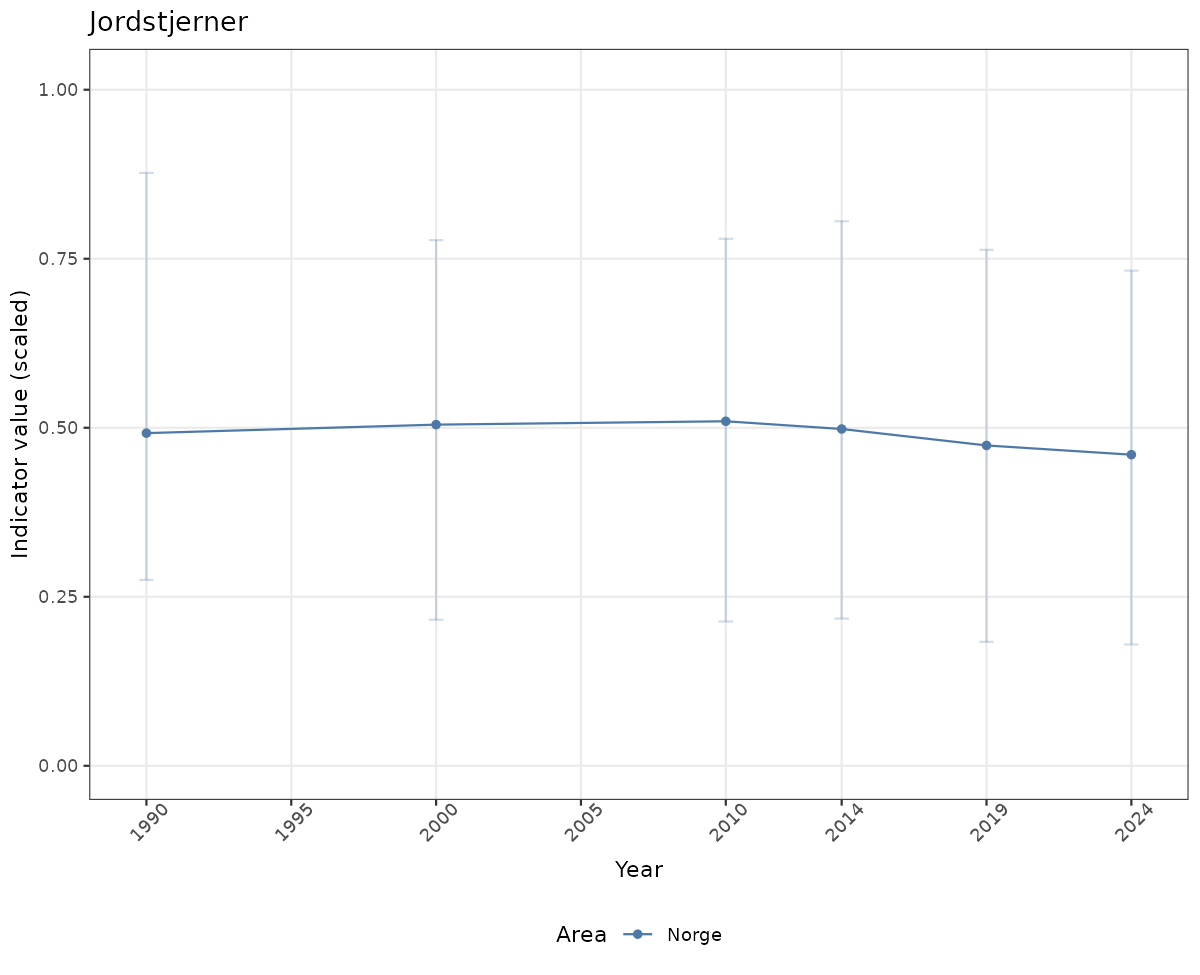
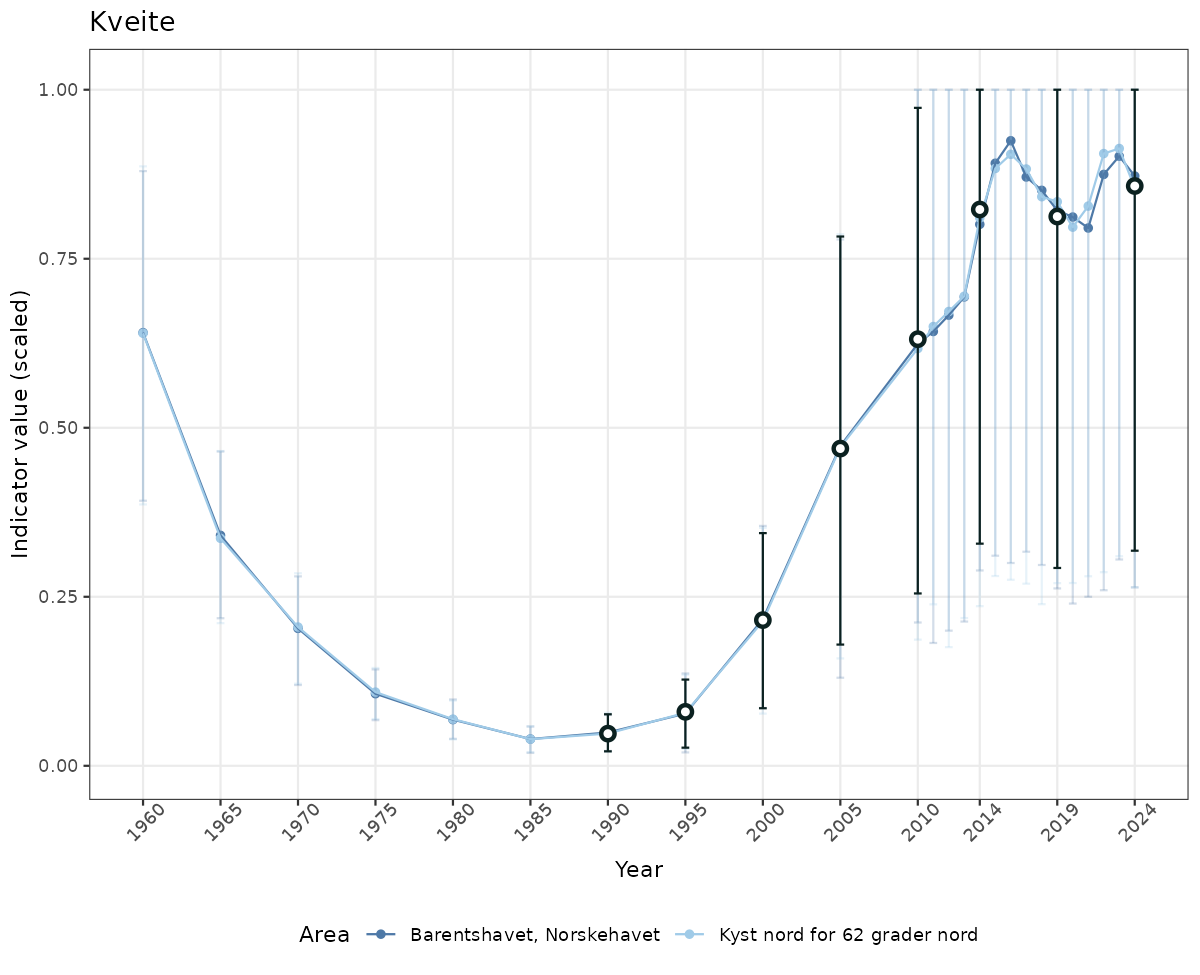
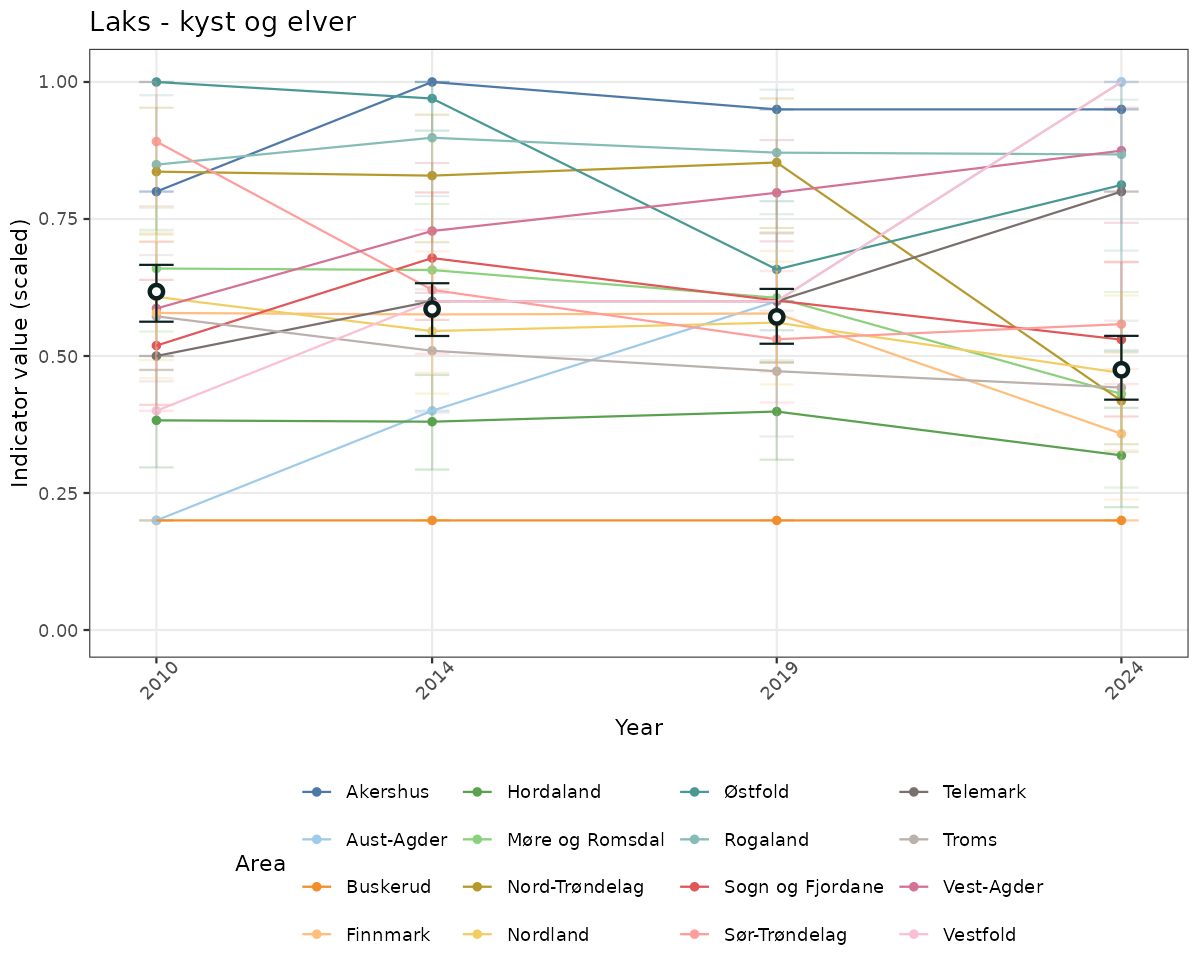
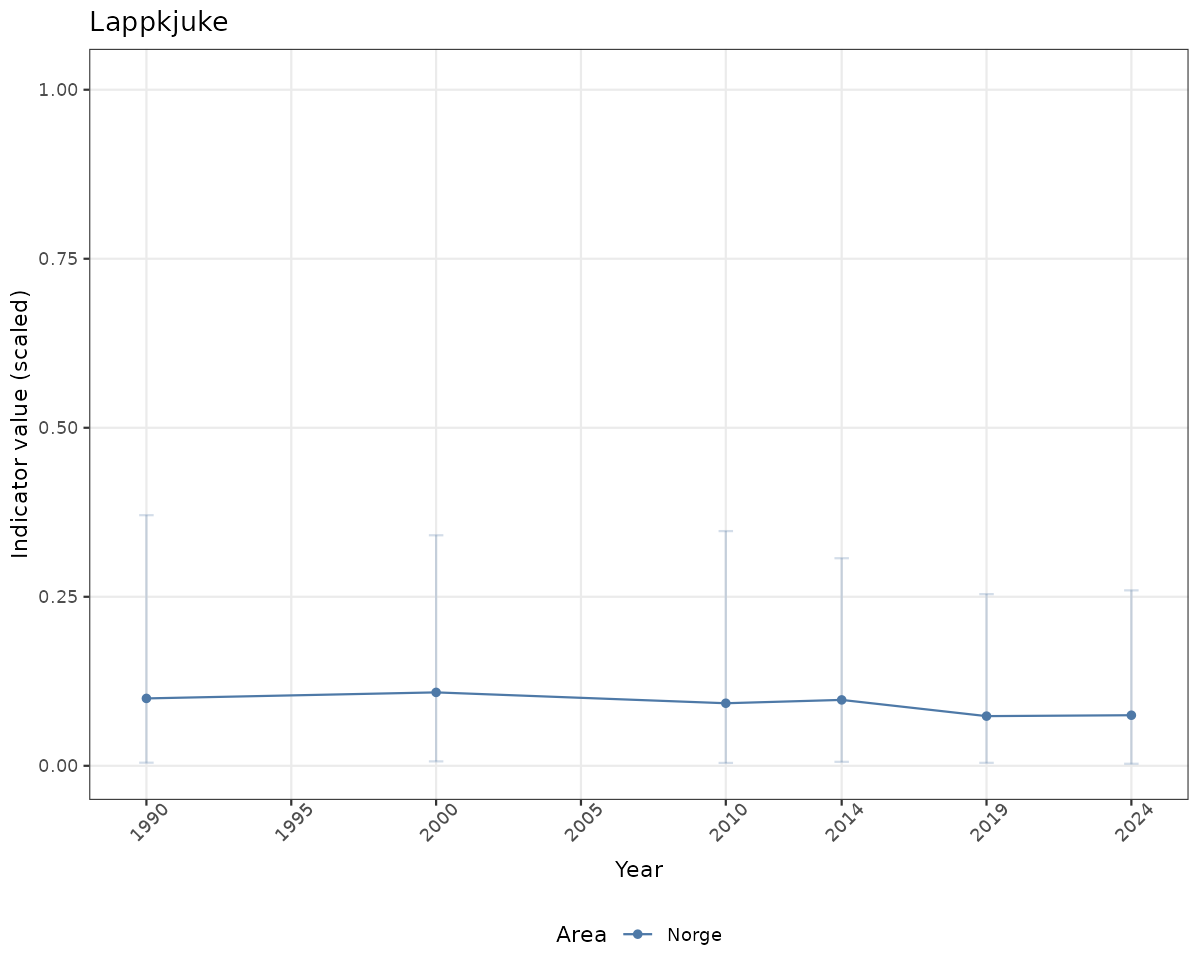
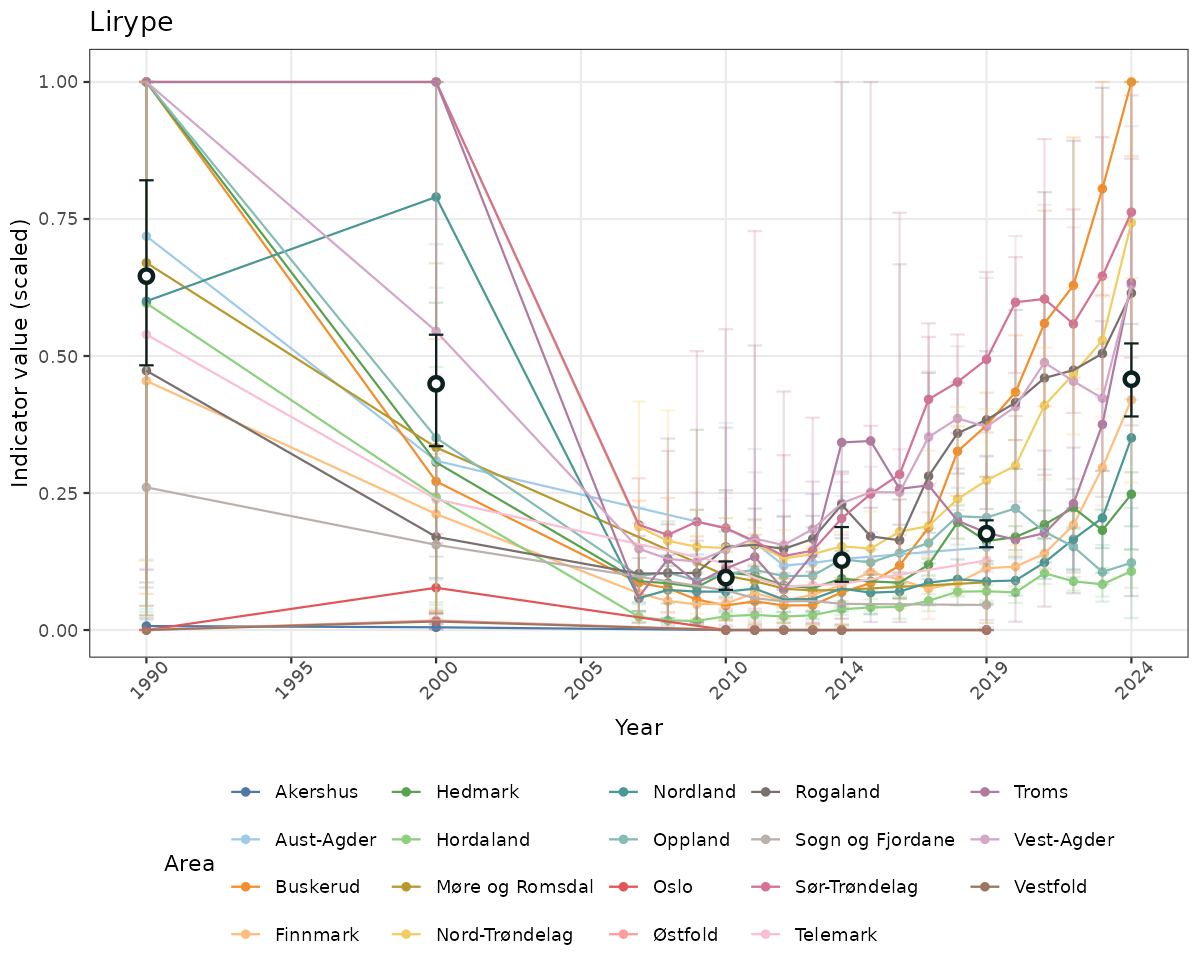
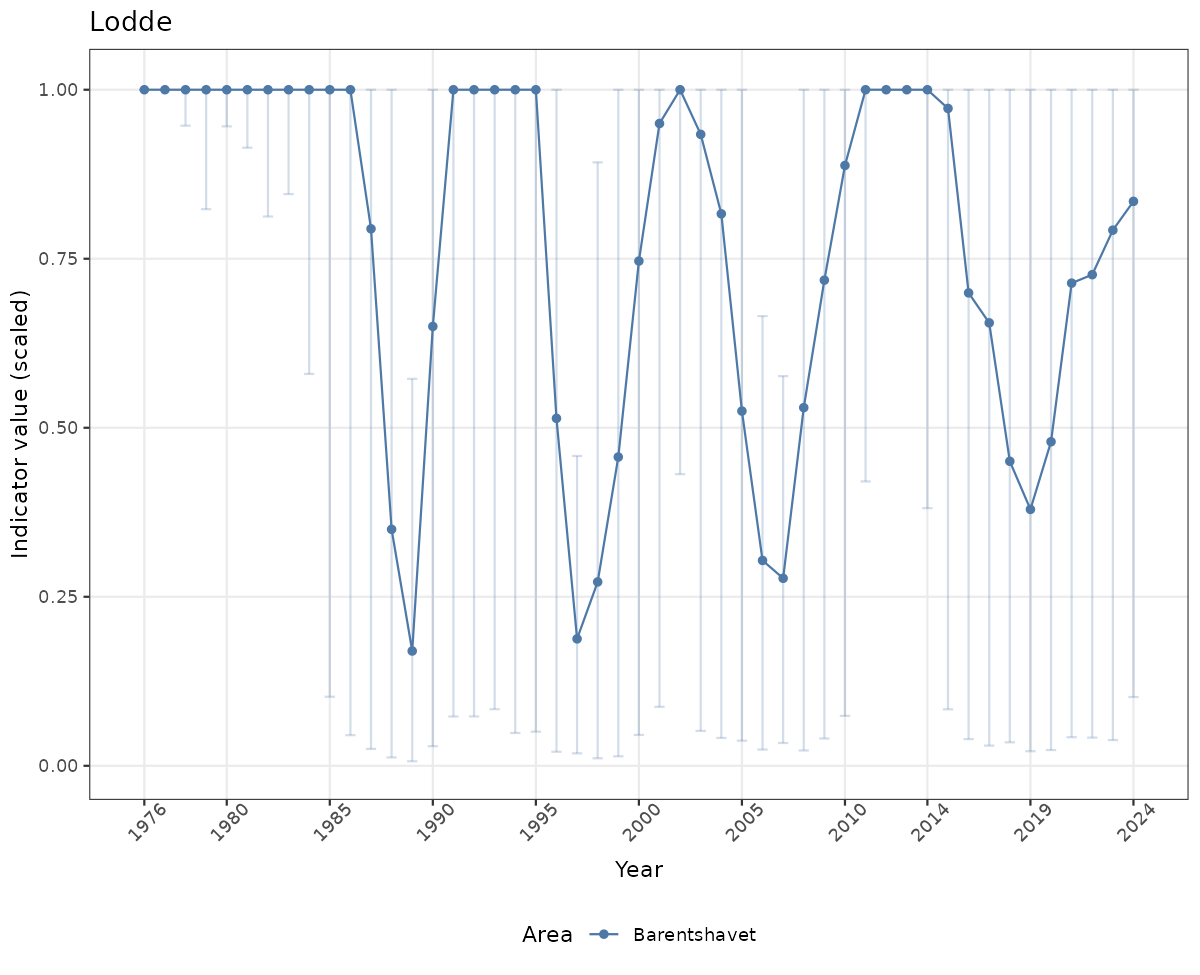
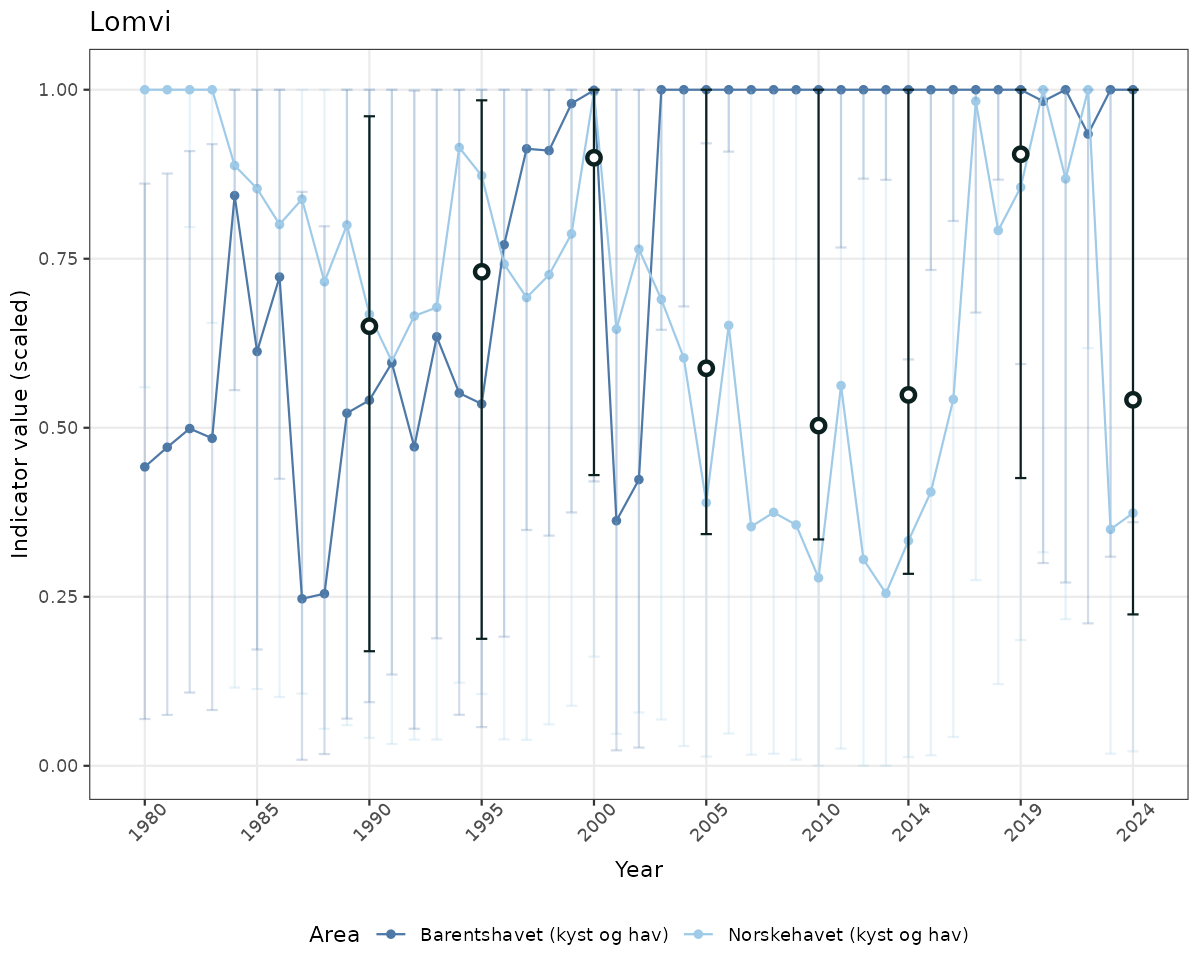
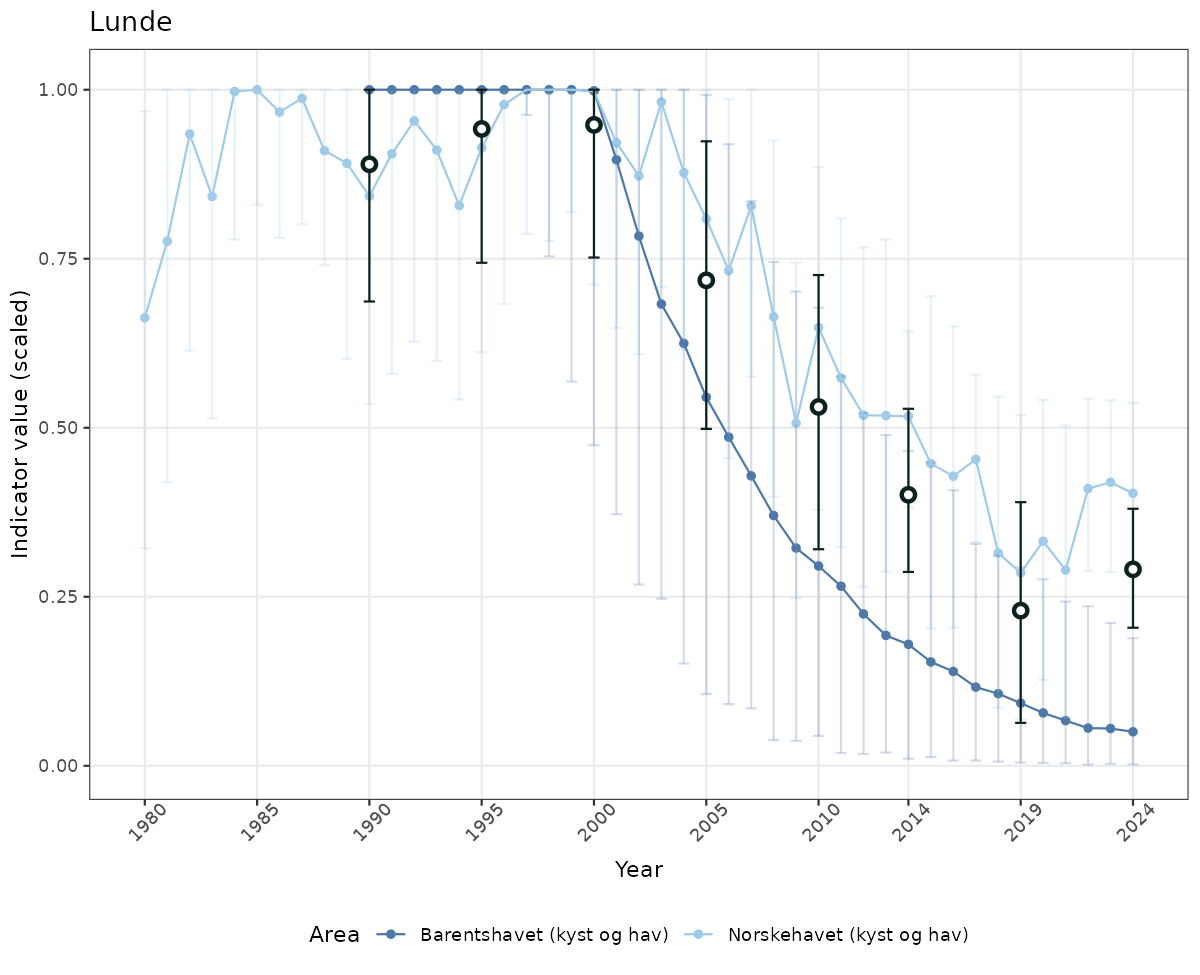
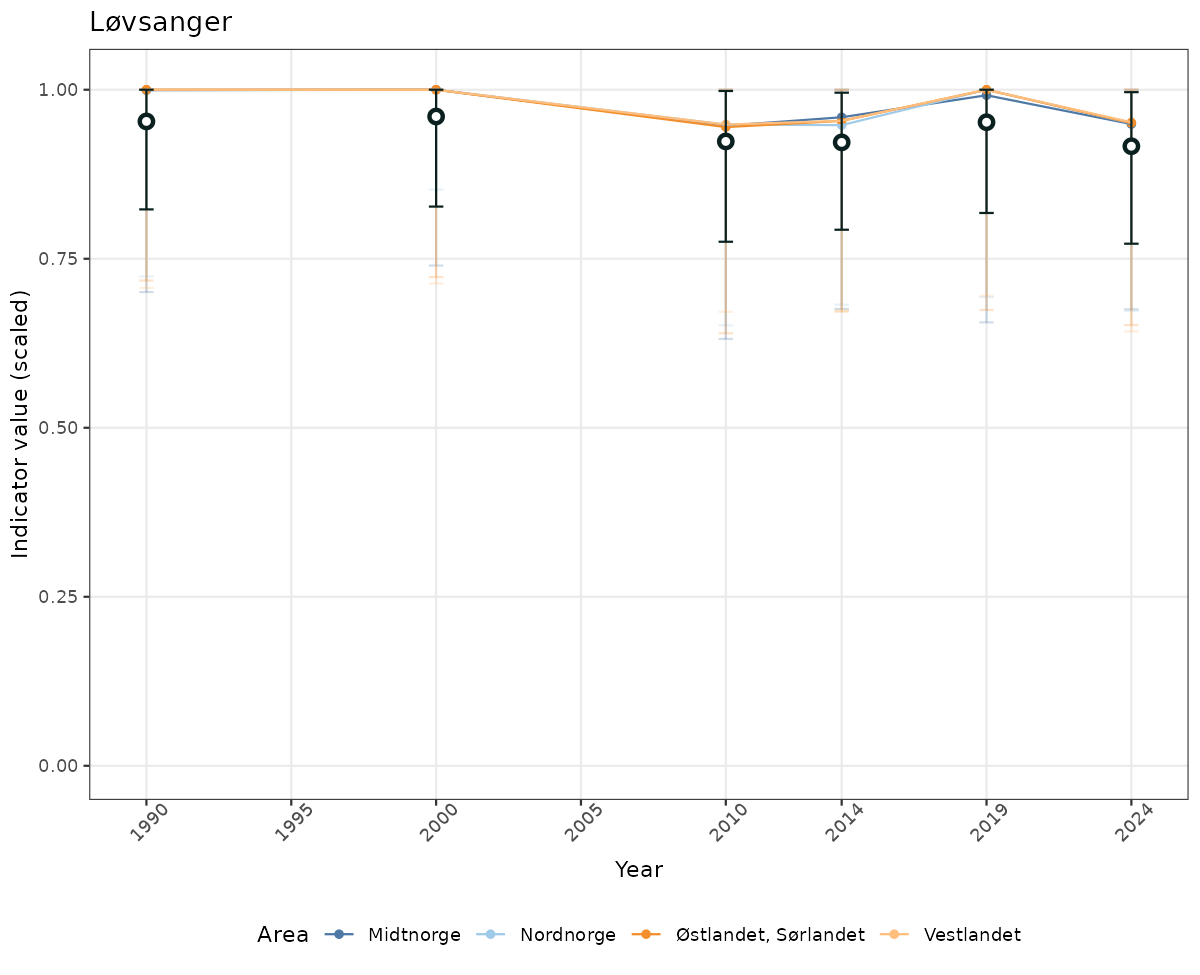
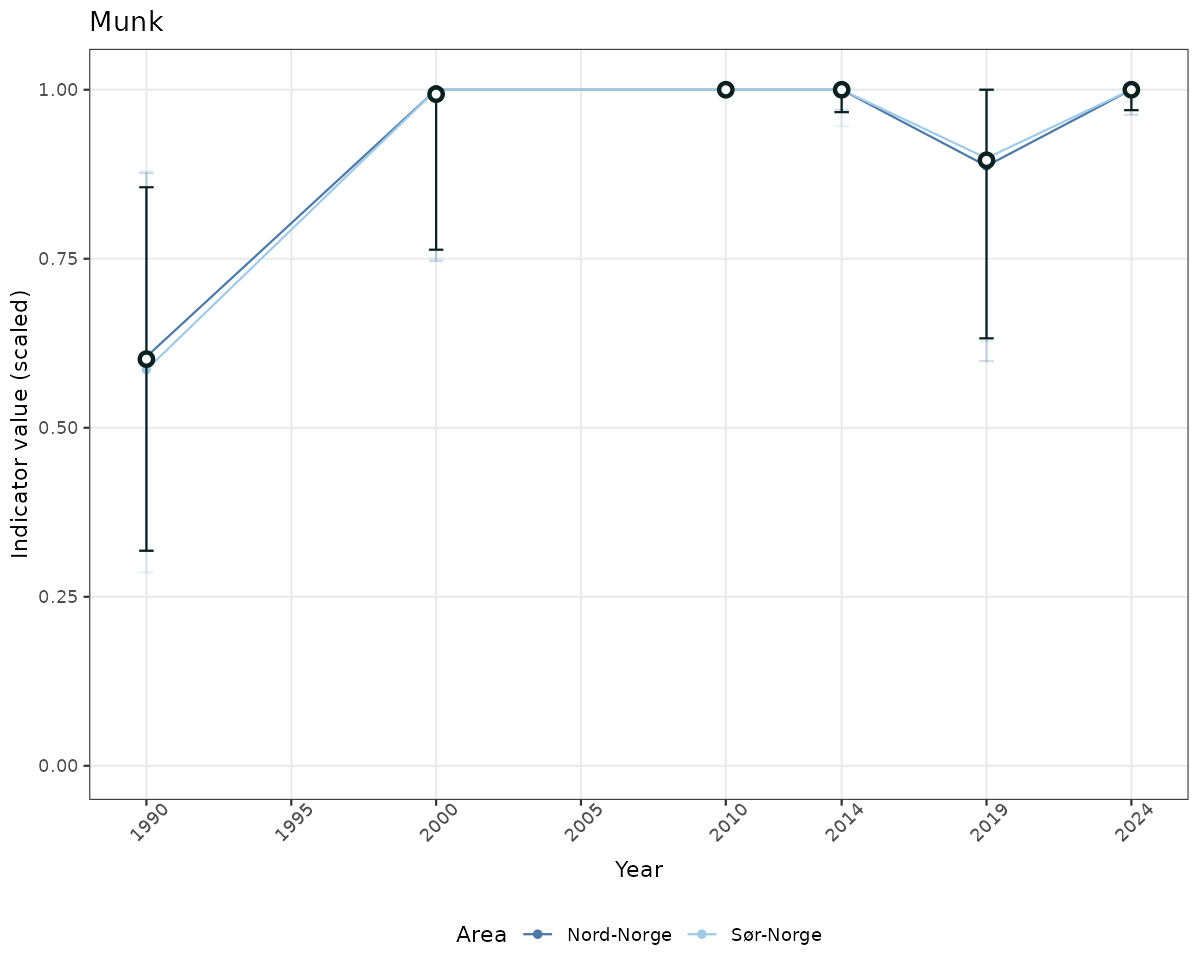
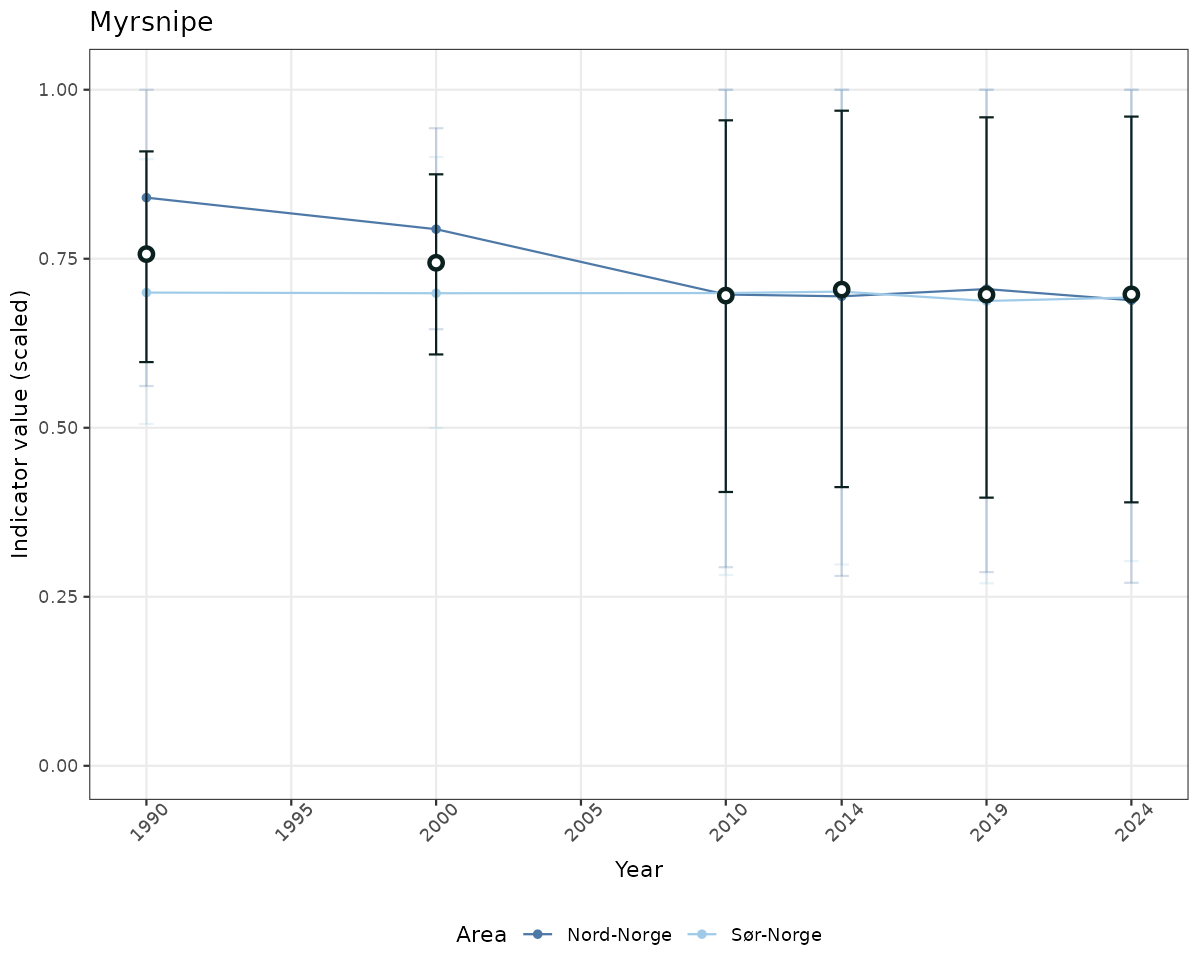
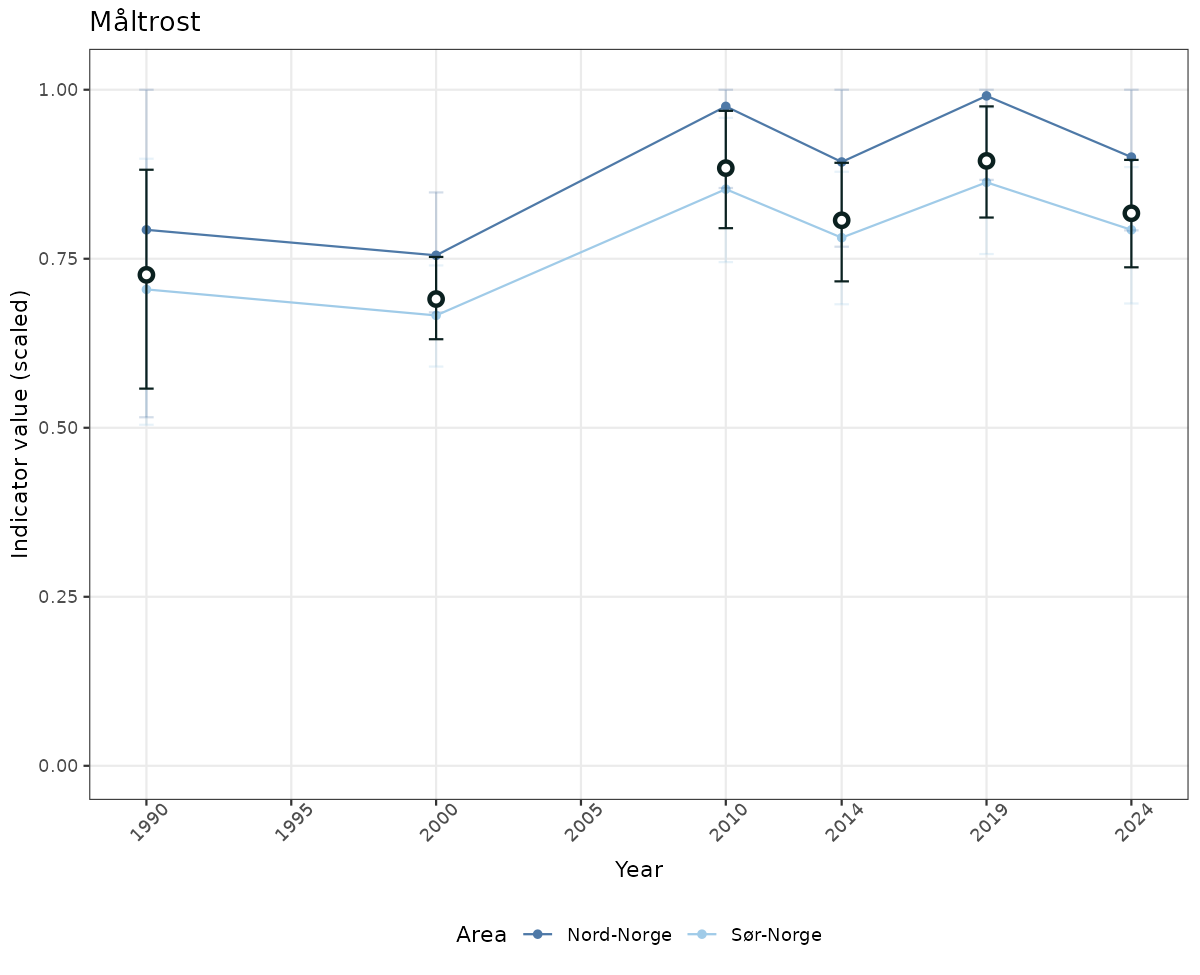
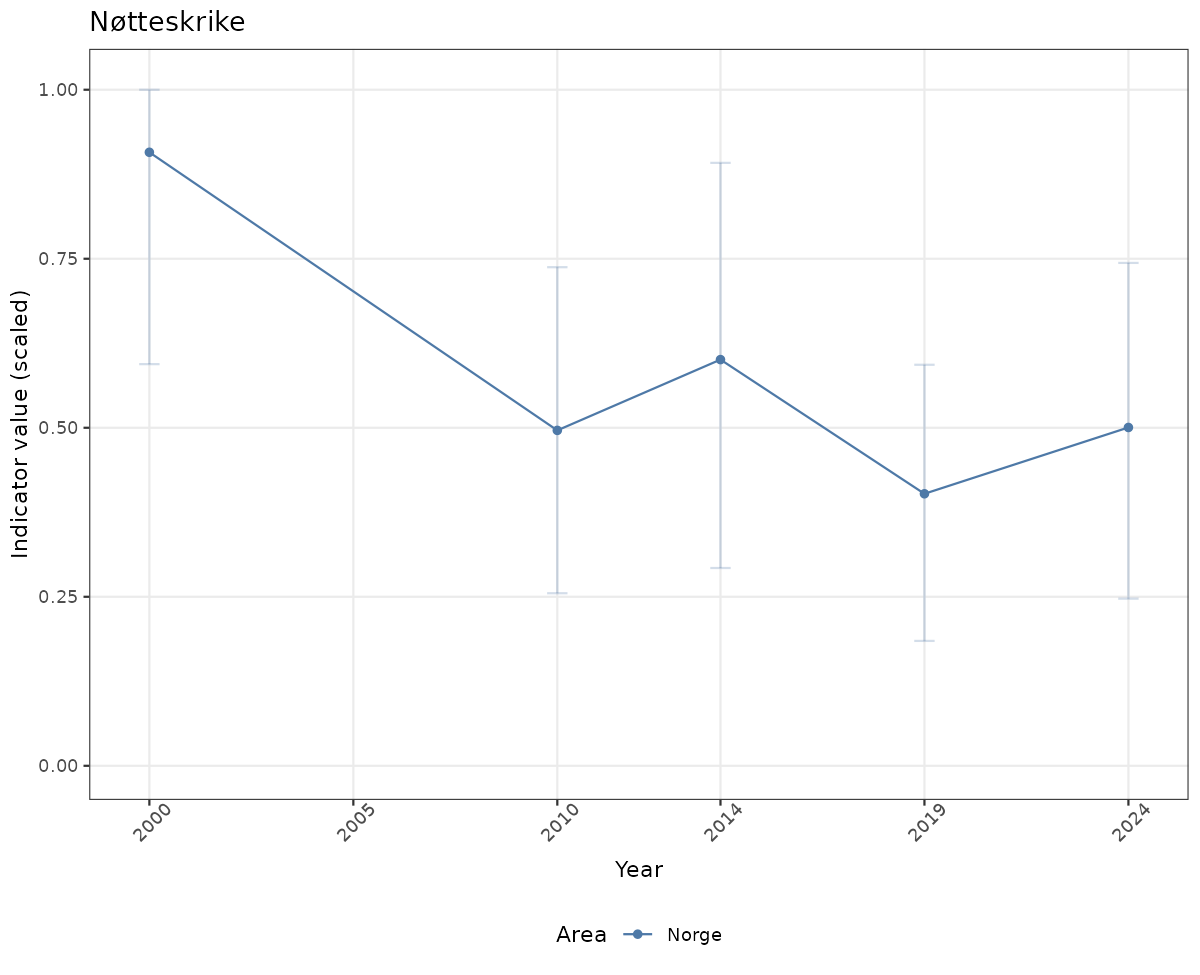
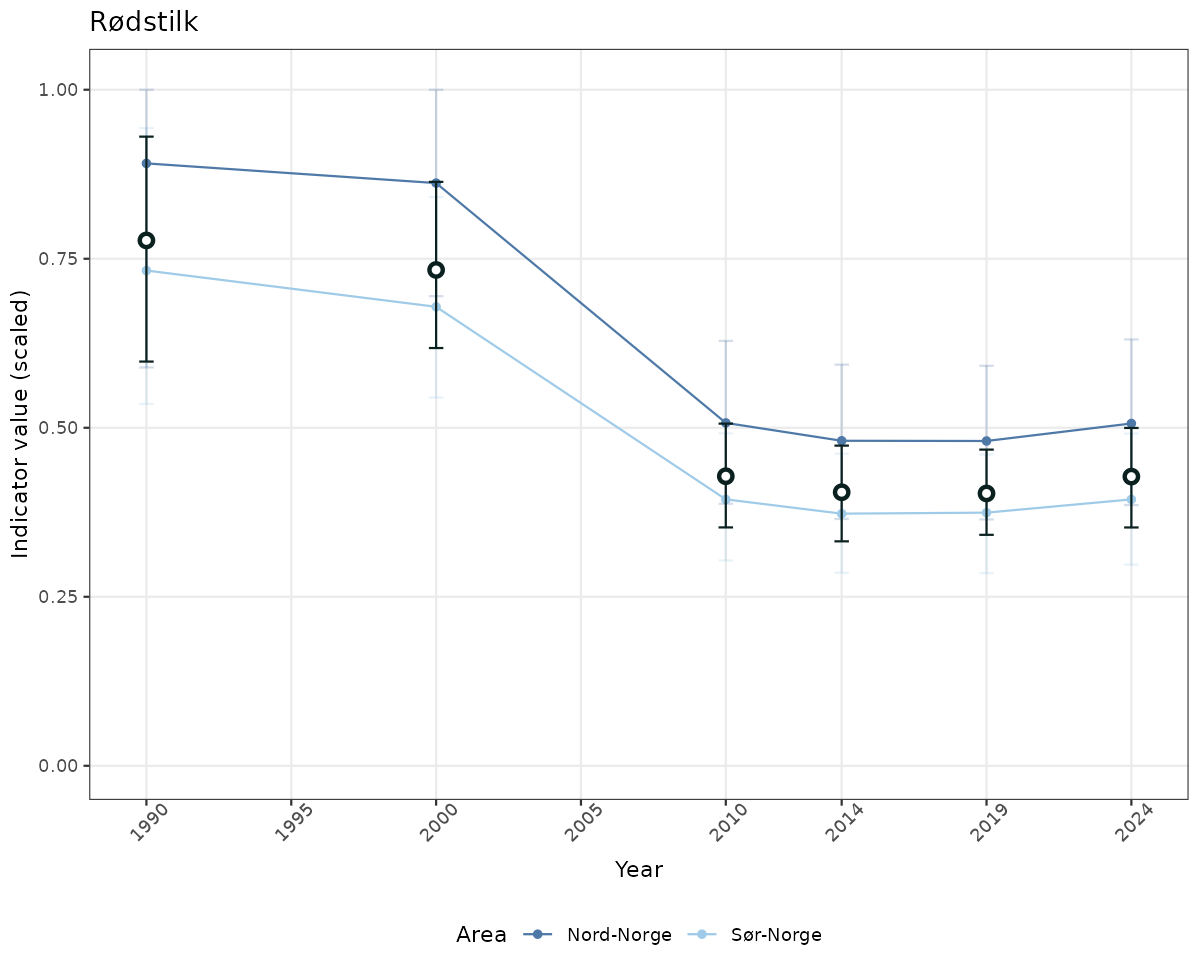
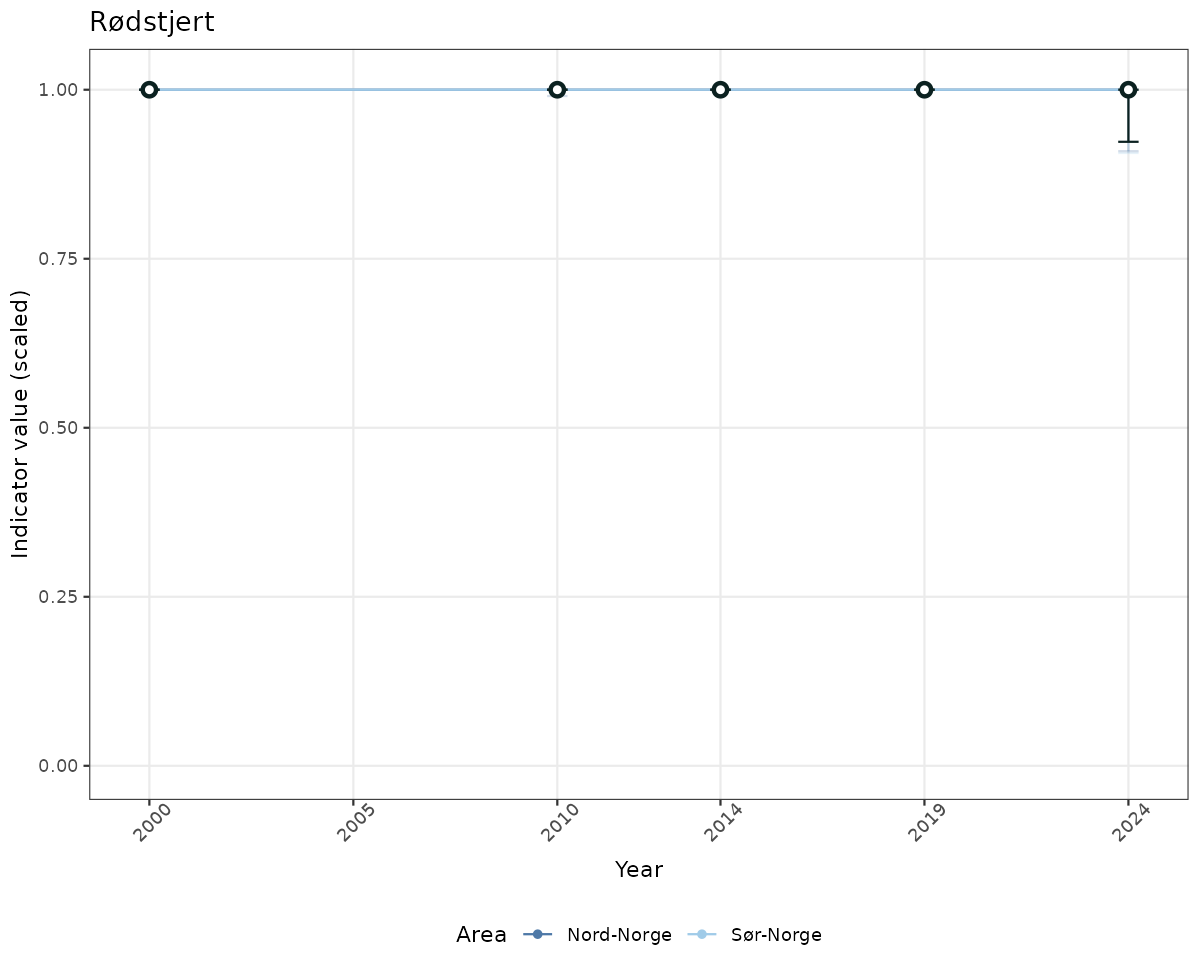
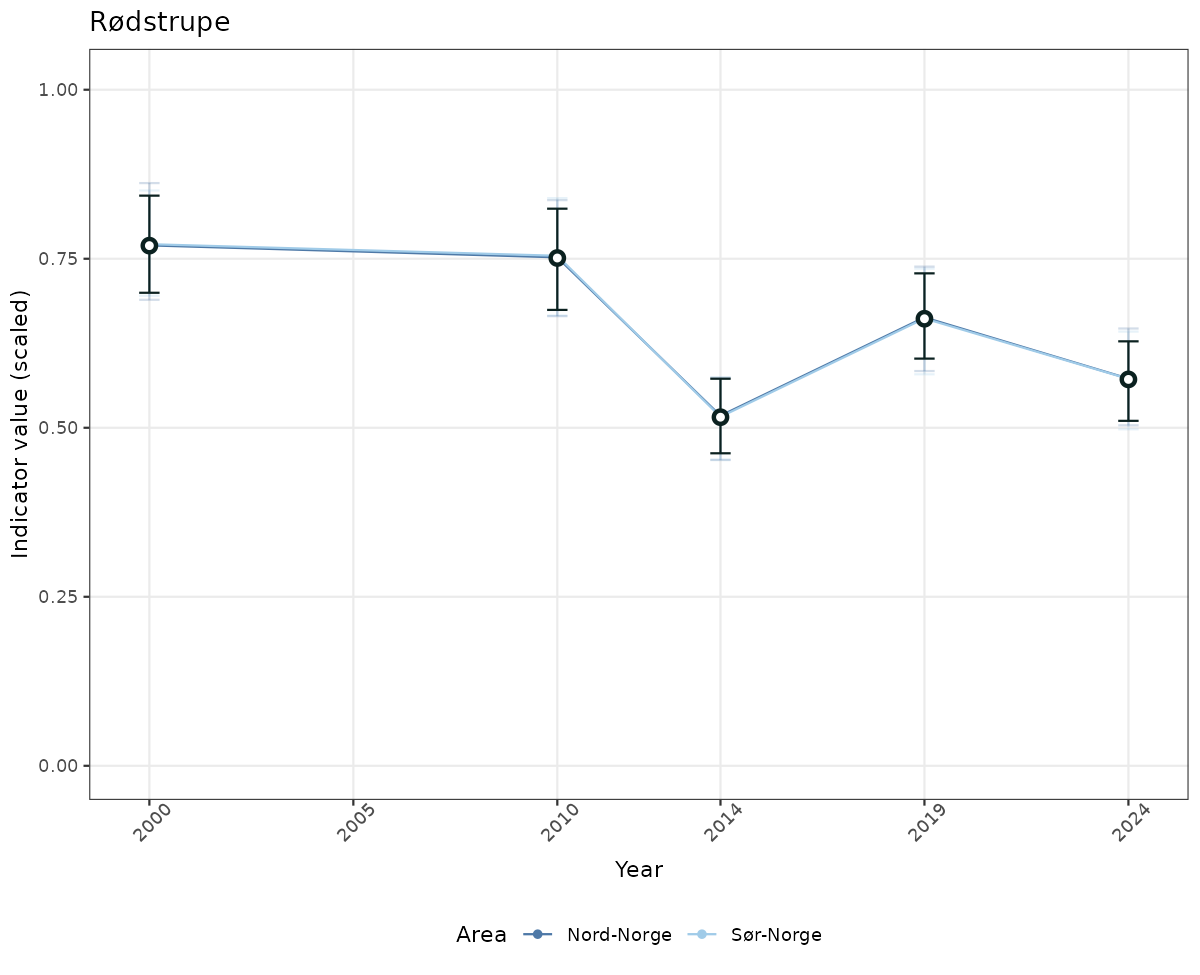
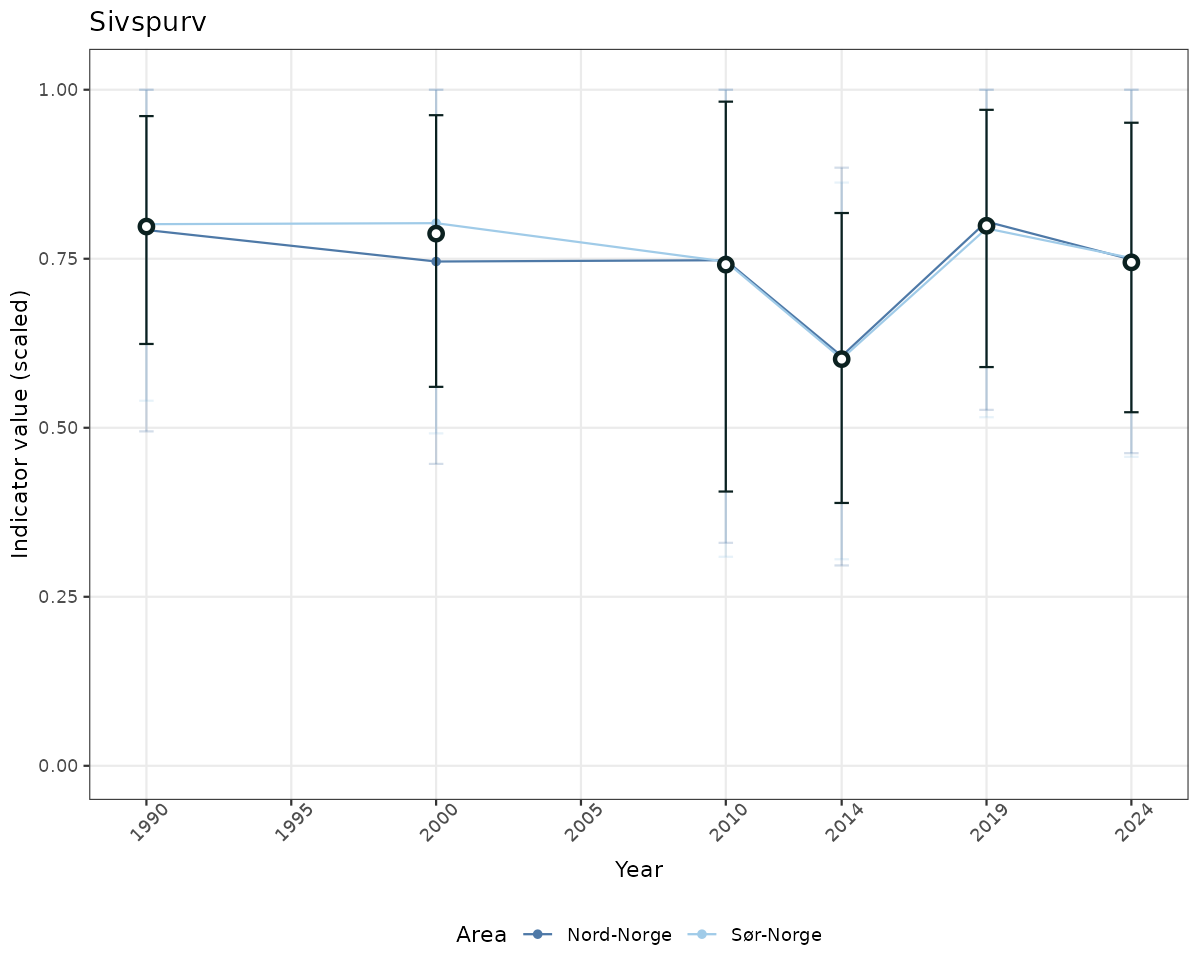
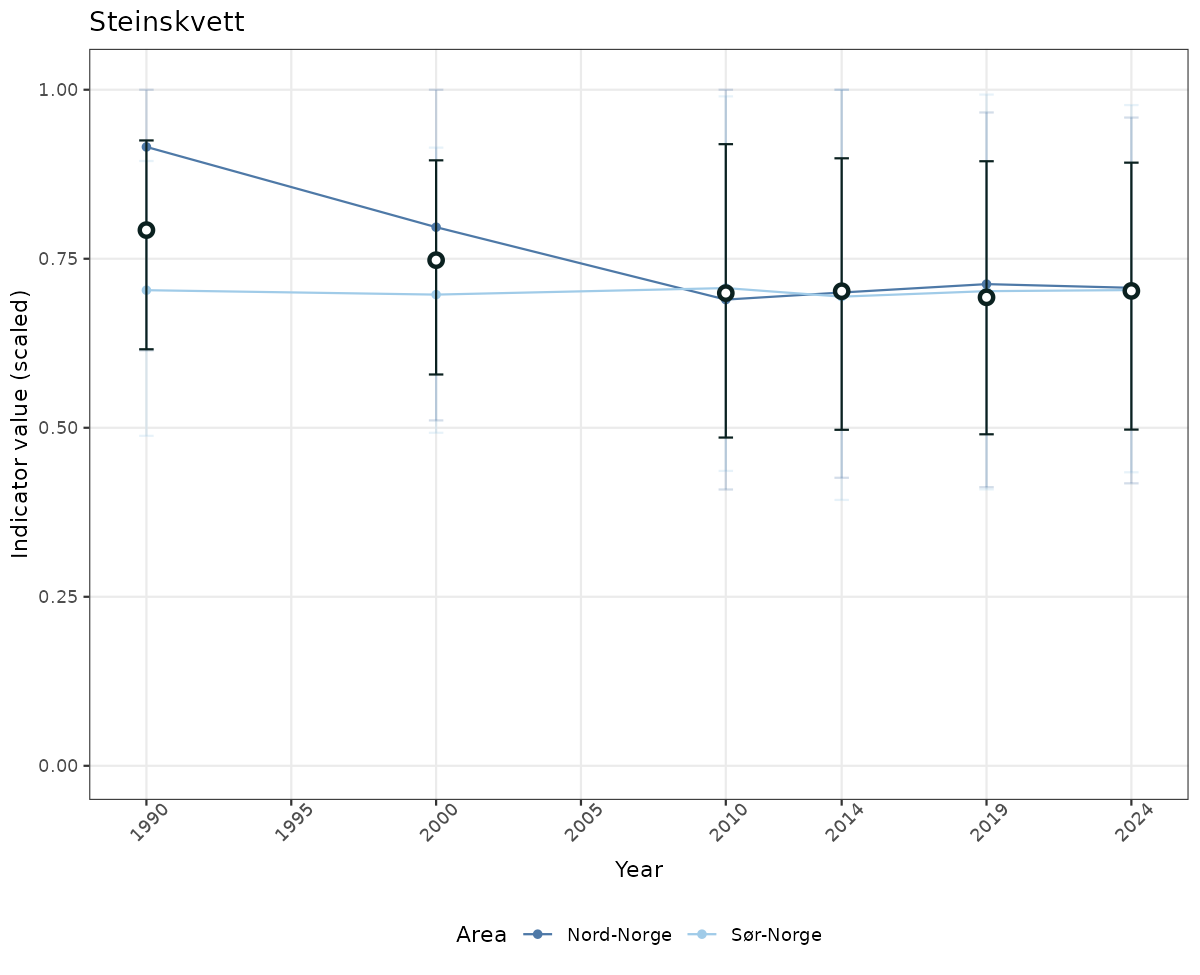
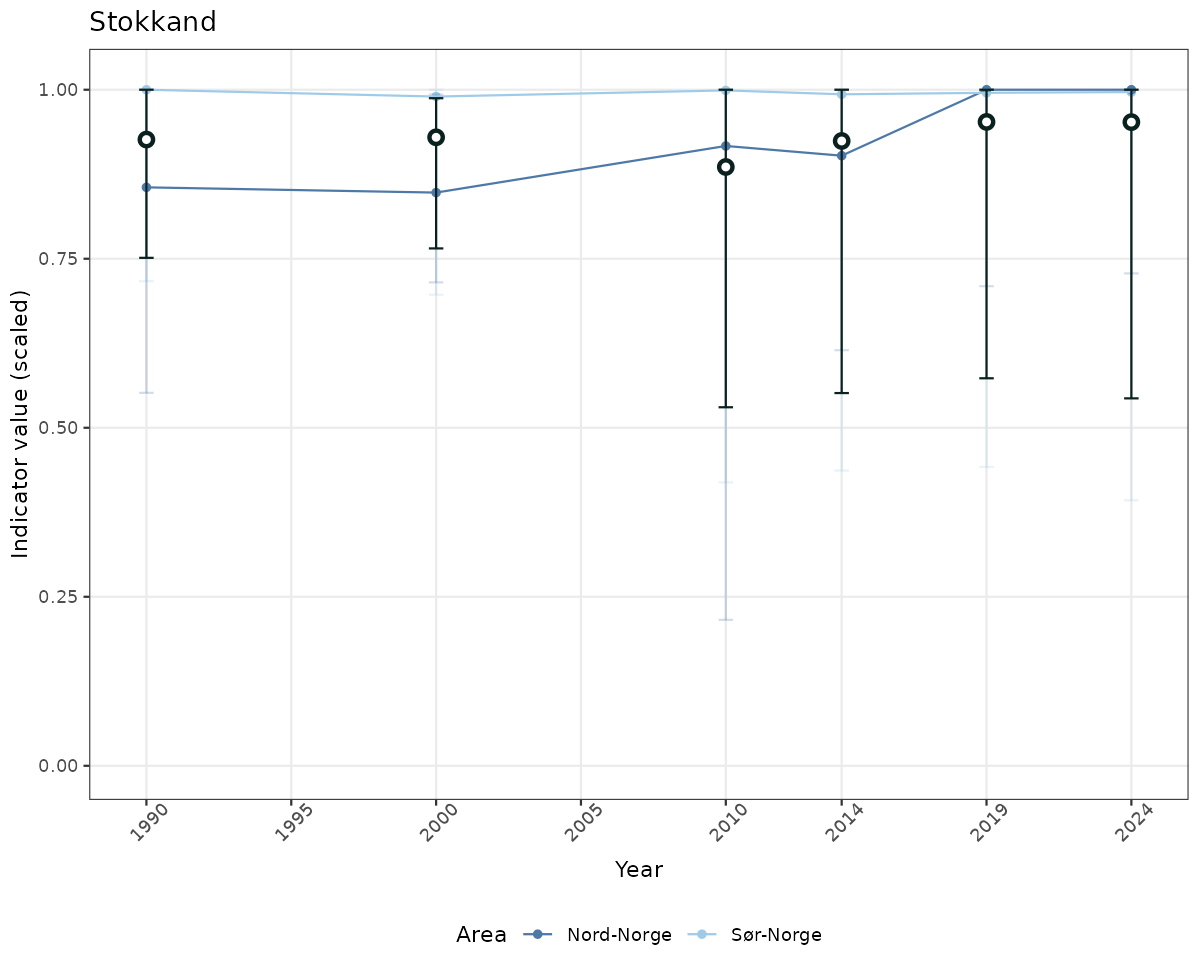
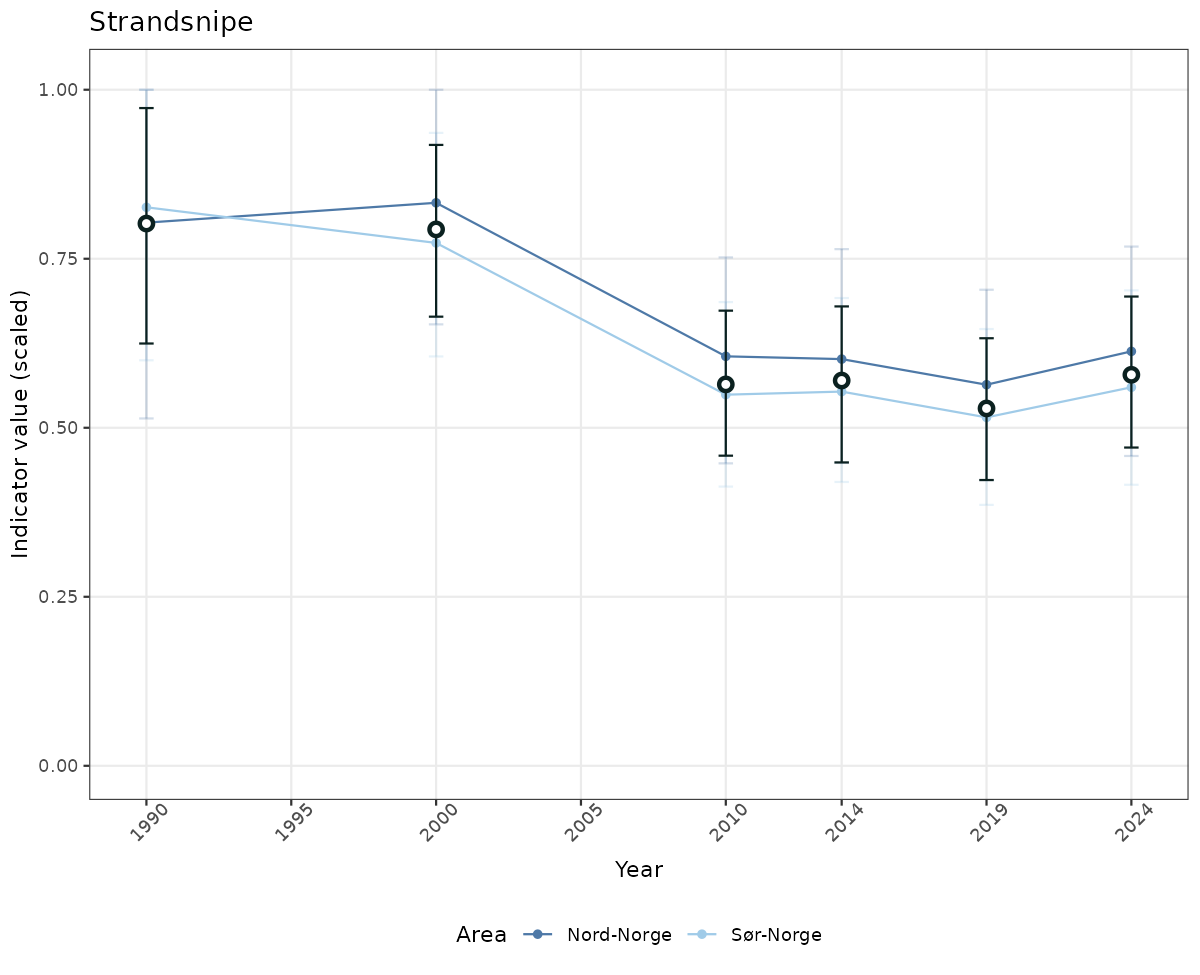
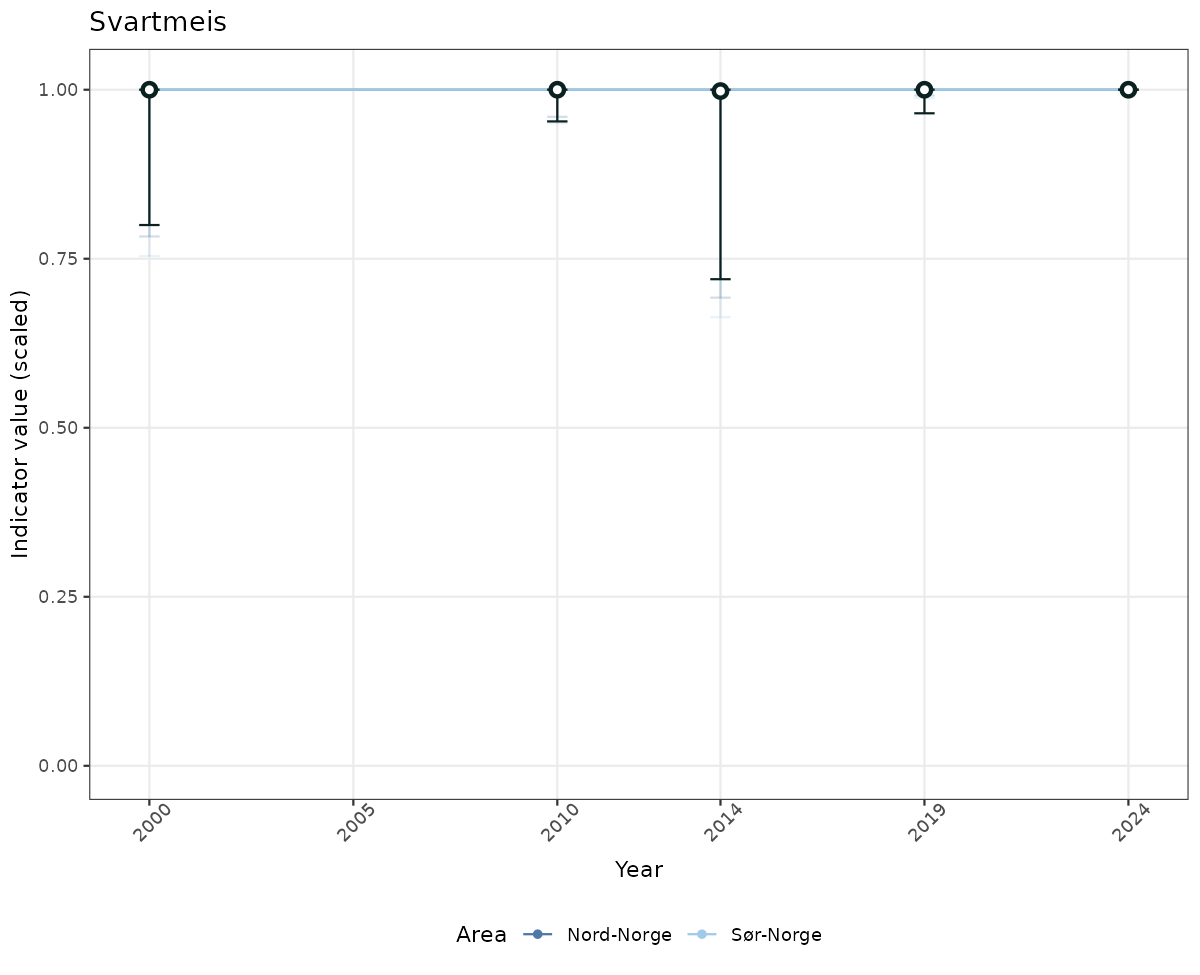
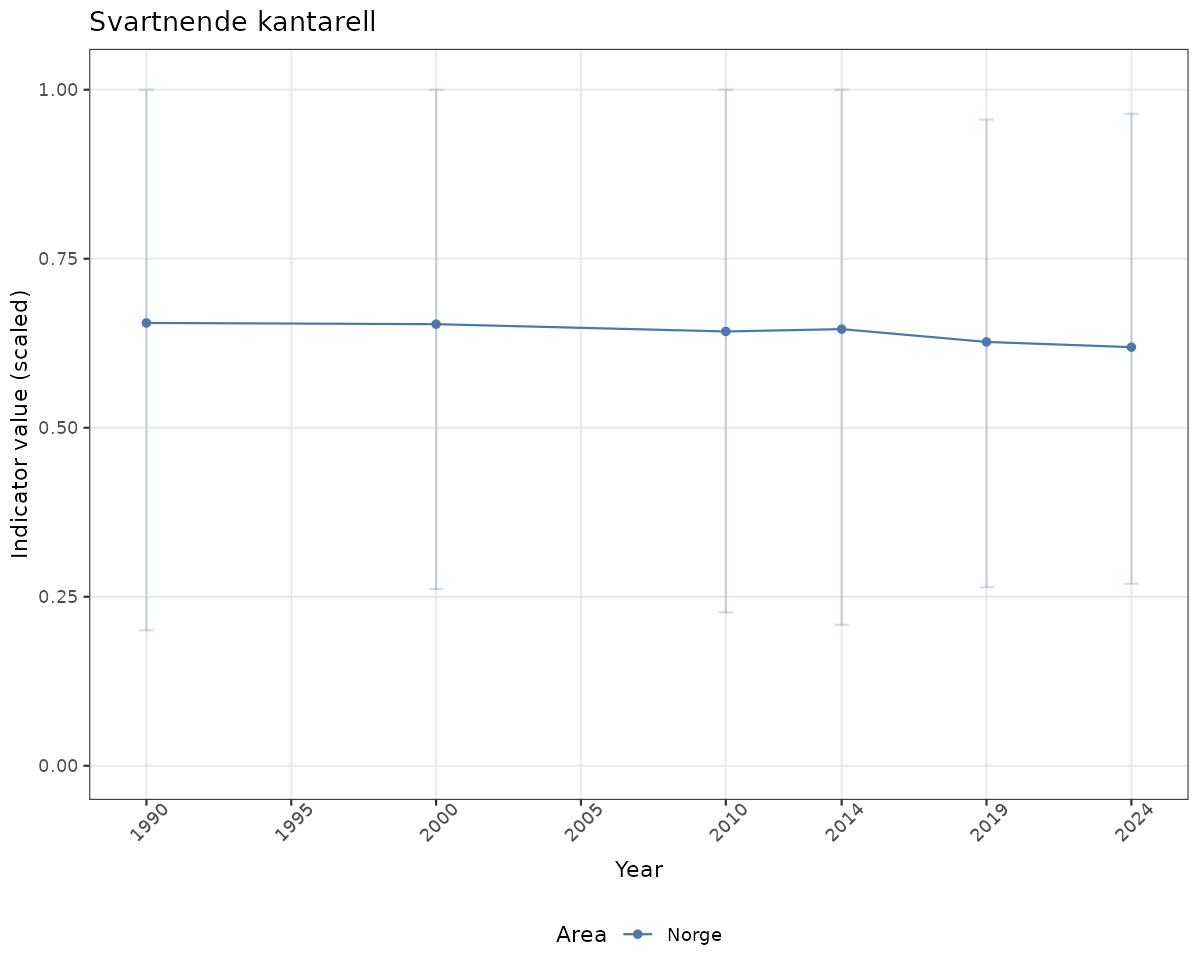
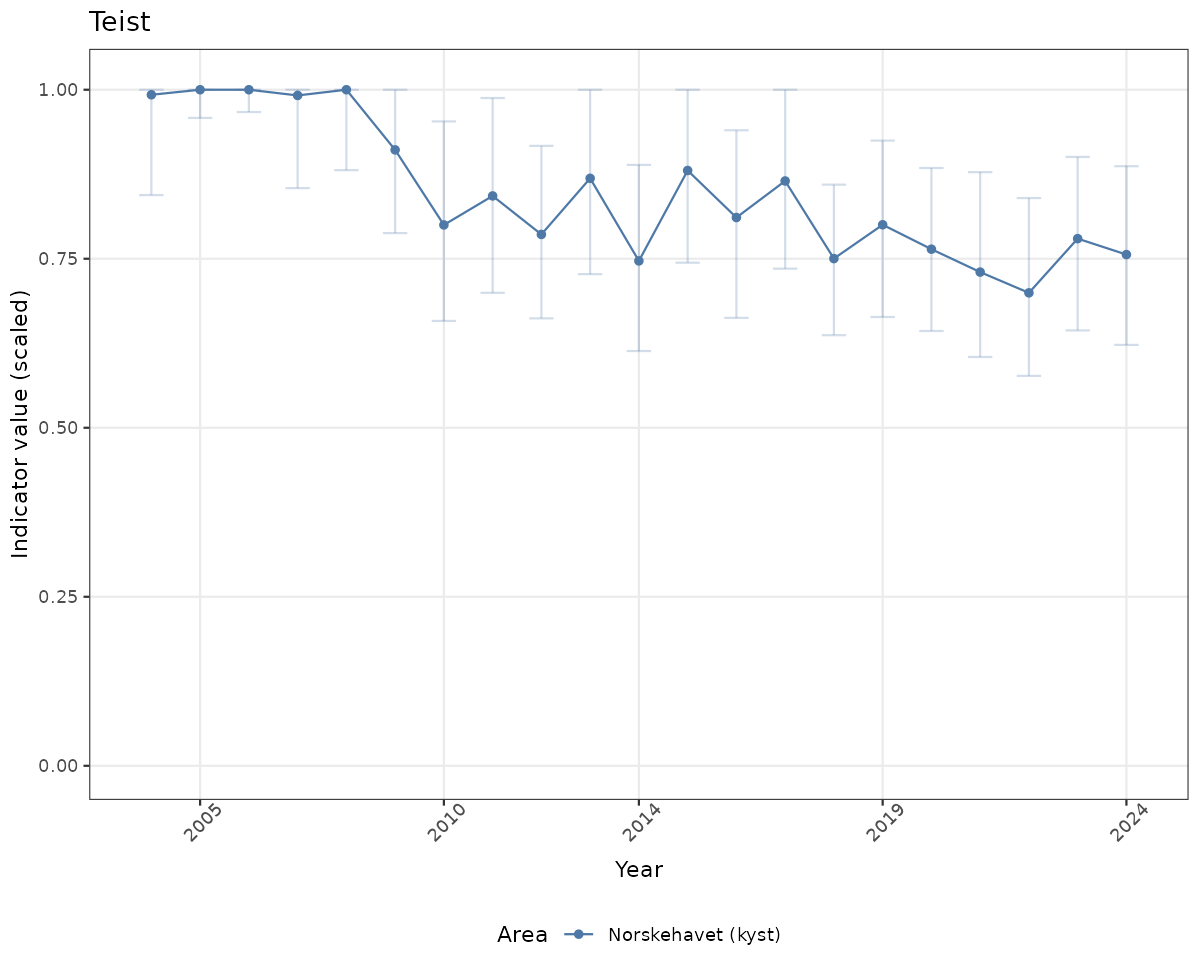
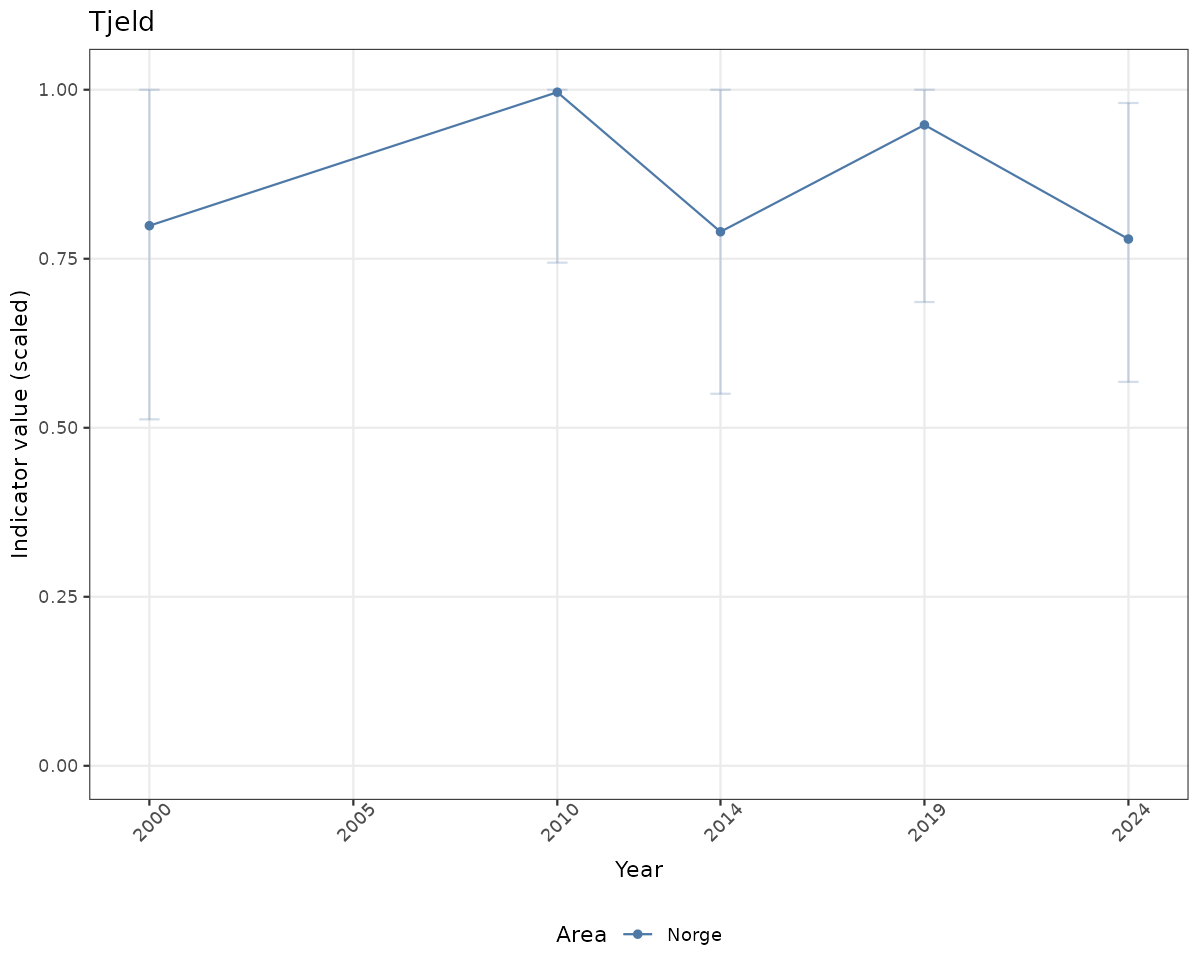
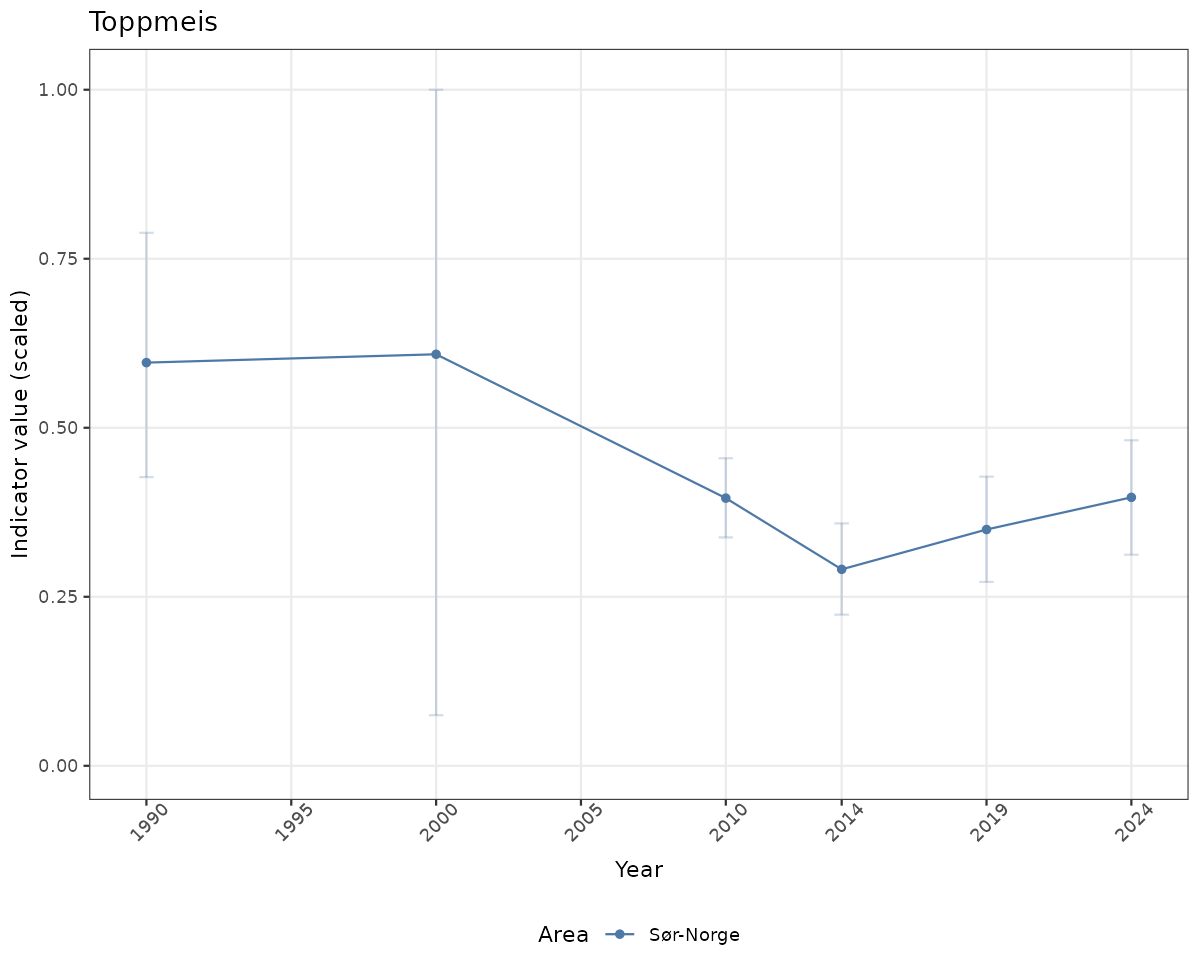
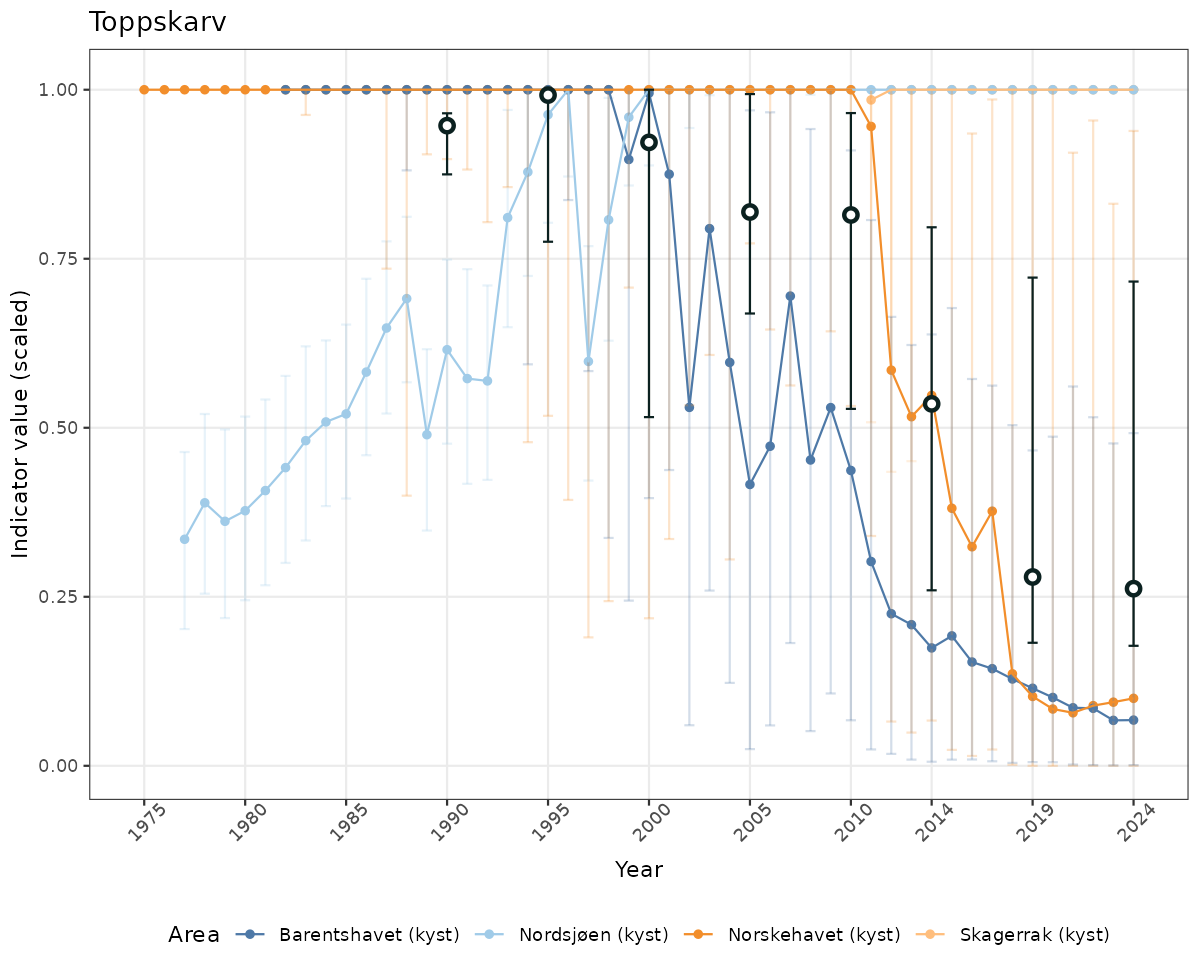
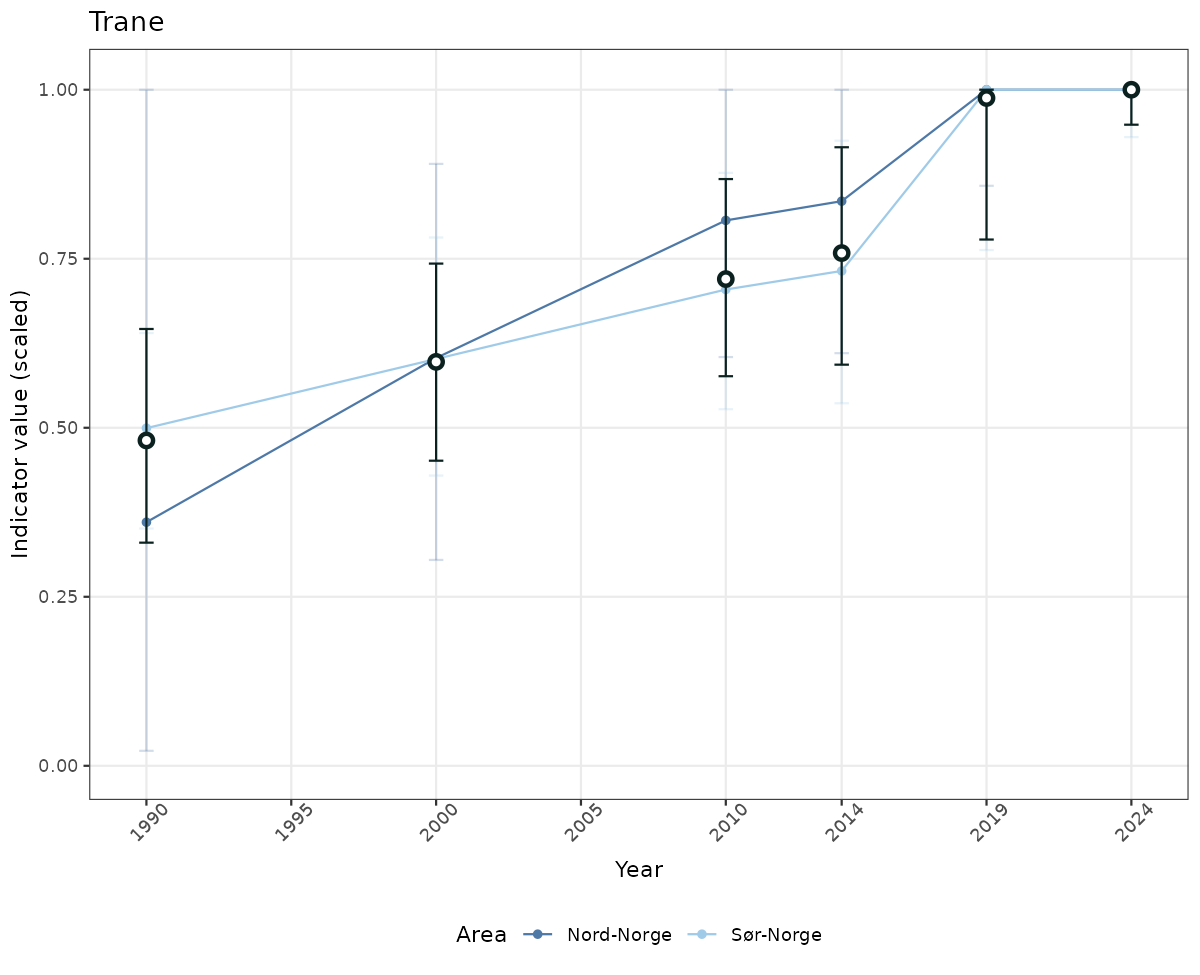
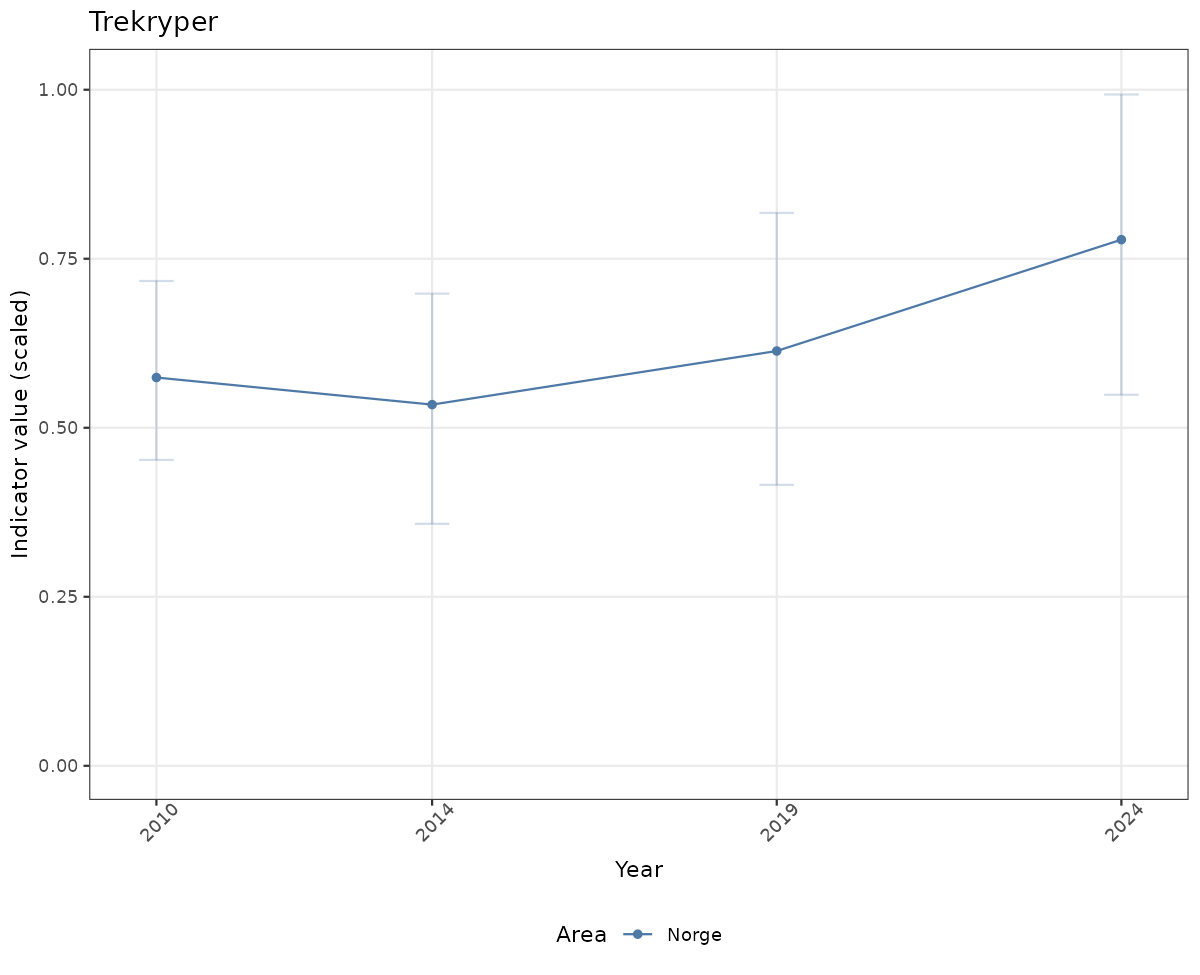
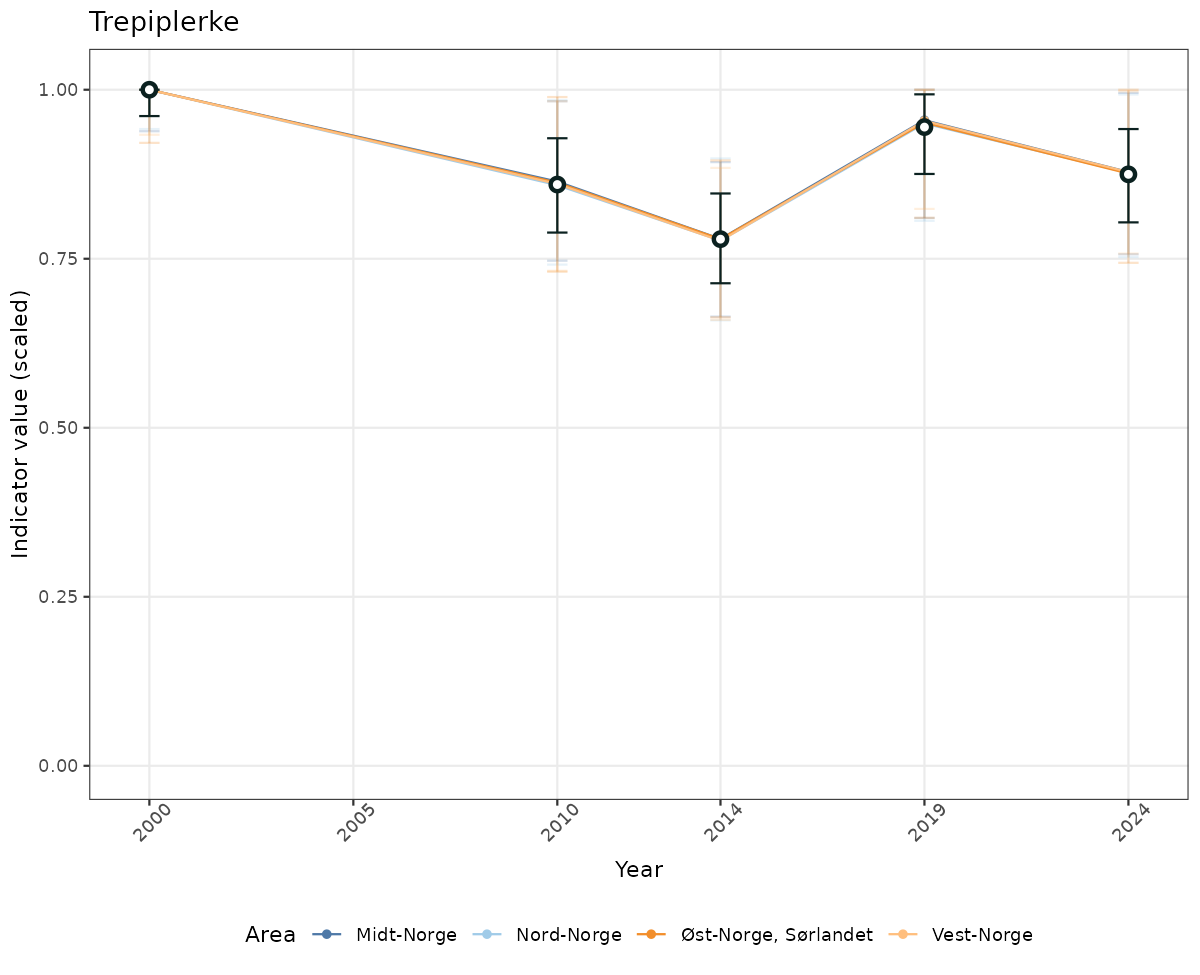
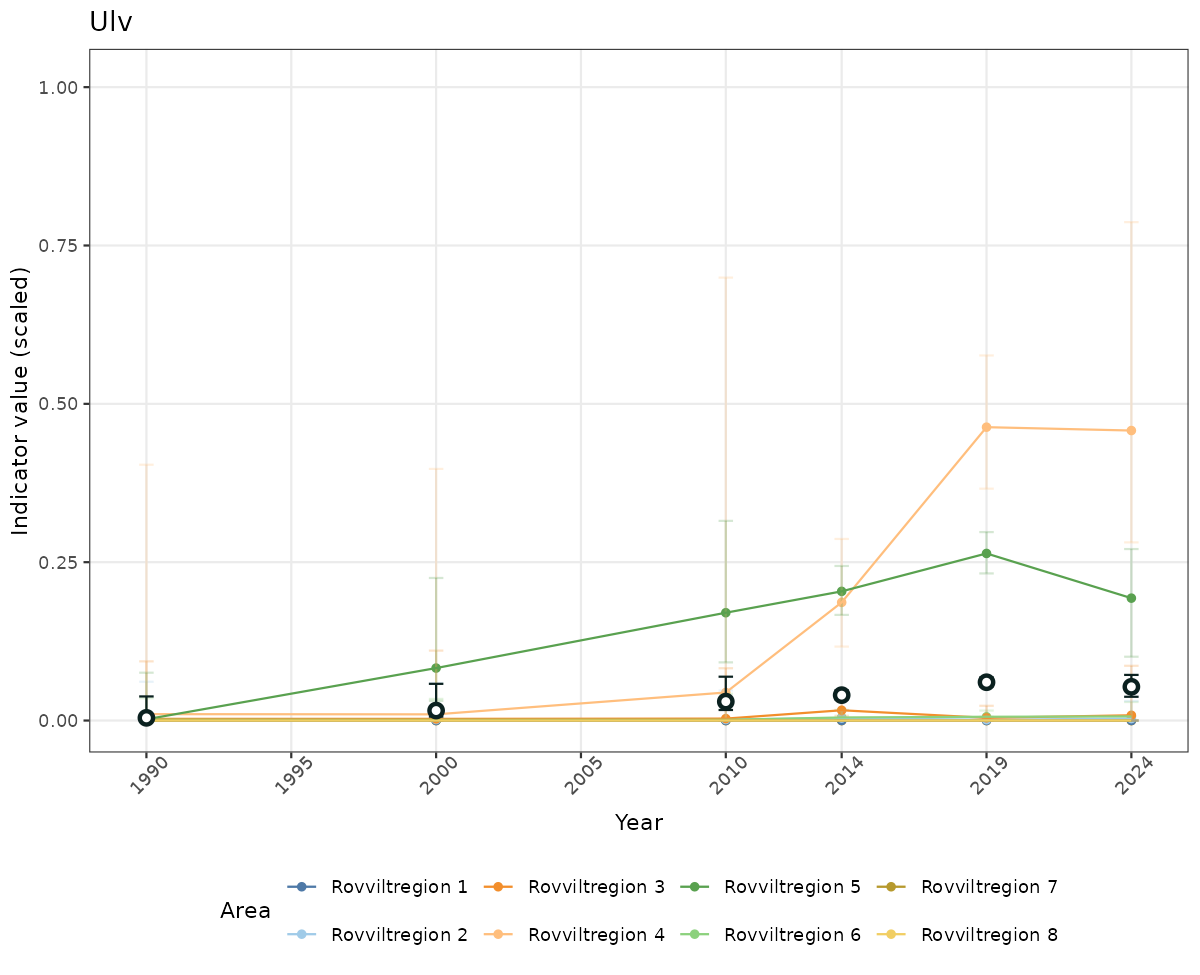
# Scaled data
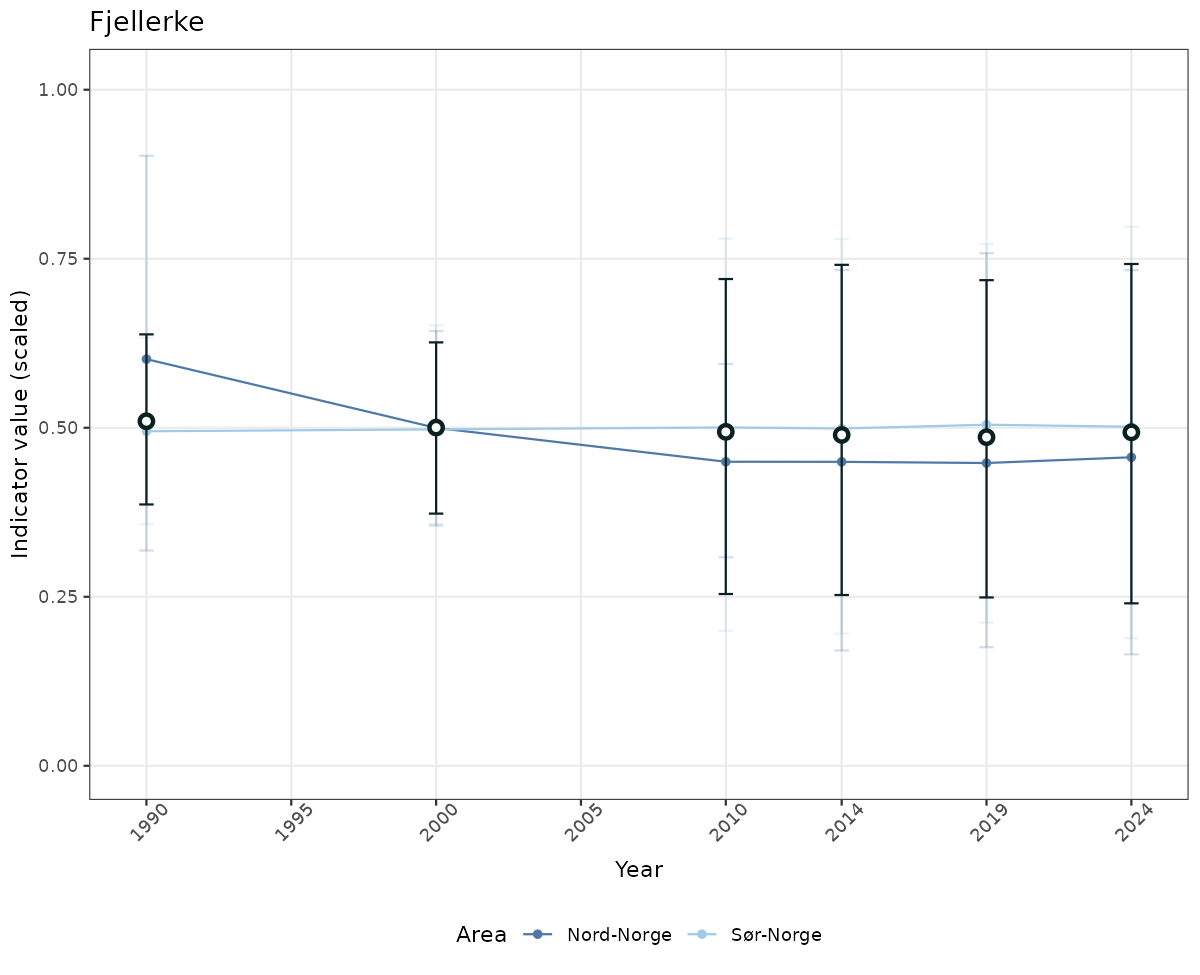
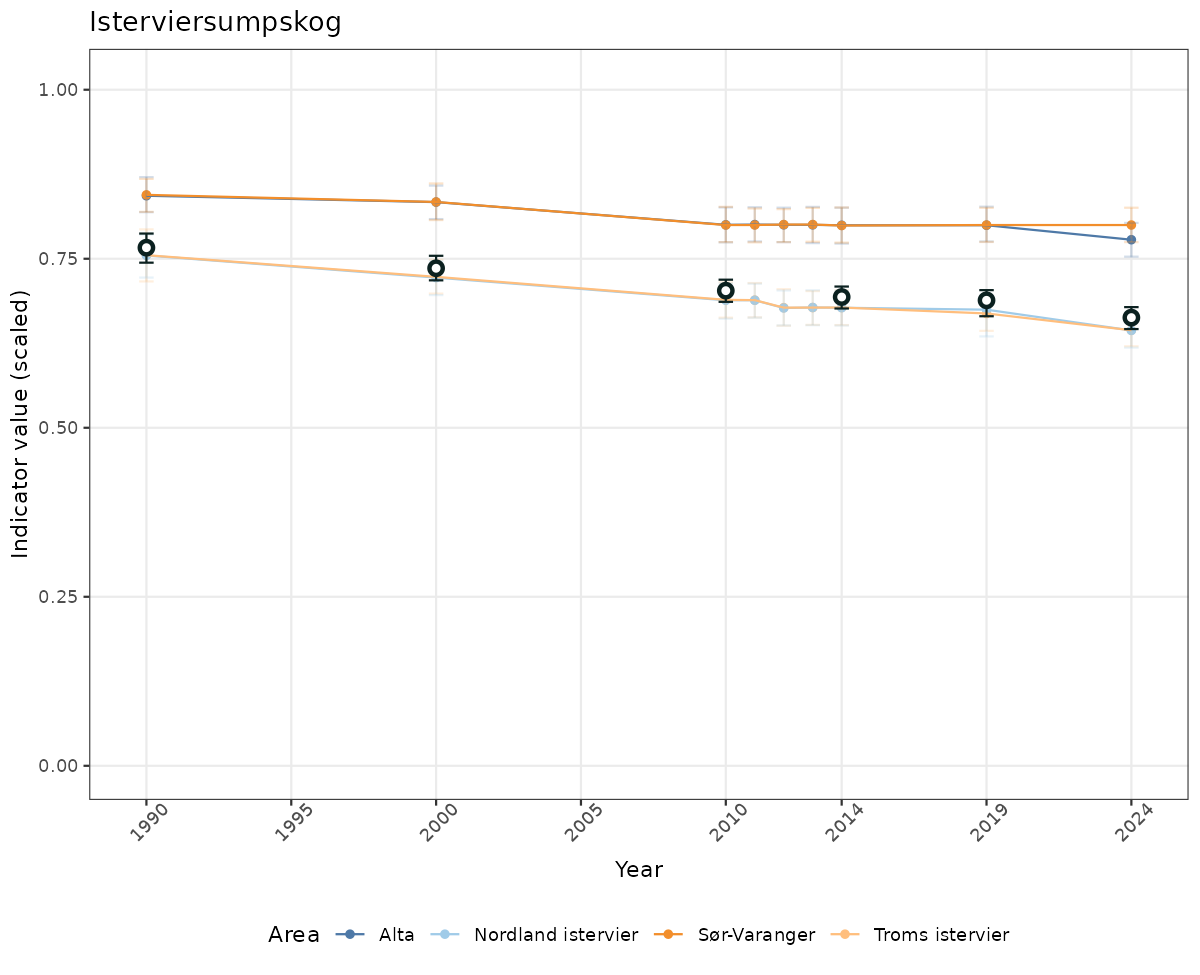
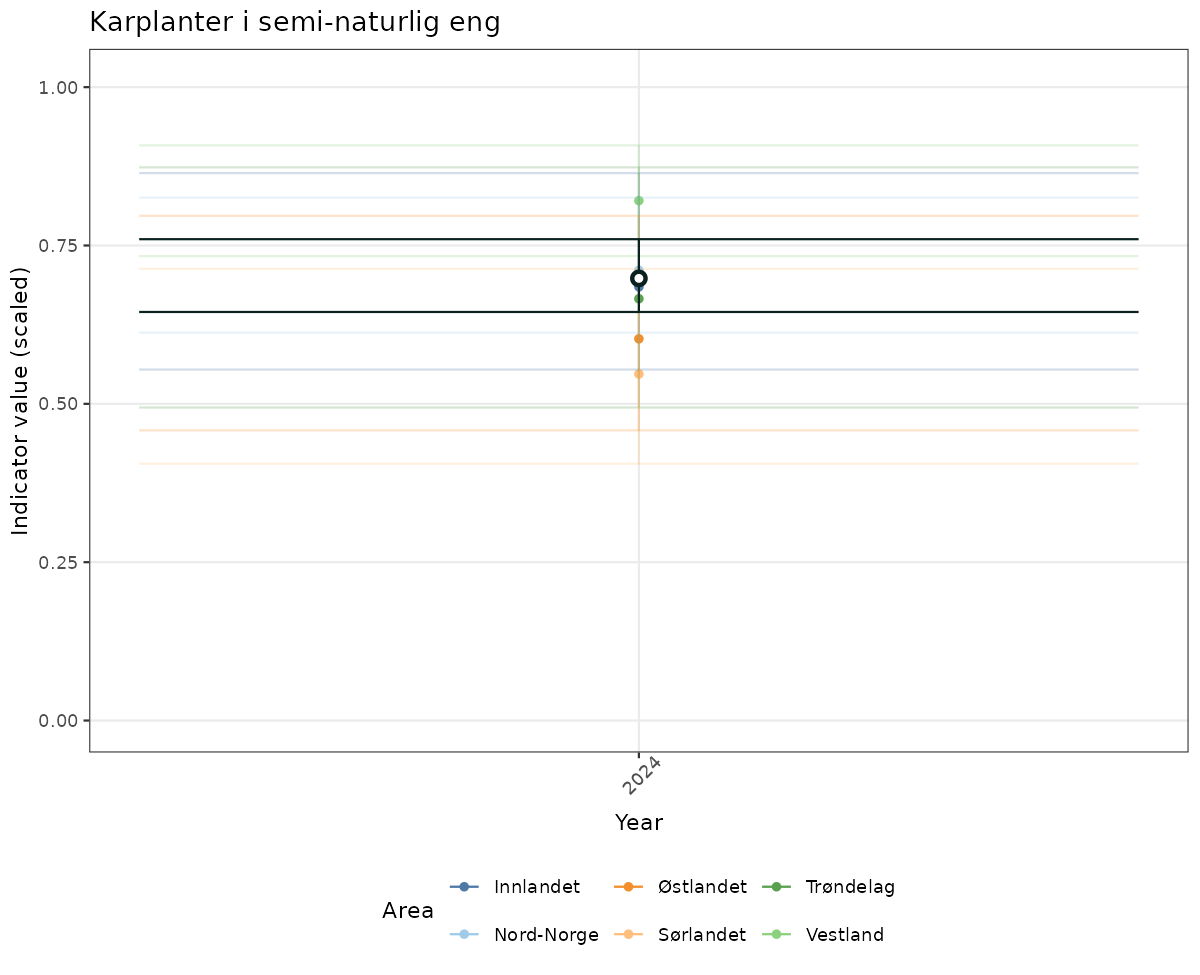
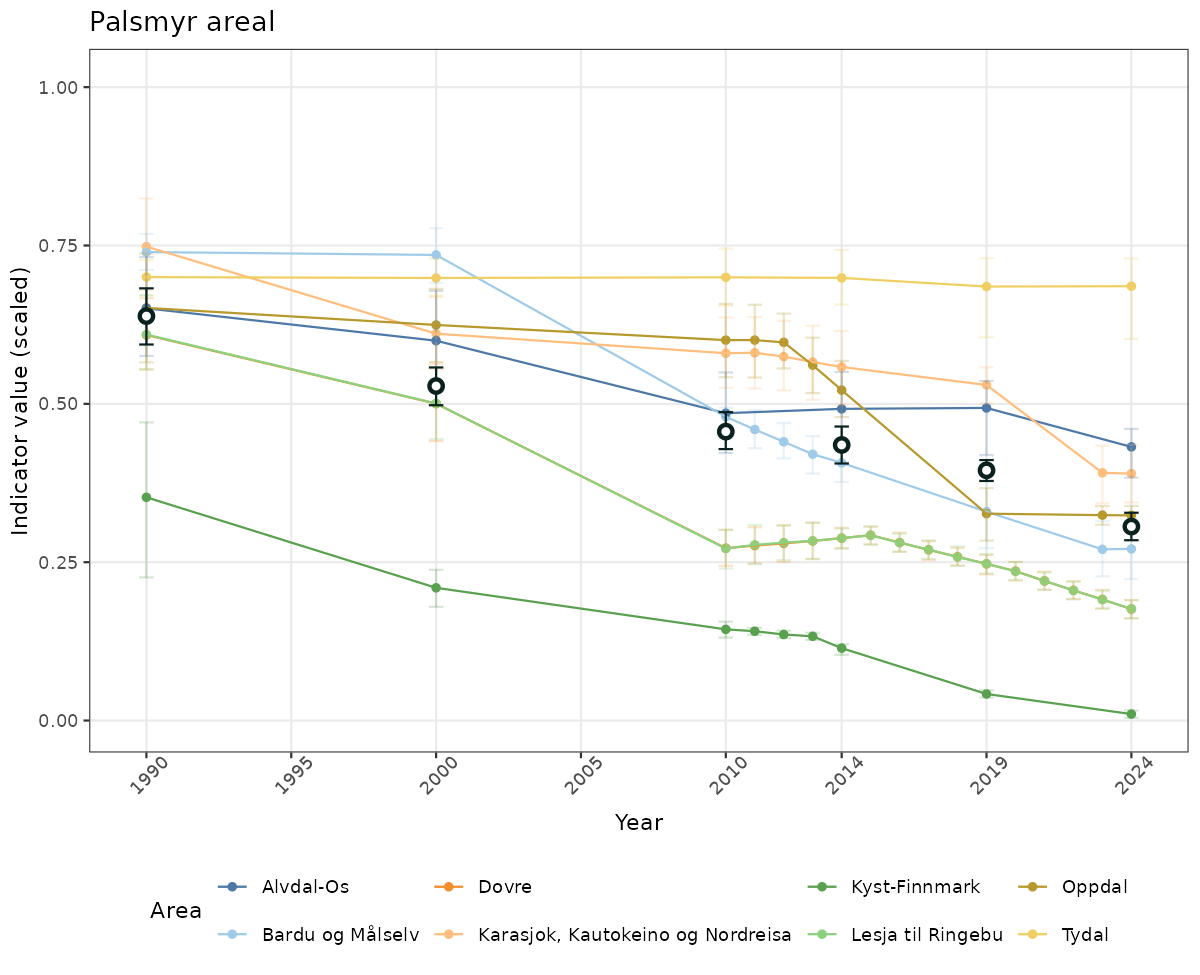
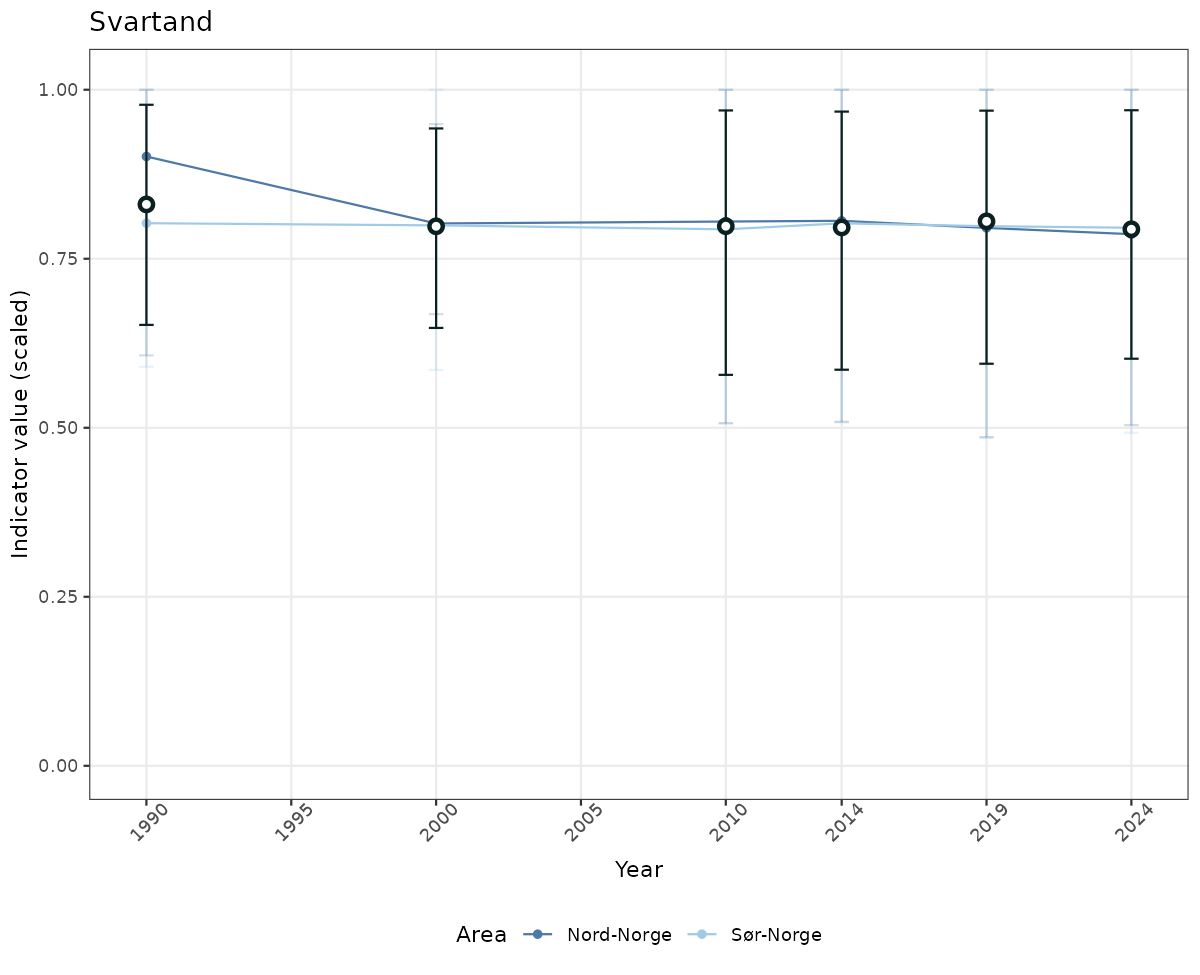
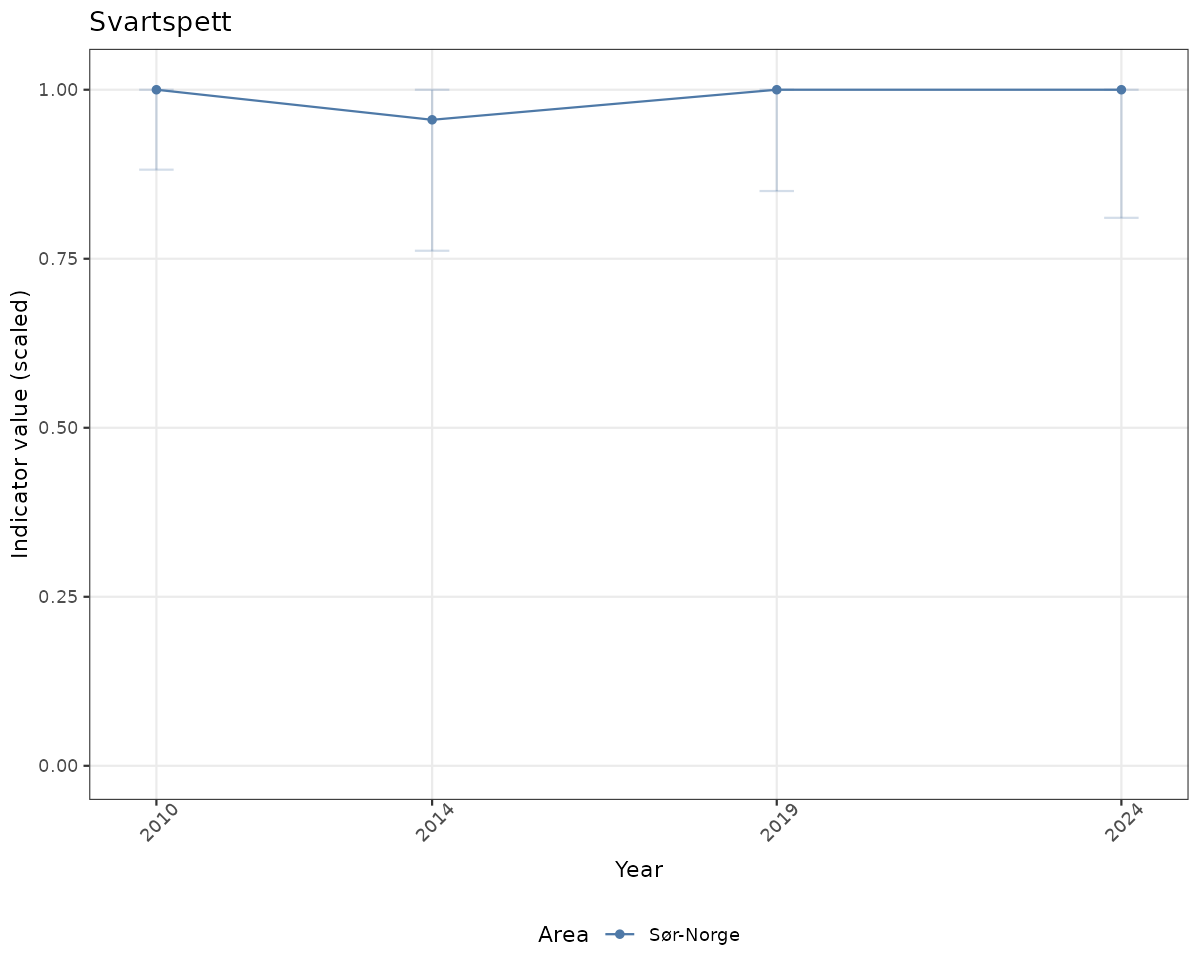
if(!manyAreas){
ts_scaled <- ggplot(data_scaled, aes(x = year_t)) +
geom_point(aes(y = median, color = ICunitName)) +
geom_errorbar(aes(ymin = q025, ymax = q975, color = ICunitName), width = 0.5, alpha = 0.25) +
geom_line(aes(y = median, color = ICunitName)) +
geom_errorbar(data = data_indIdx, aes(x = year_t, ymin = q025, ymax = q975), width = 0.5, color = plot.col2) +
geom_point(data = data_indIdx, aes(x = year_t, y = median), shape = 21, size = 2, stroke = 1.5, fill = "white", color = plot.col2) +
ylim(0, 1.01) +
xlab("Year") + ylab("Indicator value (scaled)") +
ggtitle(ind_name) +
paletteer::scale_color_paletteer_d(palette = "ggthemes::Tableau_20", name = "Area") +
scale_x_continuous(breaks = plot.yearH) +
theme_bw() +
theme(legend.position = "bottom",
panel.grid.minor = element_blank(),
axis.text.x = element_text(angle = 45, vjust = 0.8))
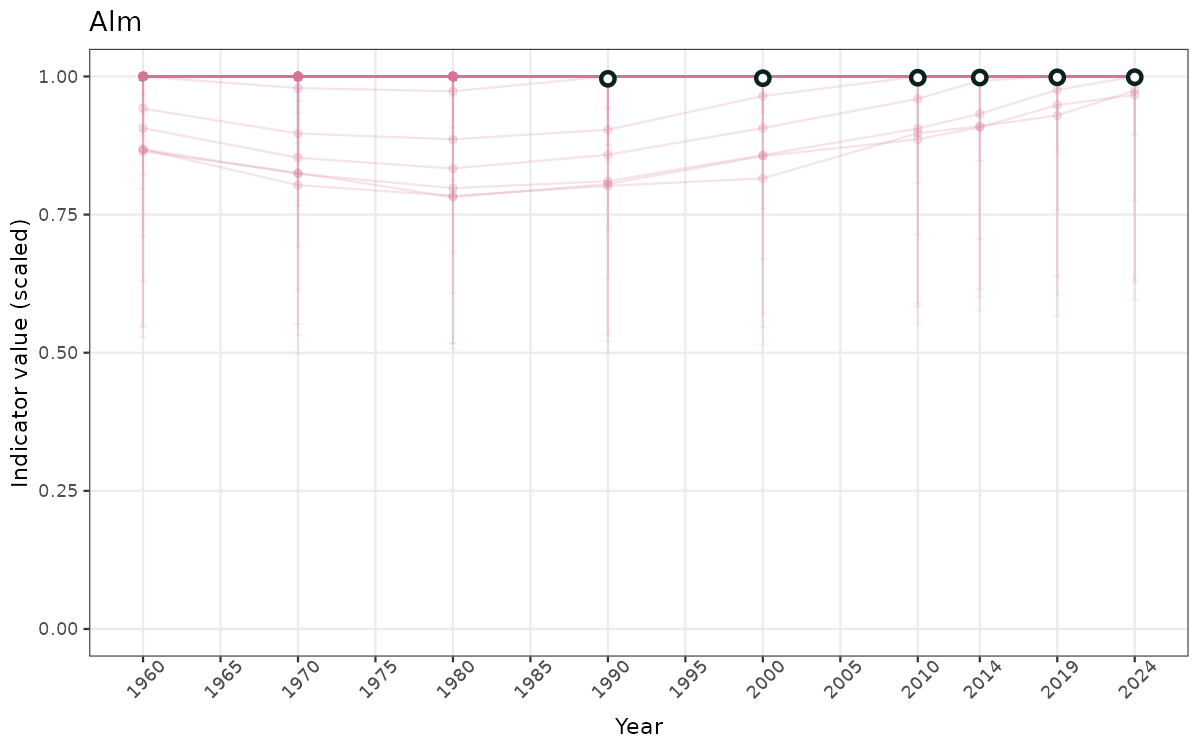
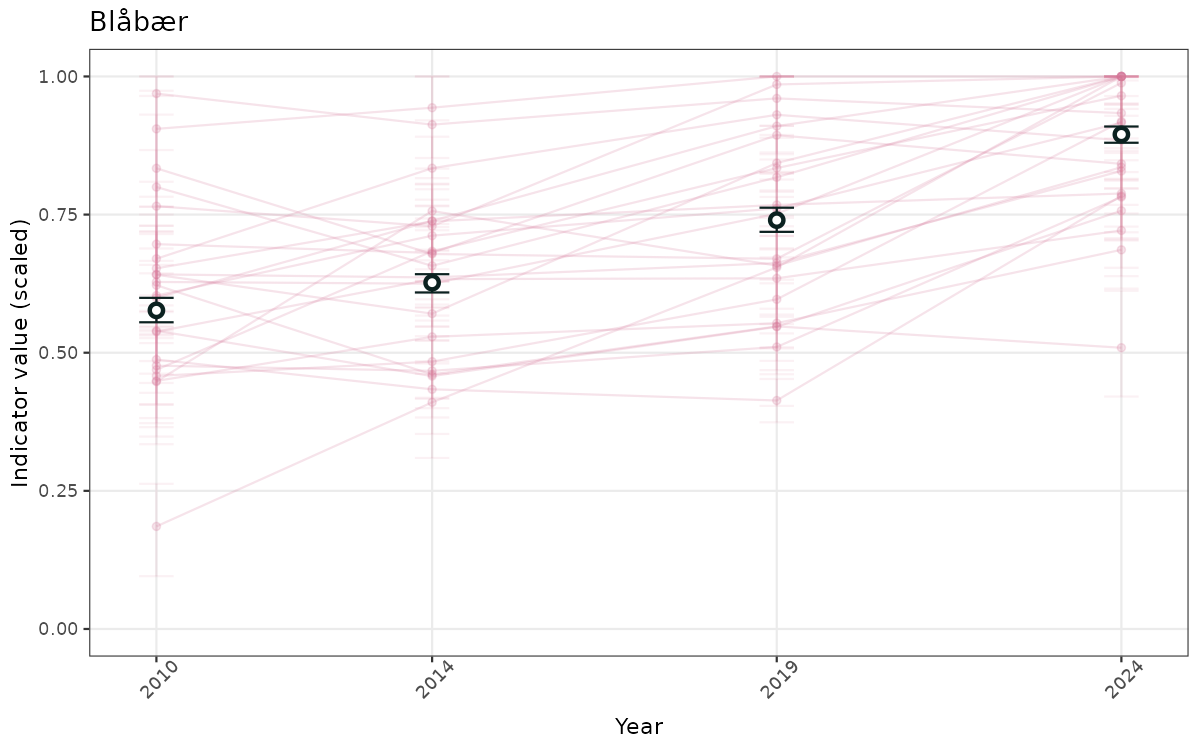
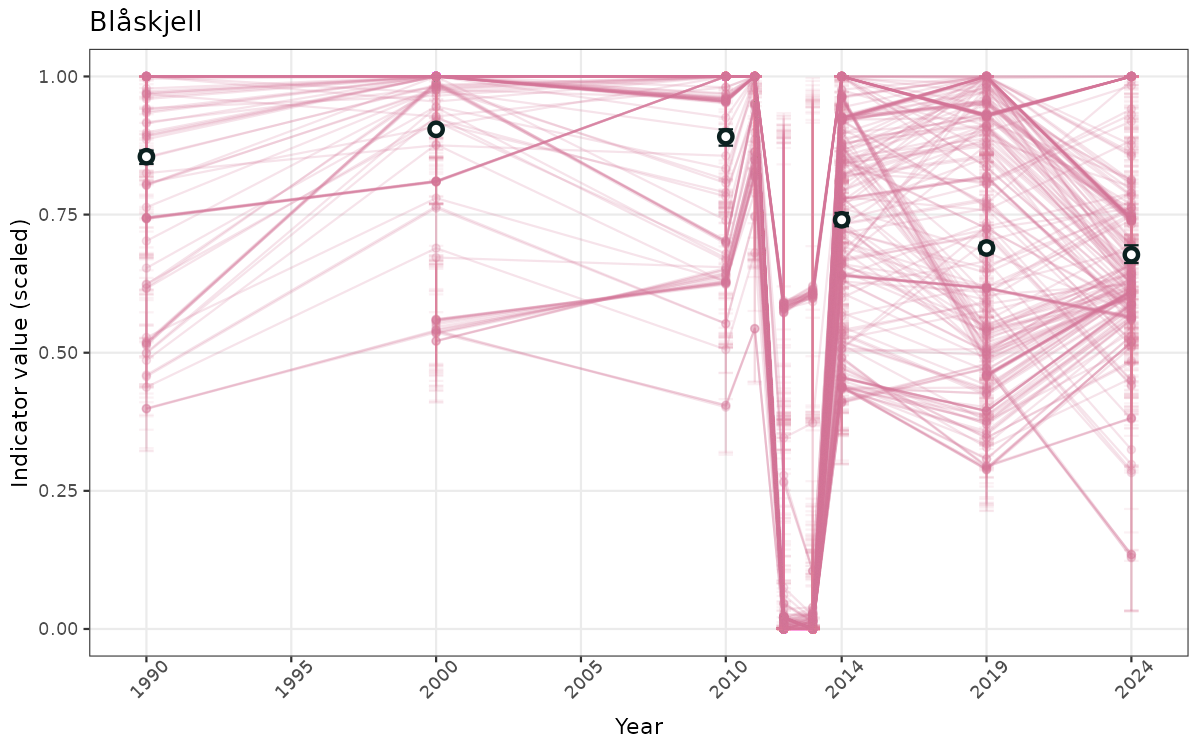
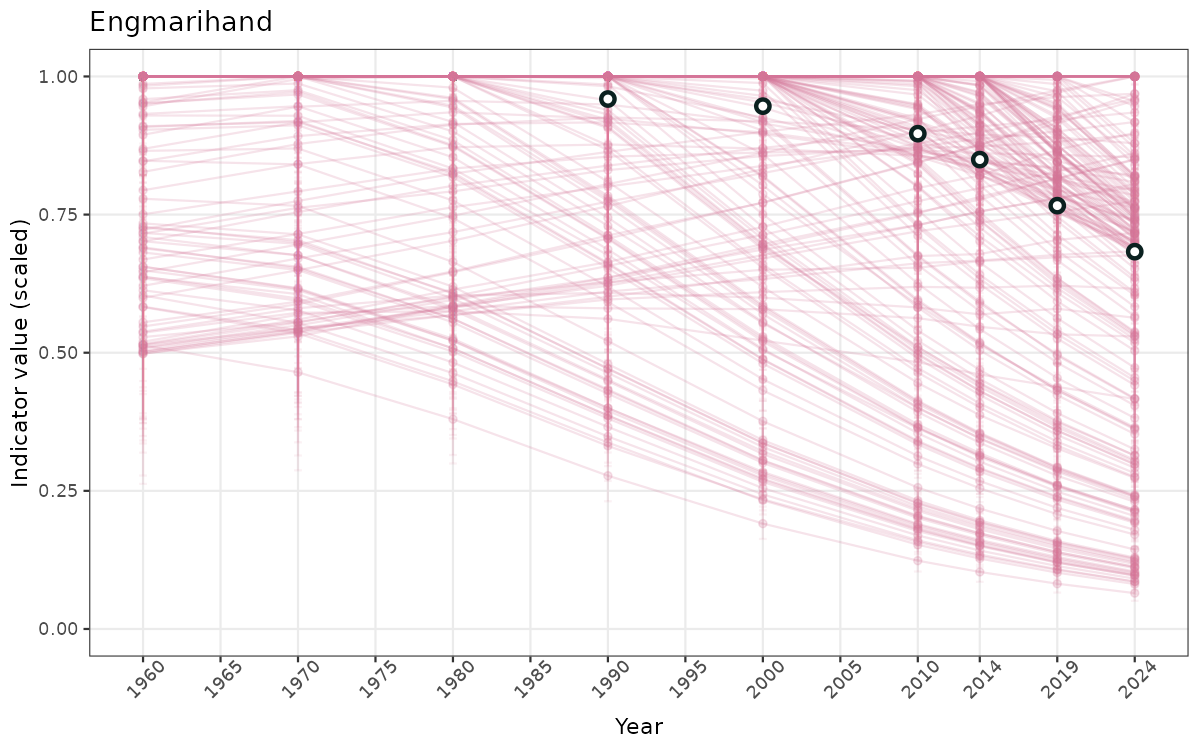
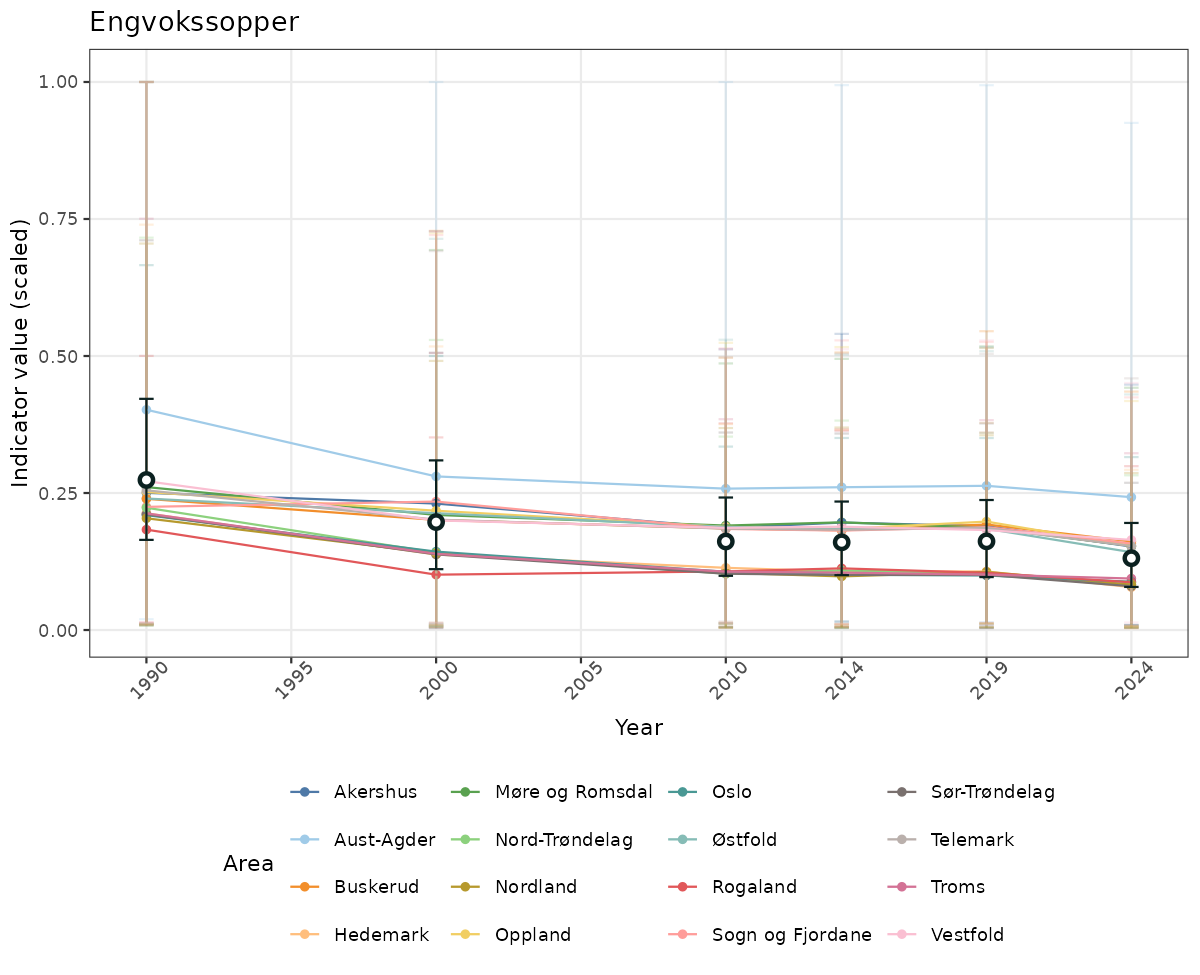
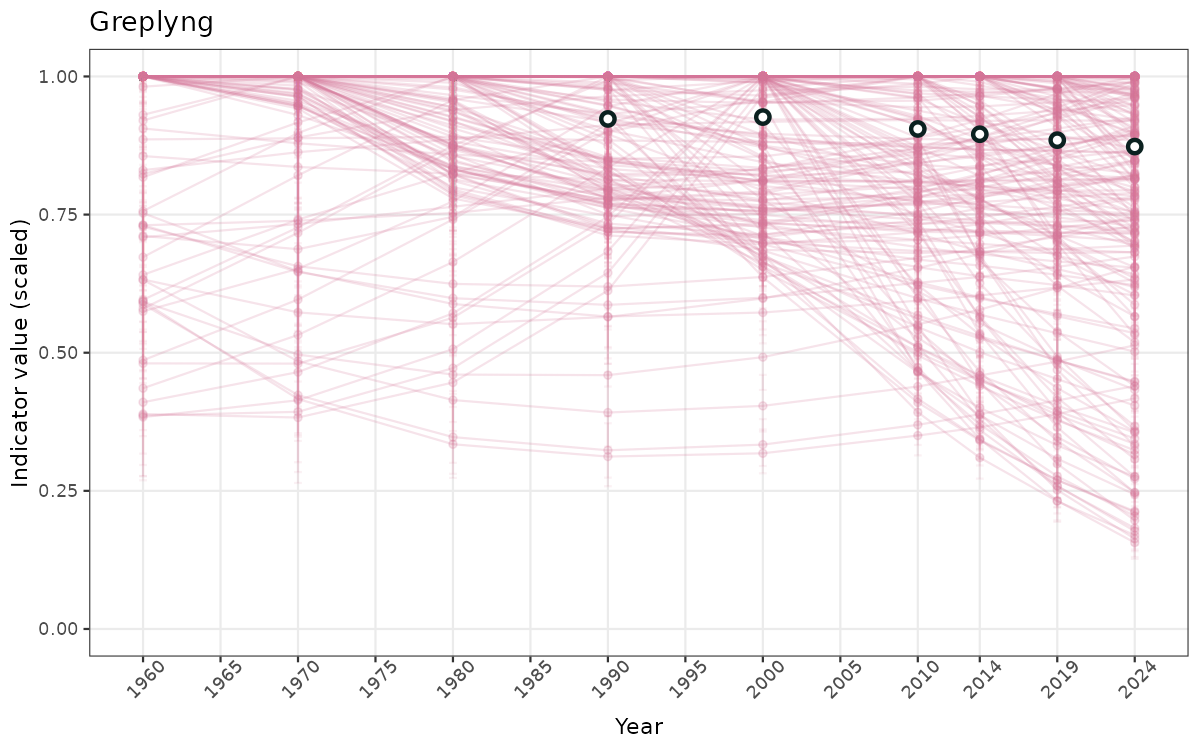
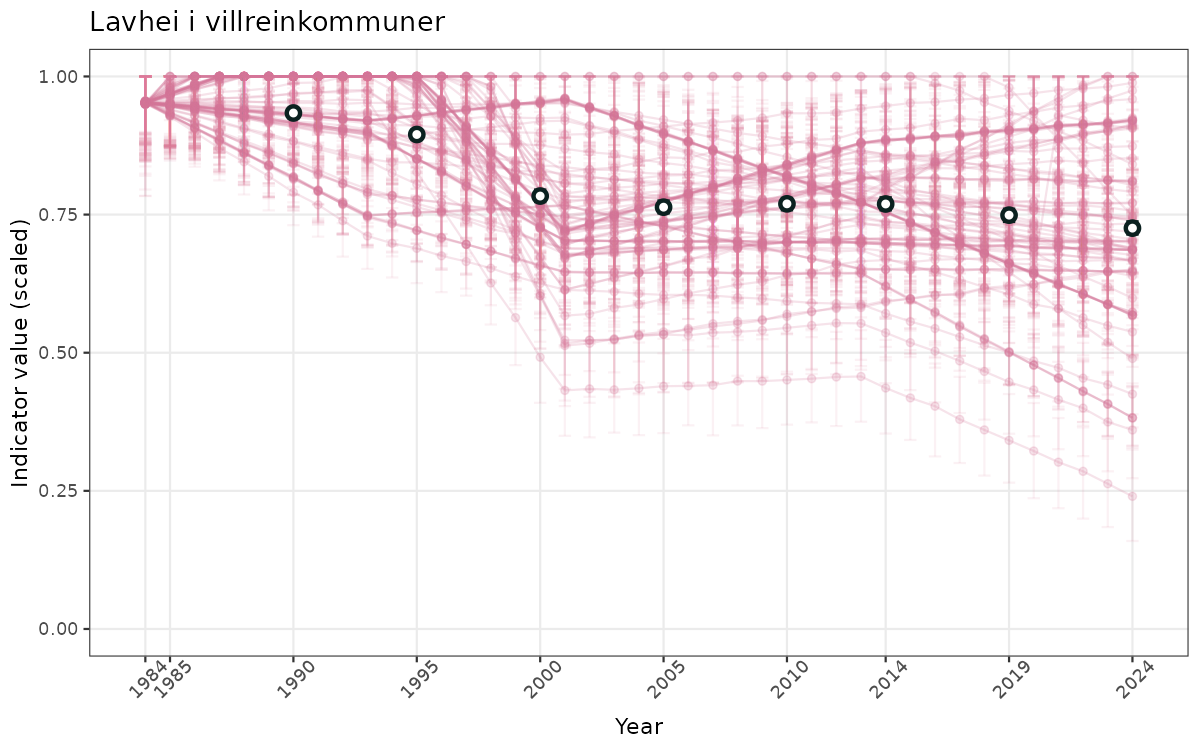
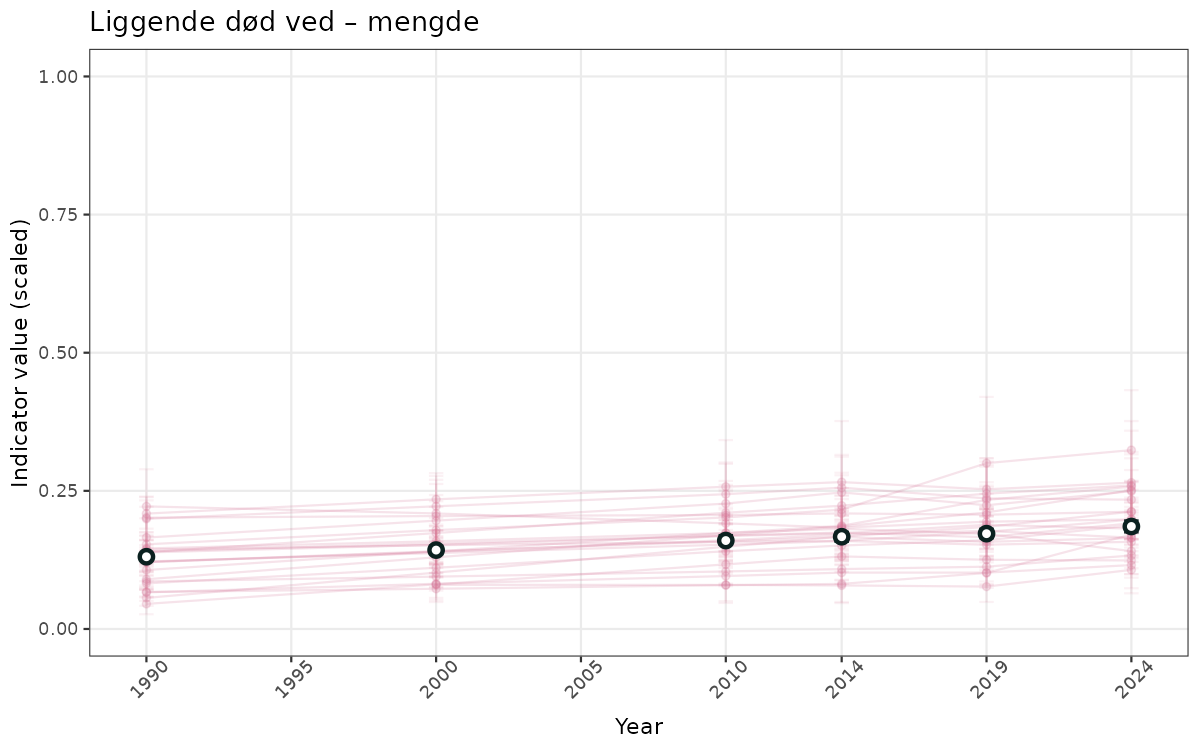
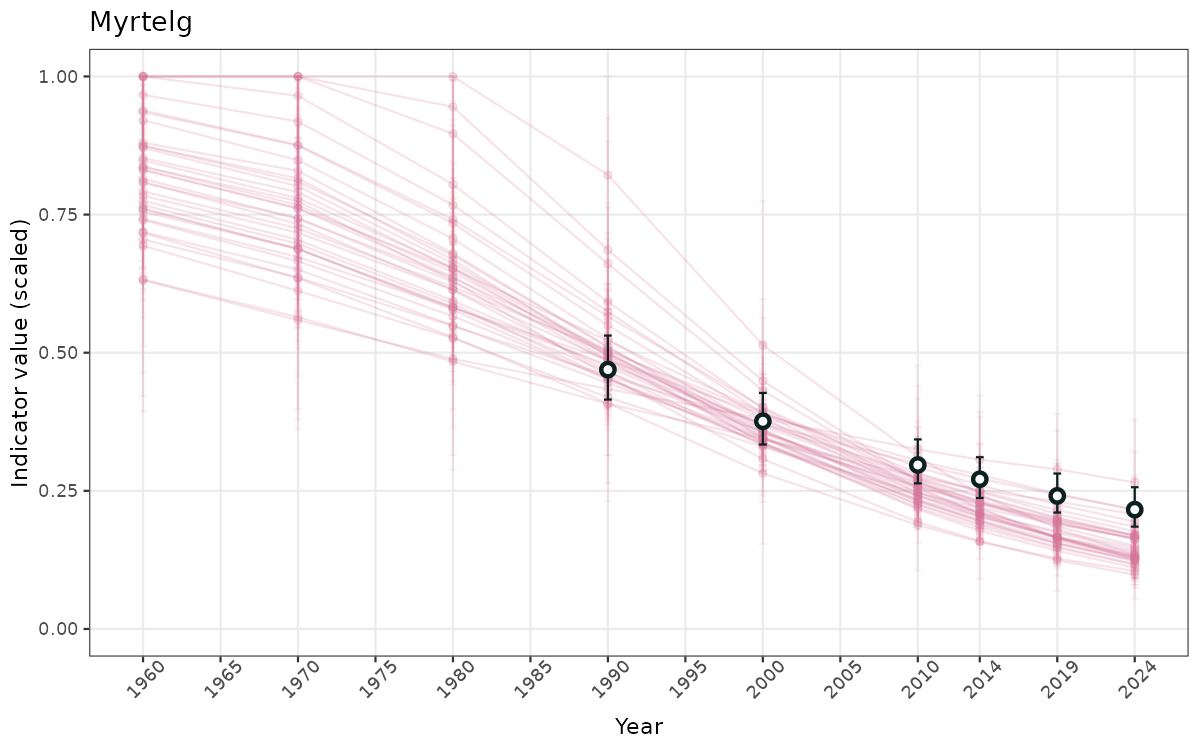
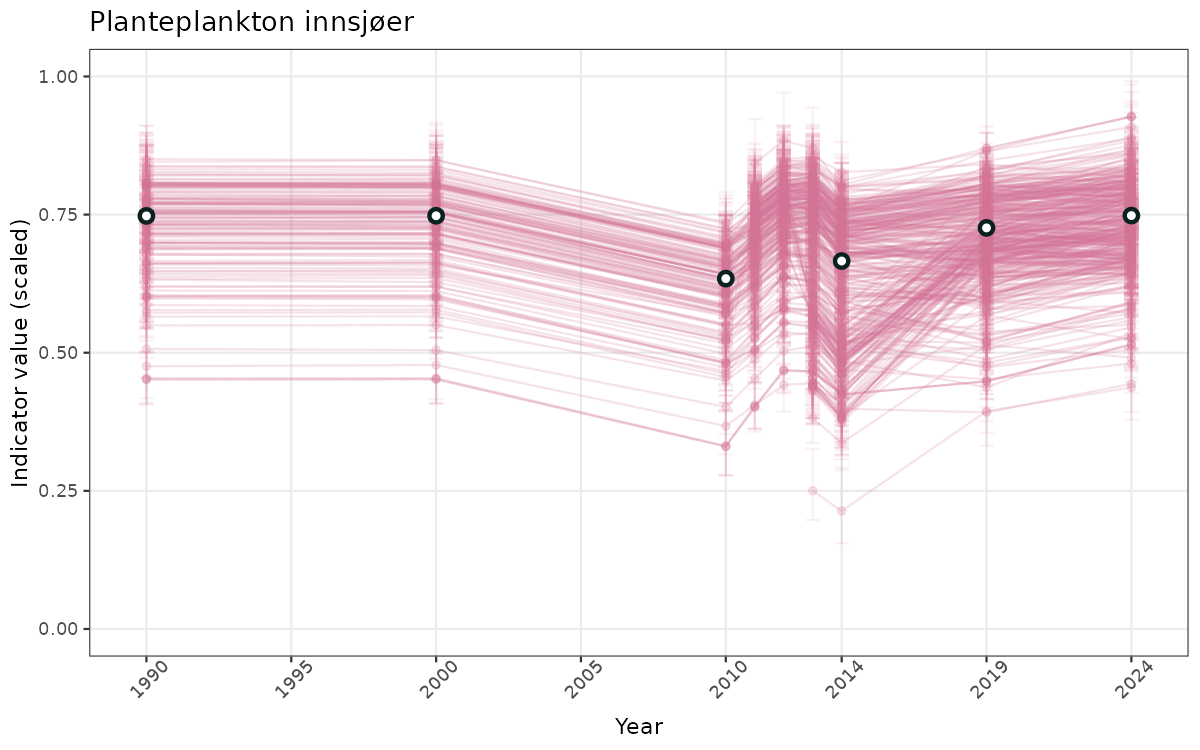
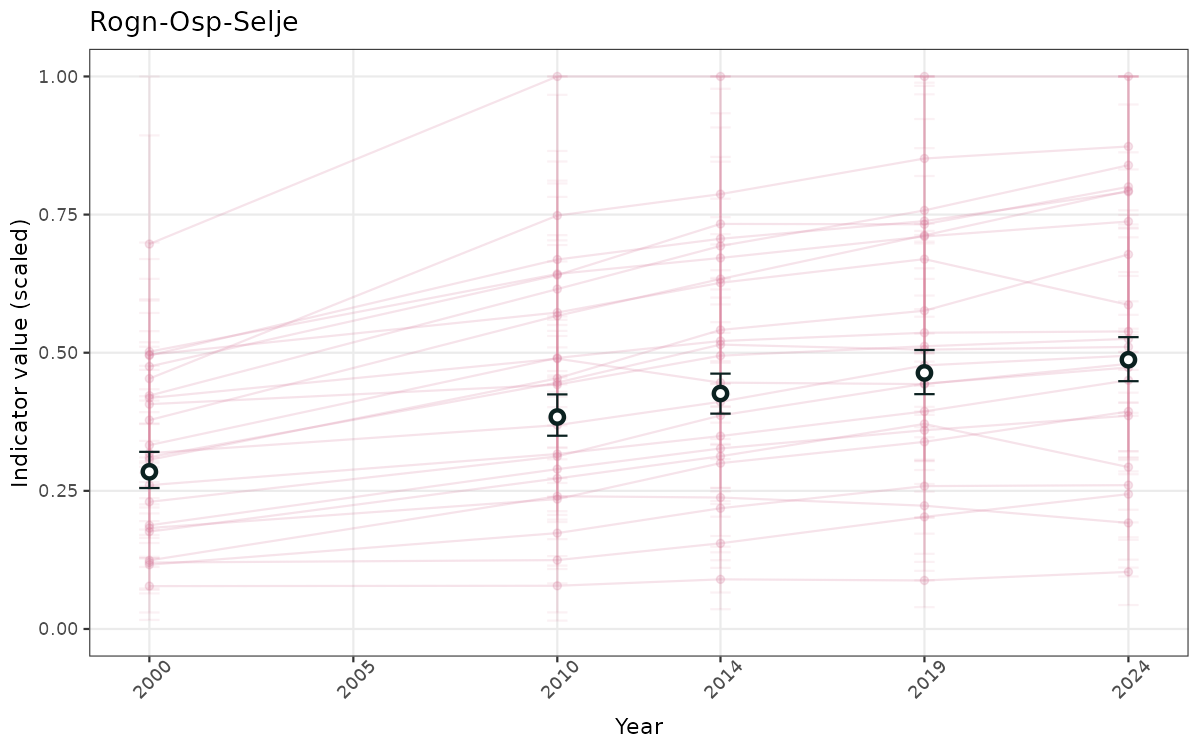
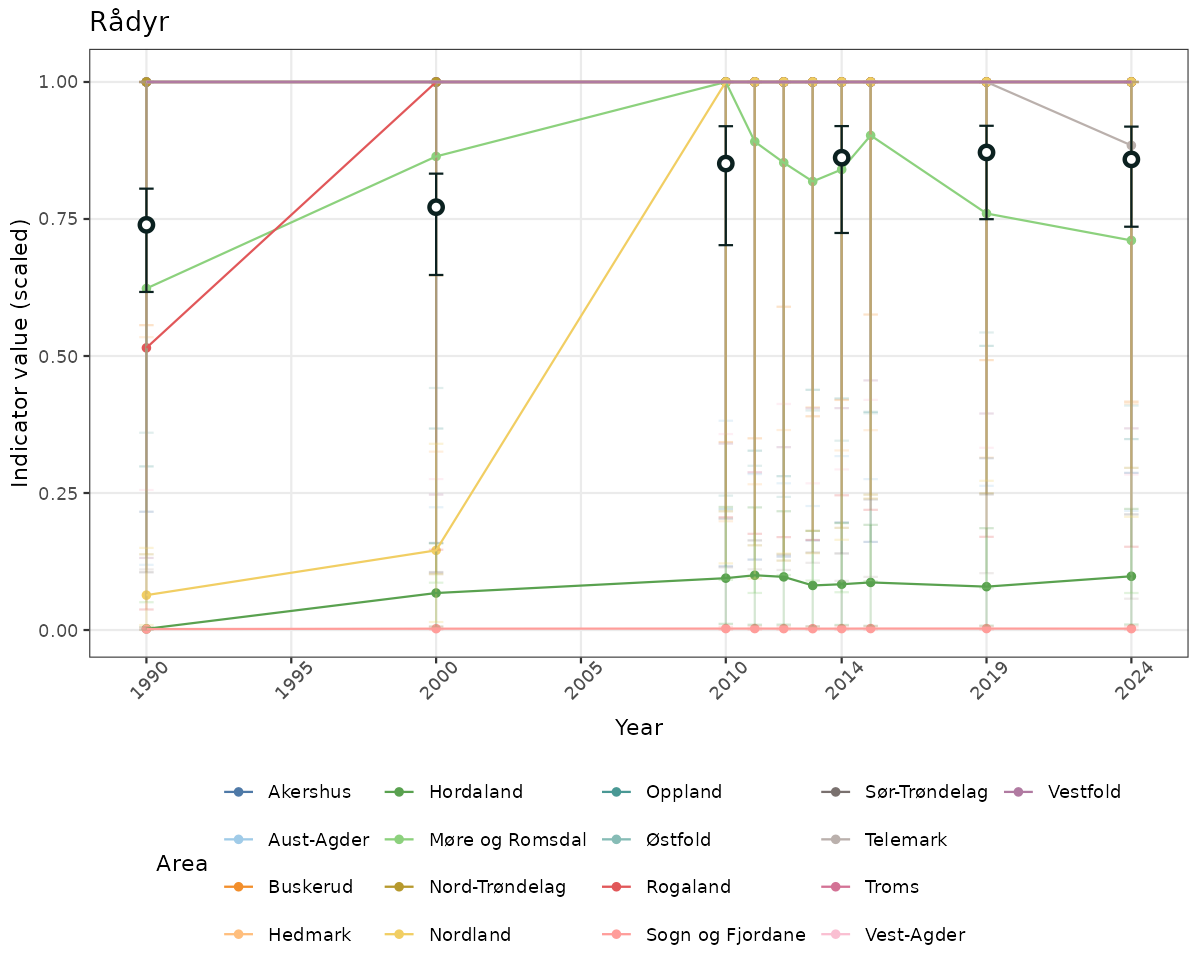
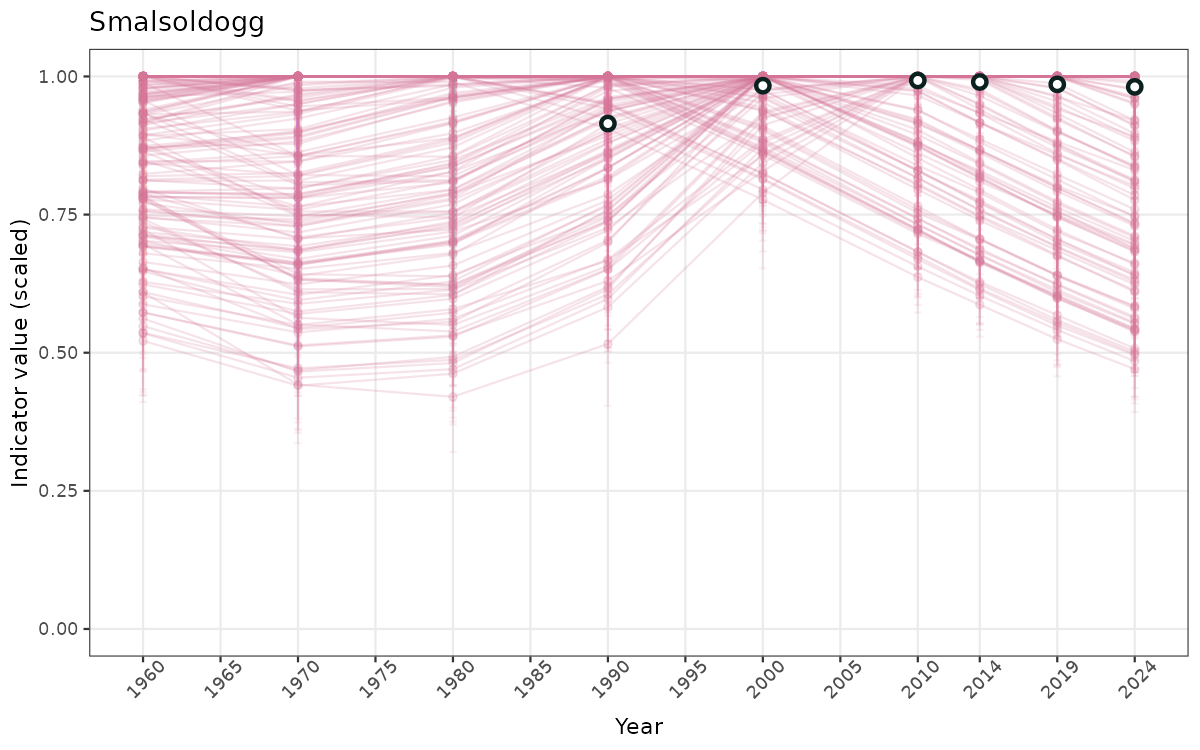
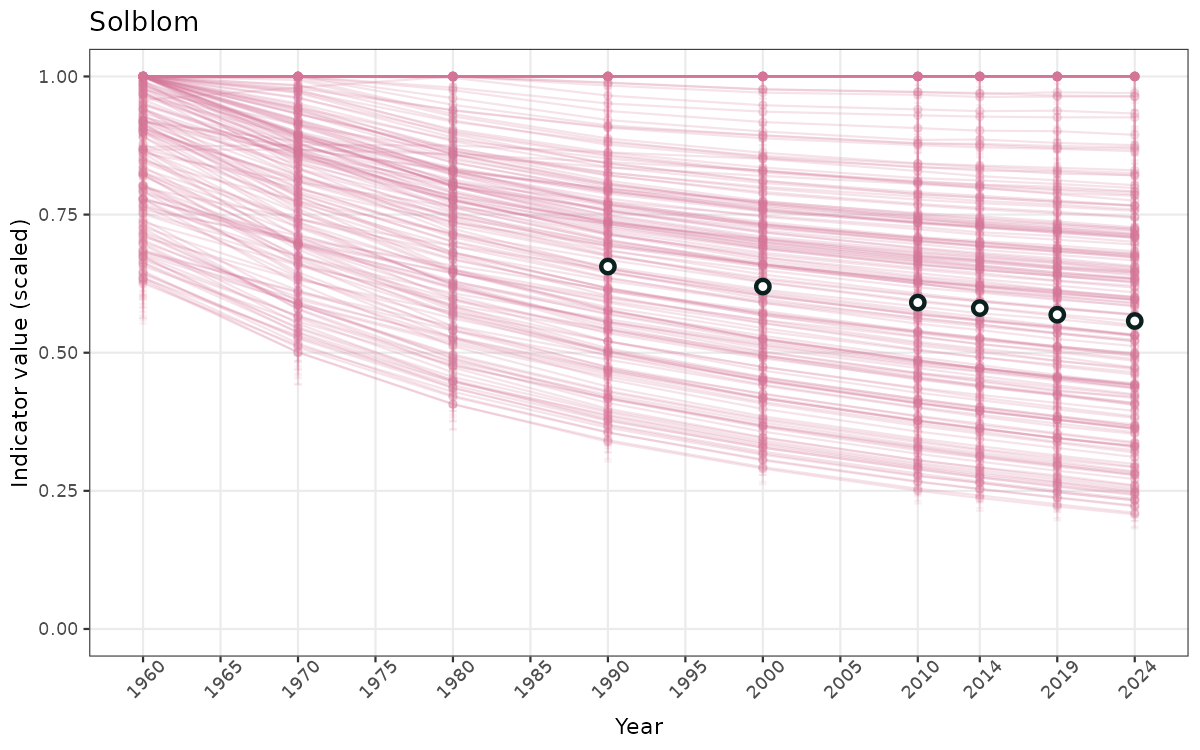
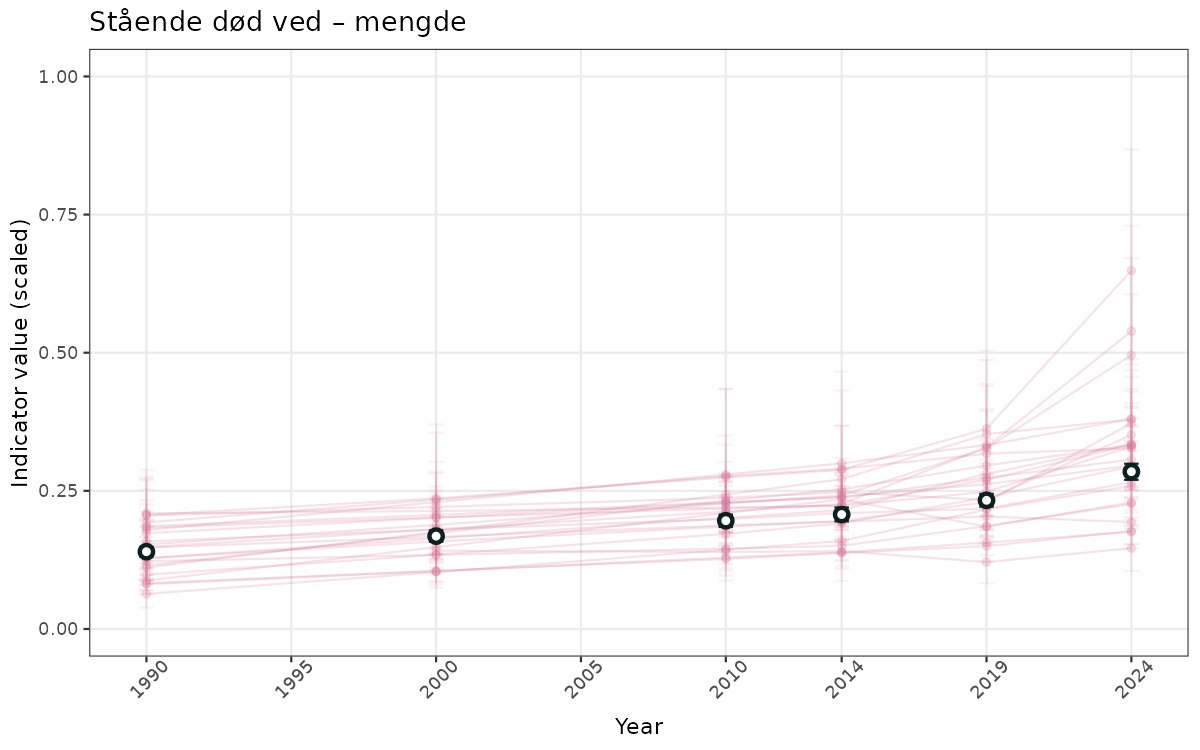
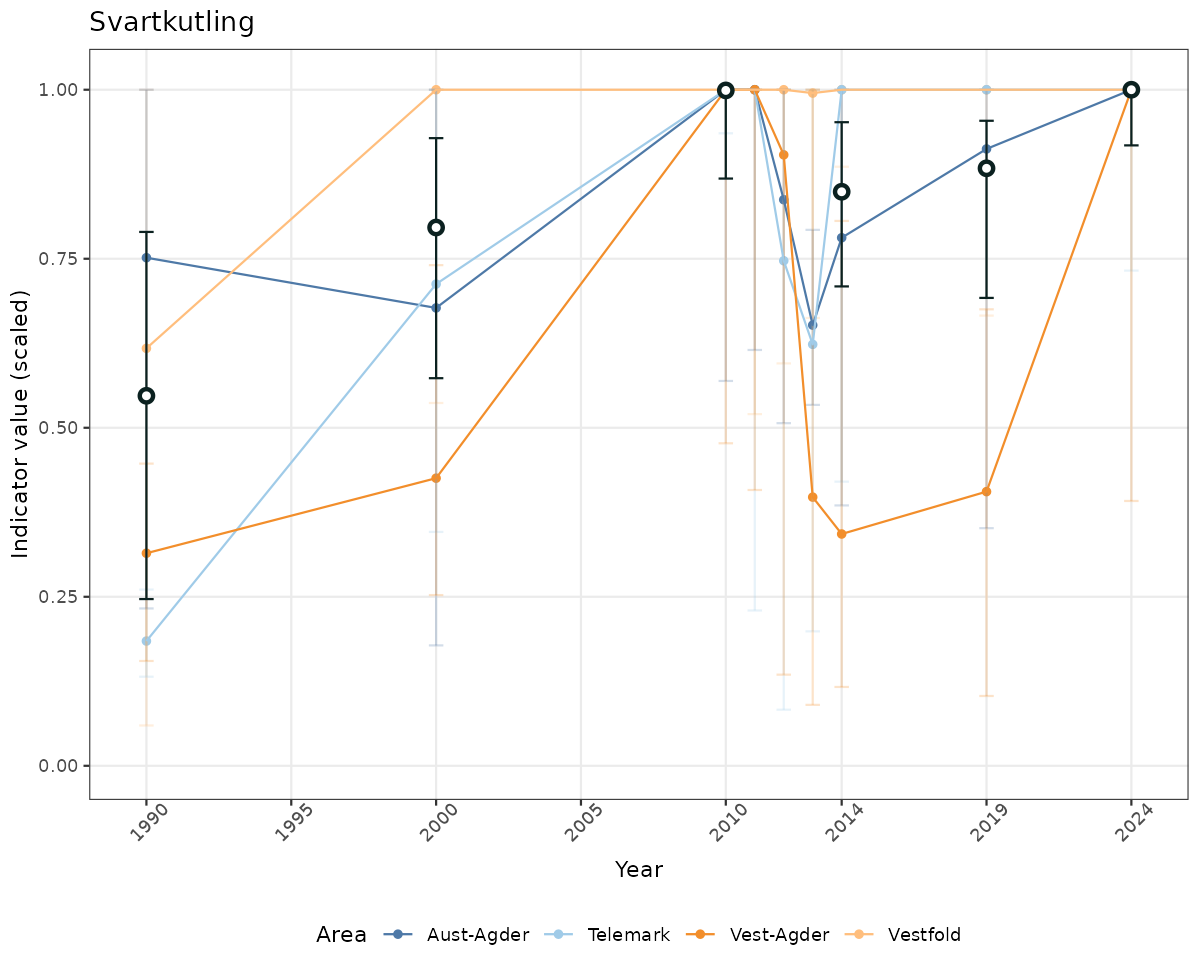
}else{
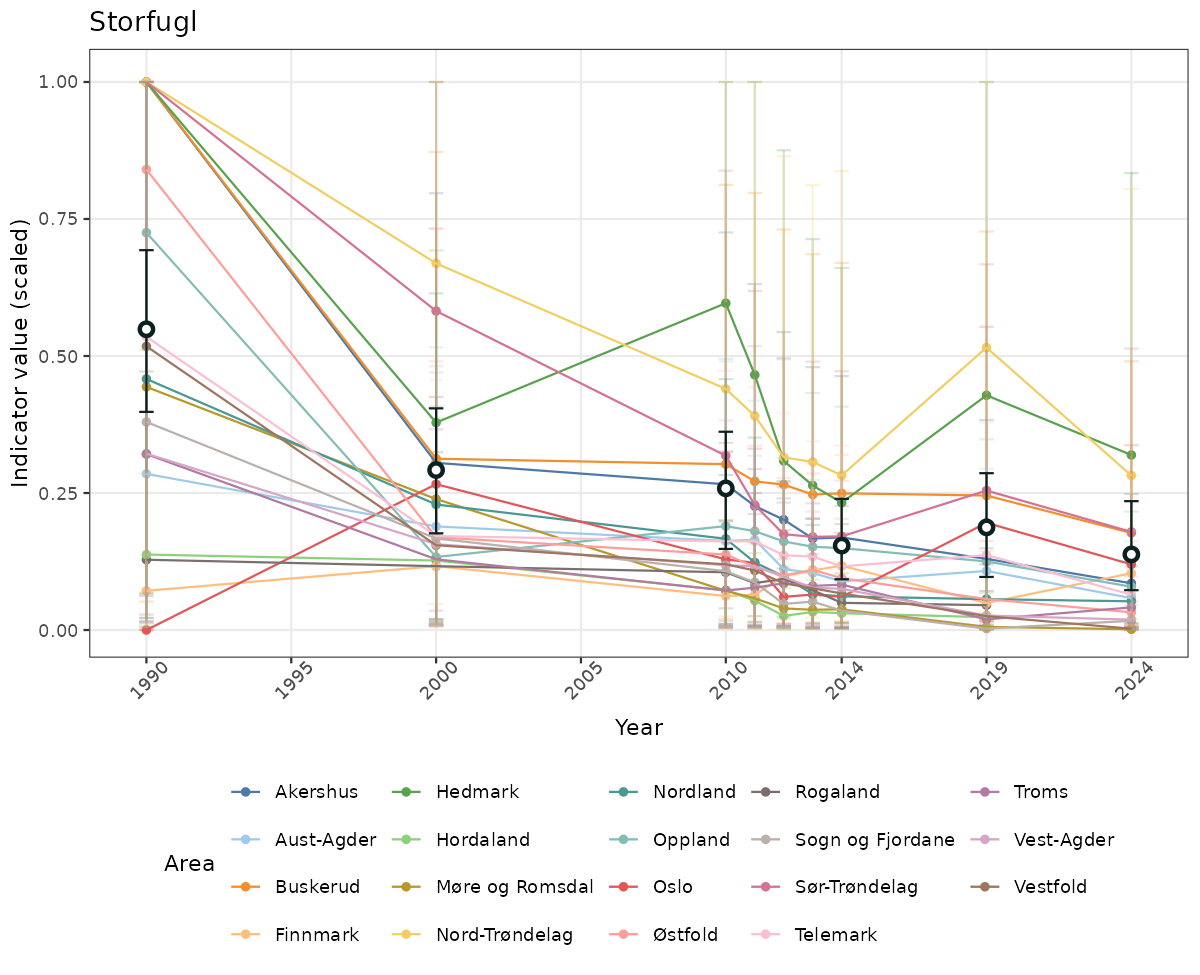
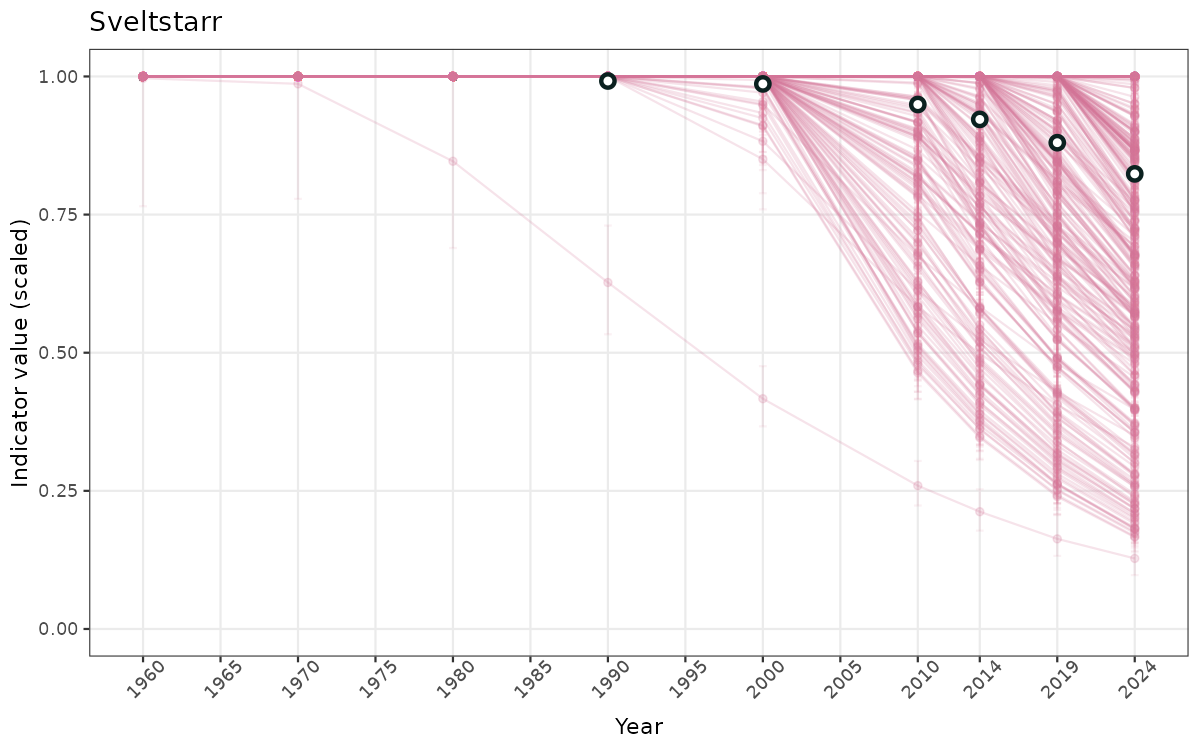
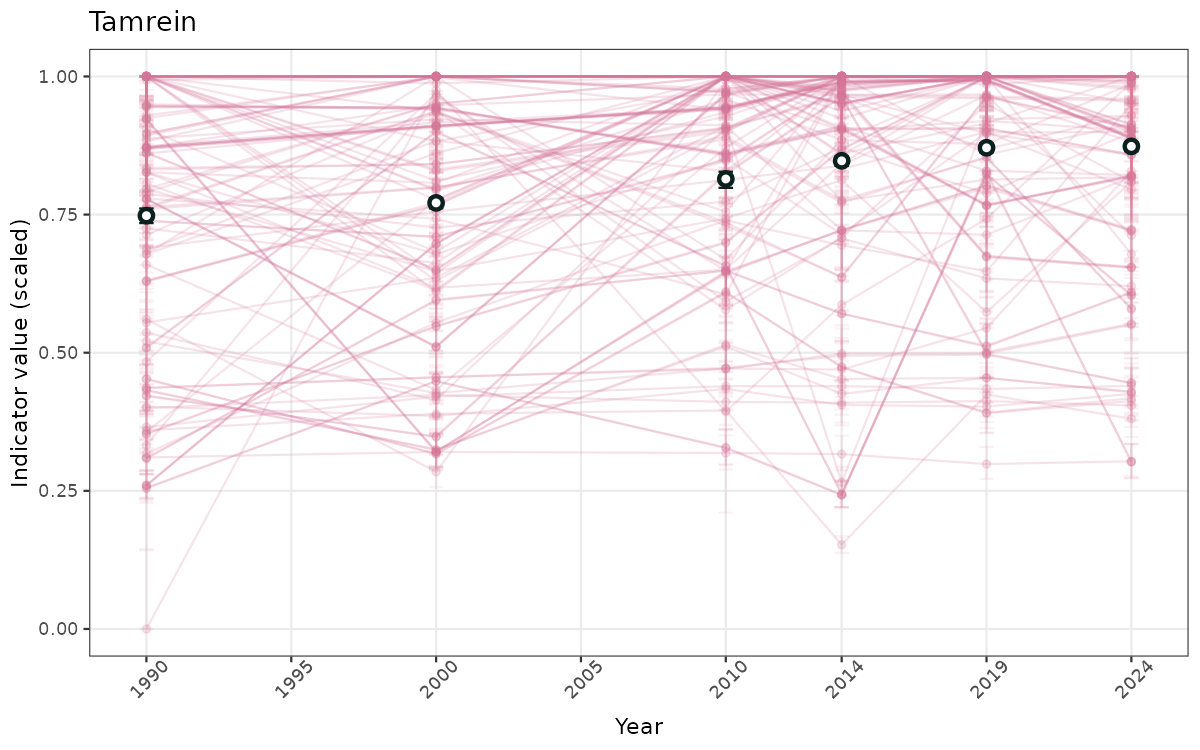
ts_scaled <- ggplot(data_scaled, aes(x = year_t)) +
geom_point(aes(y = median, group = ICunitName), color = plot.col, alpha = 0.2) +
geom_errorbar(aes(ymin = q025, ymax = q975, group = ICunitName), width = 0.5, color = plot.col, alpha = 0.1) +
geom_line(aes(y = median, group = ICunitName), color = plot.col, alpha = 0.2) +
geom_errorbar(data = data_indIdx, aes(x = year_t, ymin = q025, ymax = q975), width = 0.5, color = plot.col2) +
geom_point(data = data_indIdx, aes(x = year_t, y = median), shape = 21, size = 2, stroke = 1.5, fill = "white", color = plot.col2) +
ylim(0, 1) +
xlab("Year") + ylab("Indicator value (scaled)") +
ggtitle(ind_name) +
scale_x_continuous(breaks = plot.yearH) +
theme_bw() +
theme(legend.position = "bottom",
panel.grid.minor = element_blank(),
axis.text.x = element_text(angle = 45, vjust = 0.8))
}
TS_scaled[[i]] <- ts_scaled
ggsave(file = paste0(savePath_i, "/TimeSeries_scaled.svg"), plot = ts_scaled, width = 8, height = ifelse(manyAreas, 5, 6.5))
ggsave(file = paste0(savePath_i, "/TimeSeries_scaled.pdf"), plot = ts_scaled, width = 8, height = ifelse(manyAreas, 5, 6.5))
ggsave(file = paste0(savePath_i, "/TimeSeries_scaled.png"), plot = ts_scaled, width = 8, height = ifelse(manyAreas, 5, 6.5), dpi = 150)
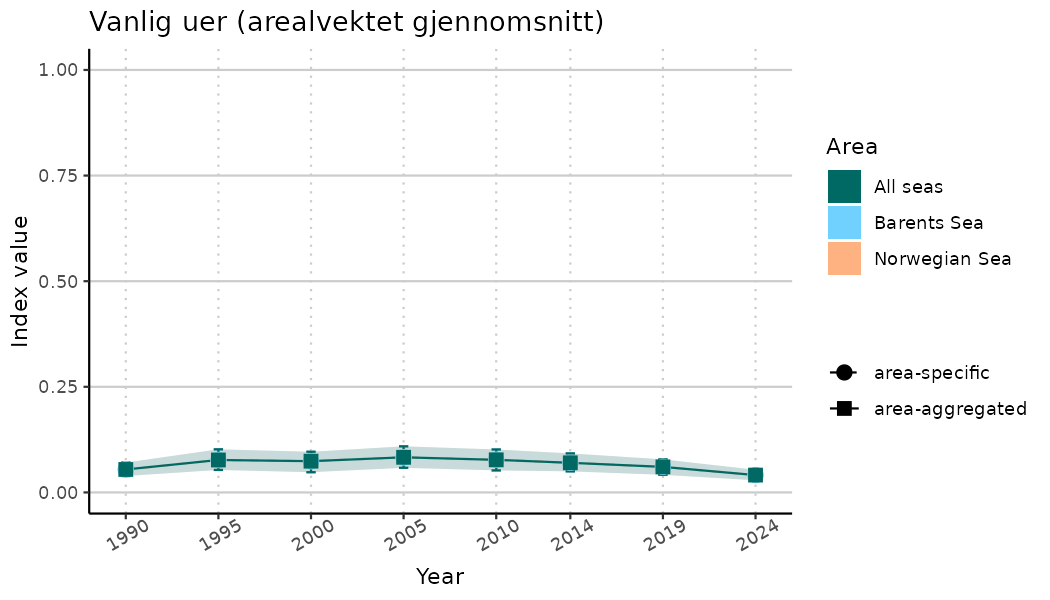
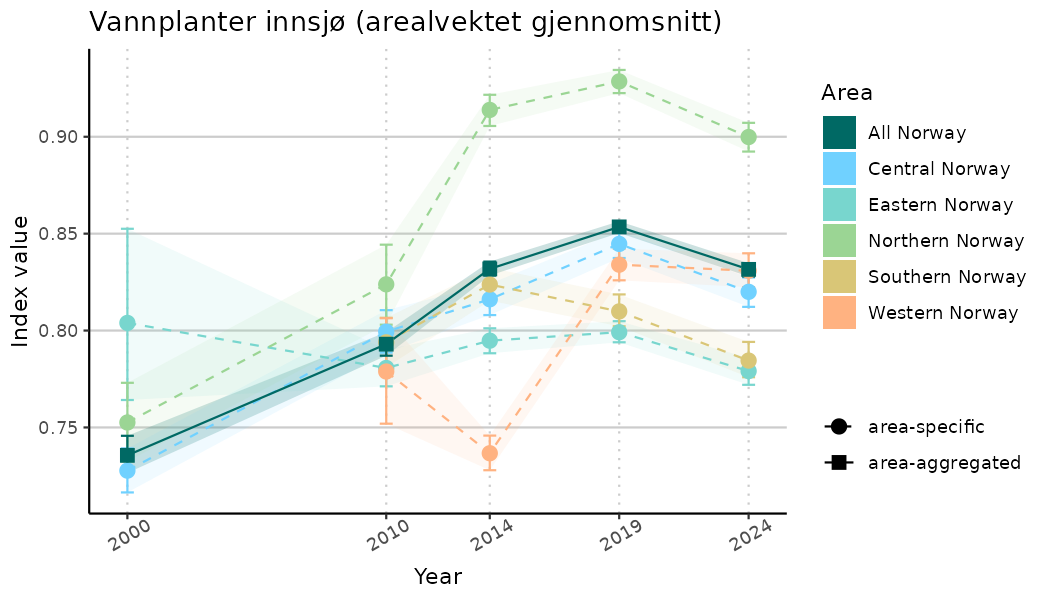
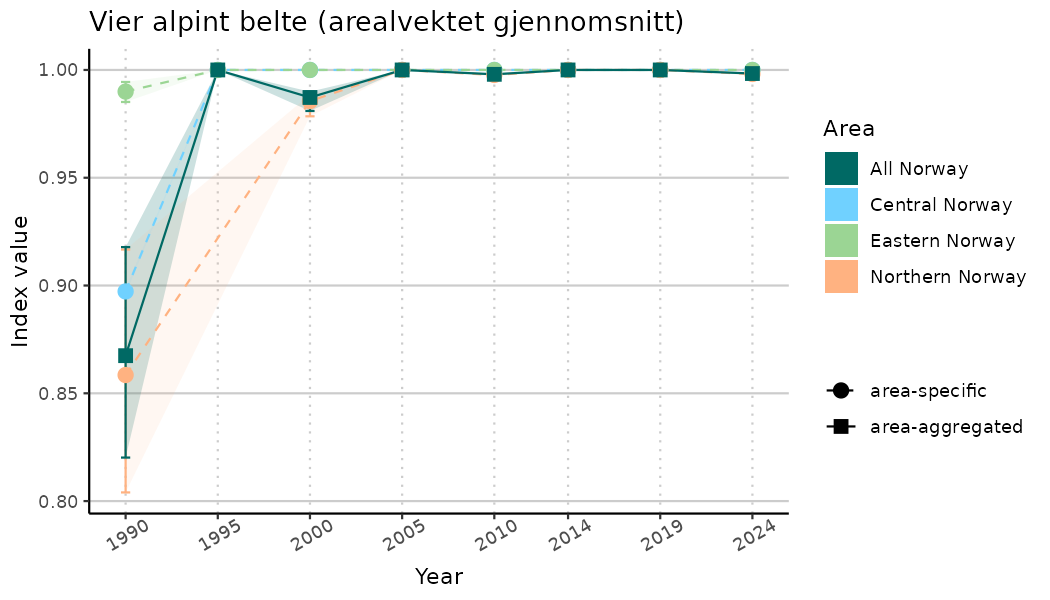
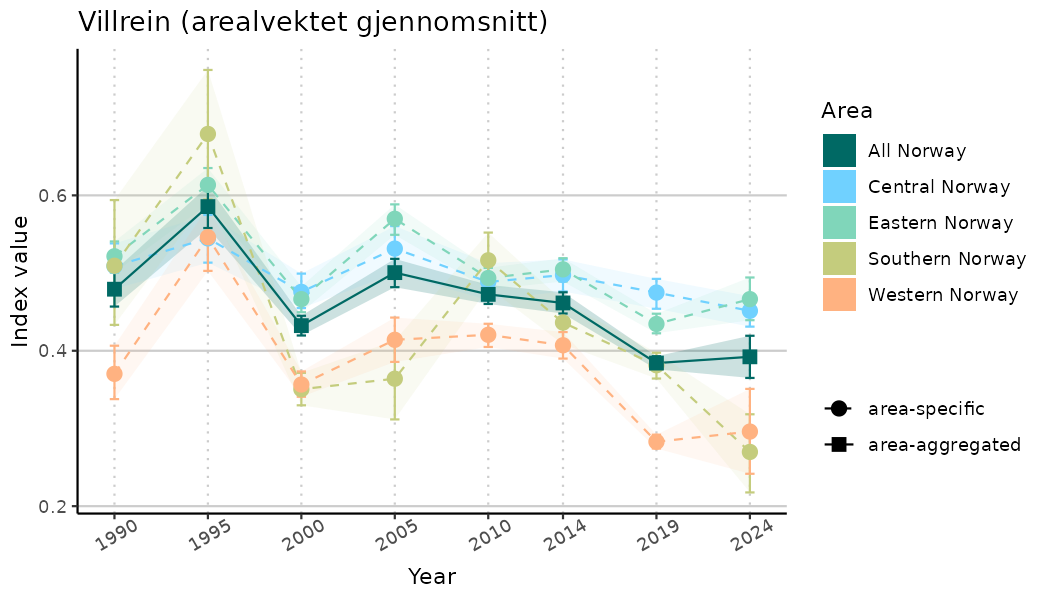
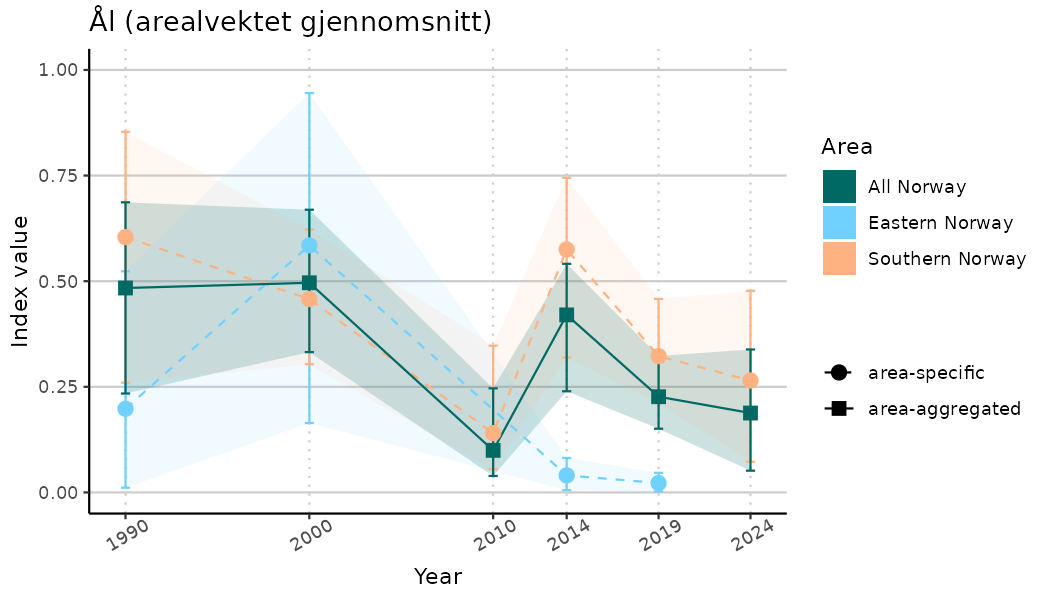
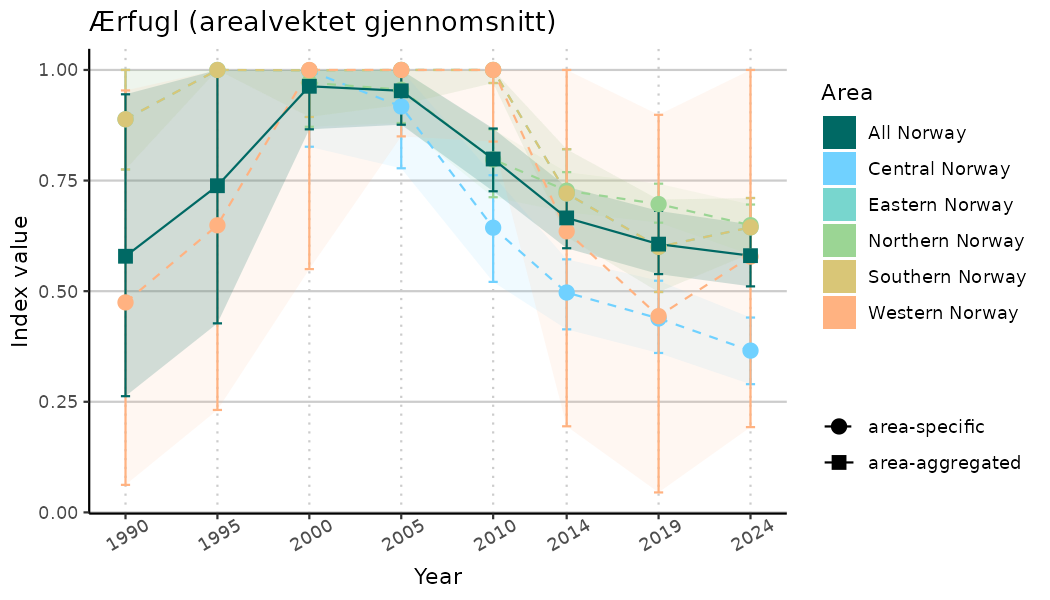
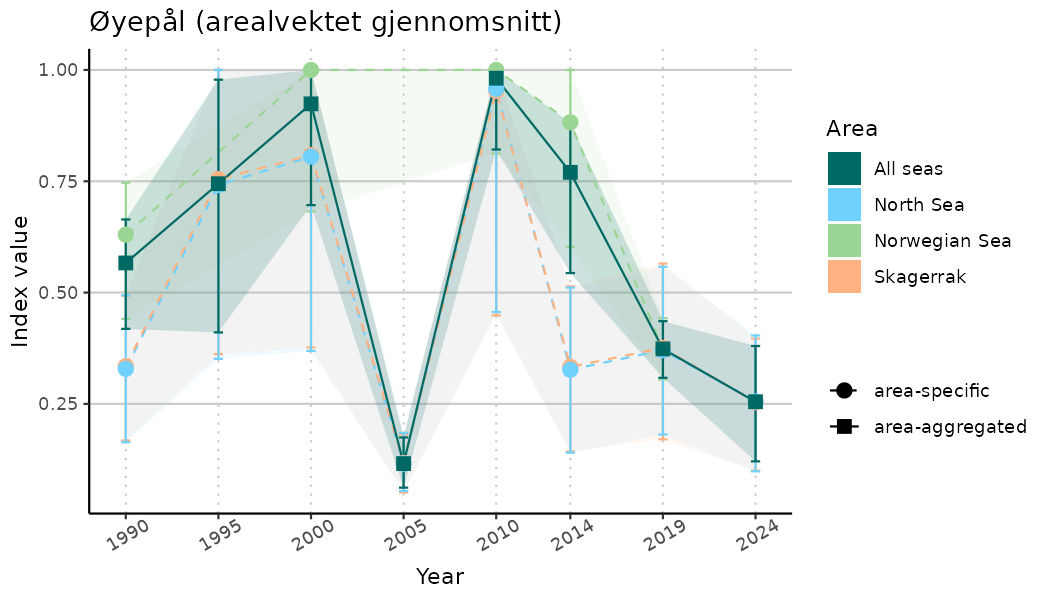
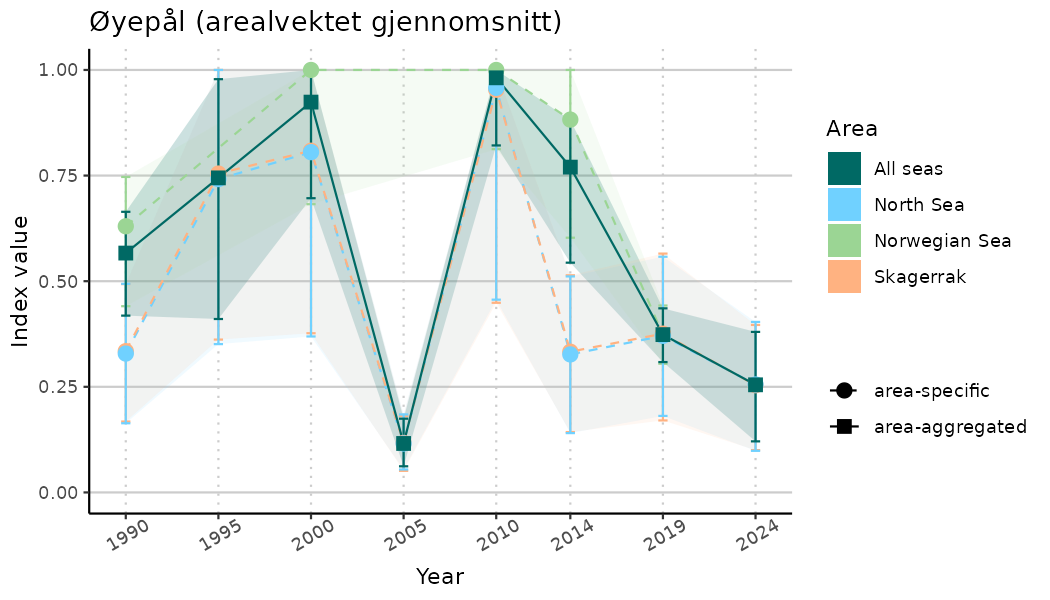
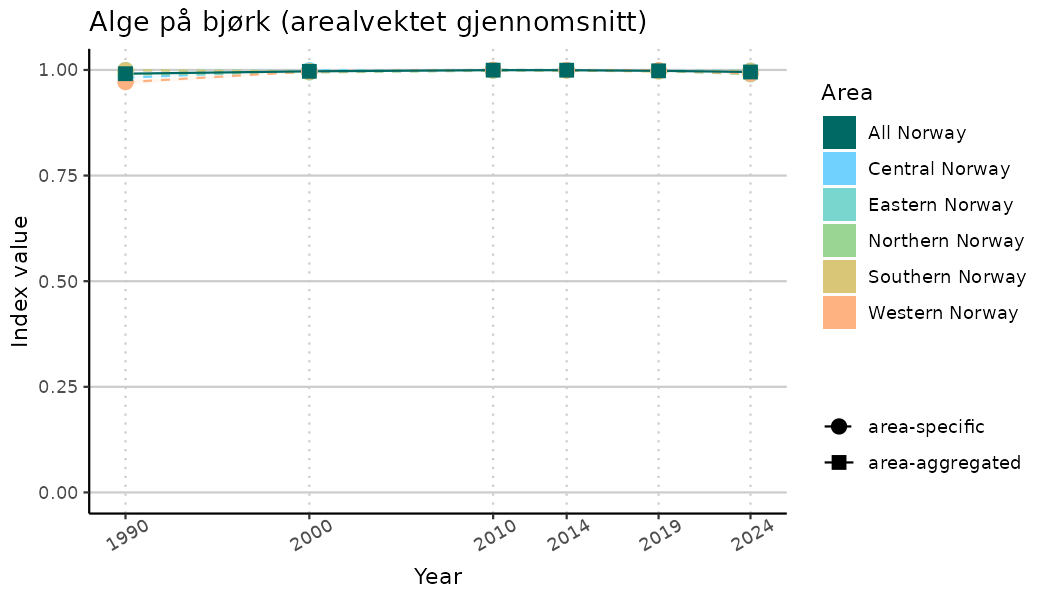
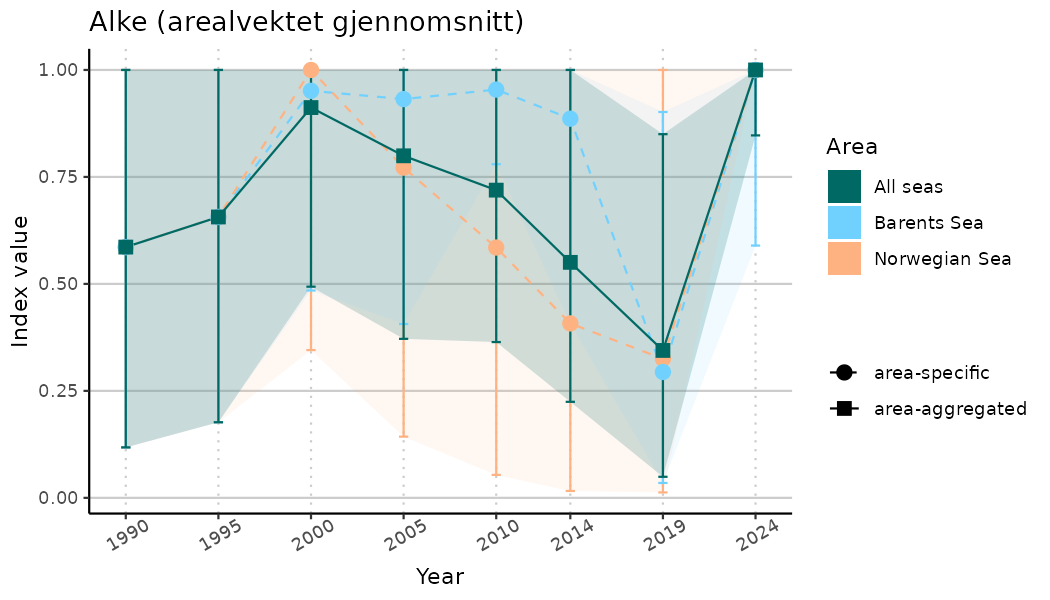
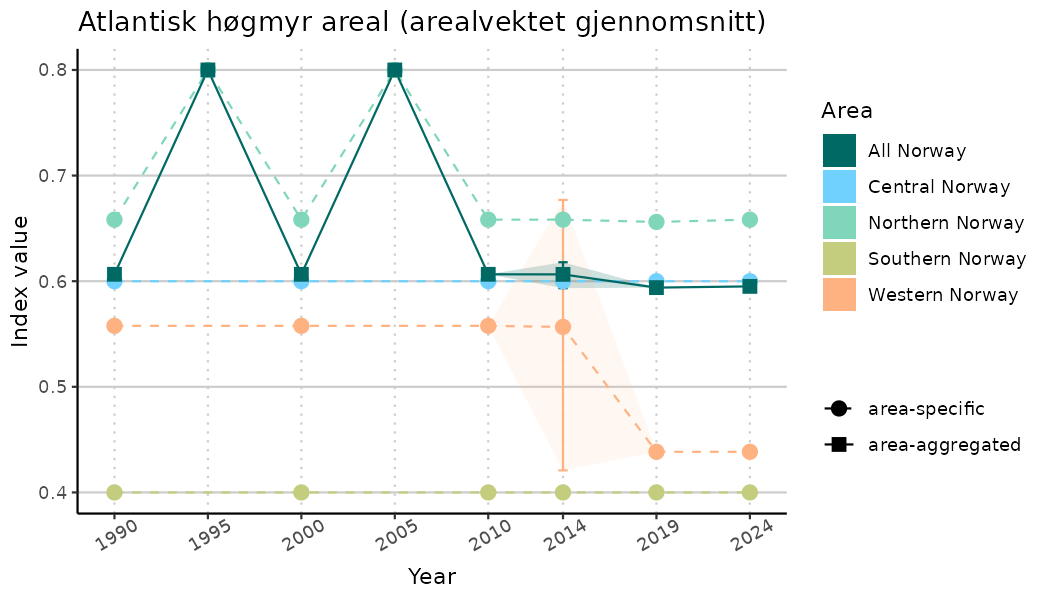
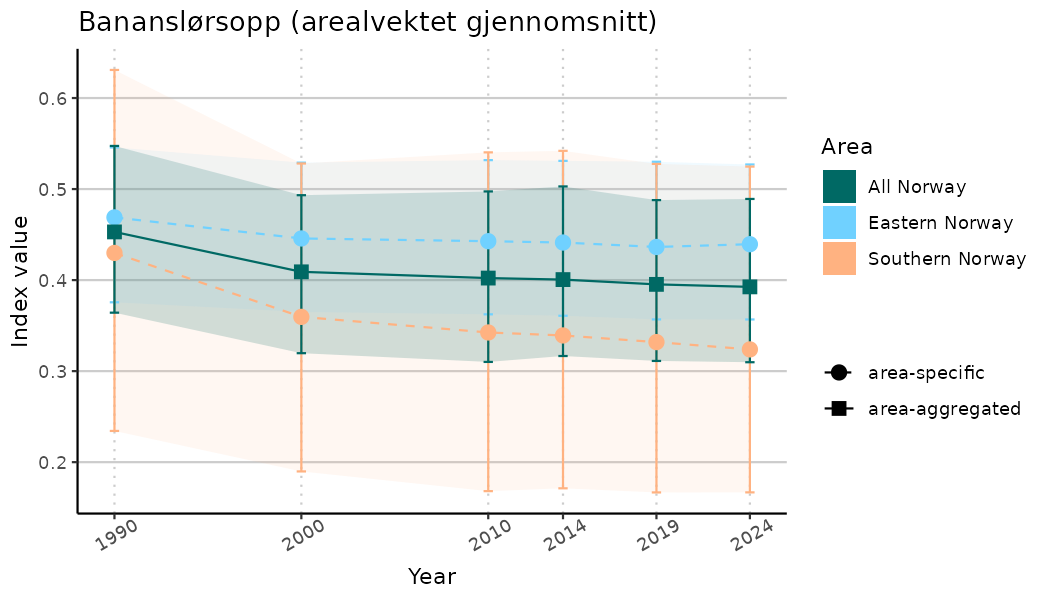
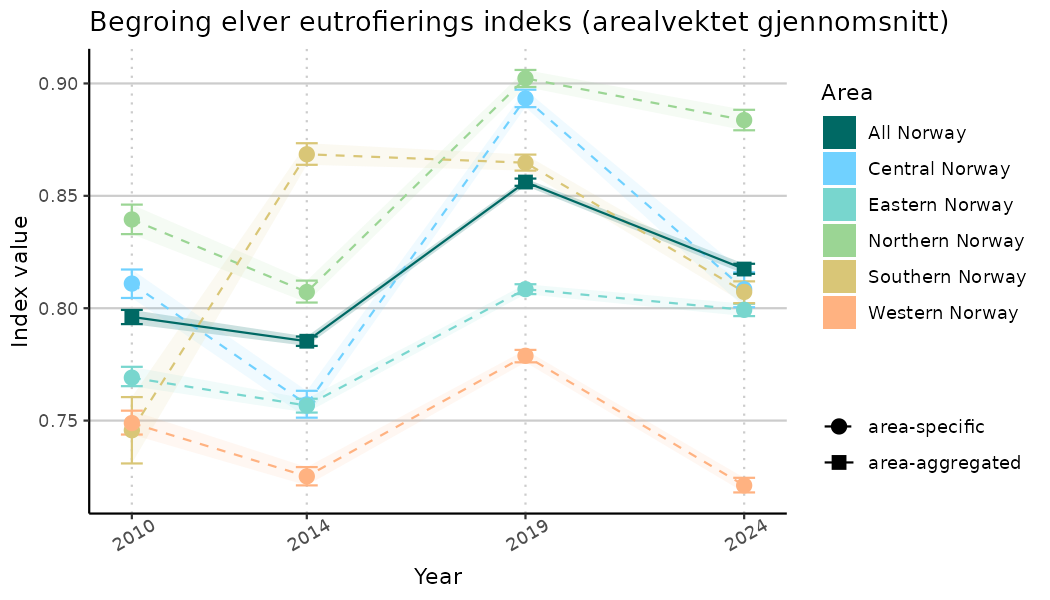
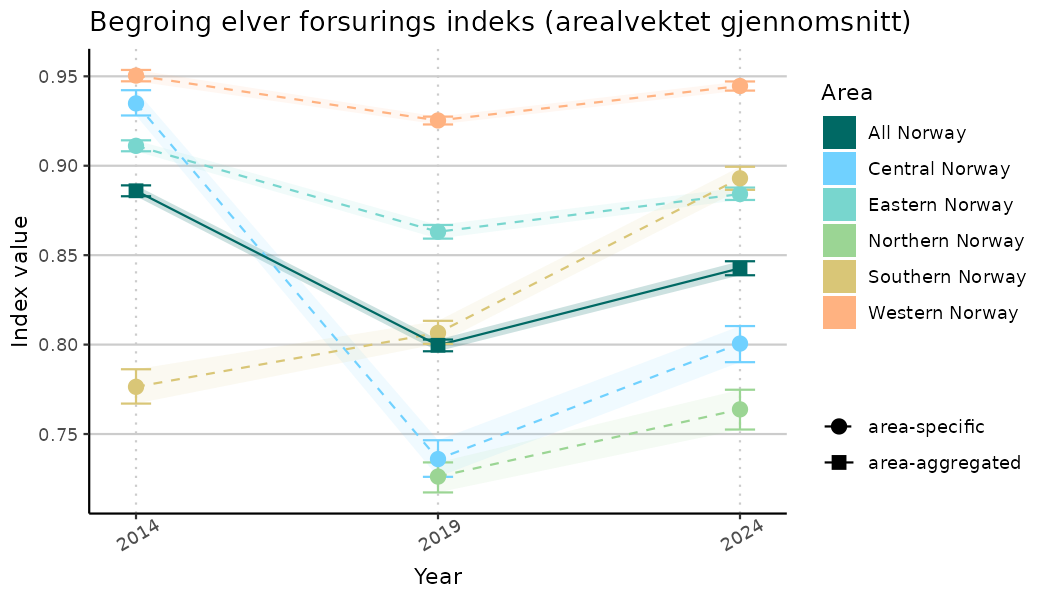
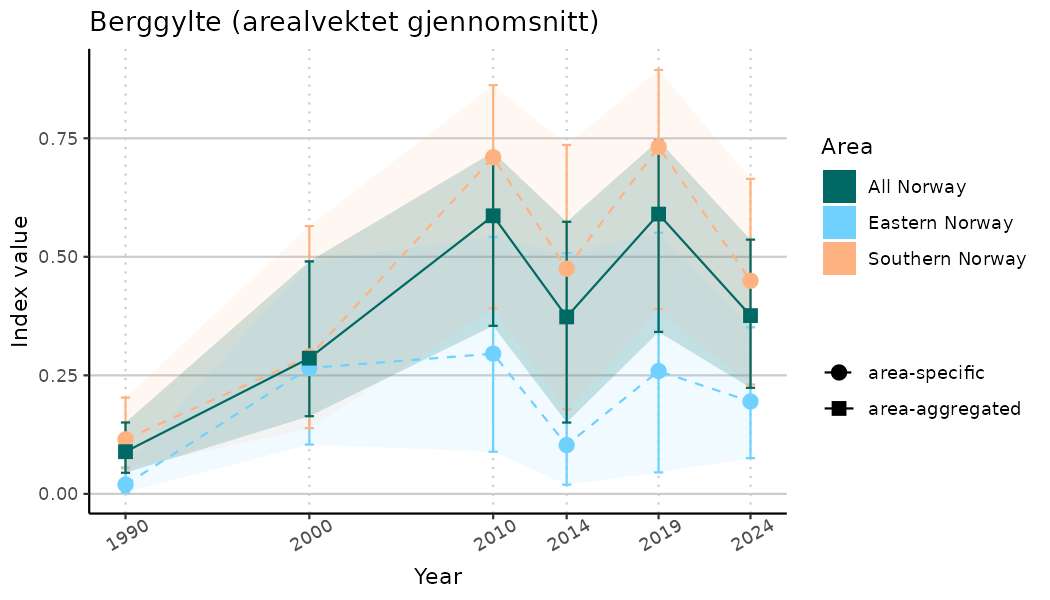
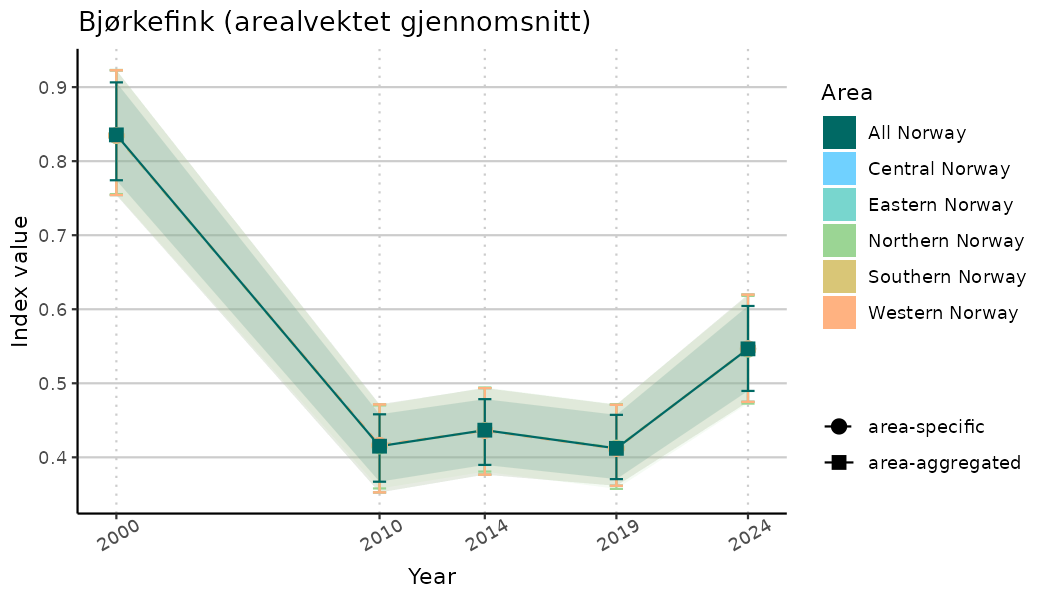
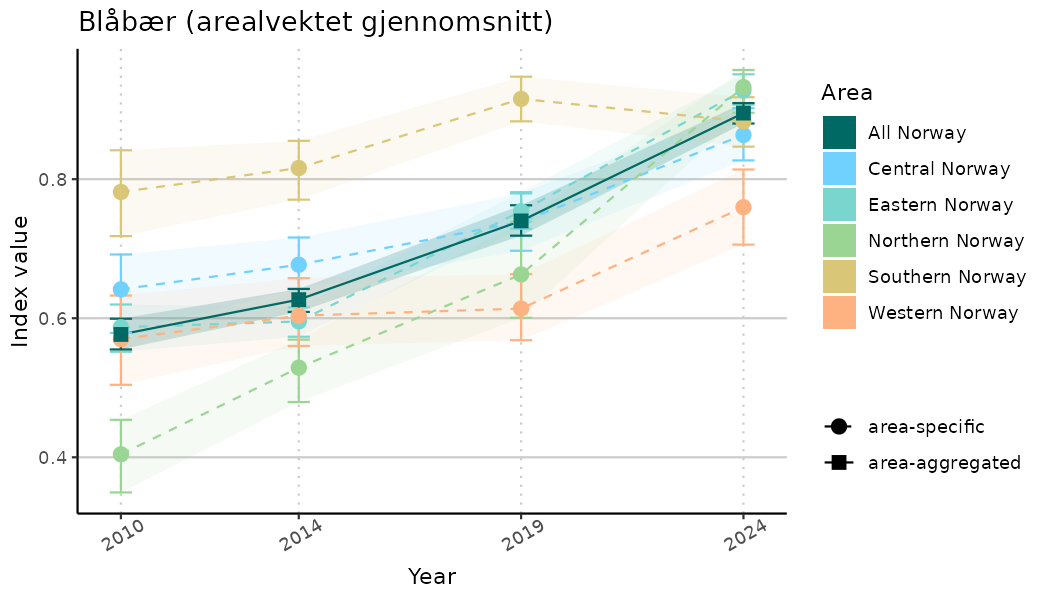
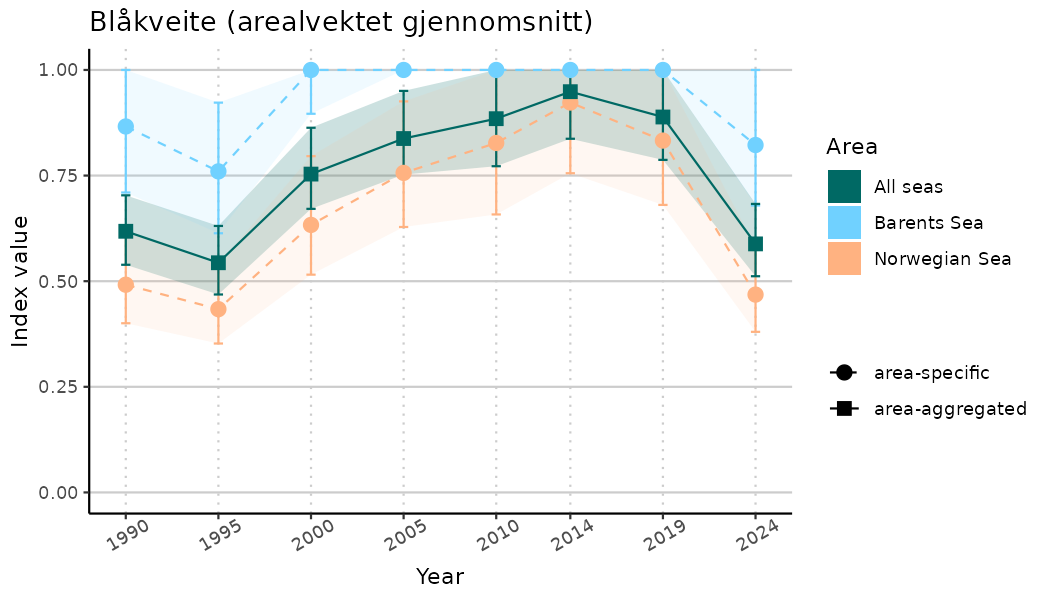
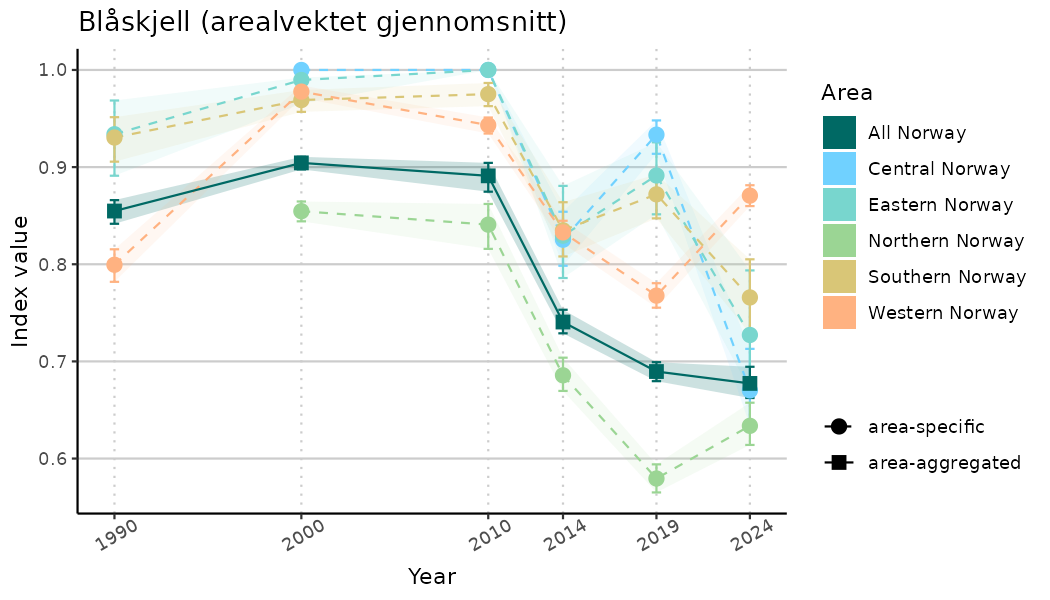
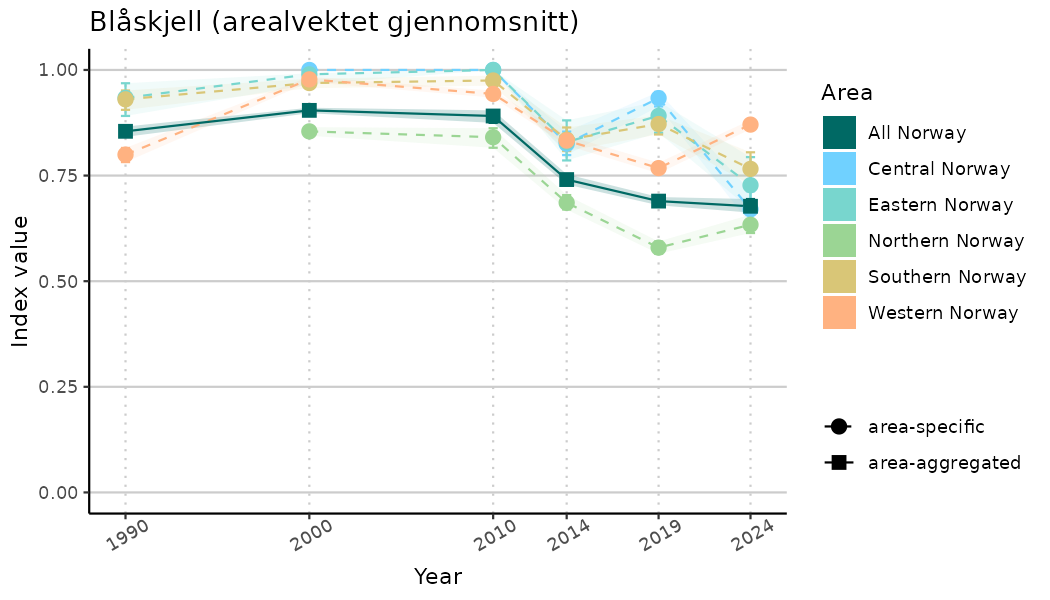
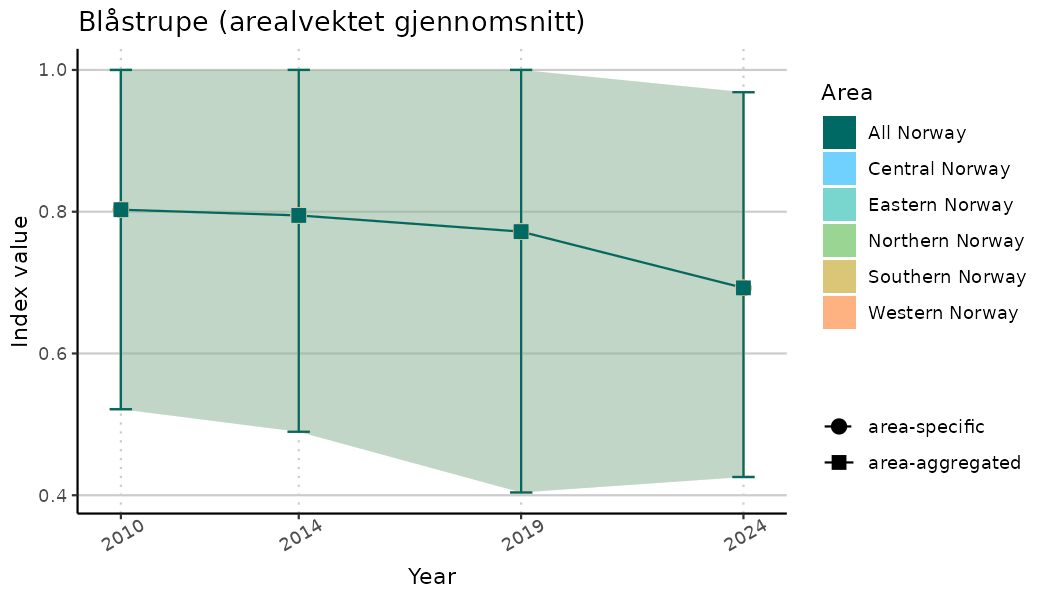
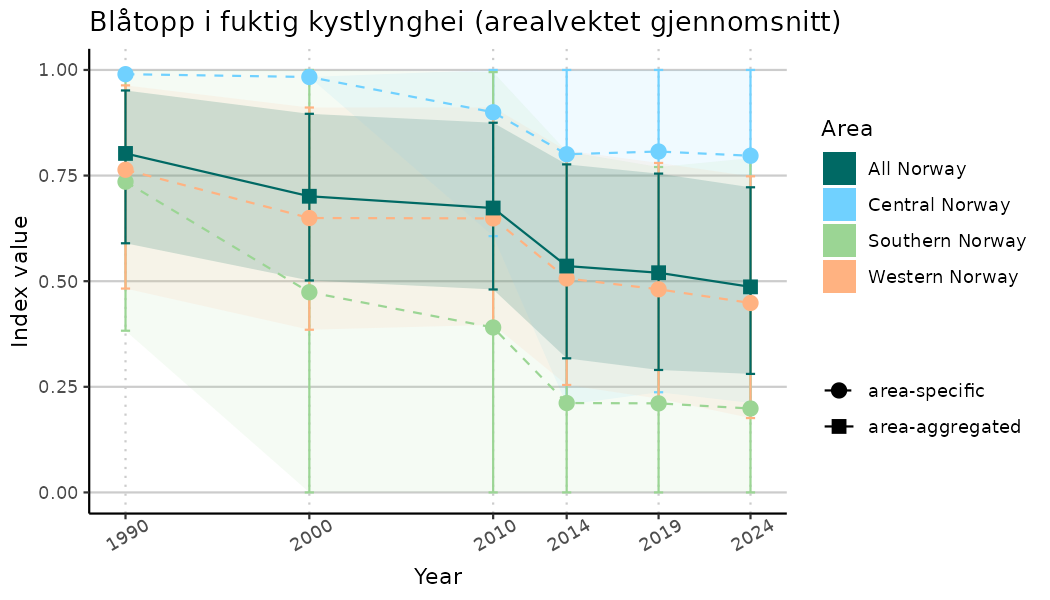
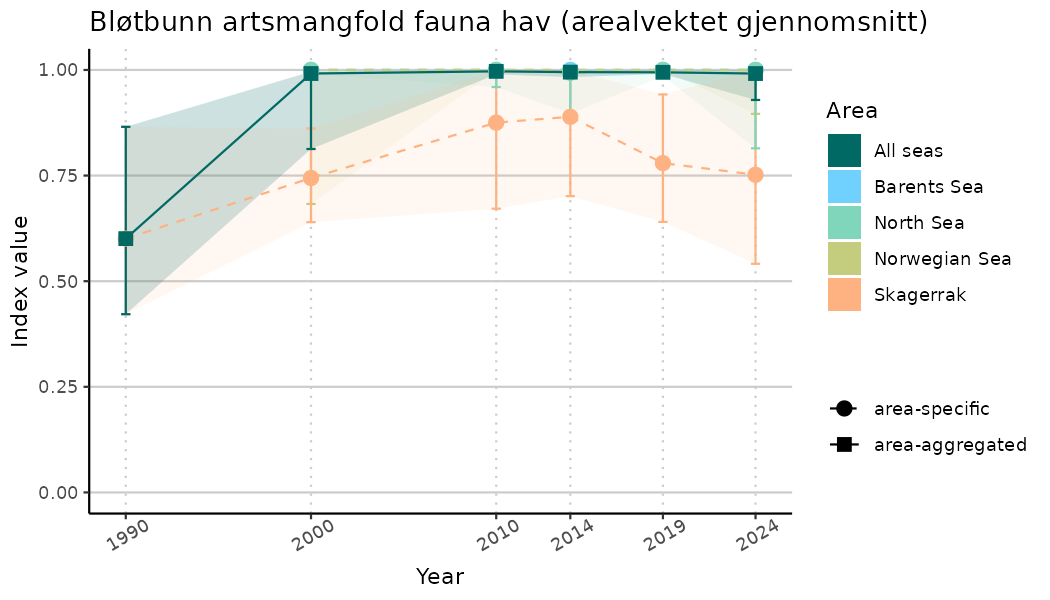
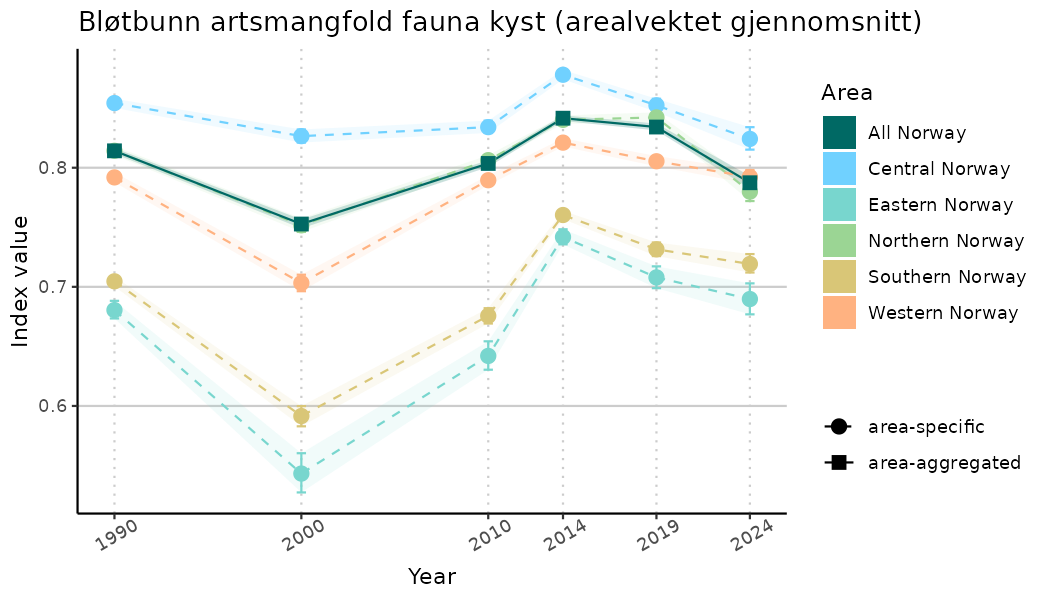
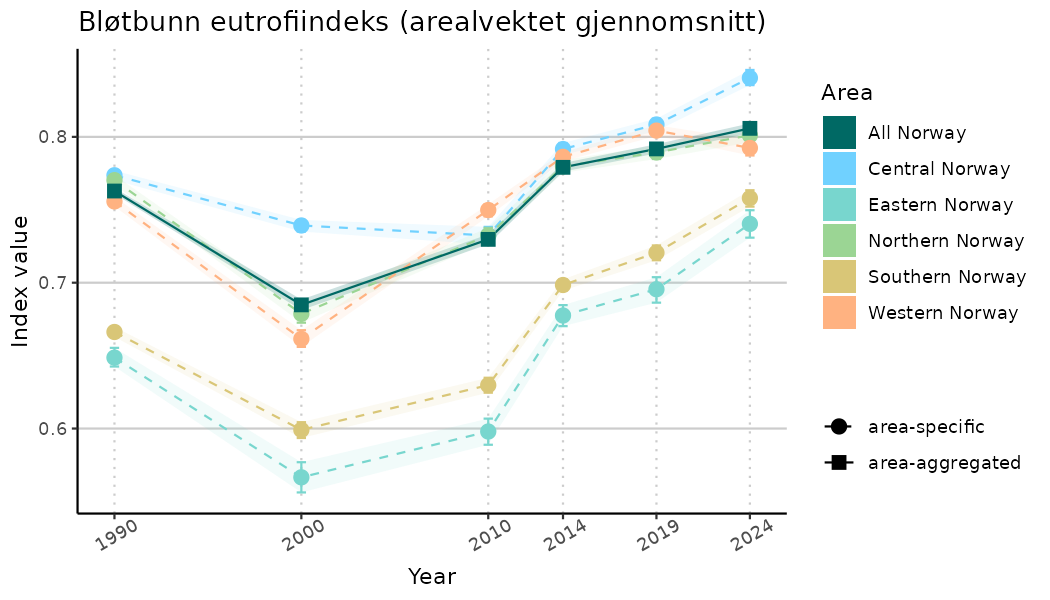
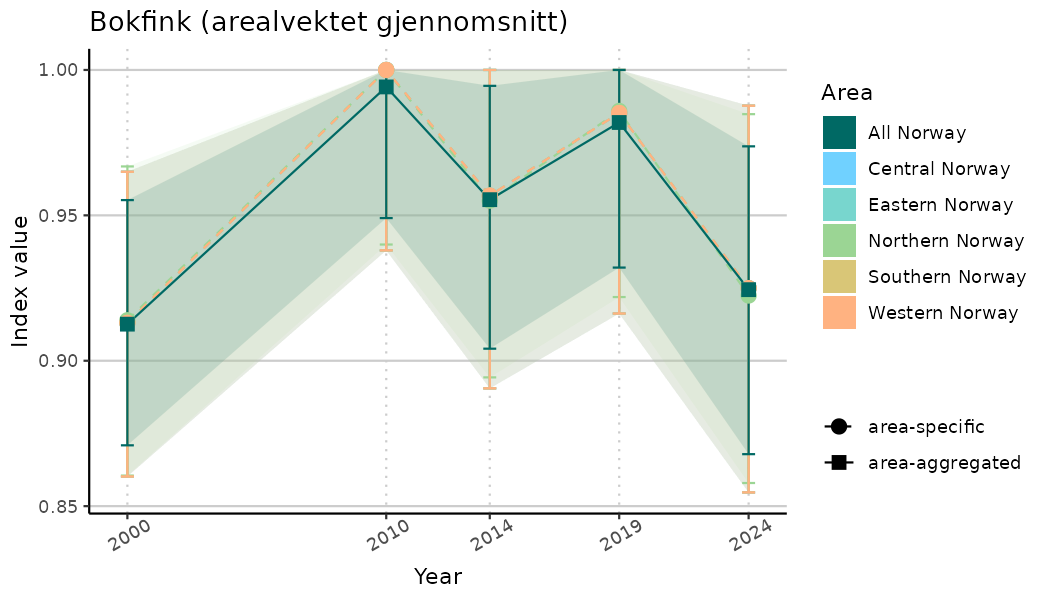
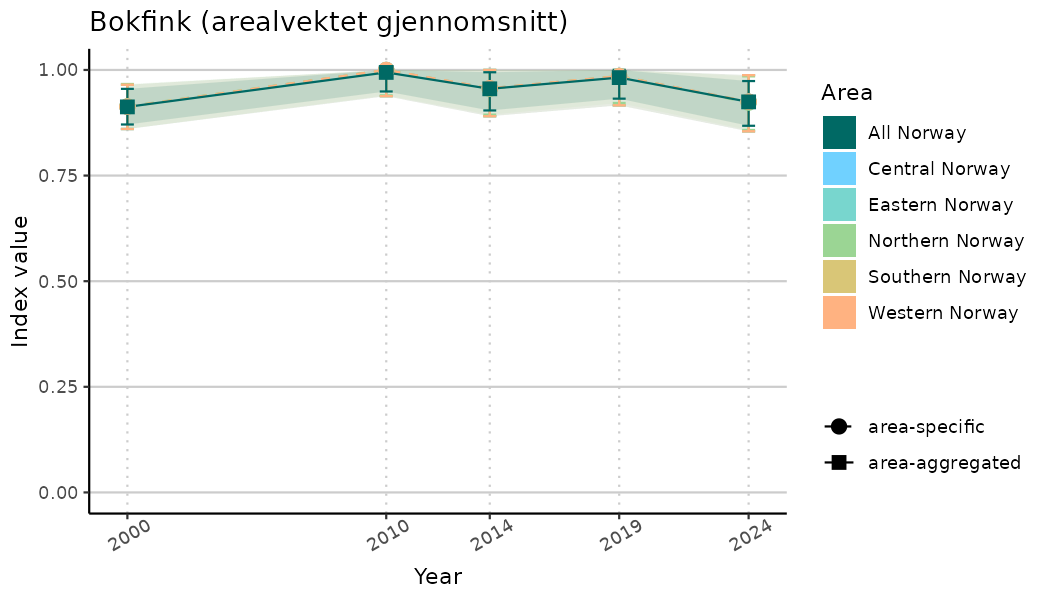
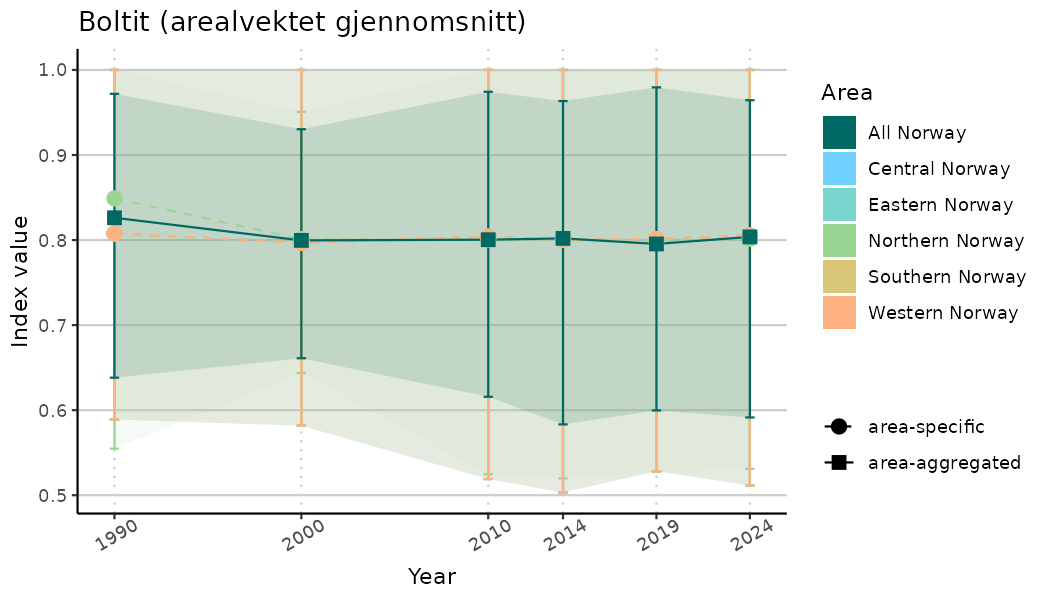
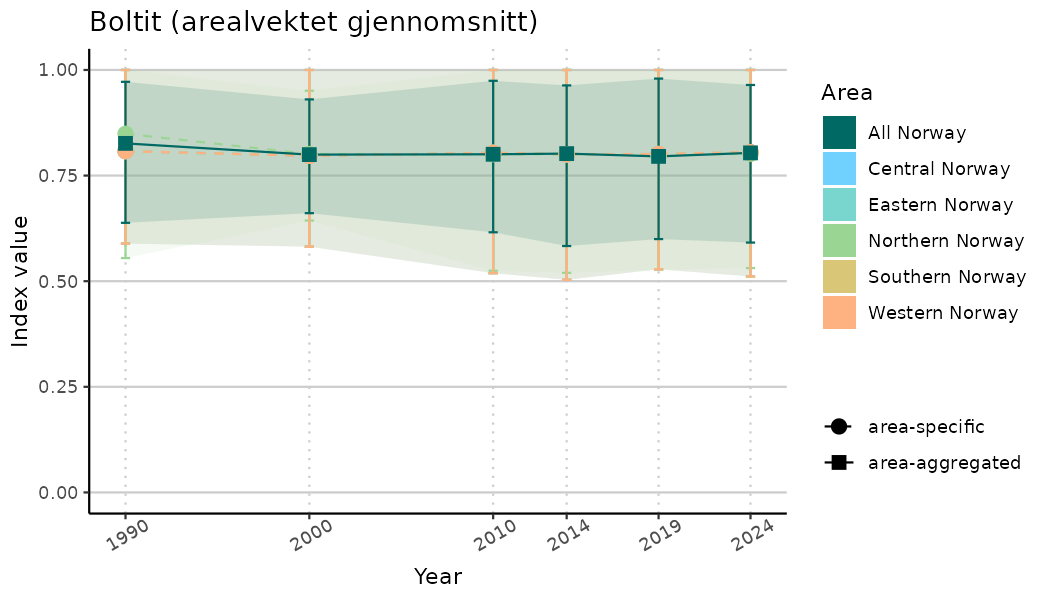
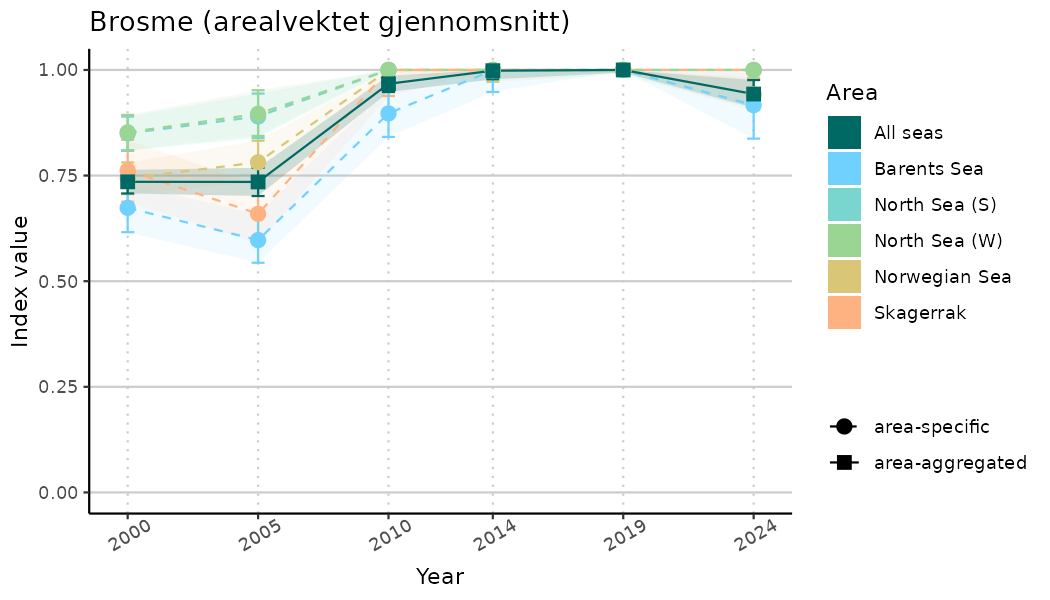
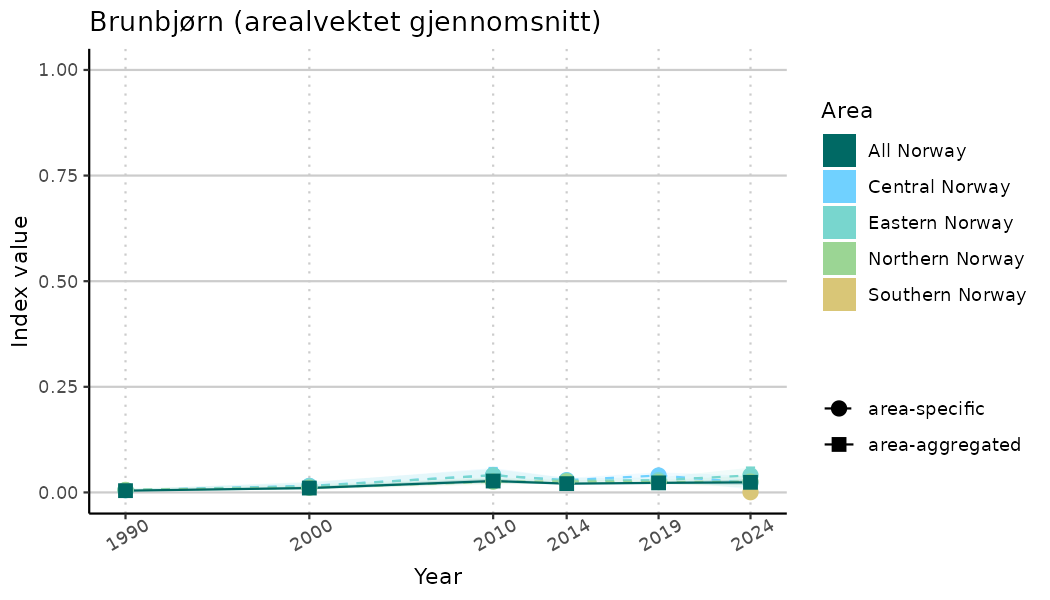
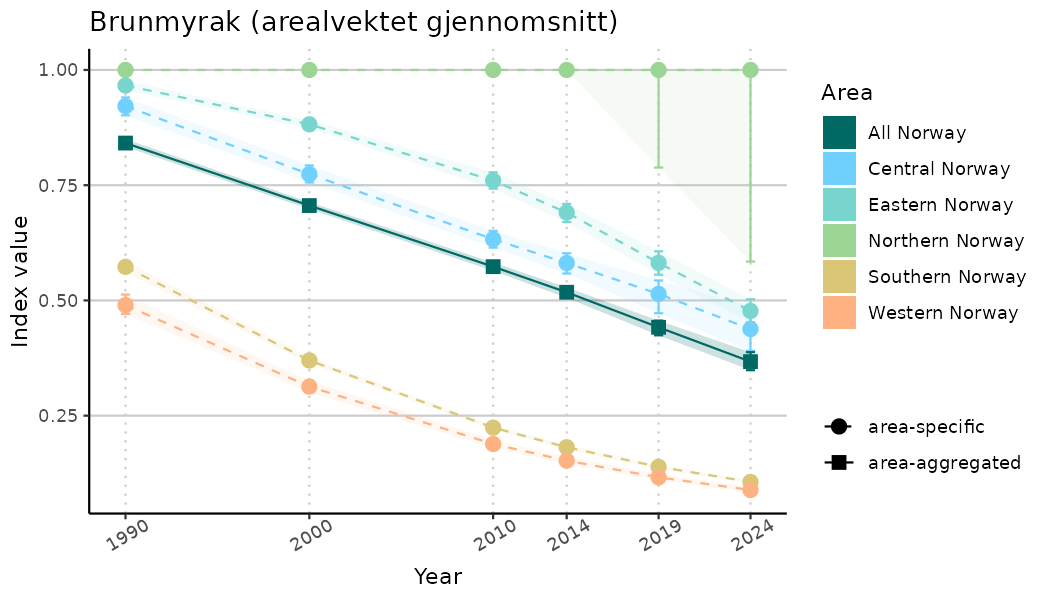
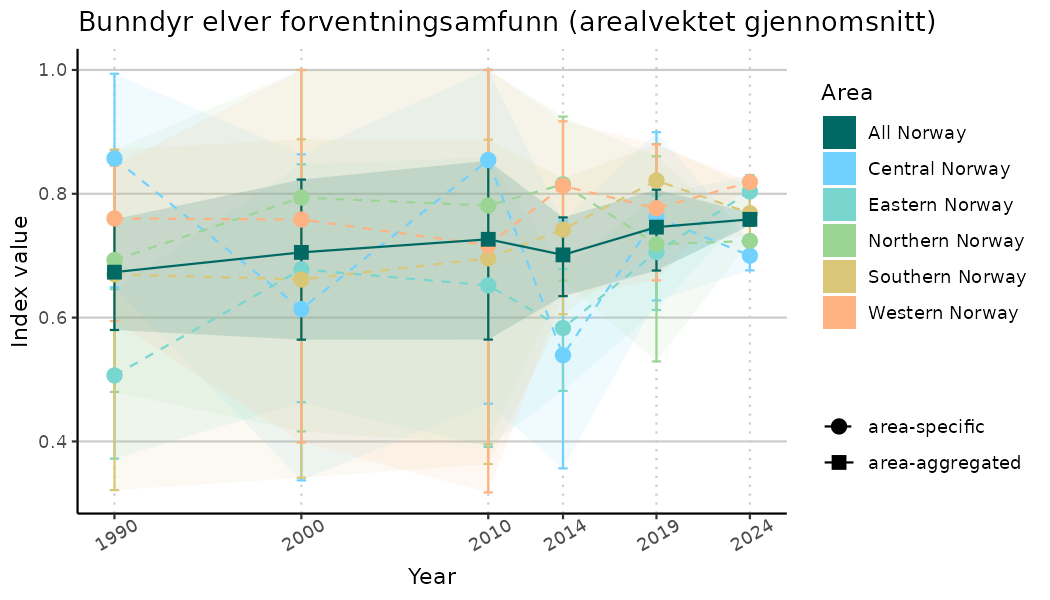
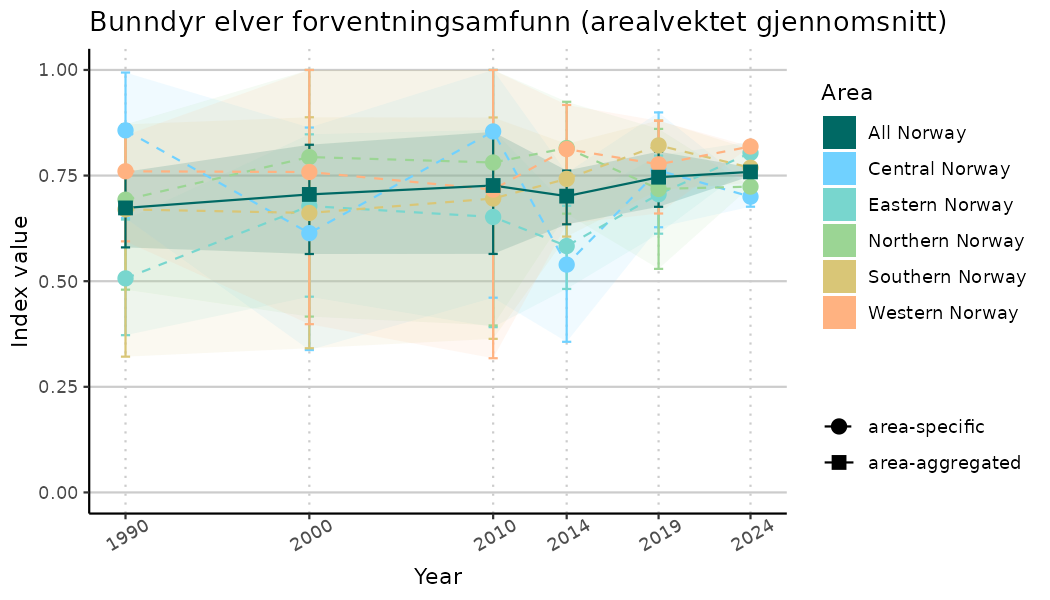
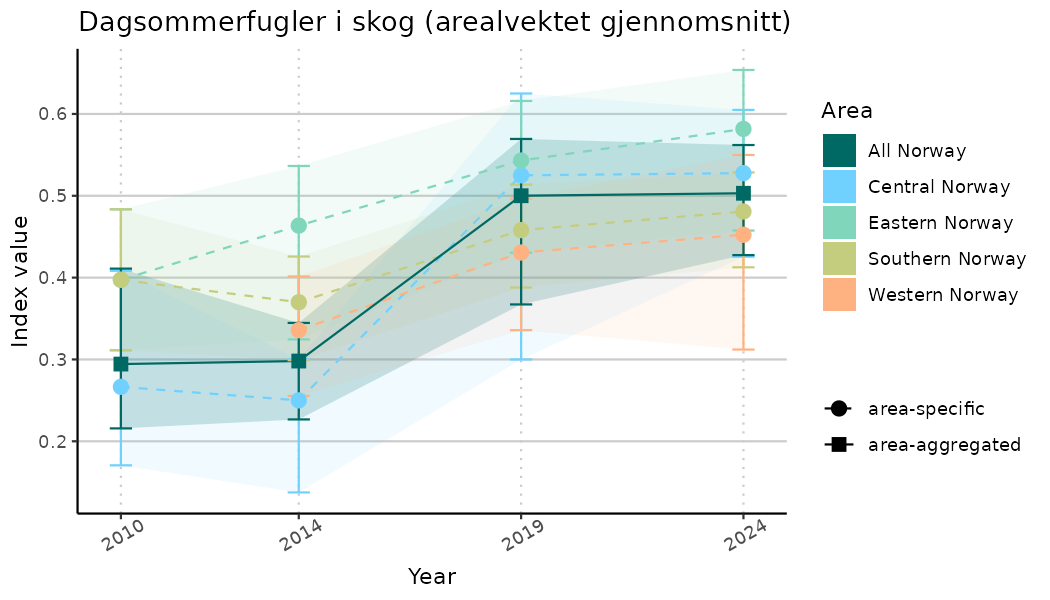
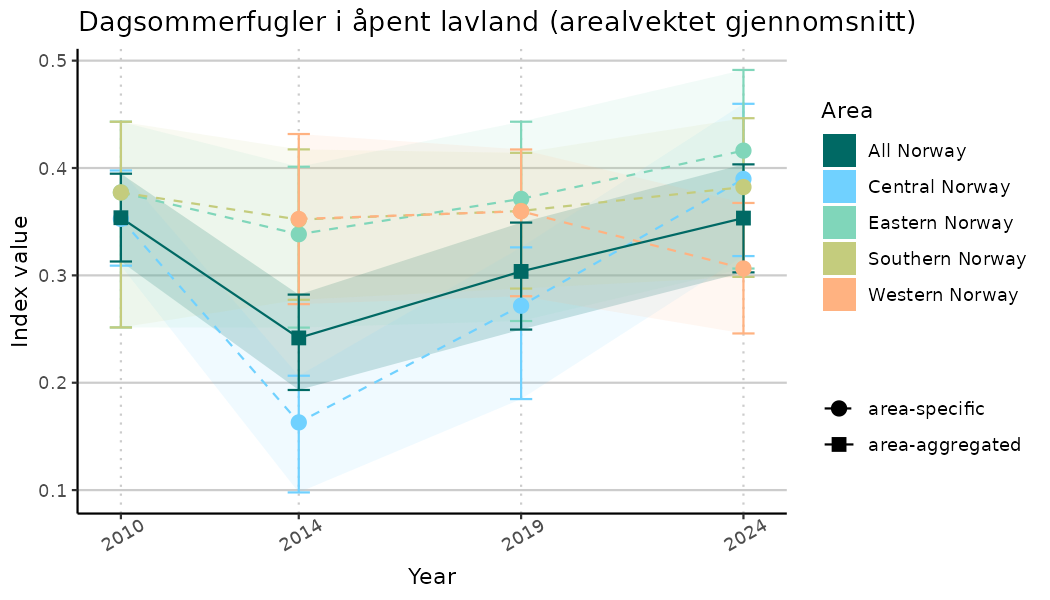
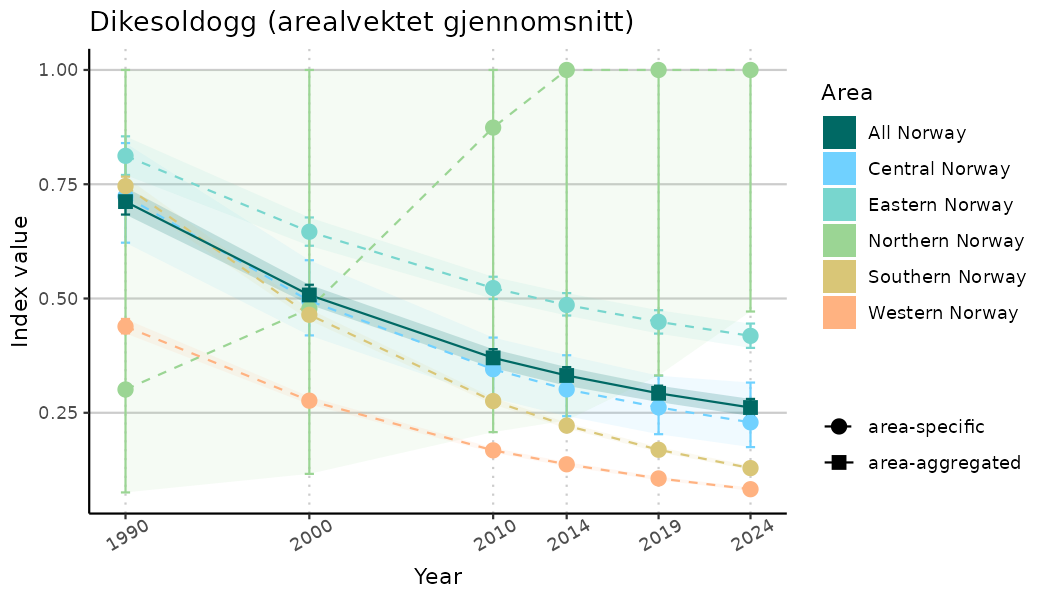
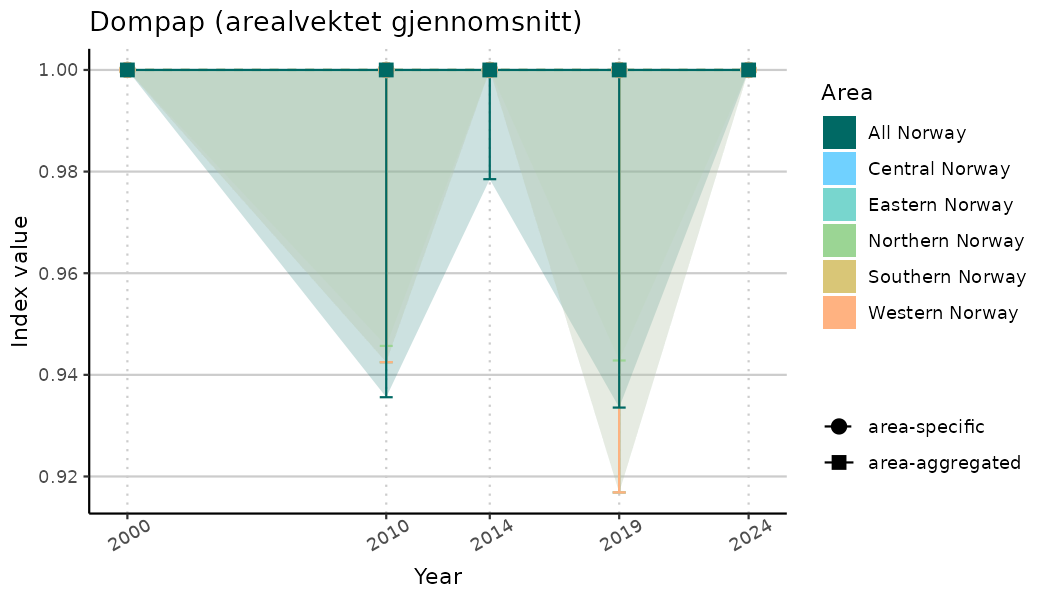
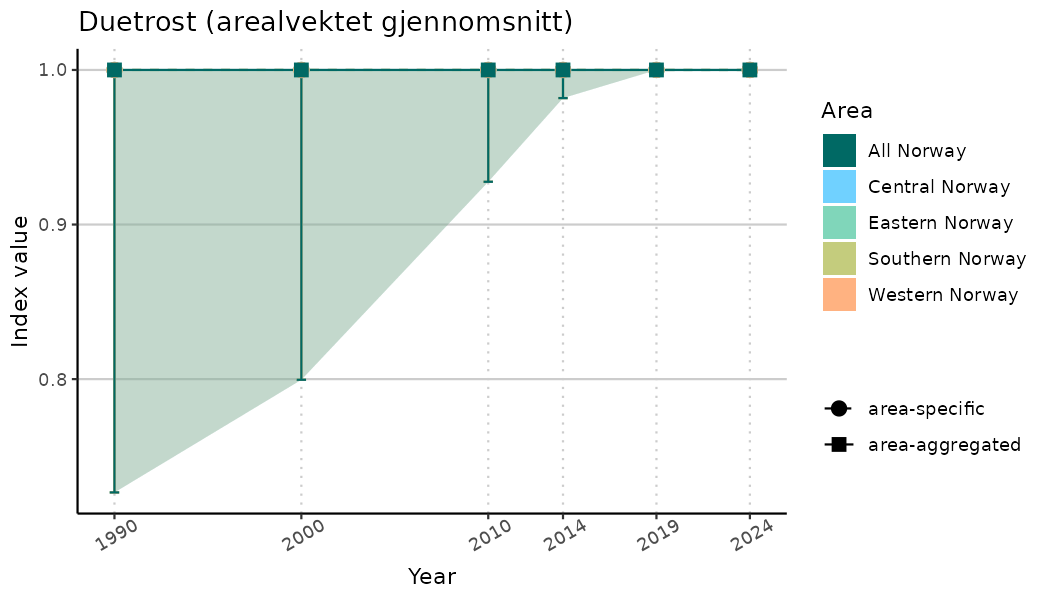
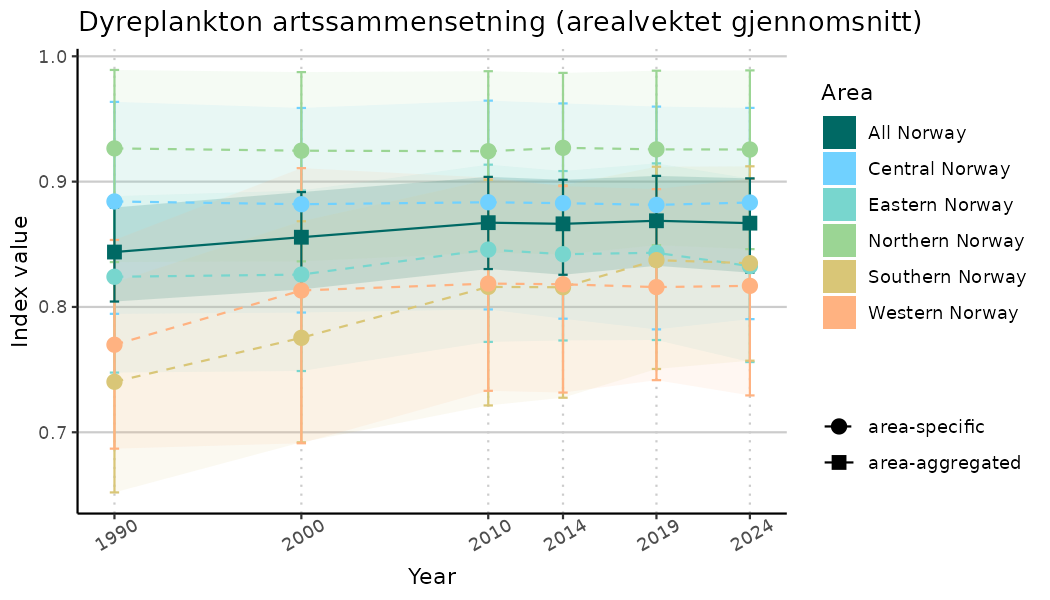
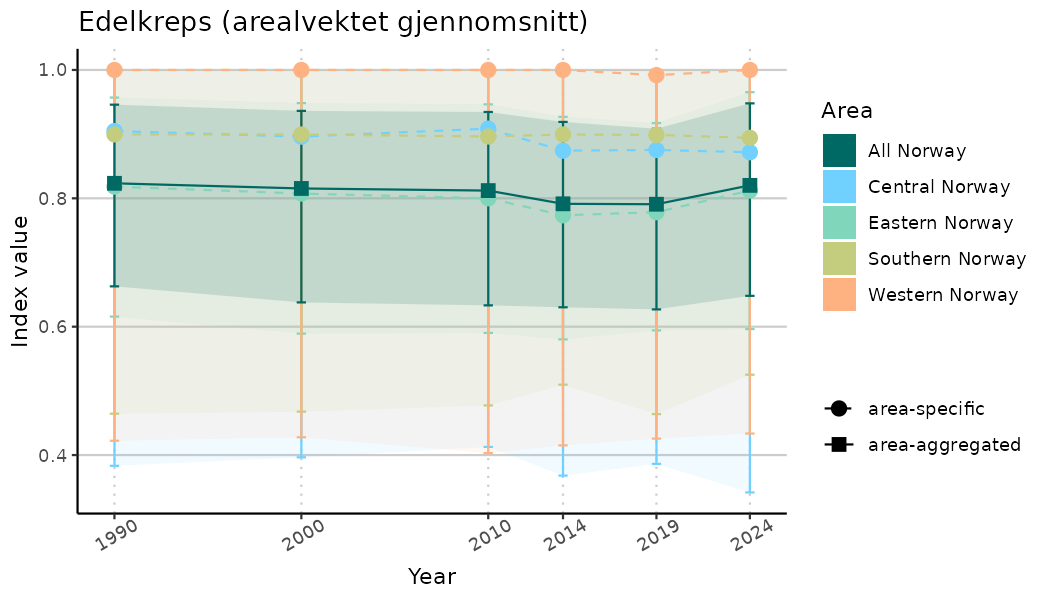
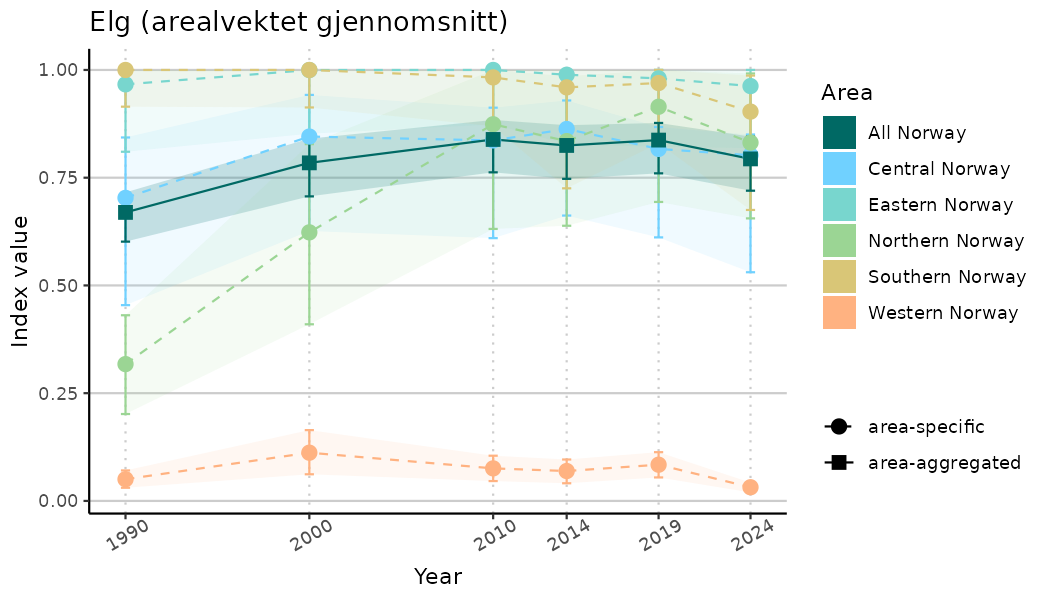
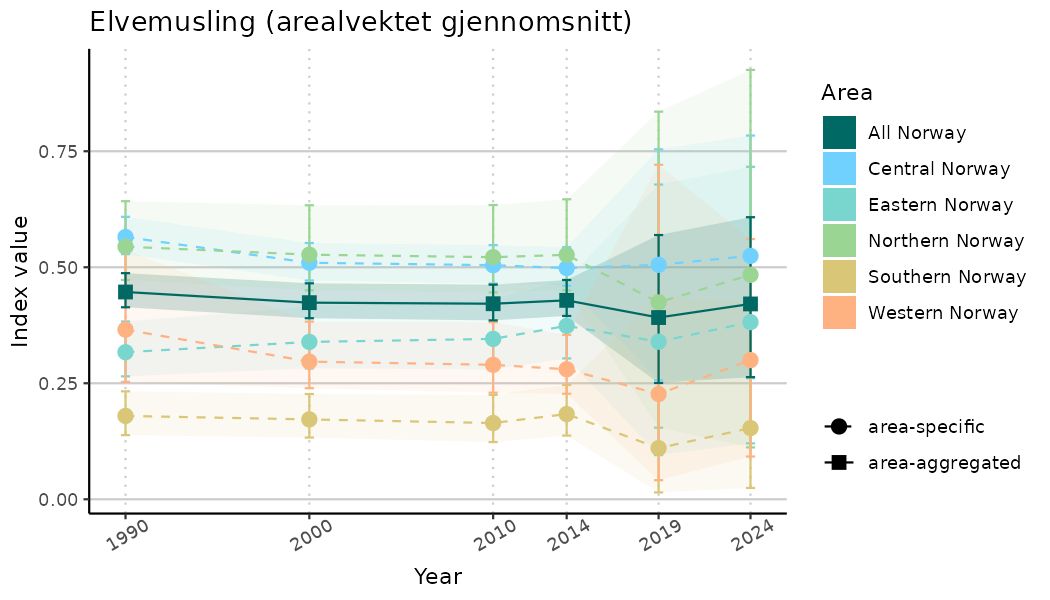
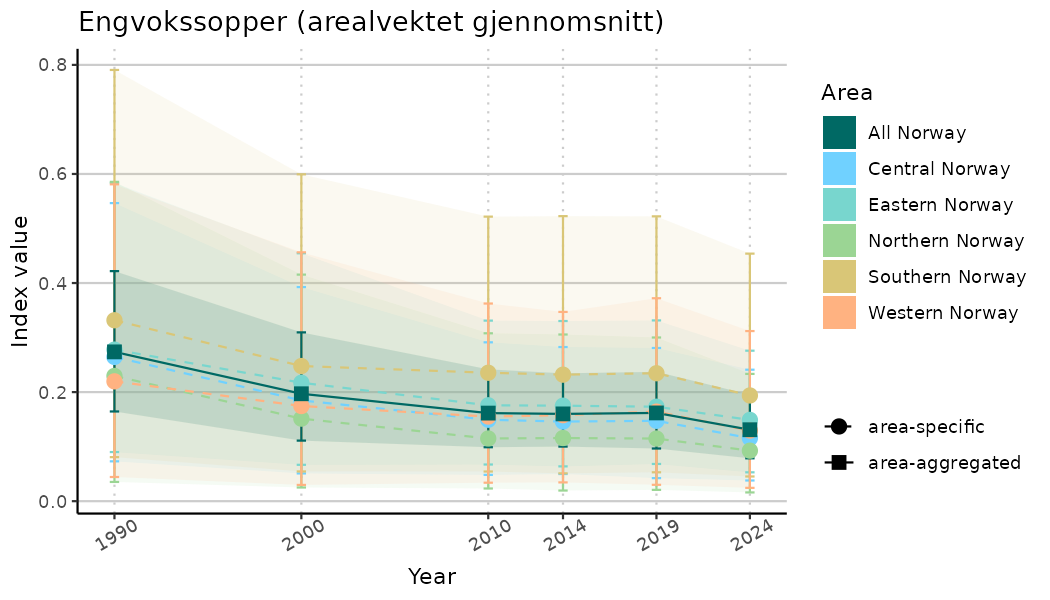
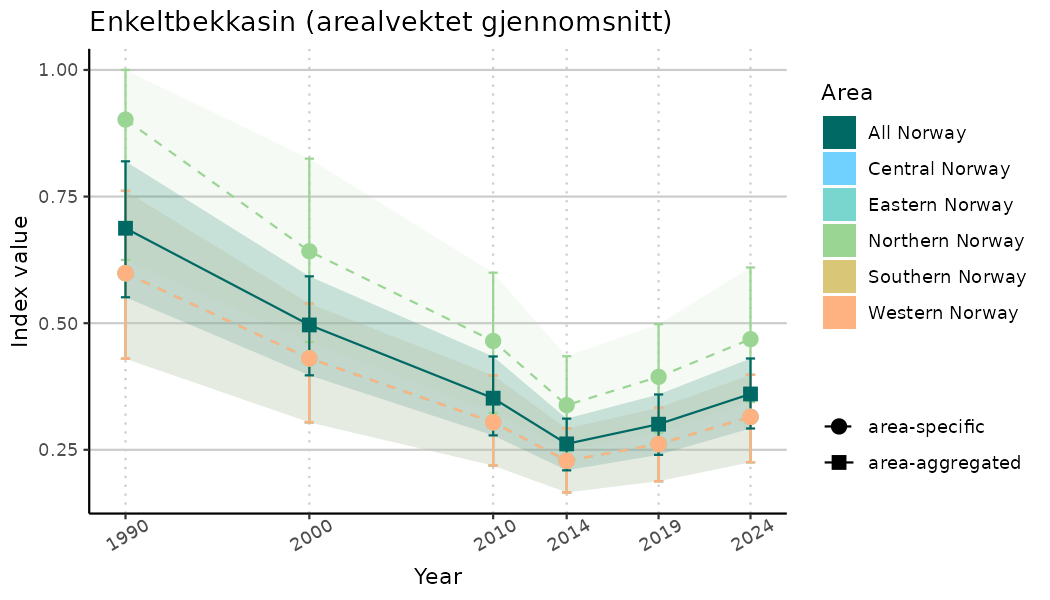
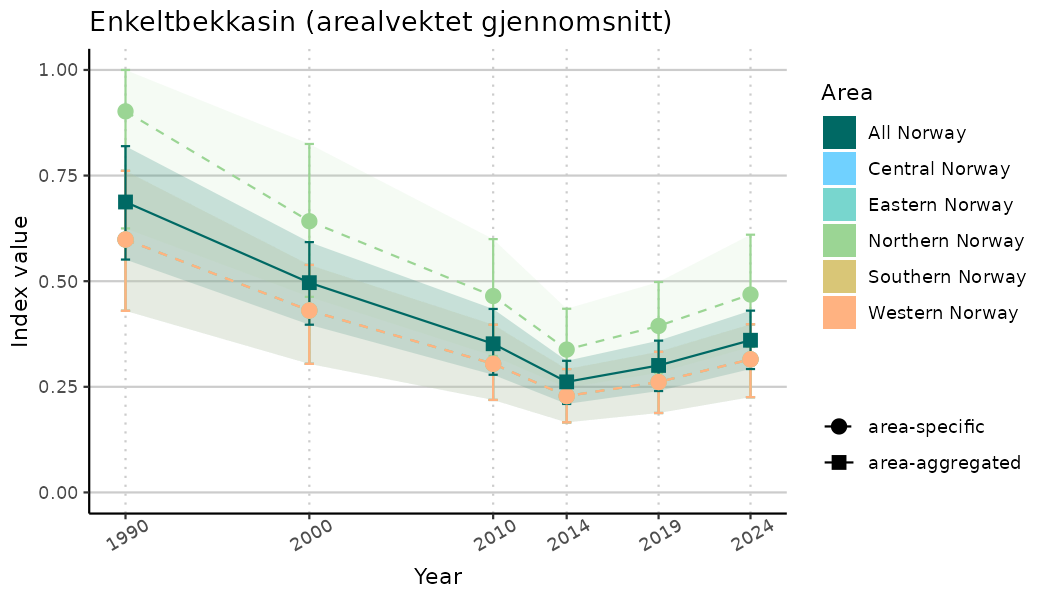
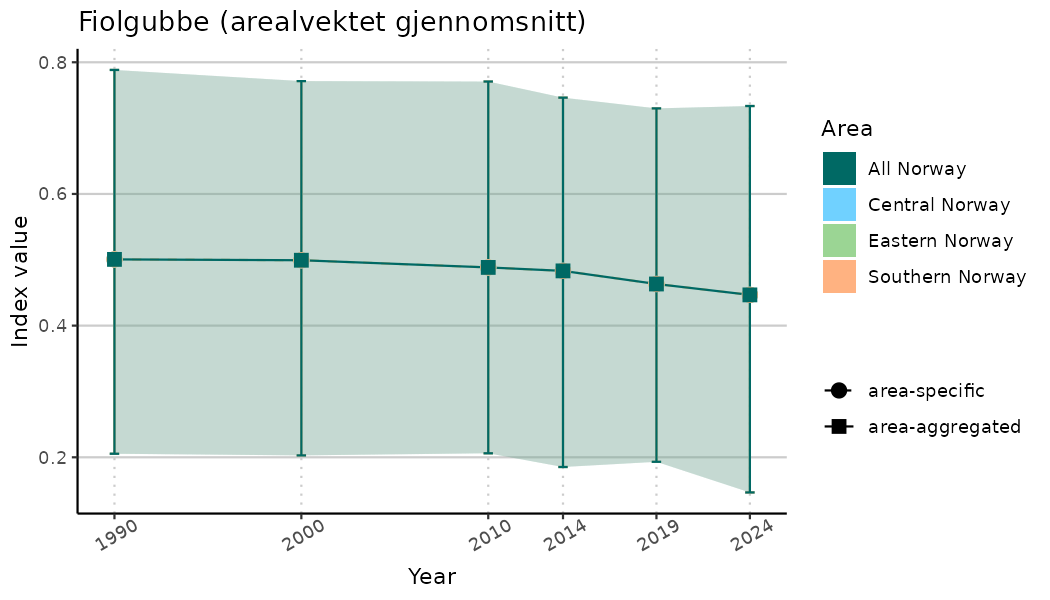
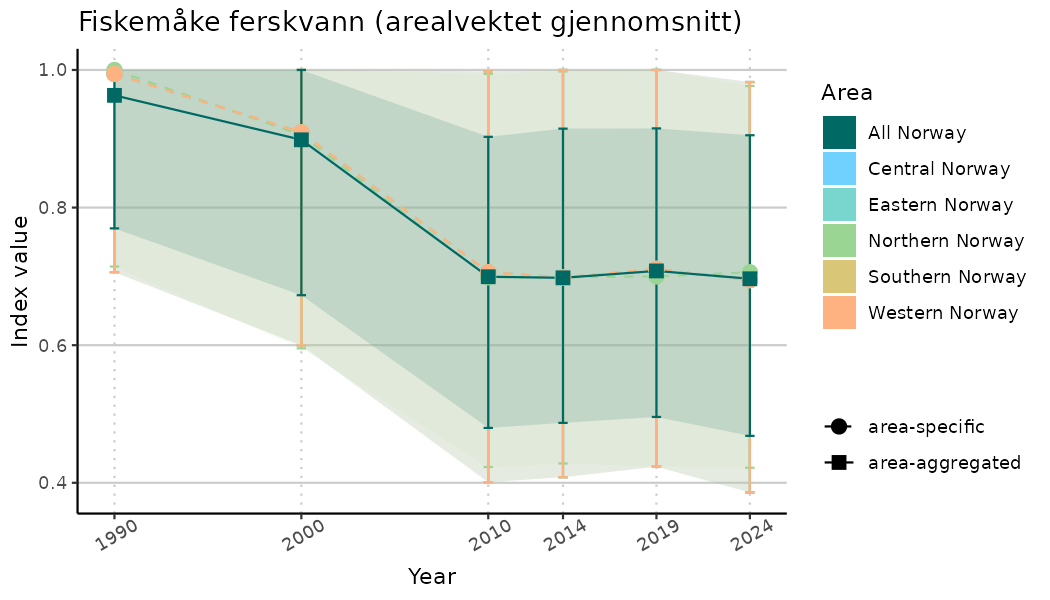
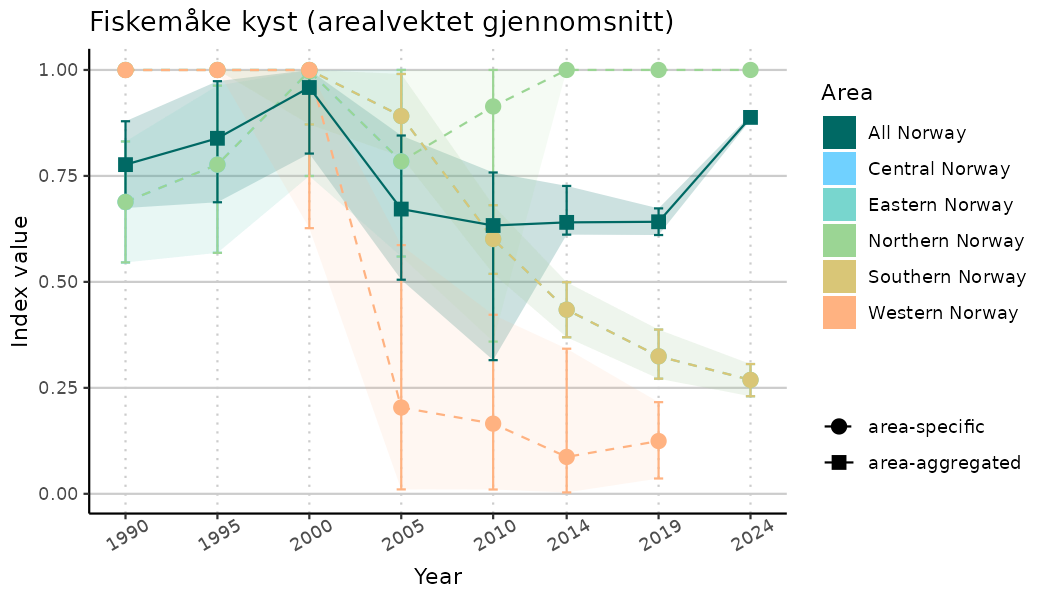
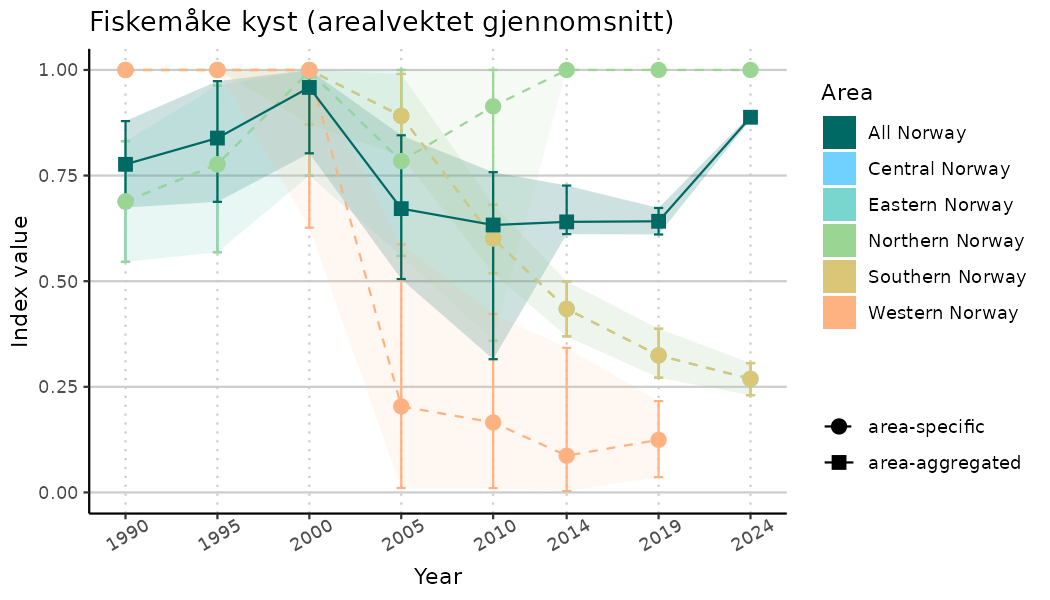
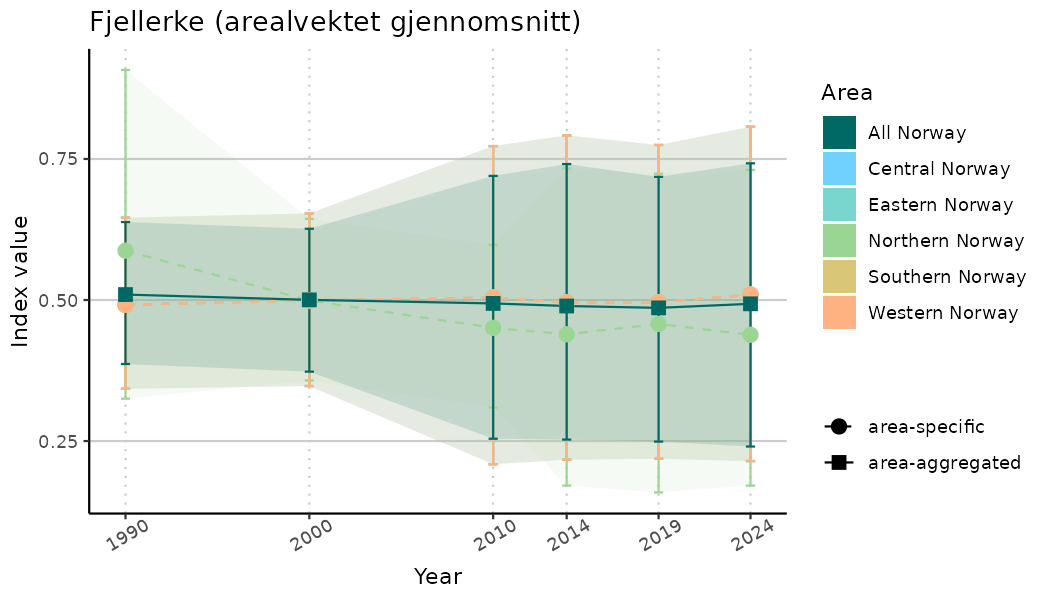
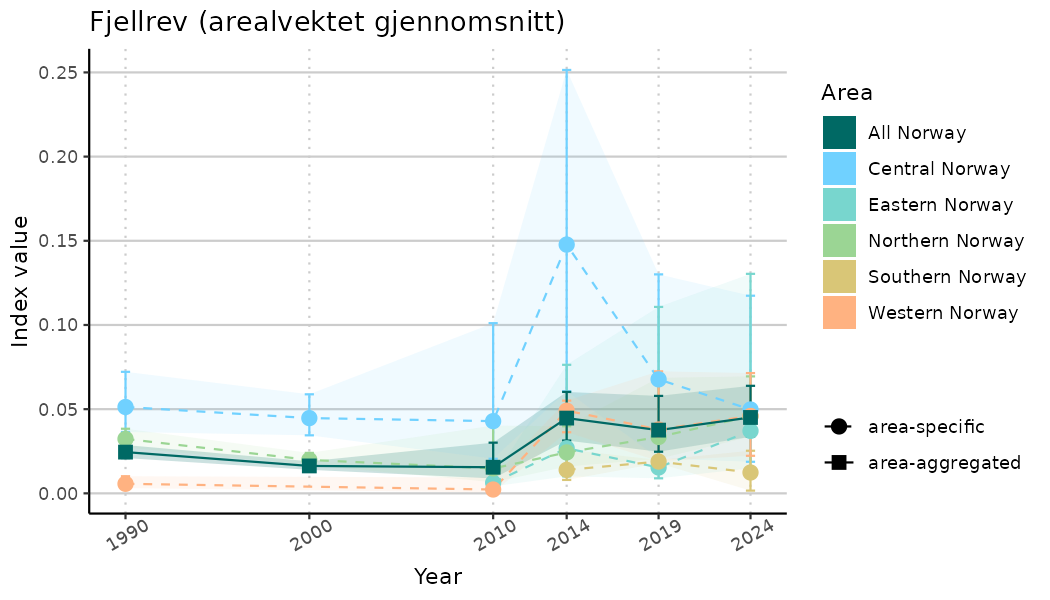
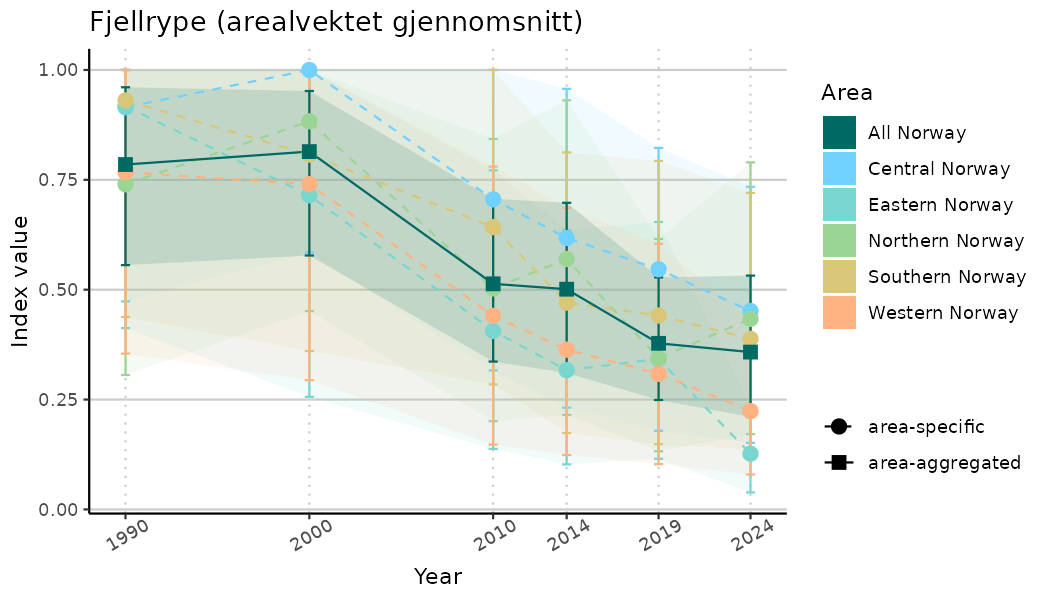
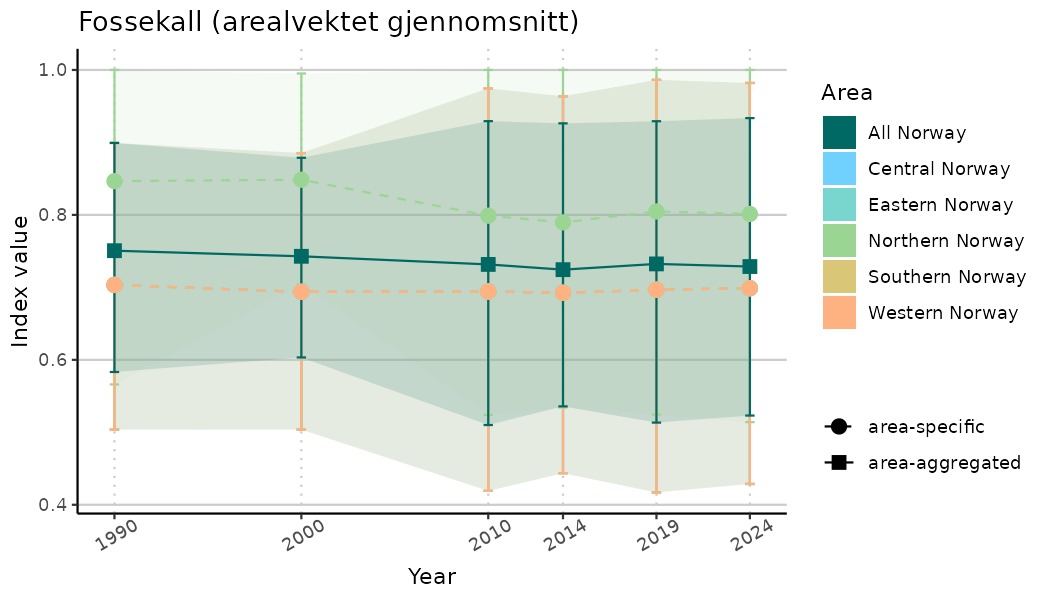
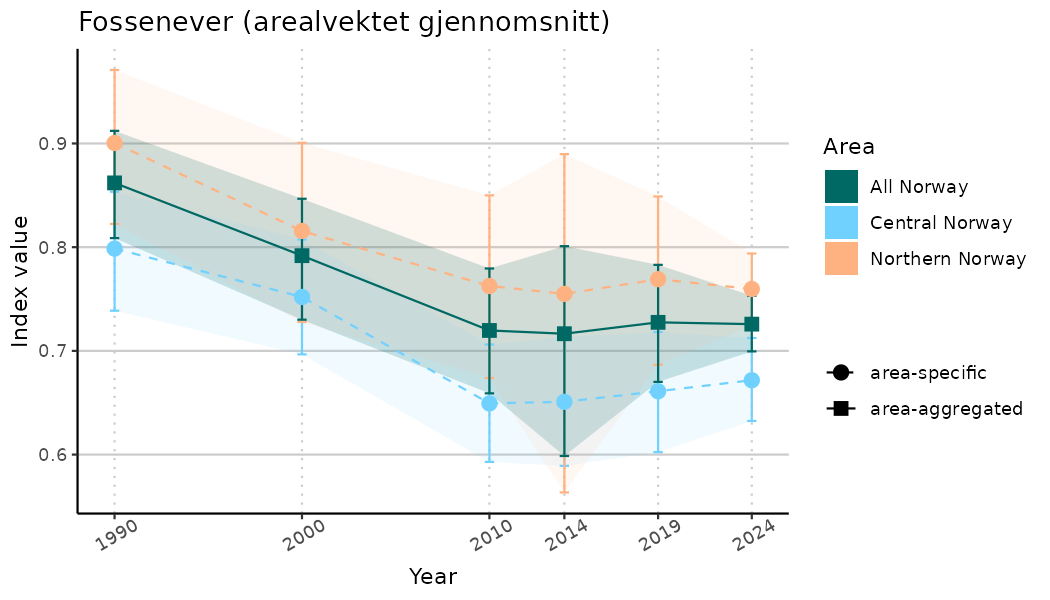
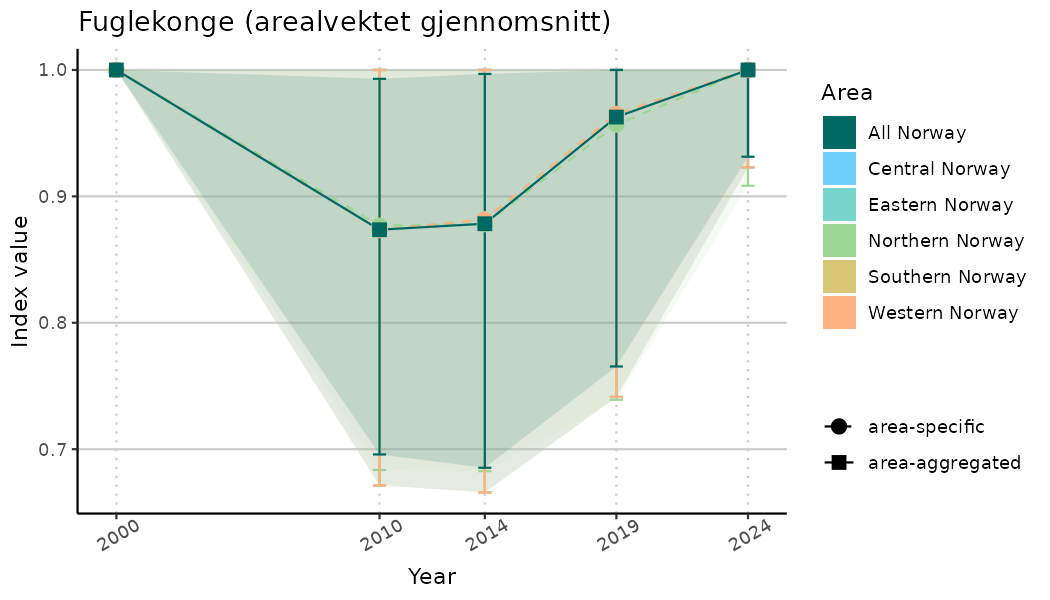
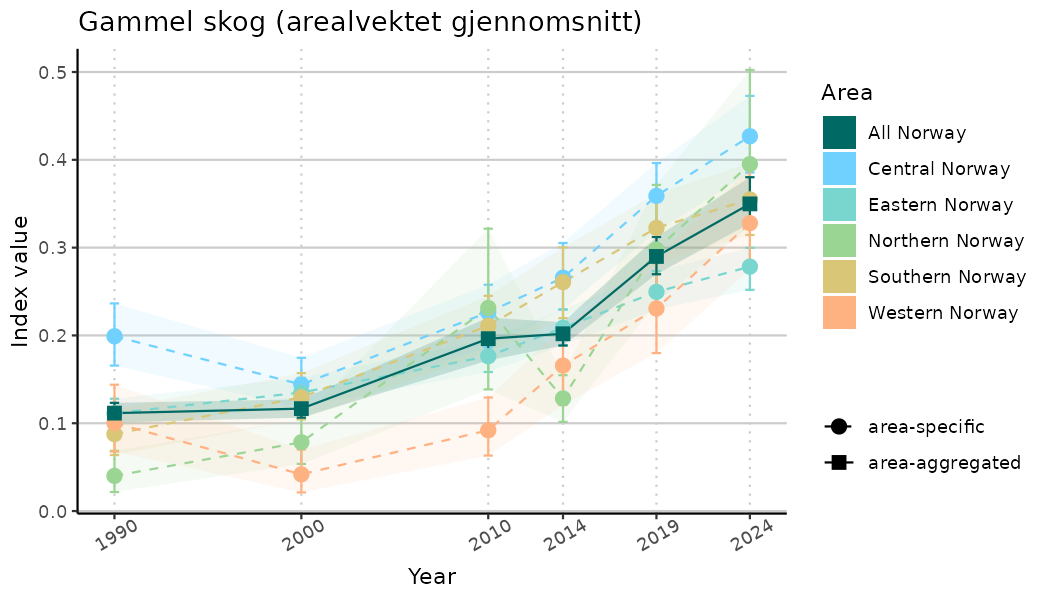
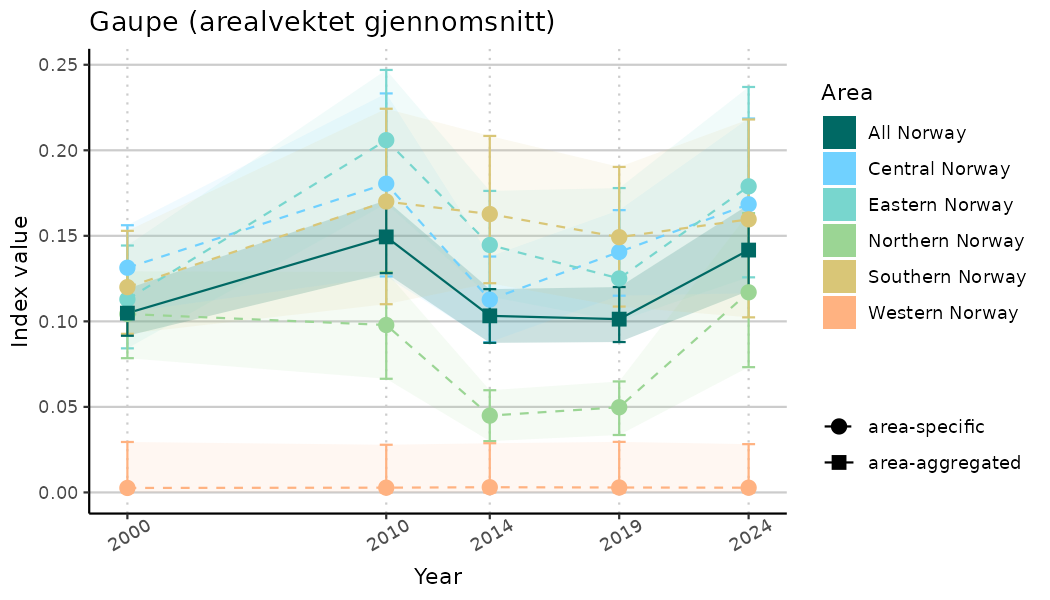
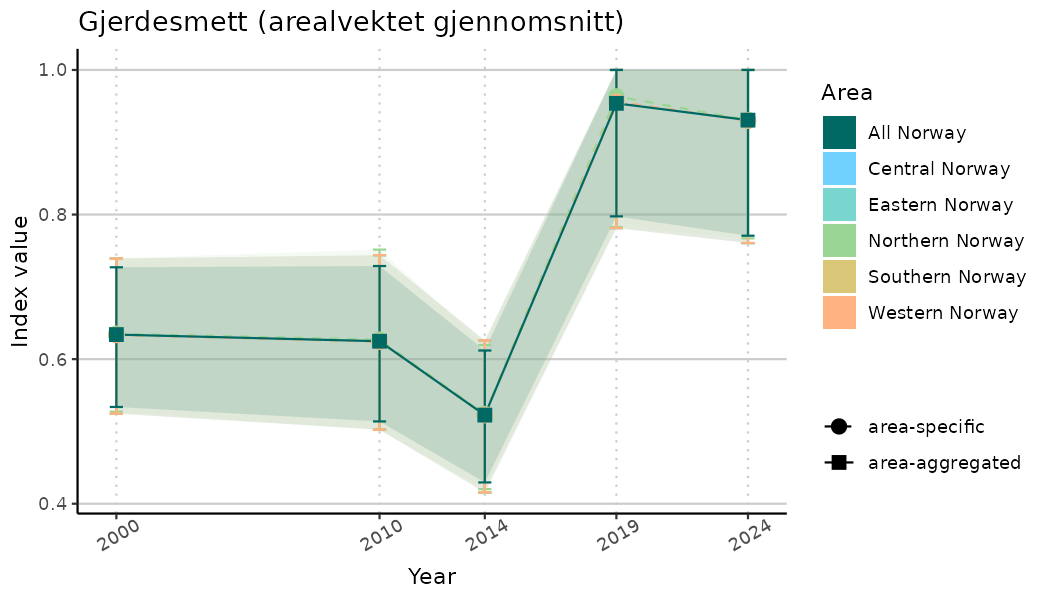
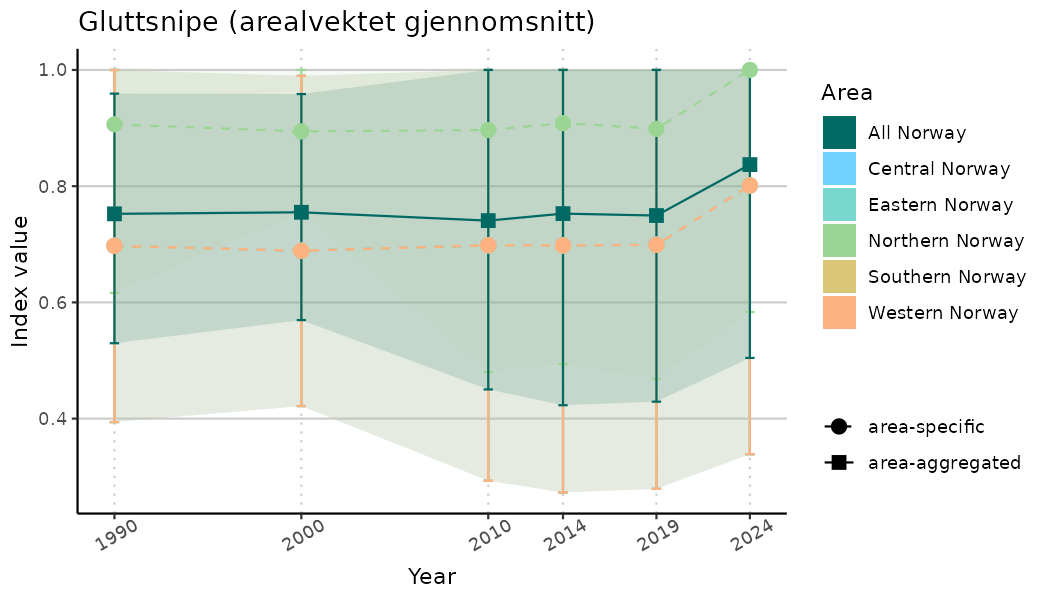
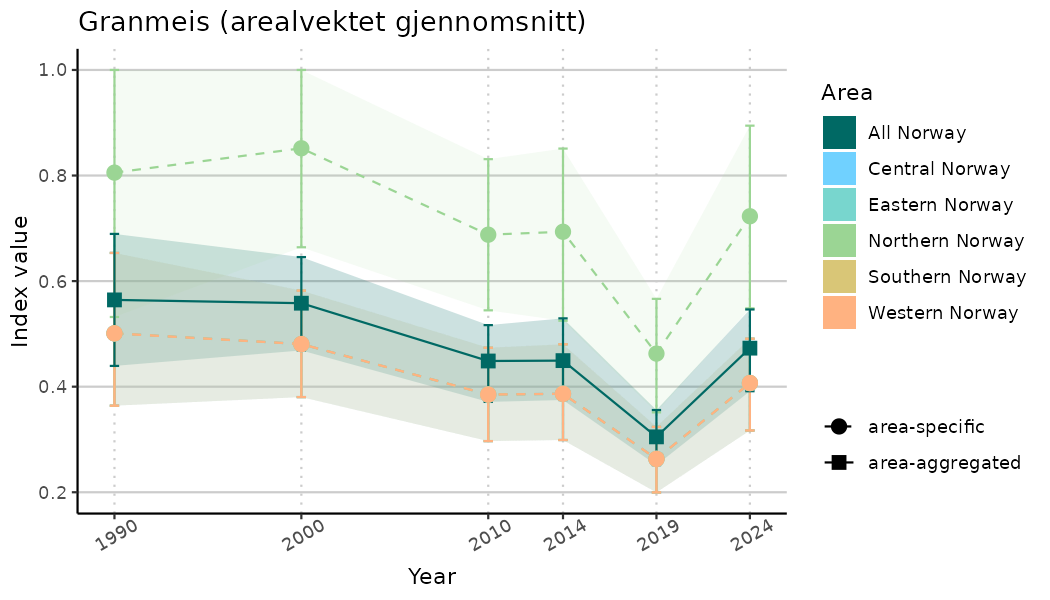
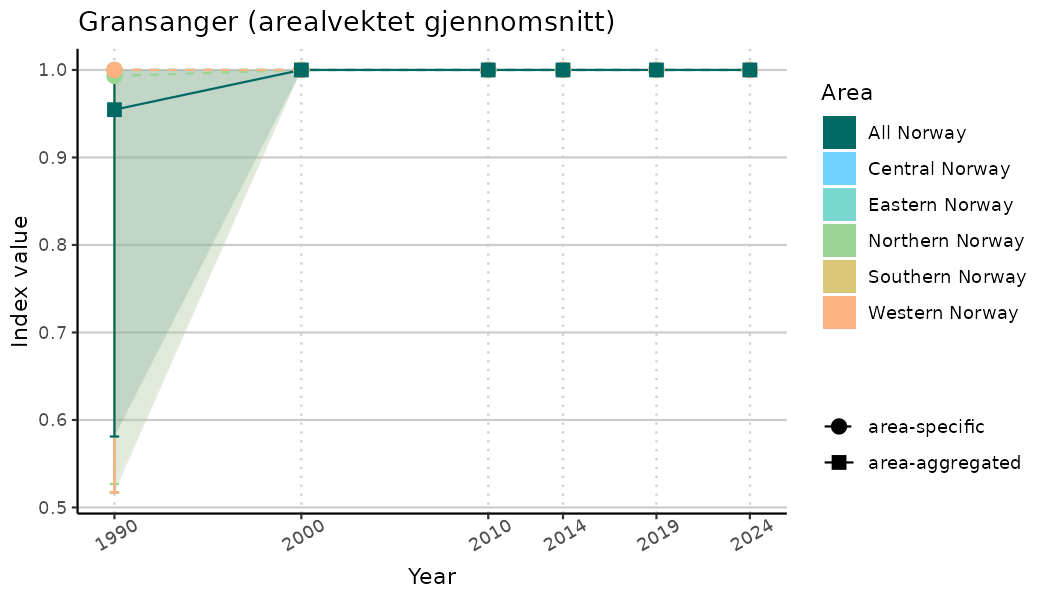
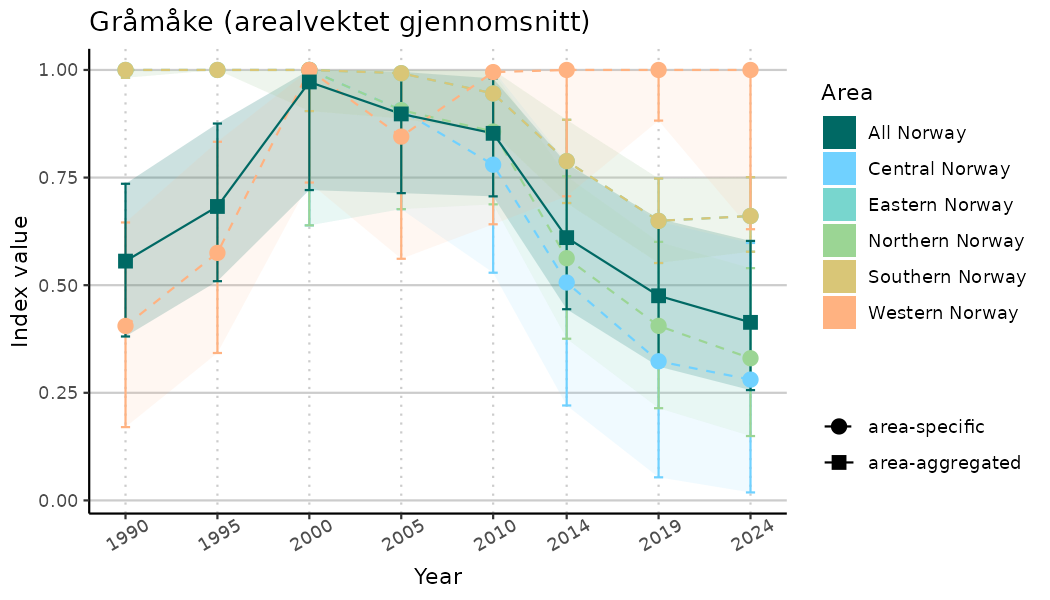
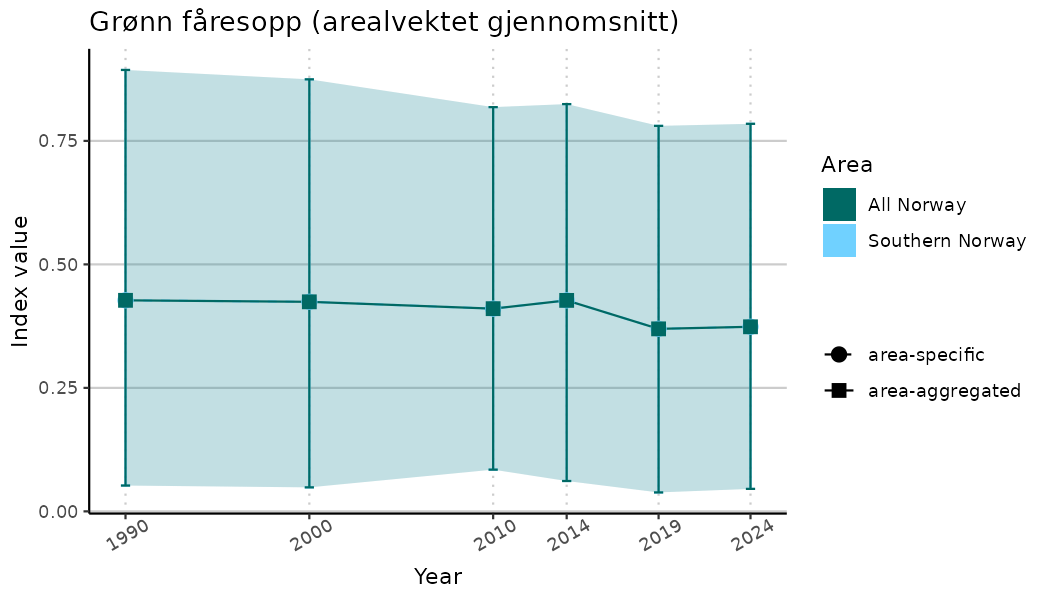
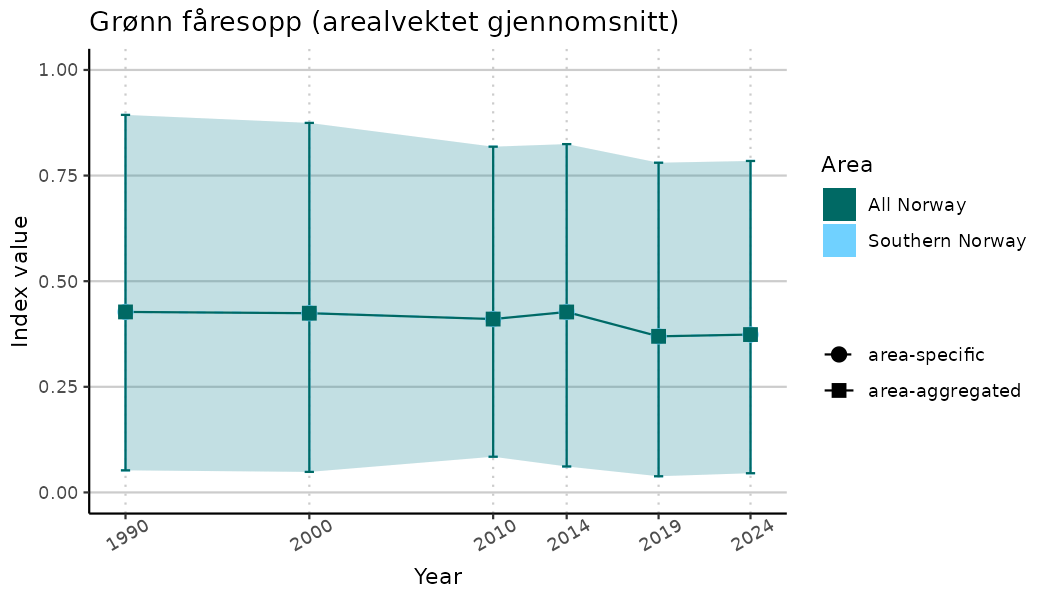
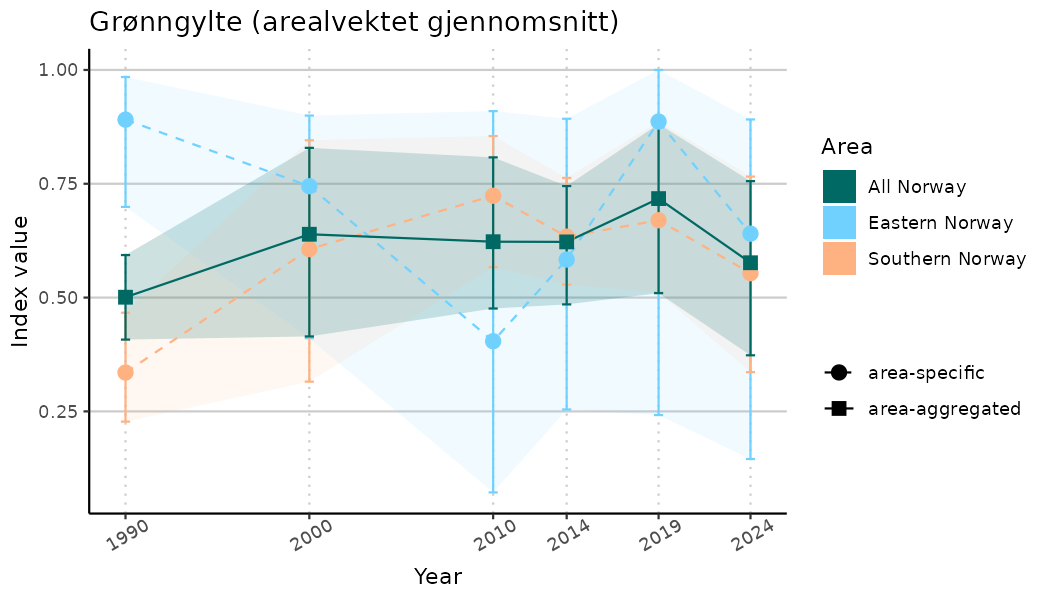
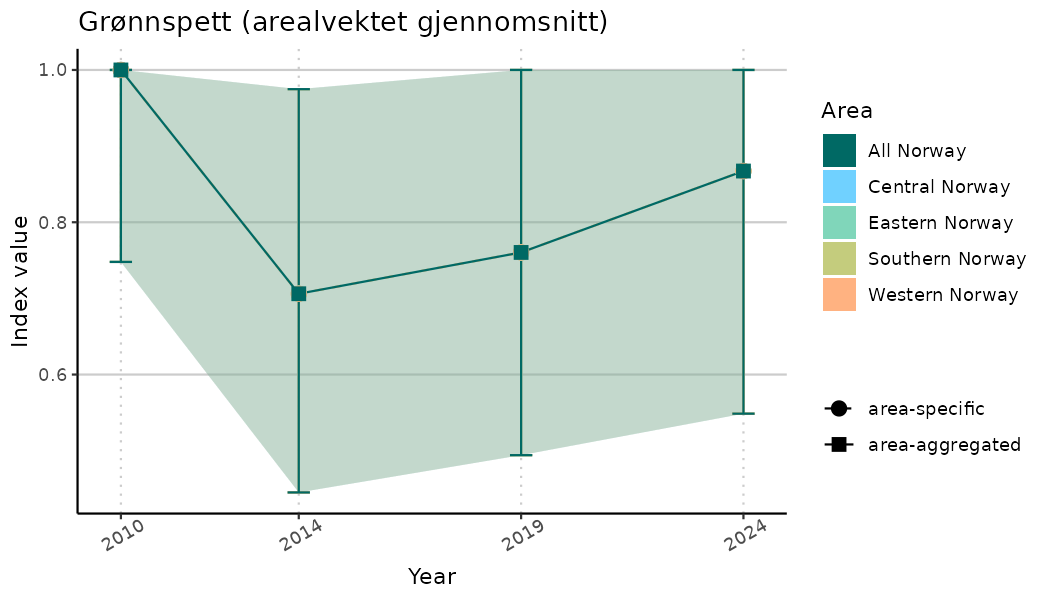
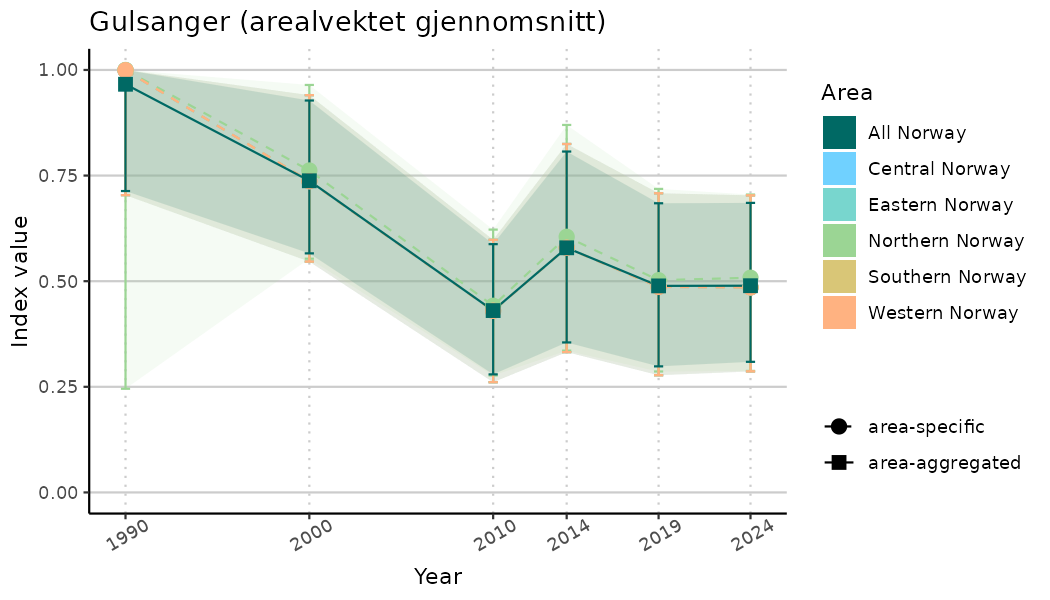
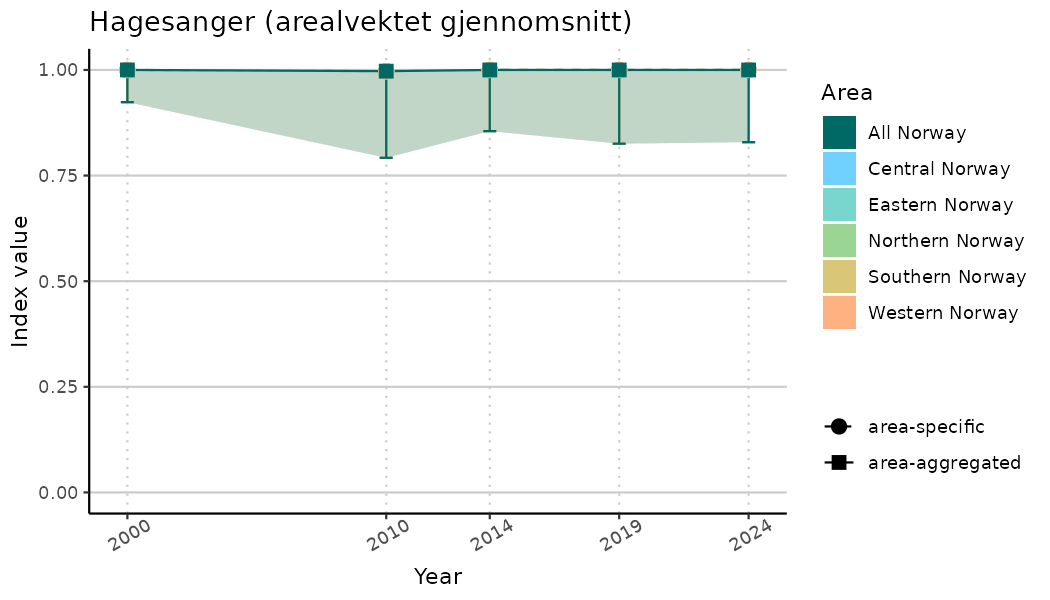
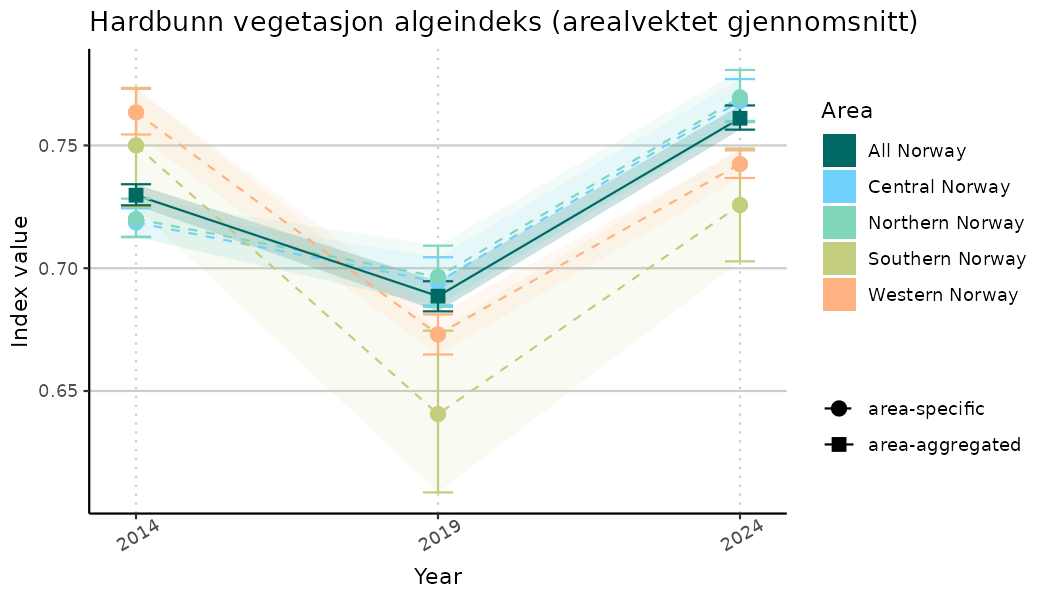
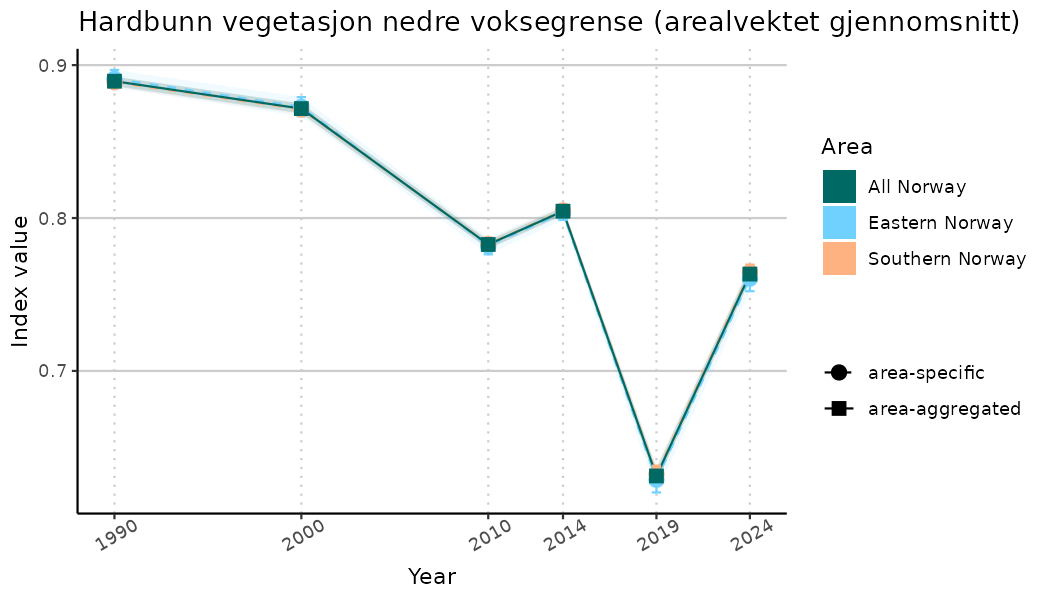
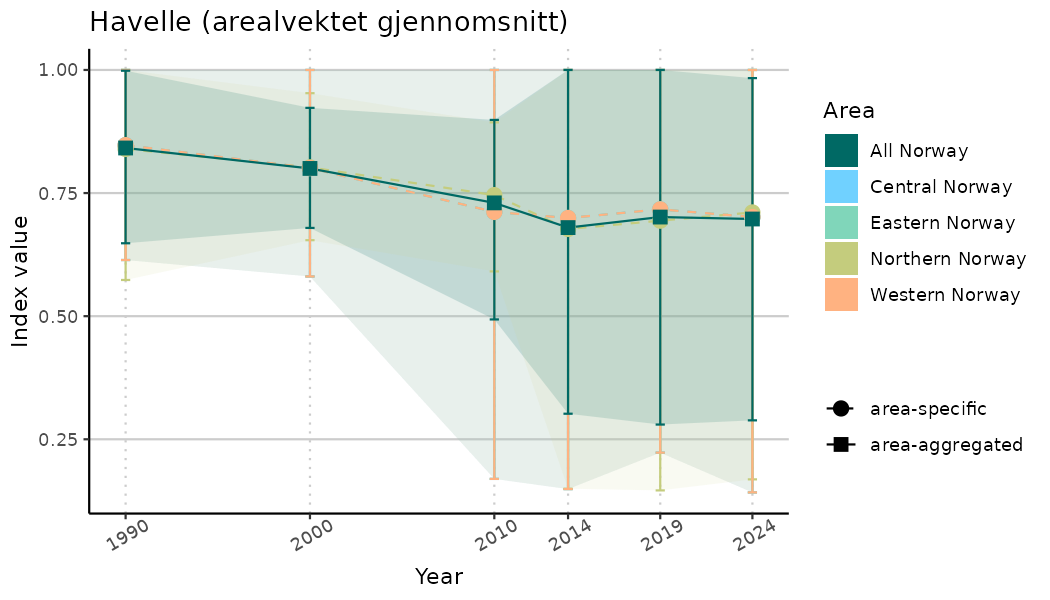
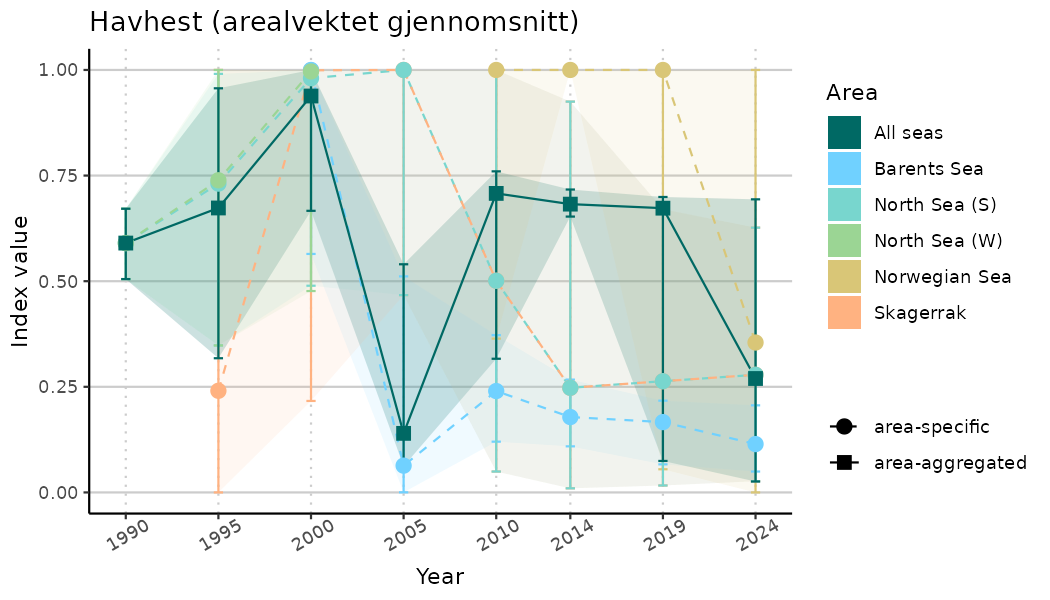
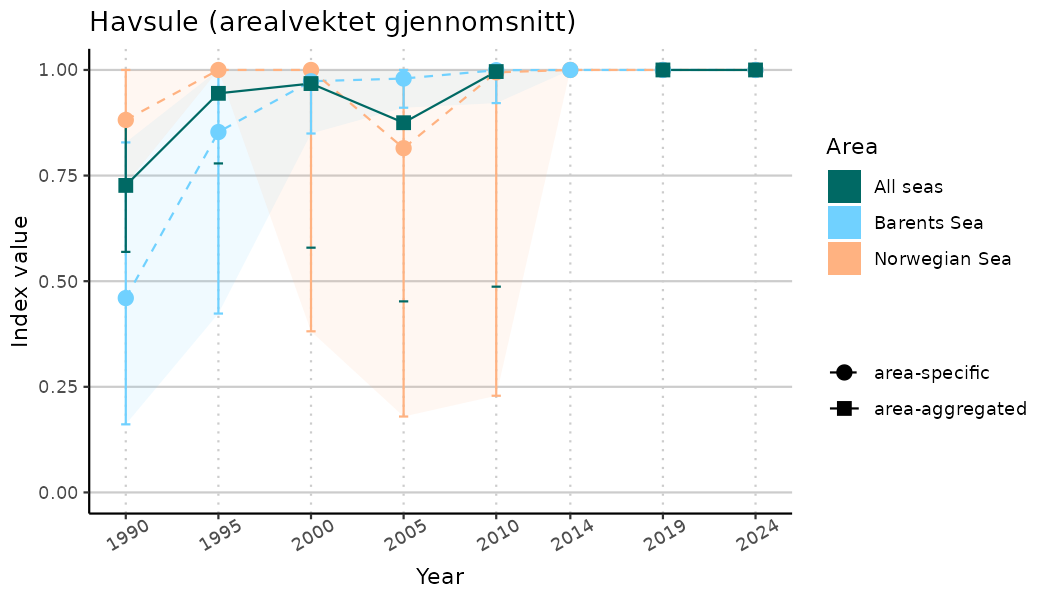
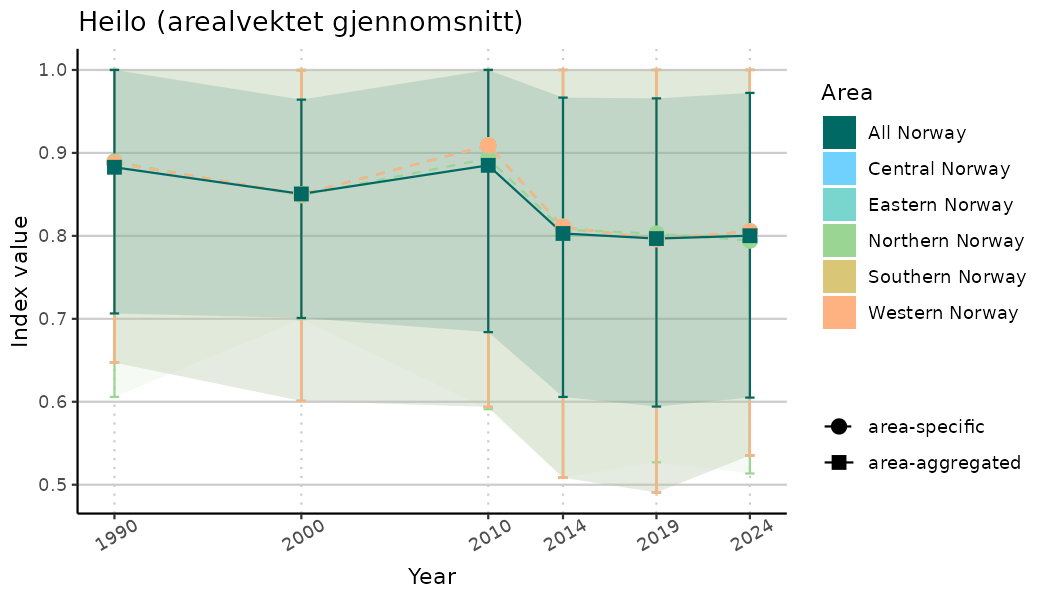
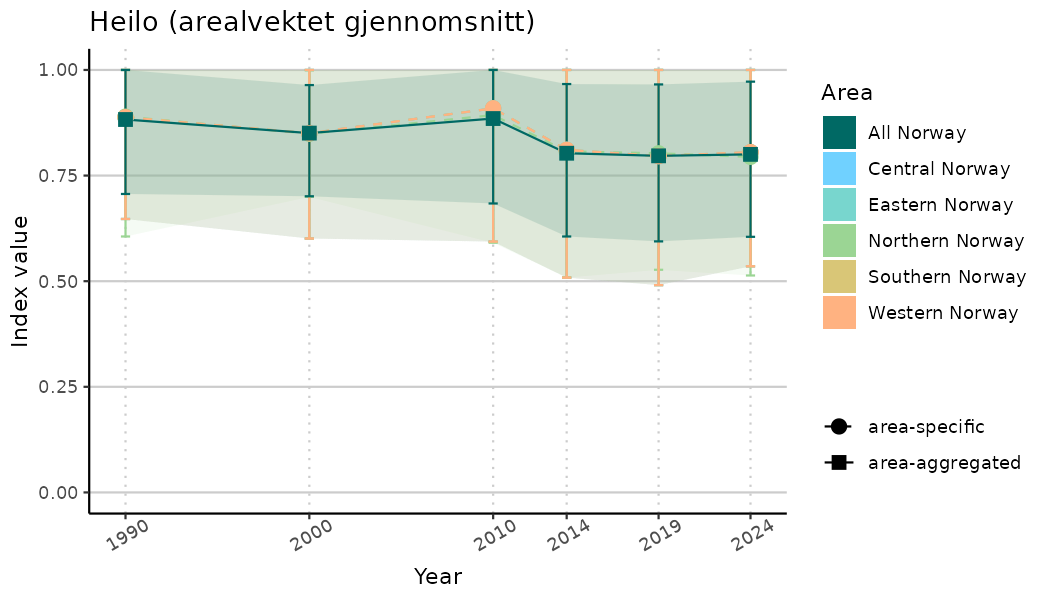
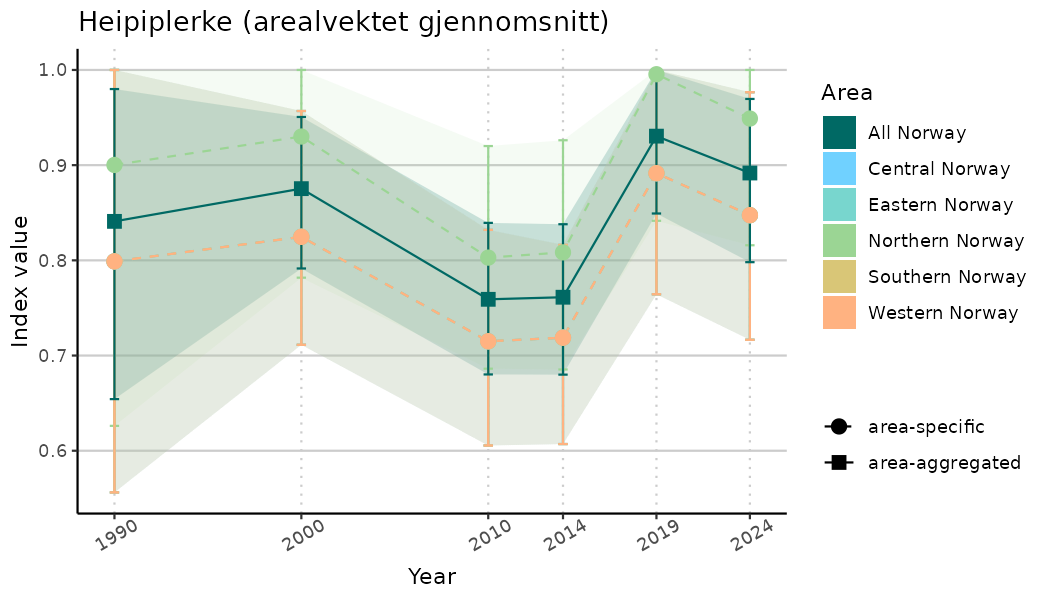
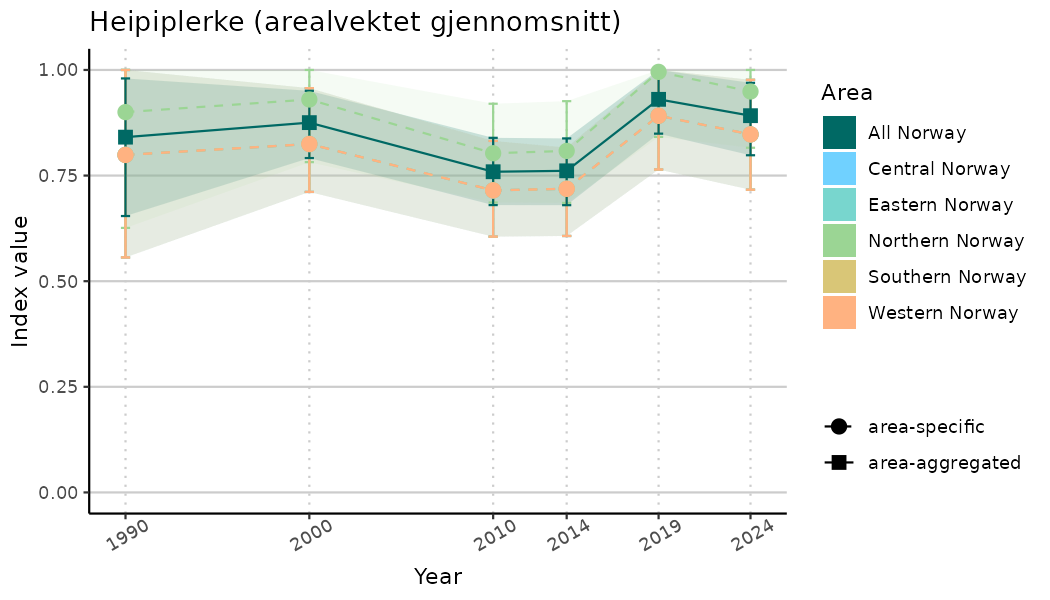
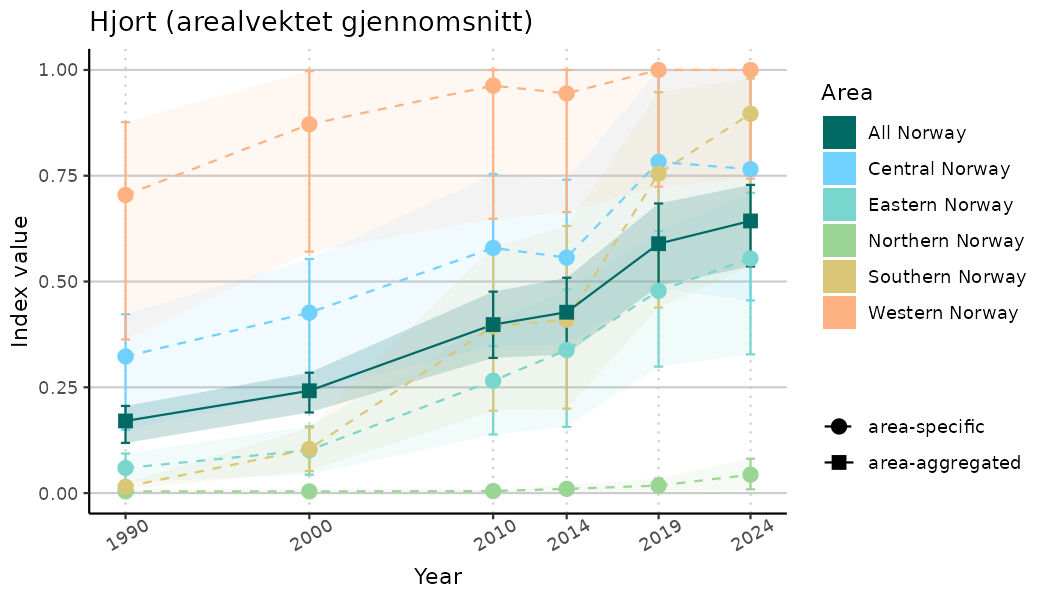
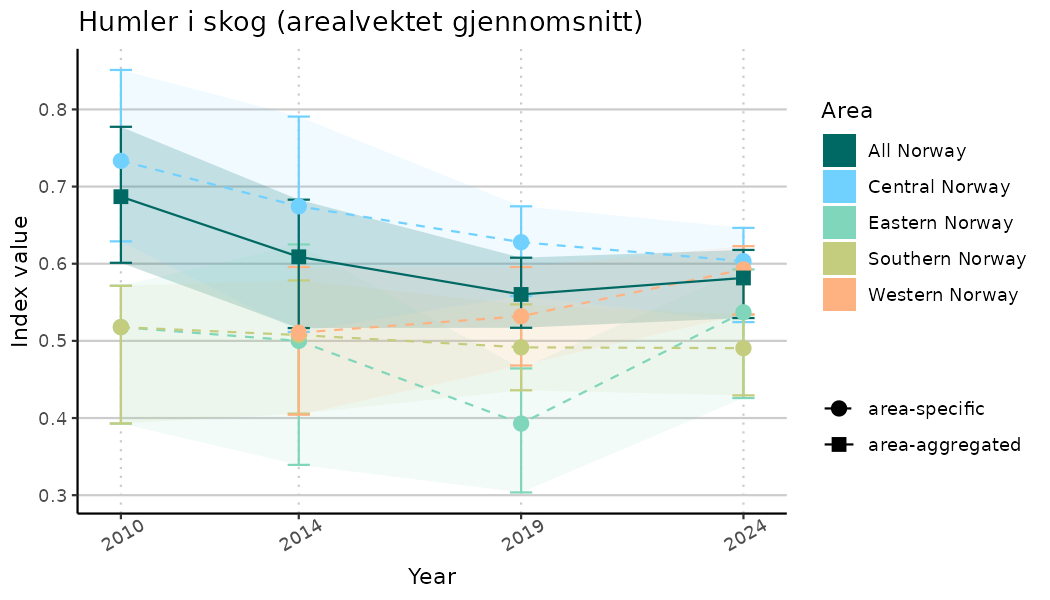
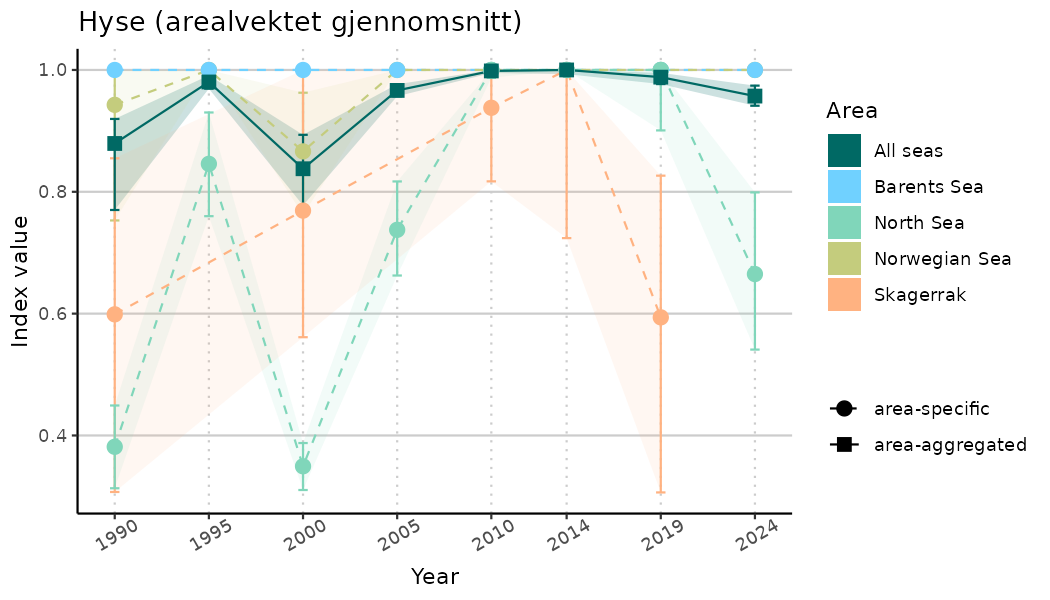
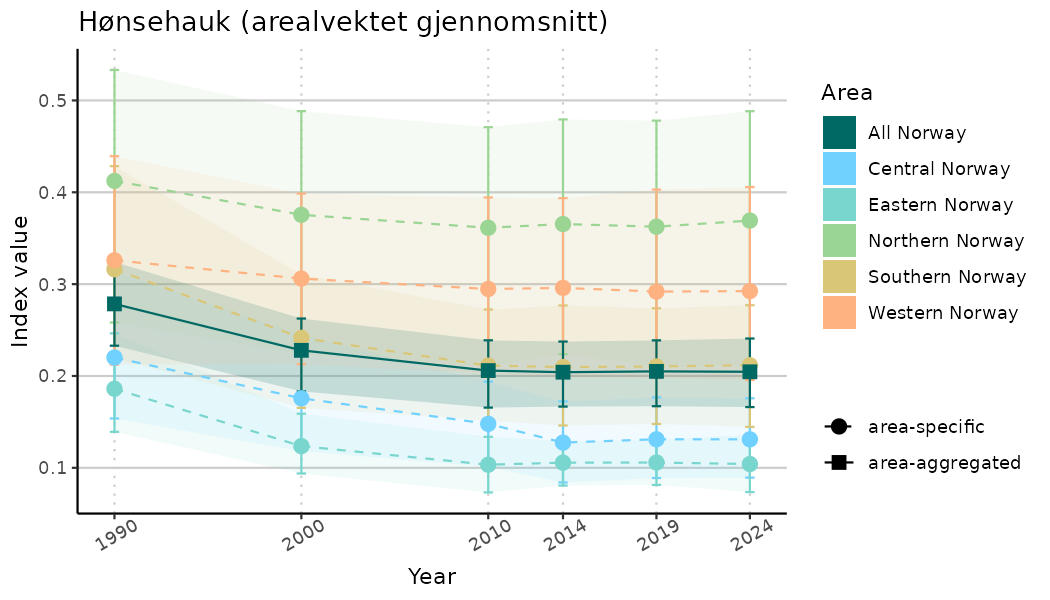
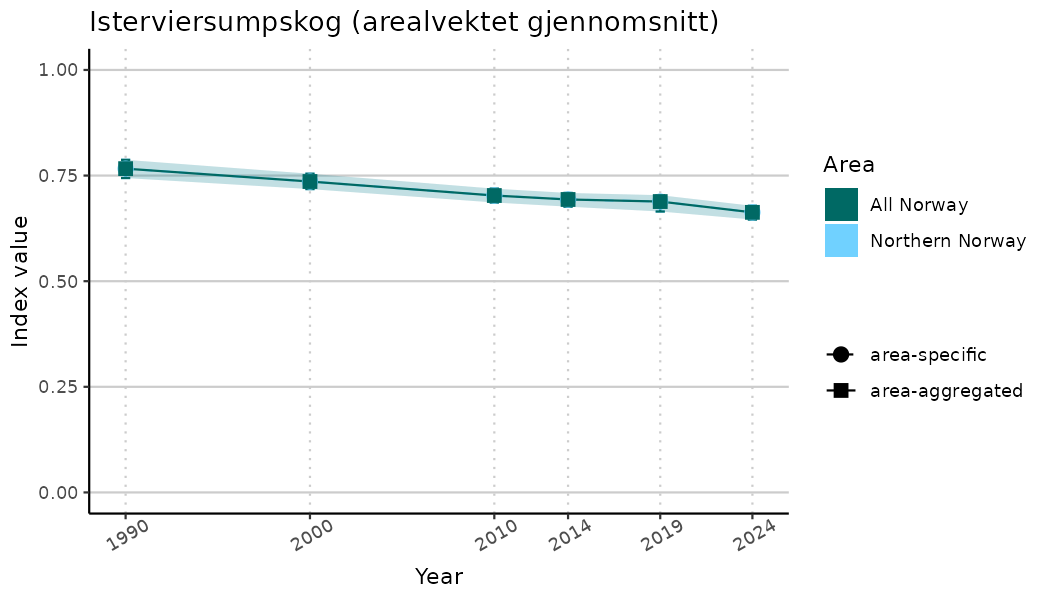
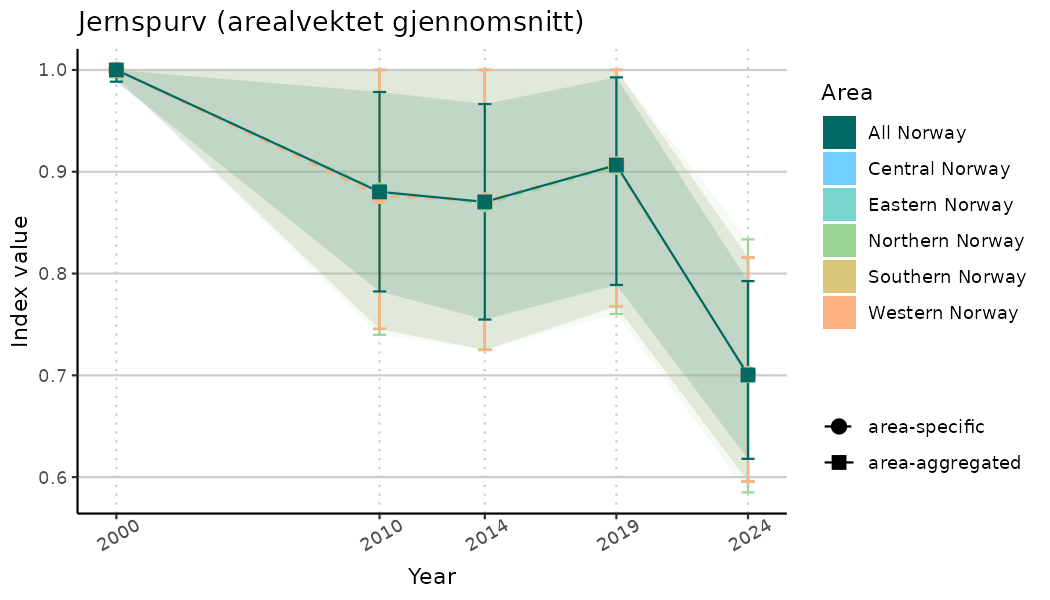
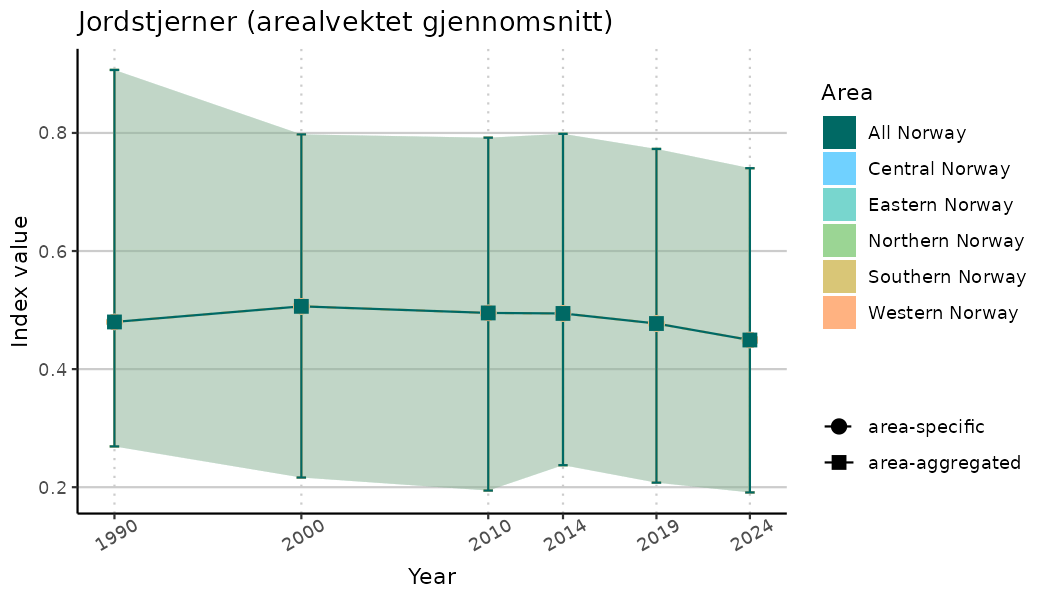
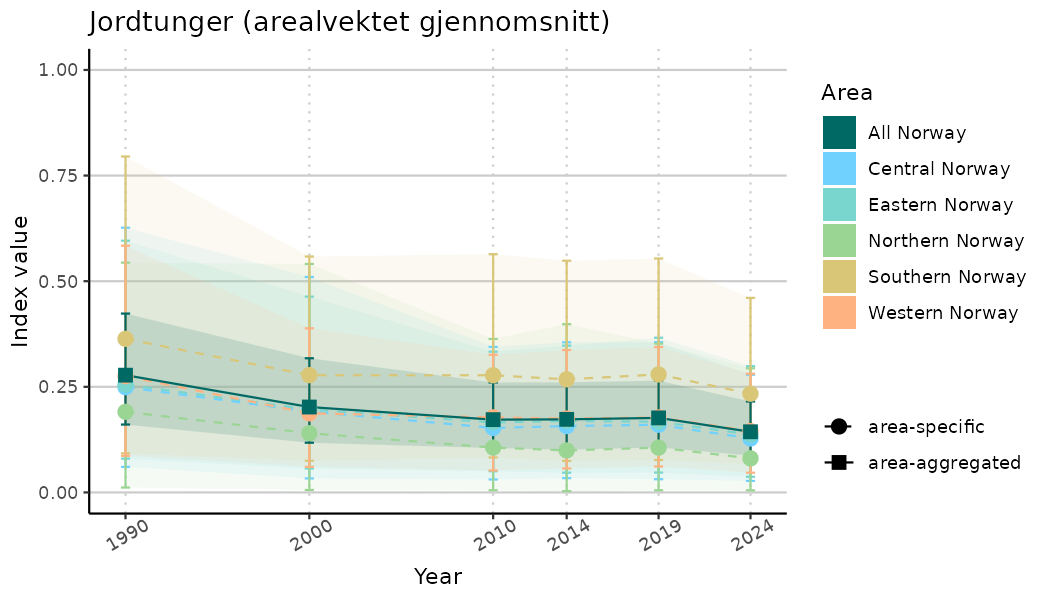
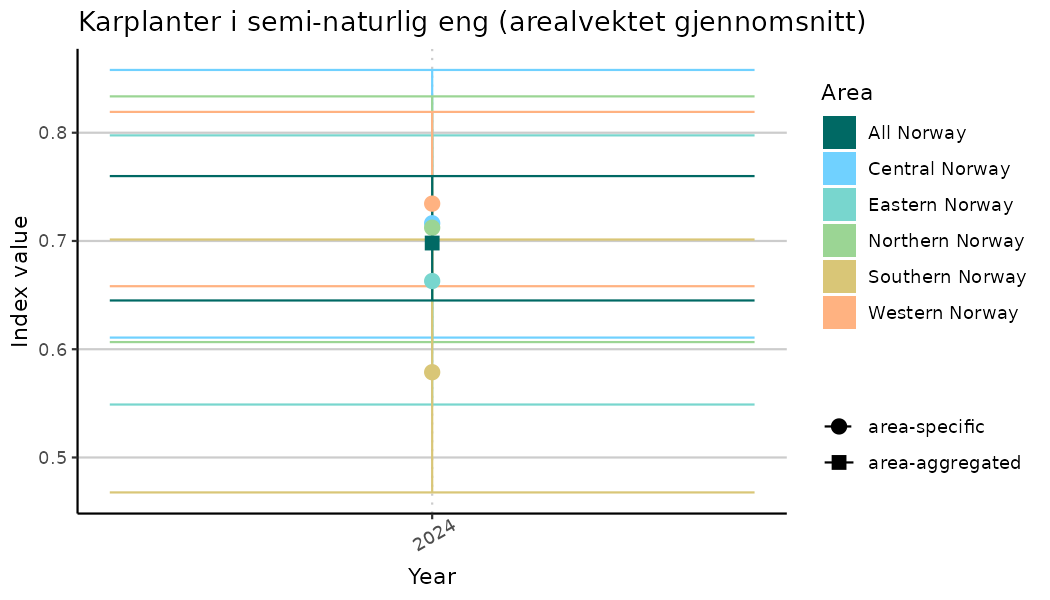
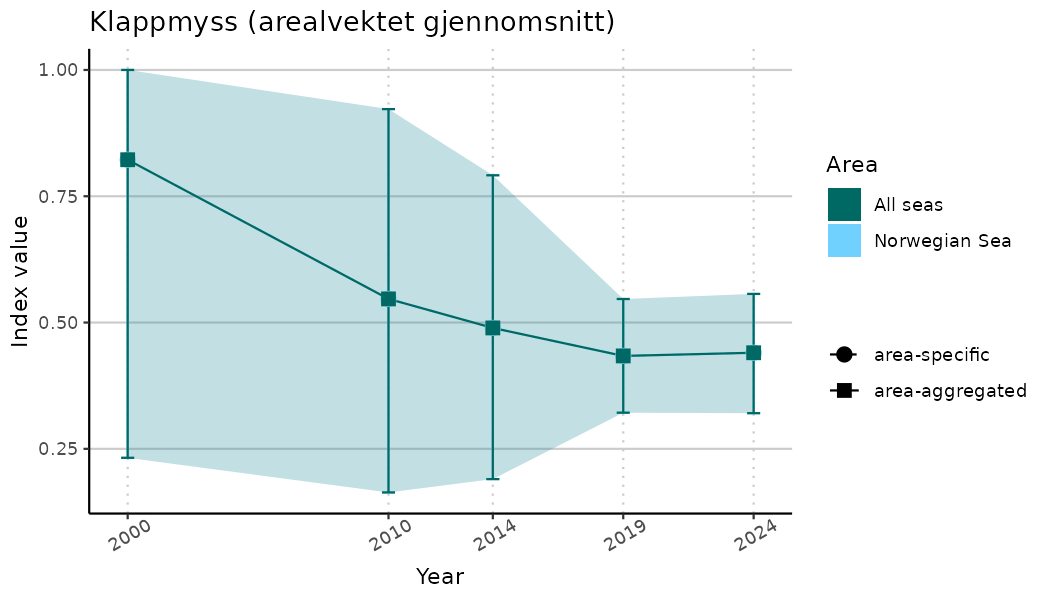
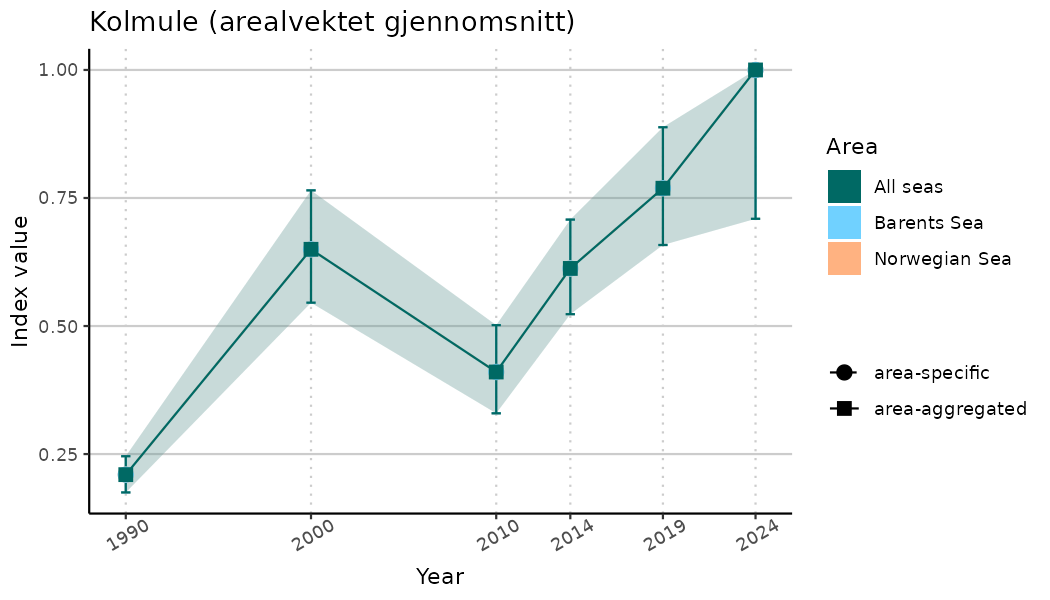
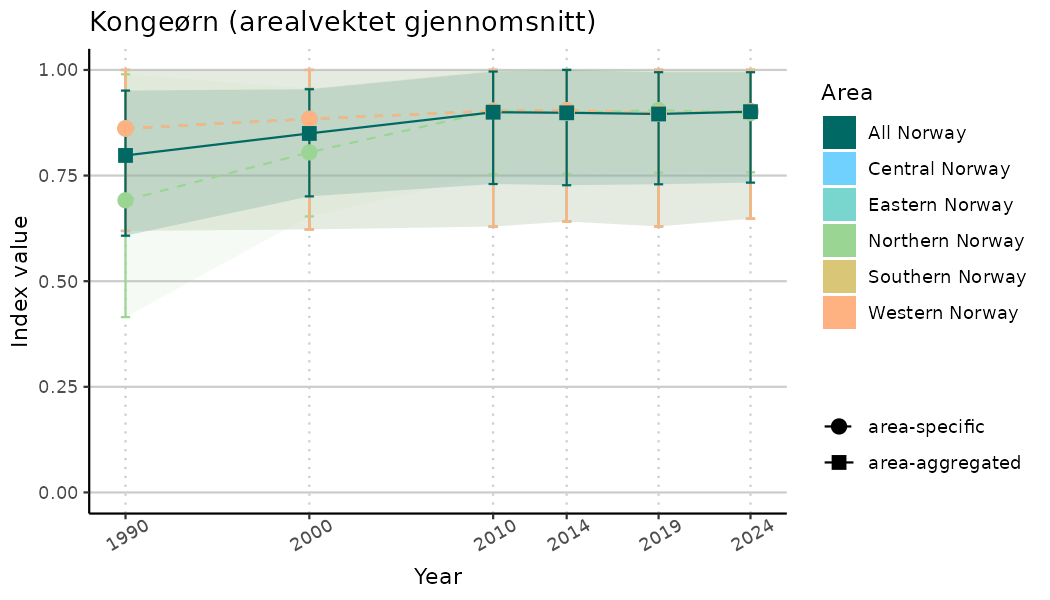
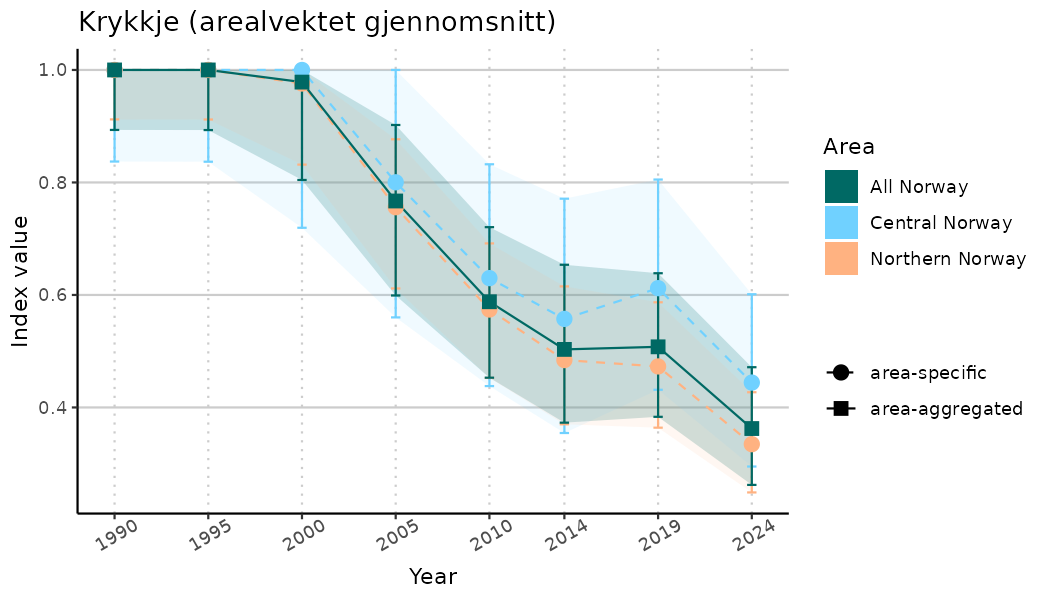
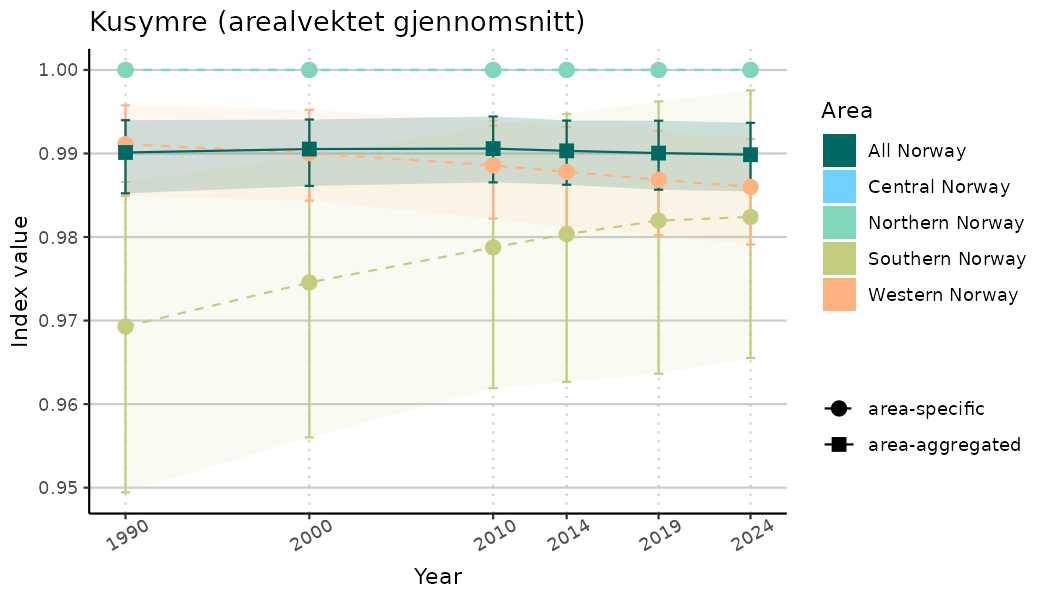
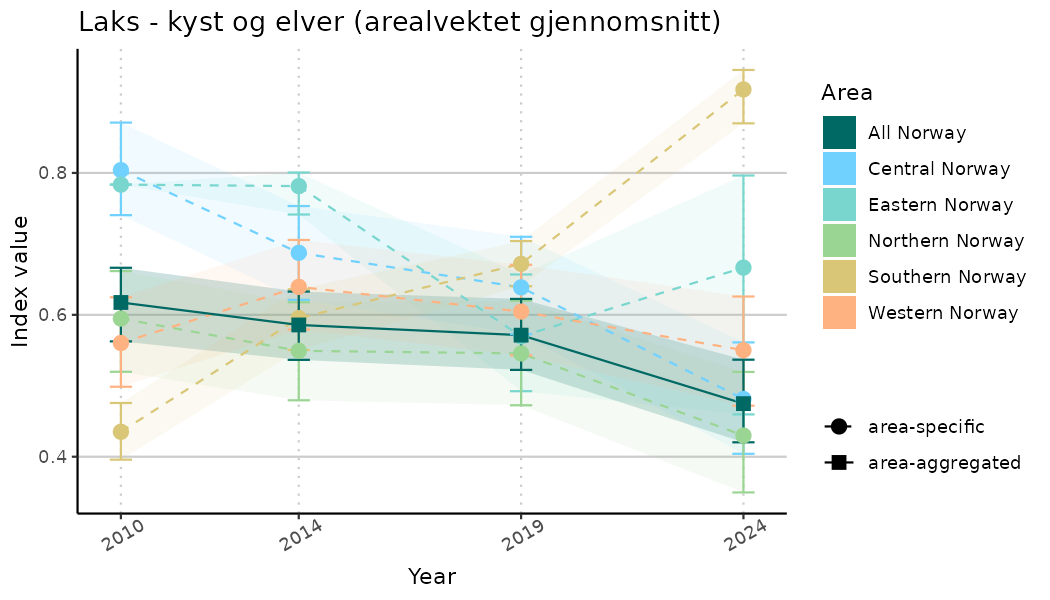
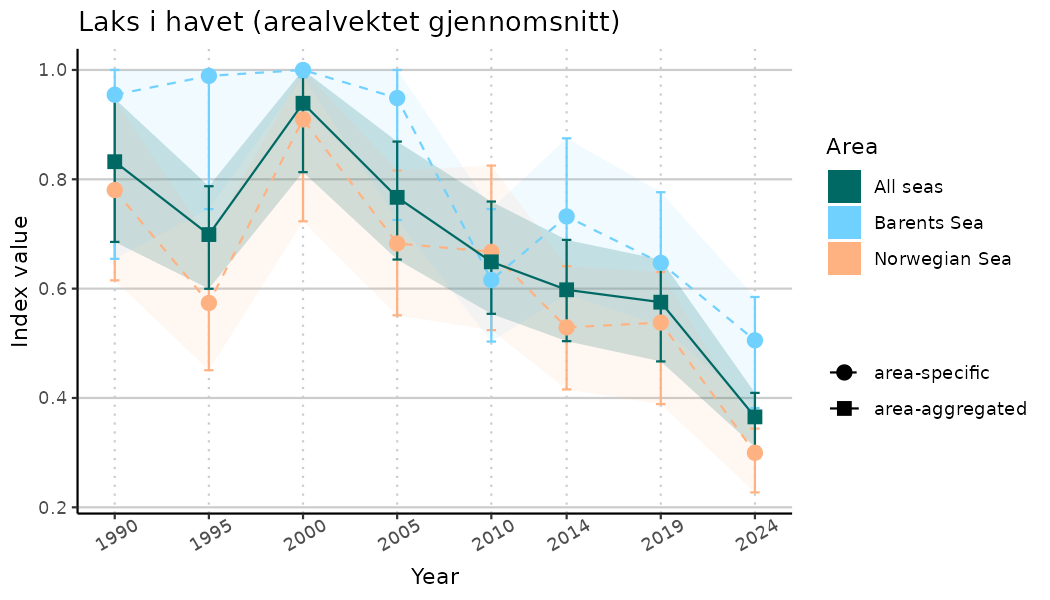
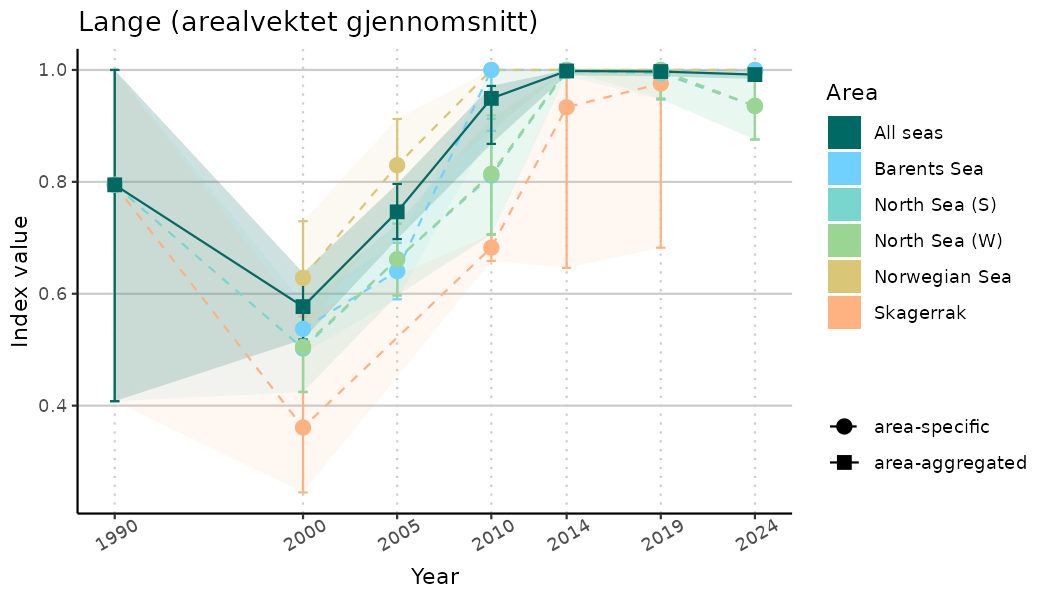
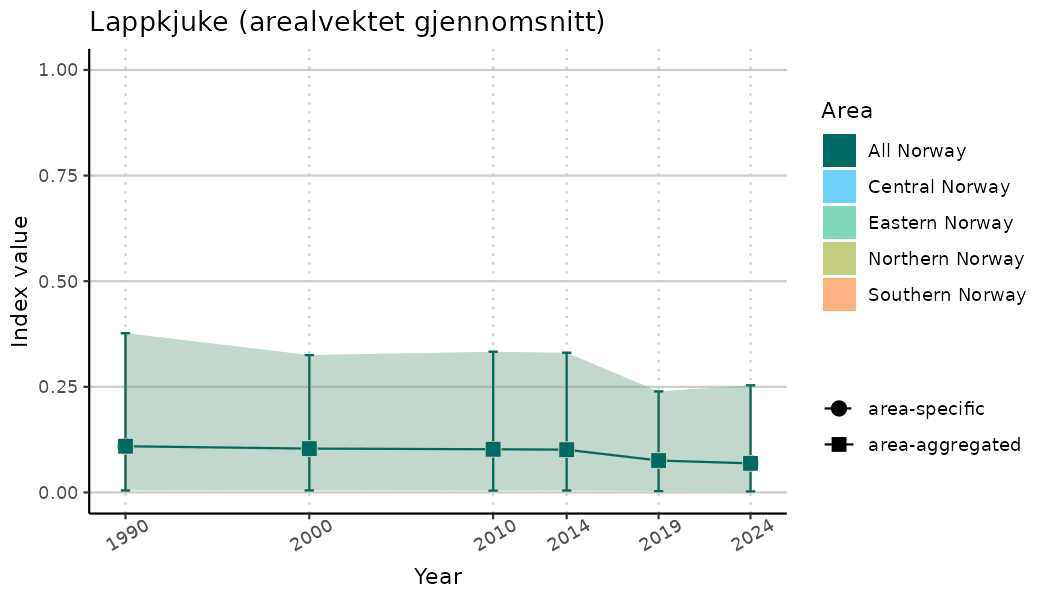
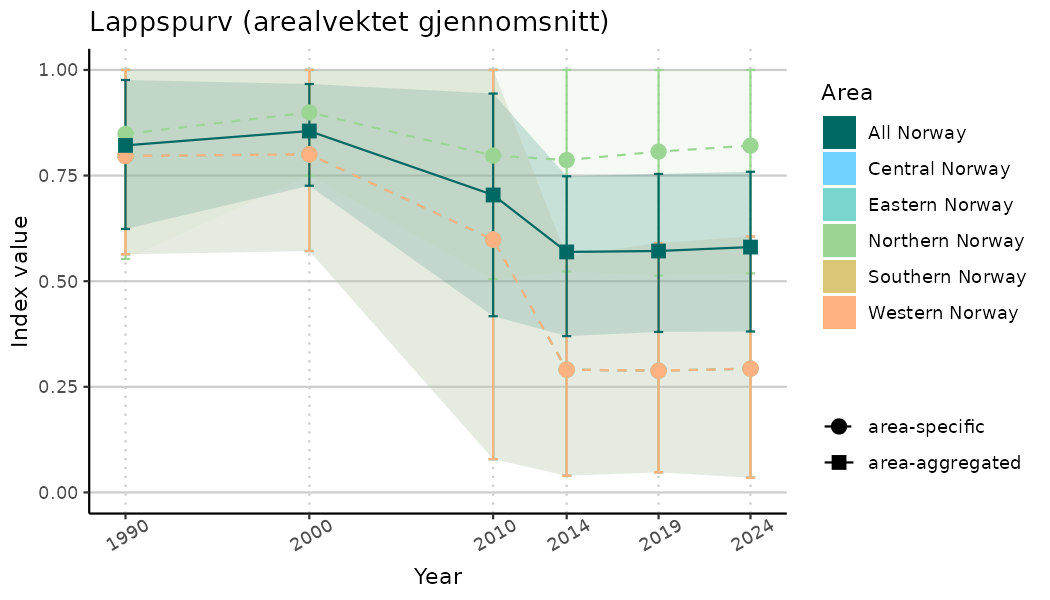
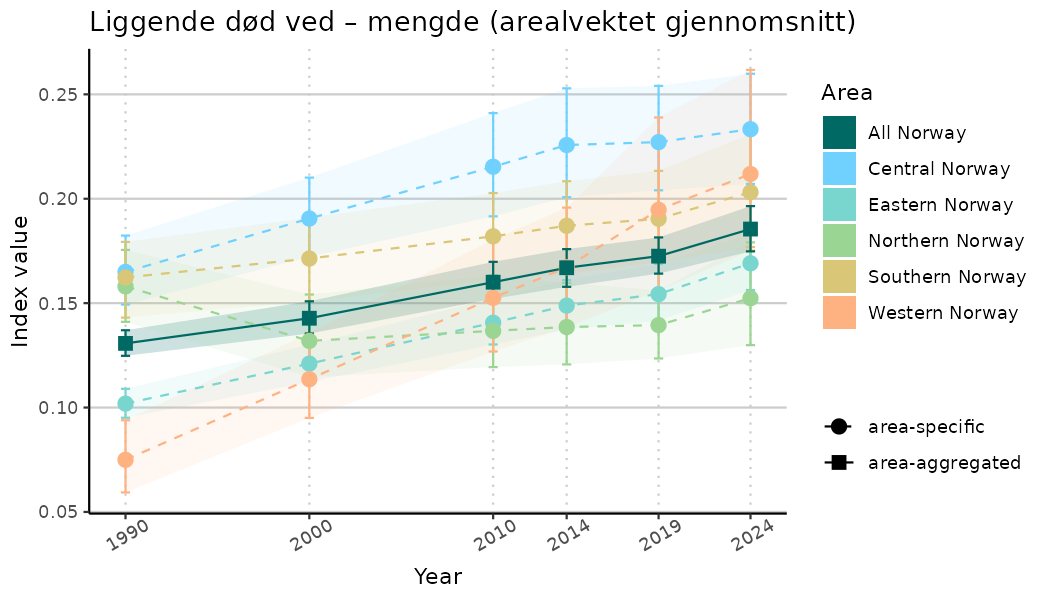
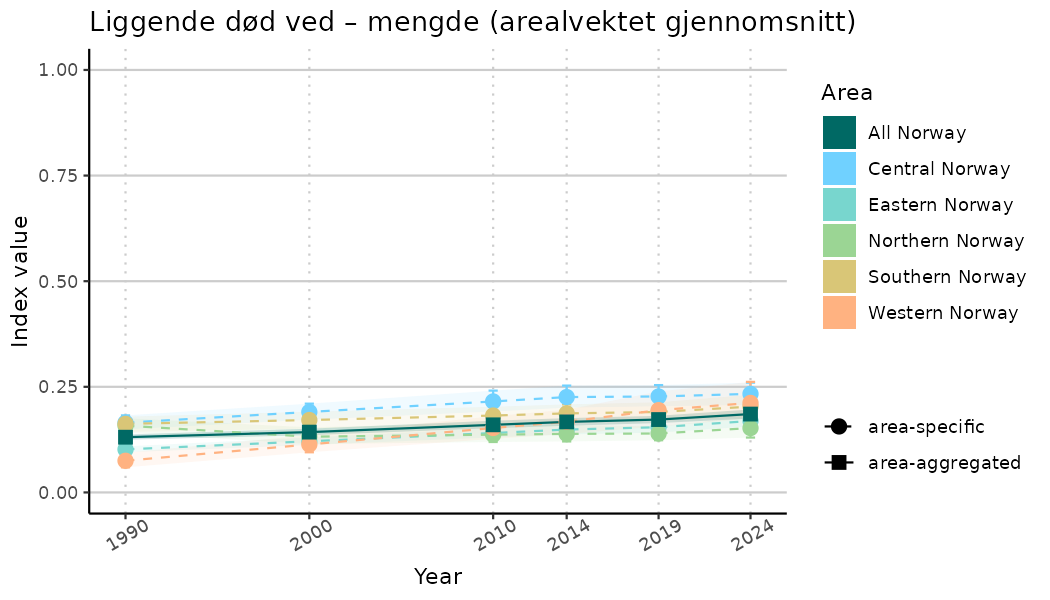
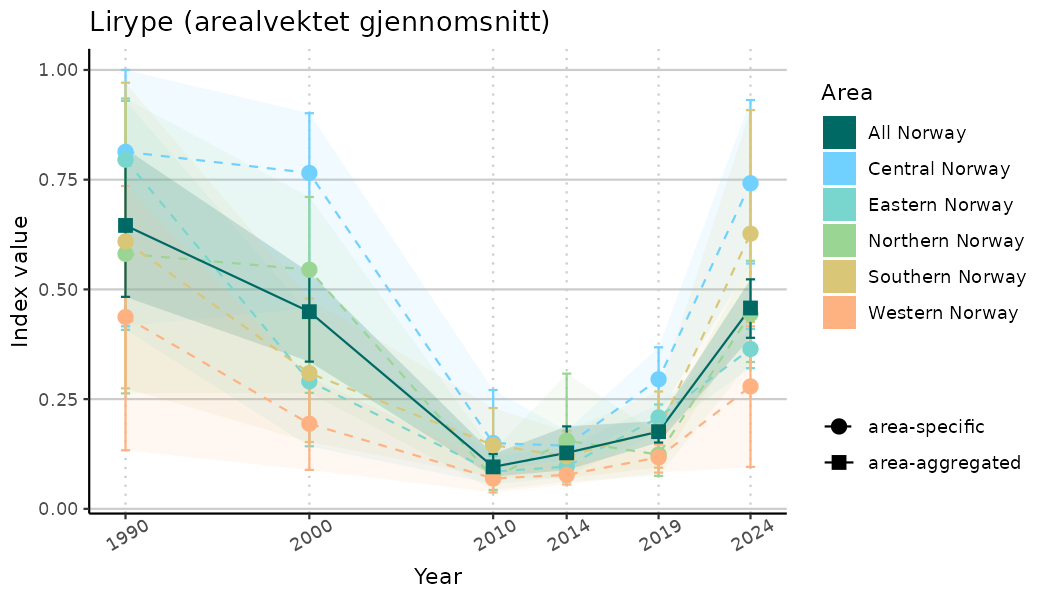
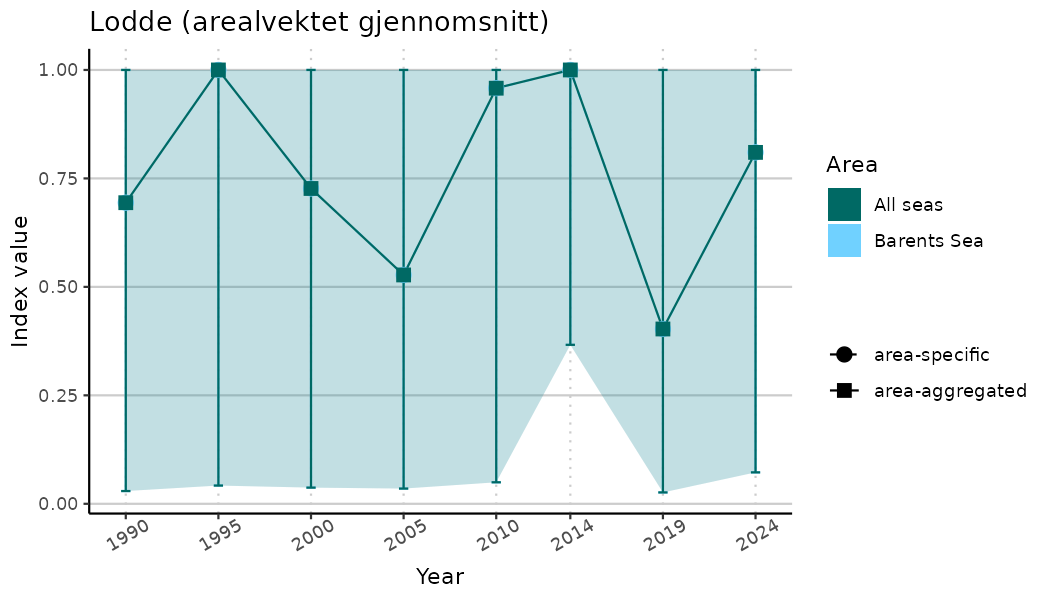
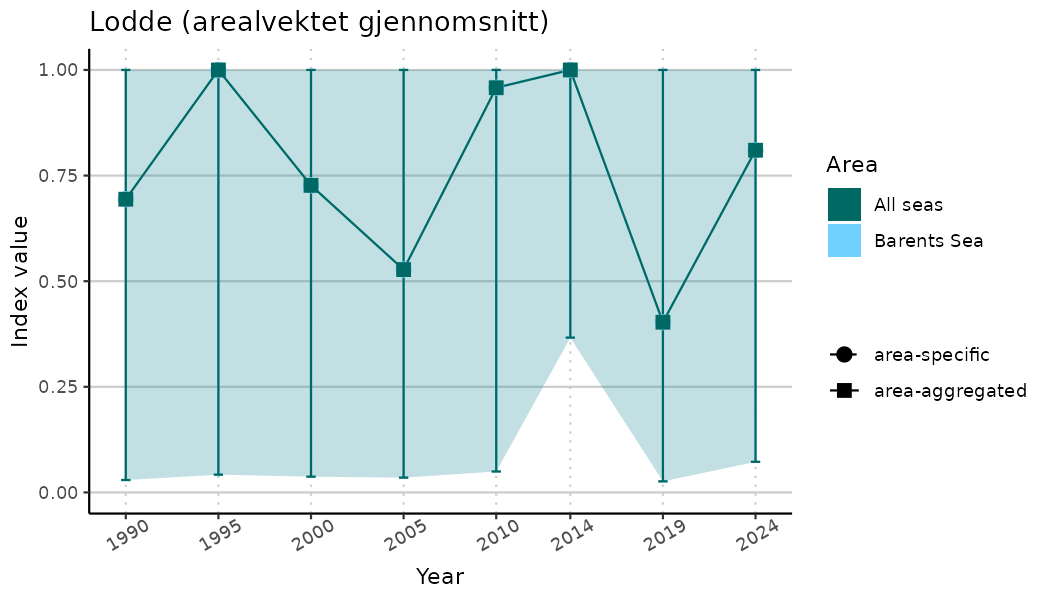
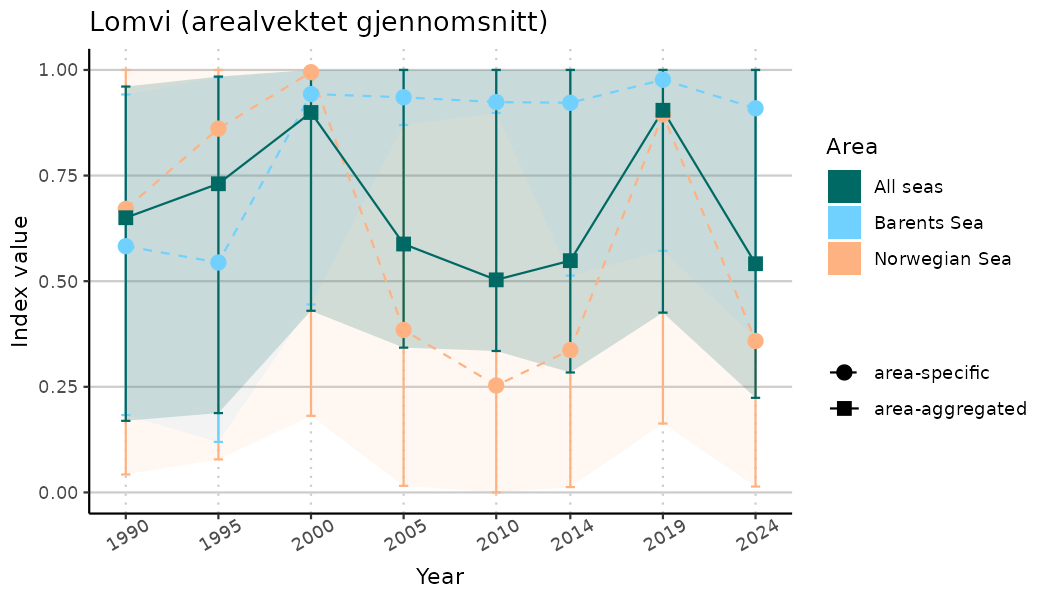
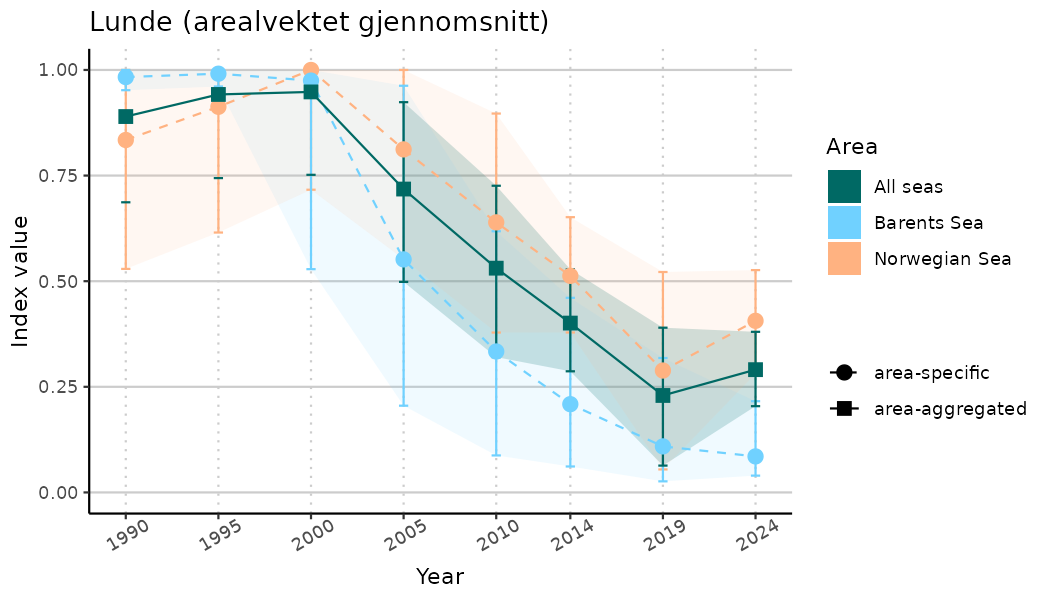
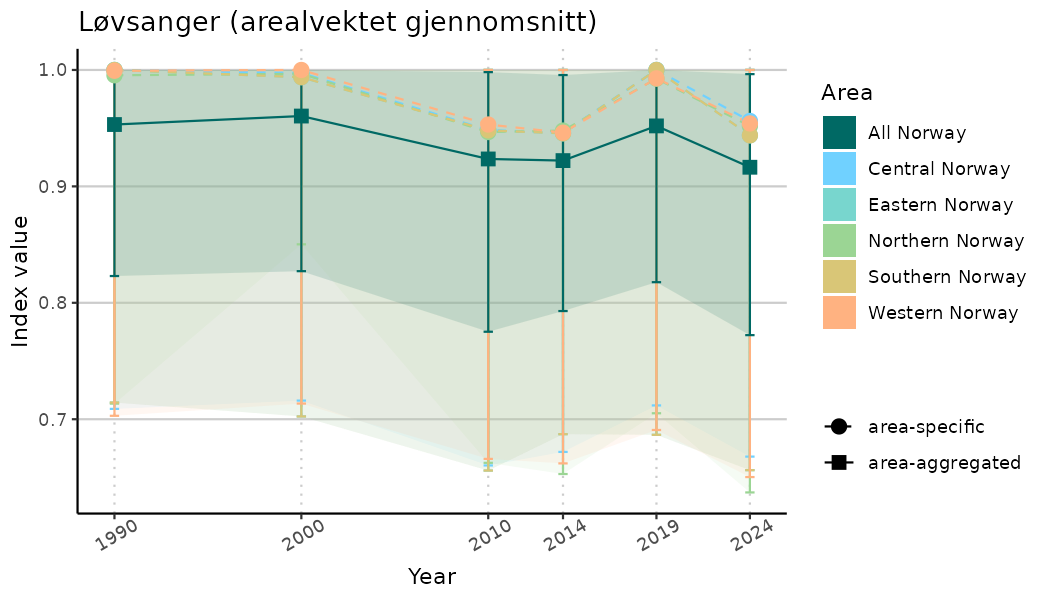
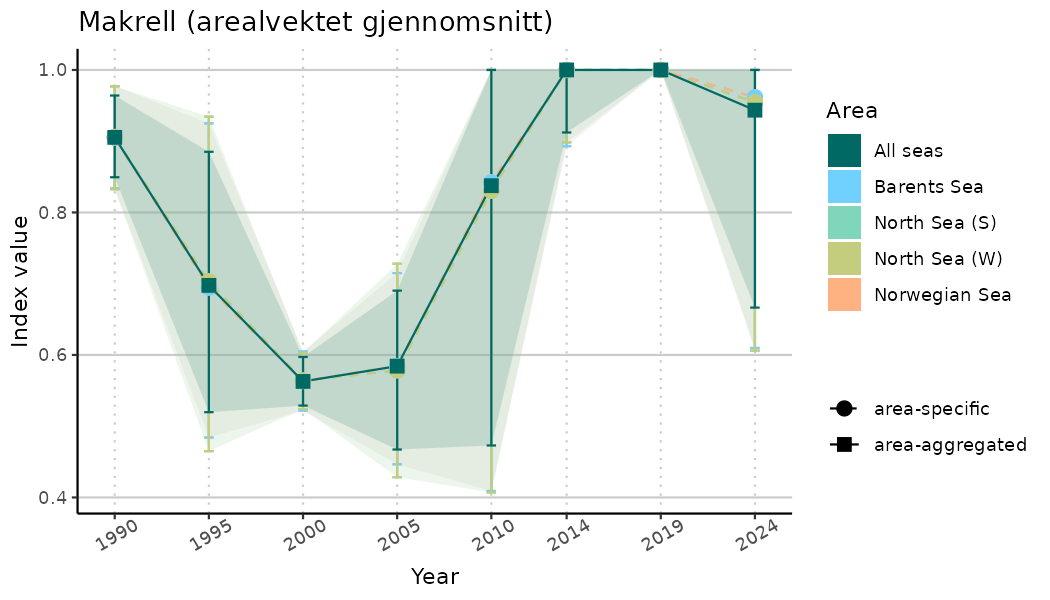
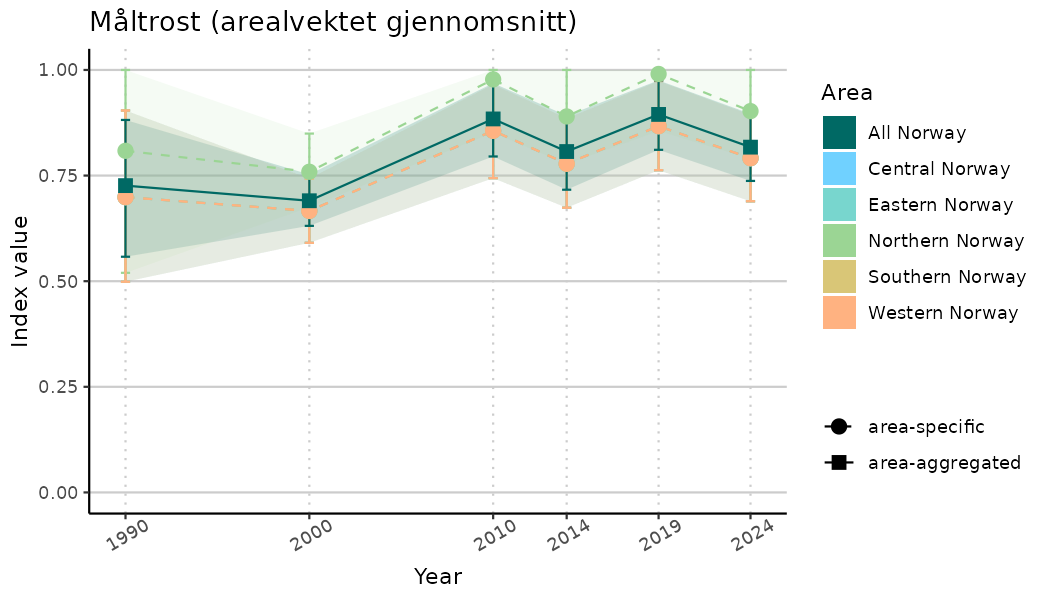
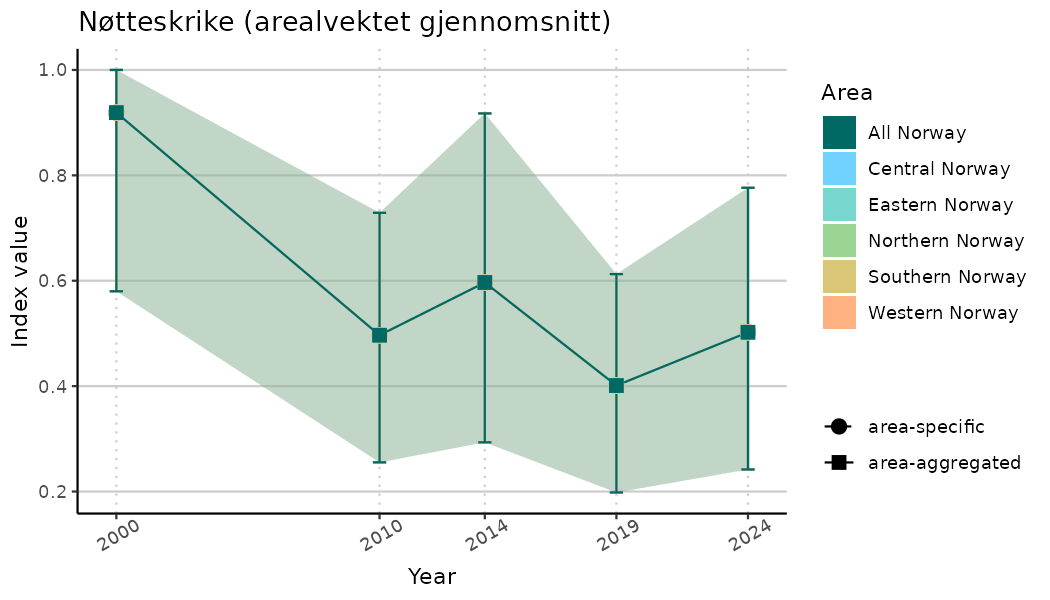
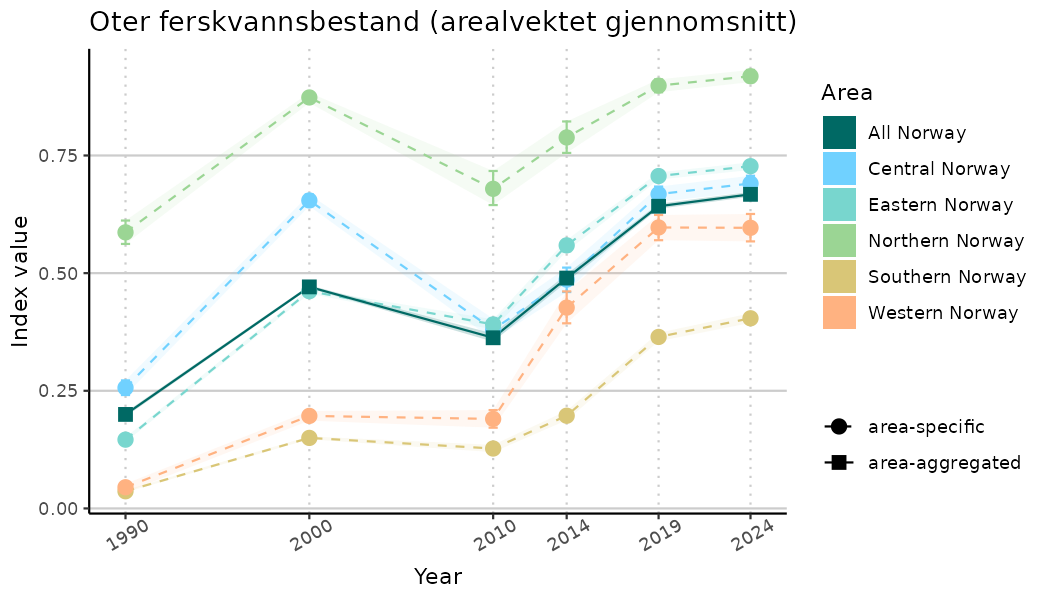
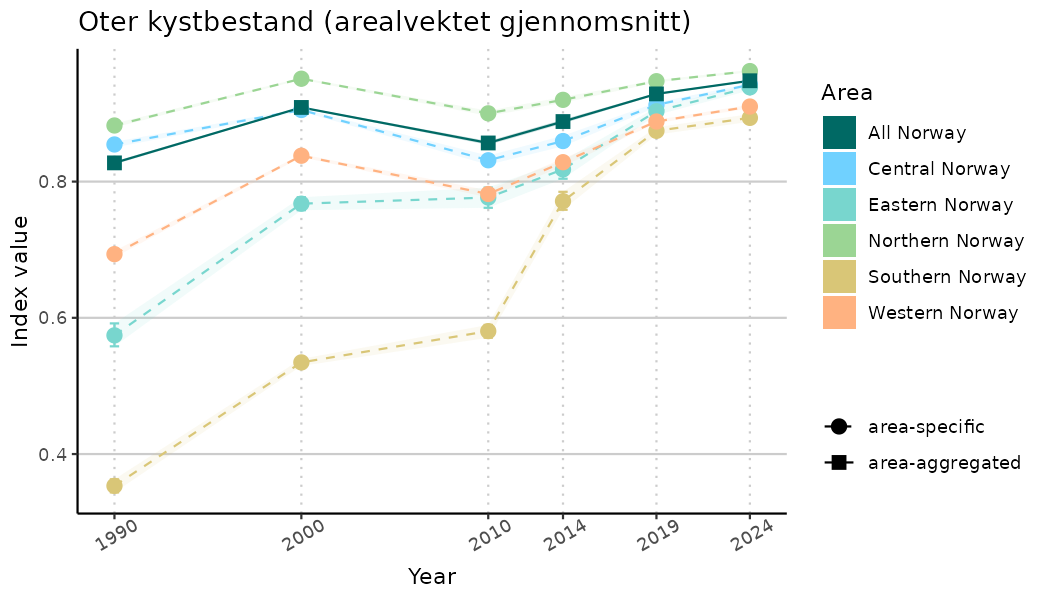
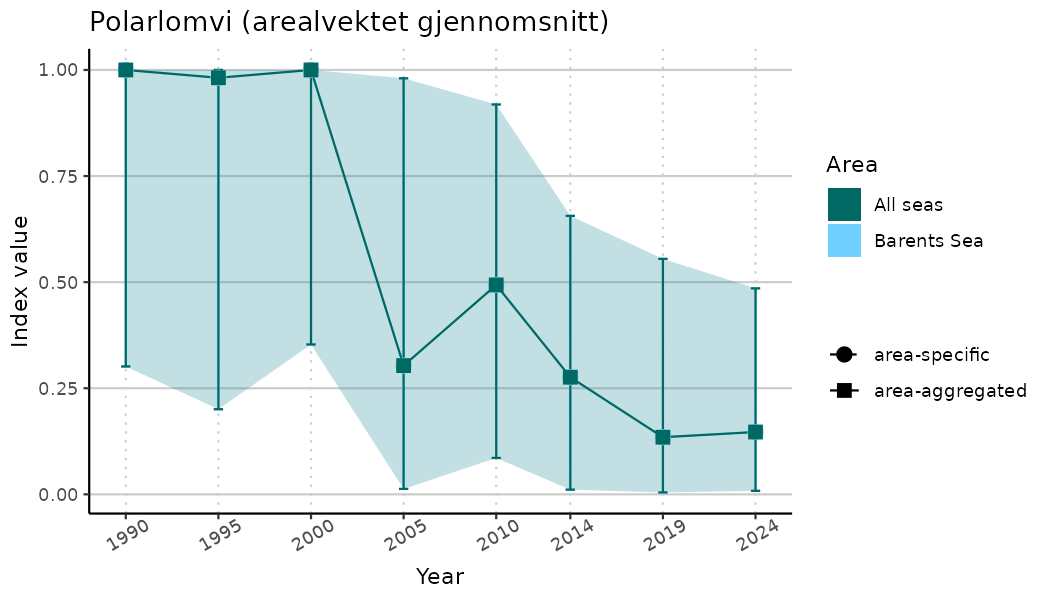
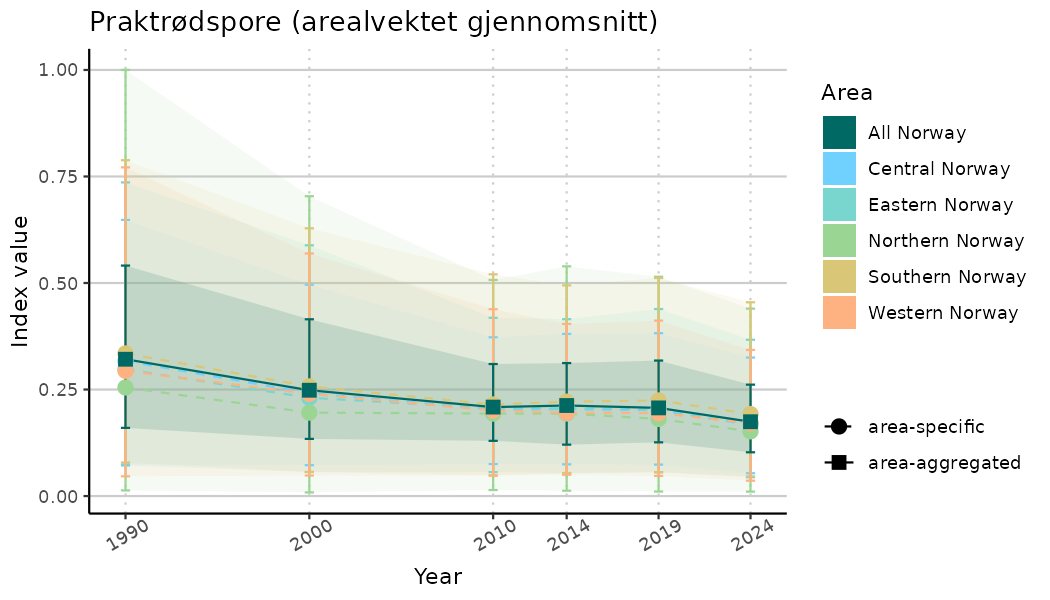
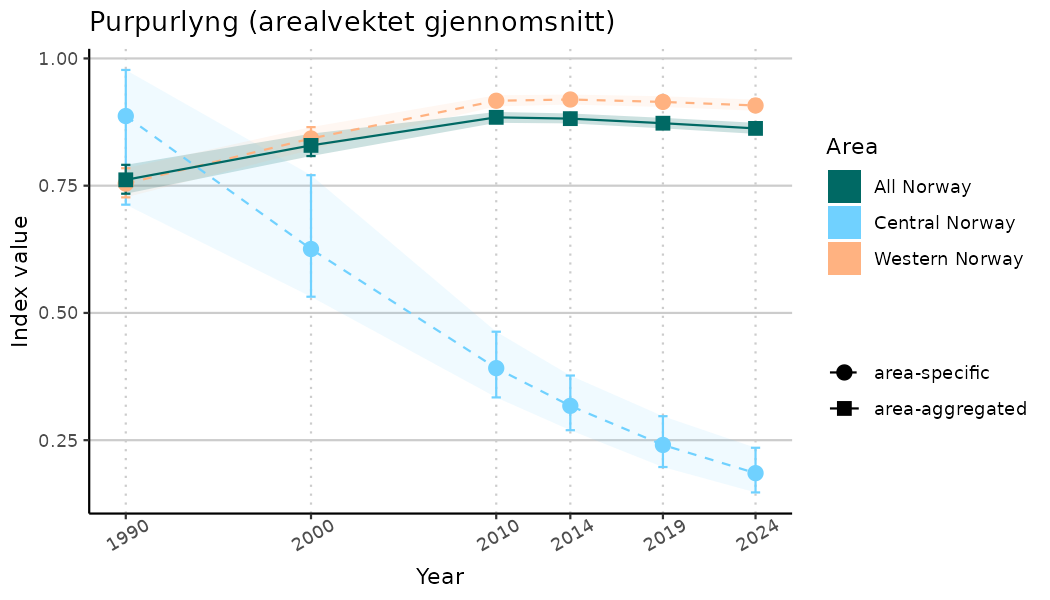
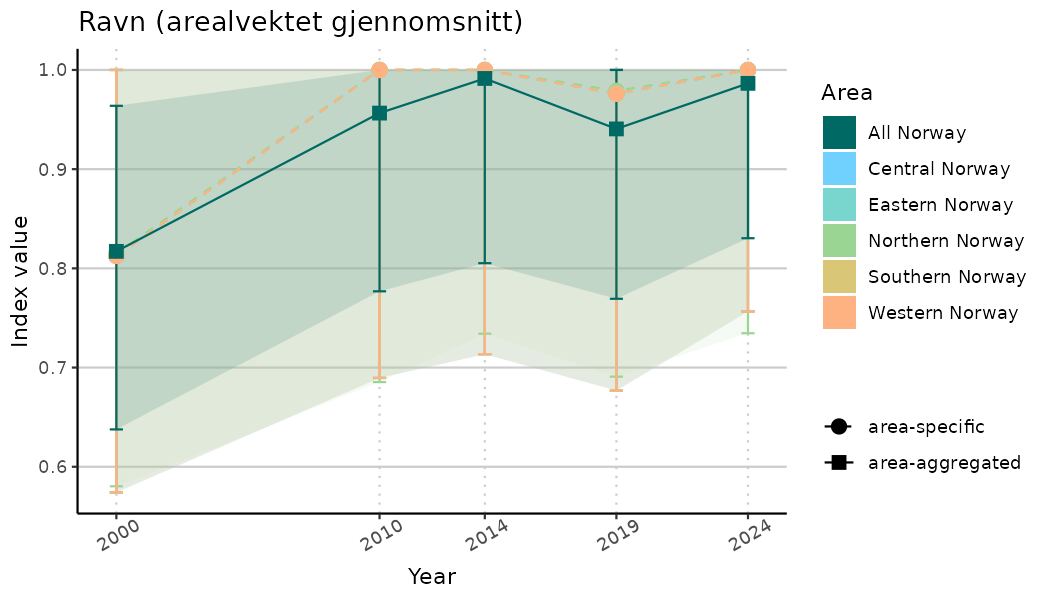
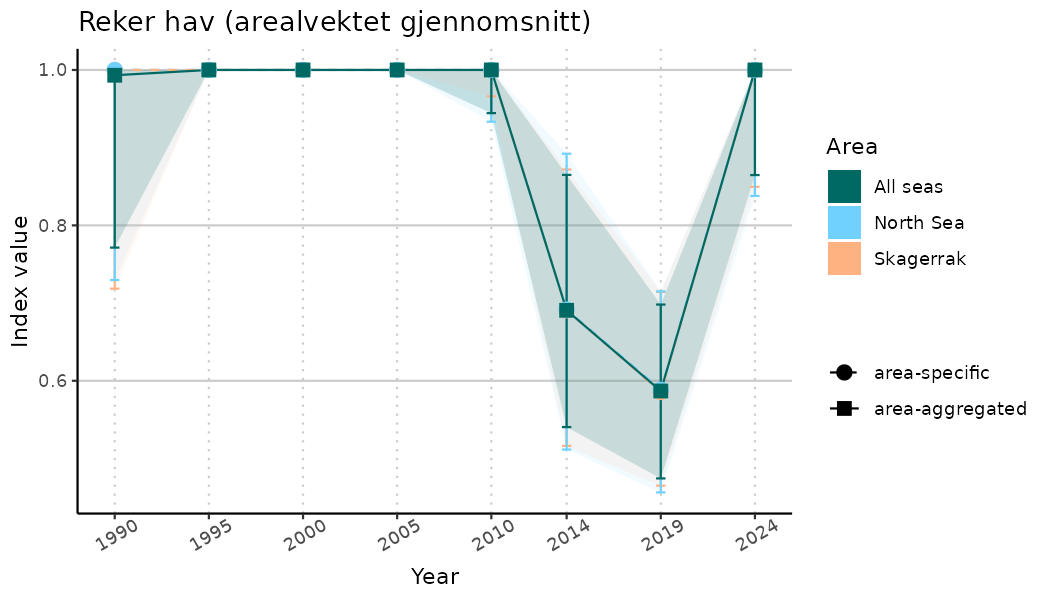
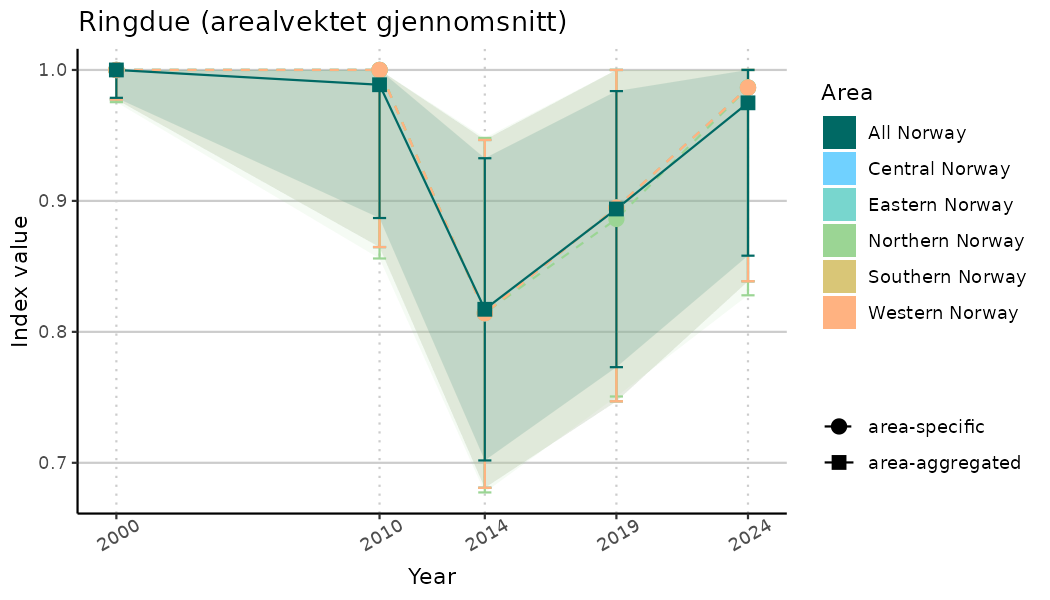
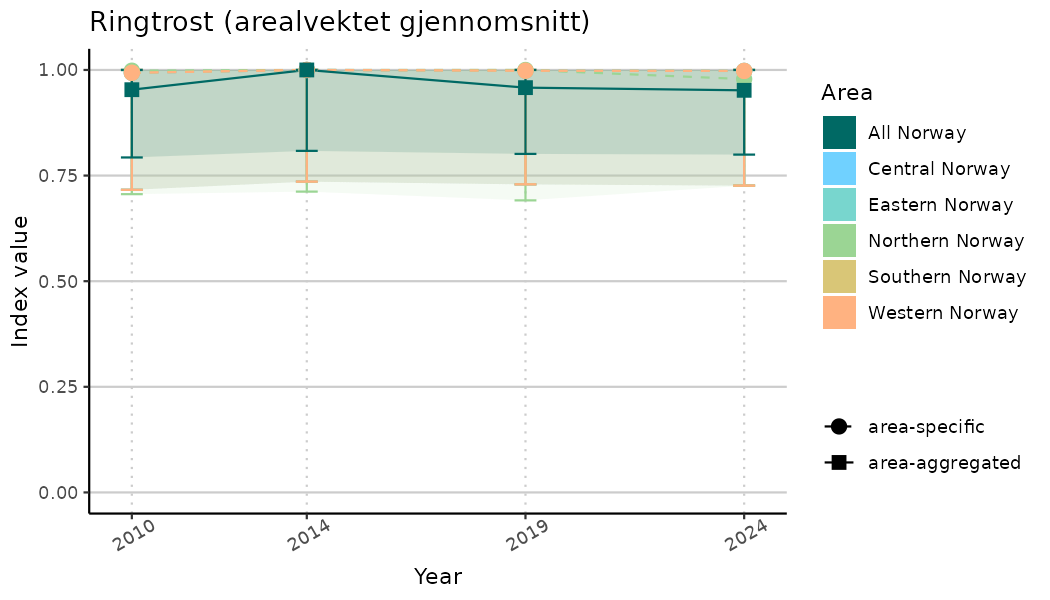
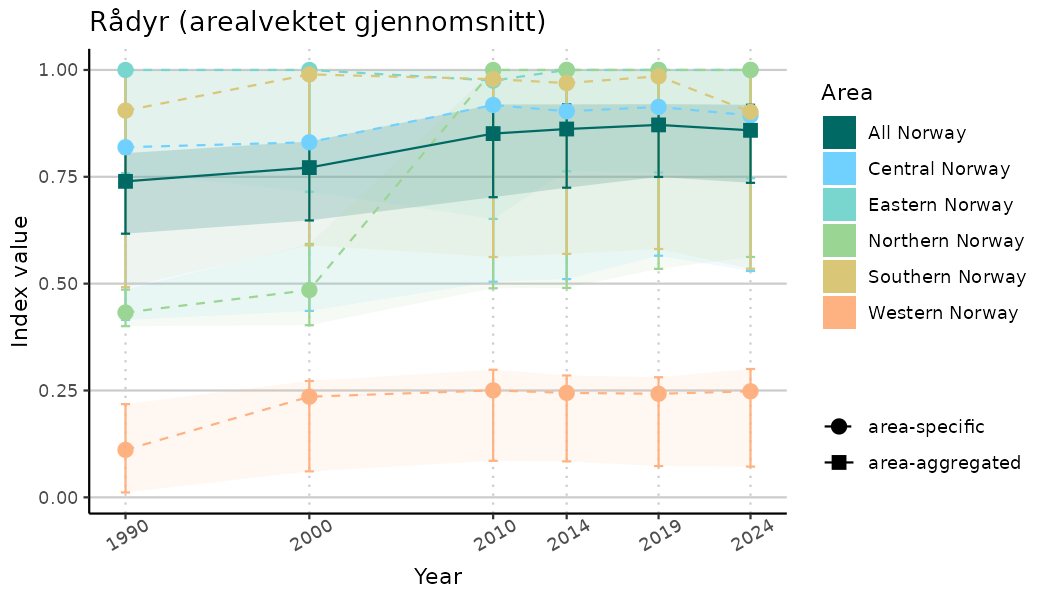
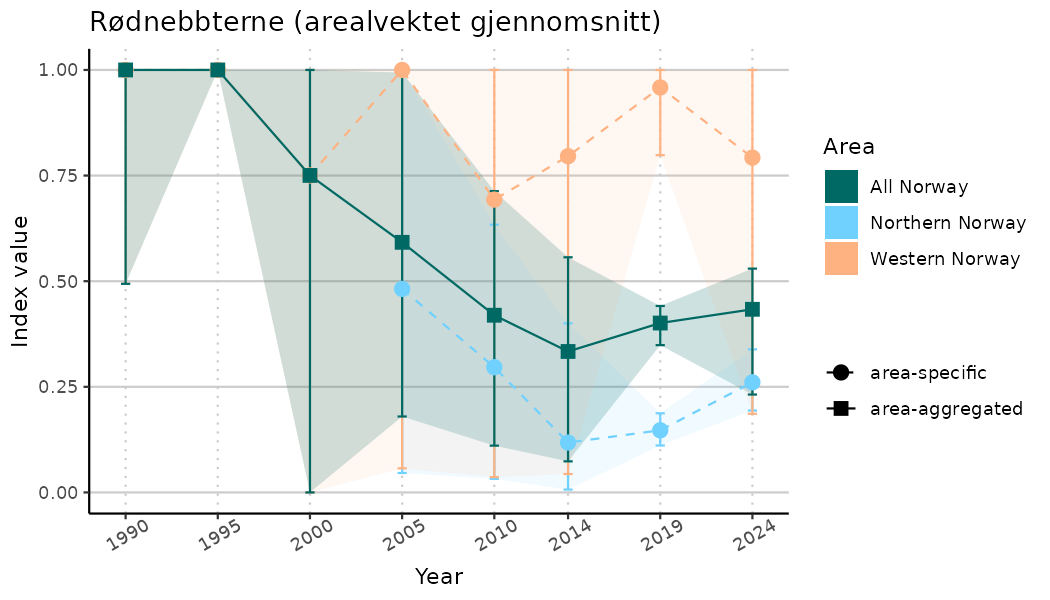
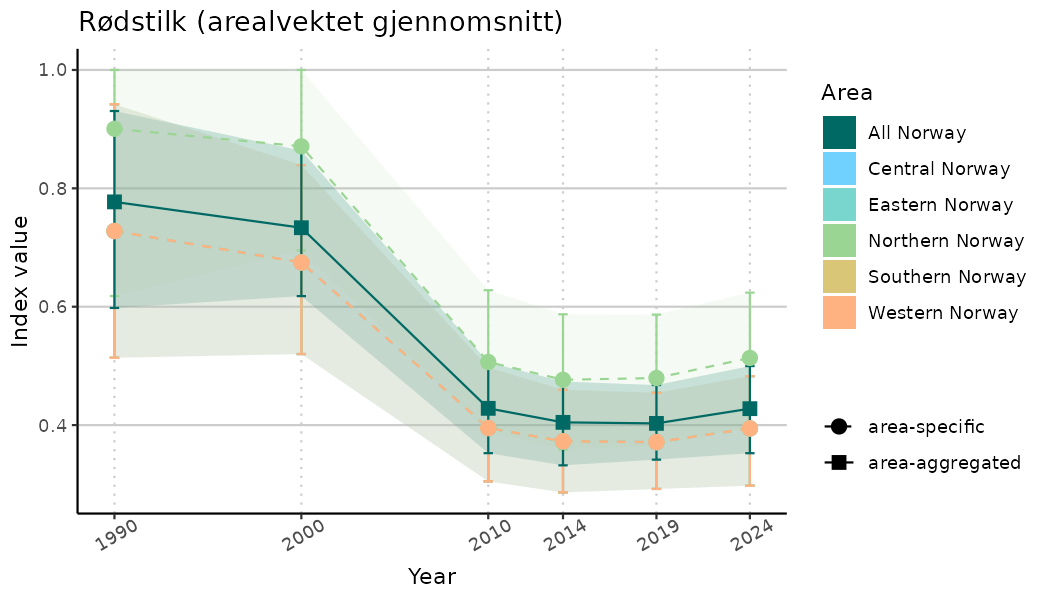
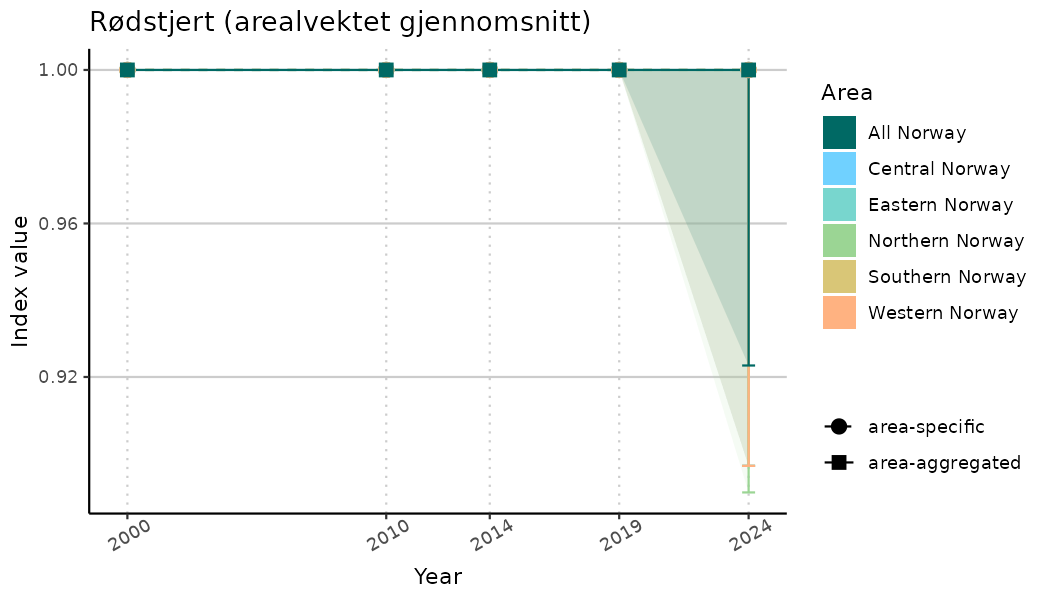
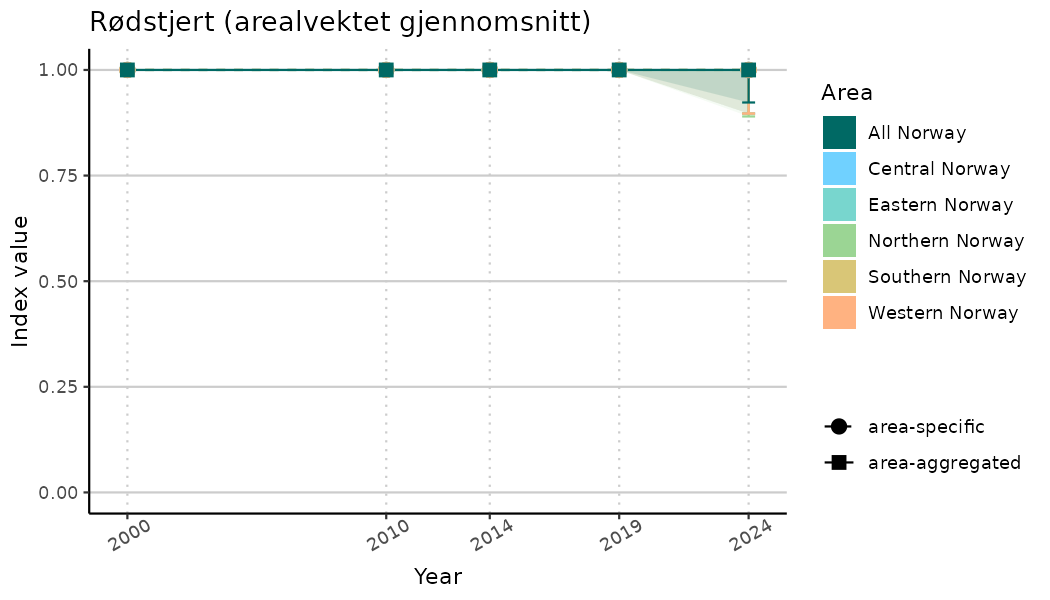
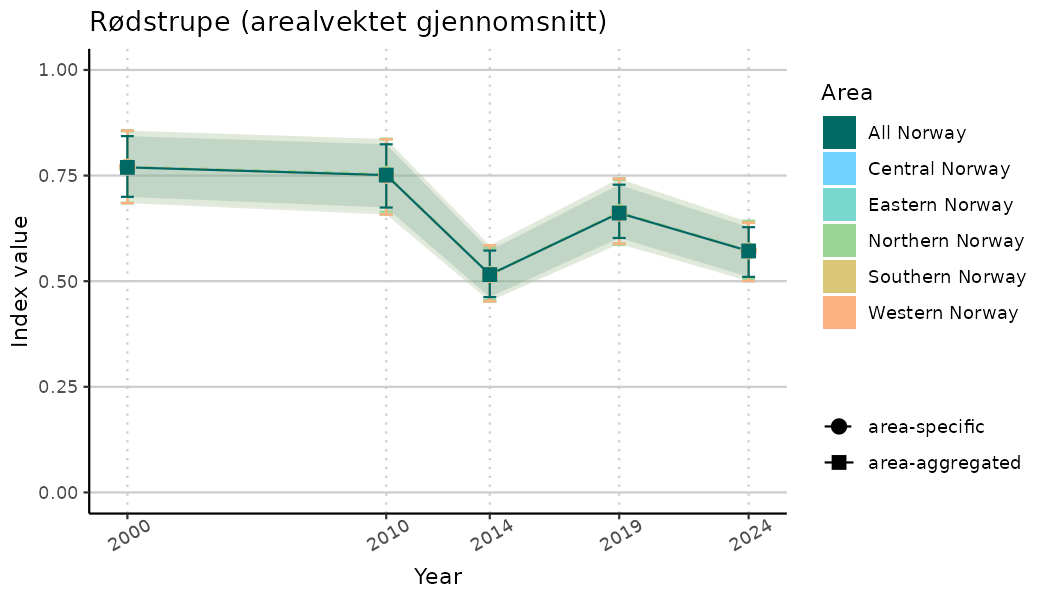
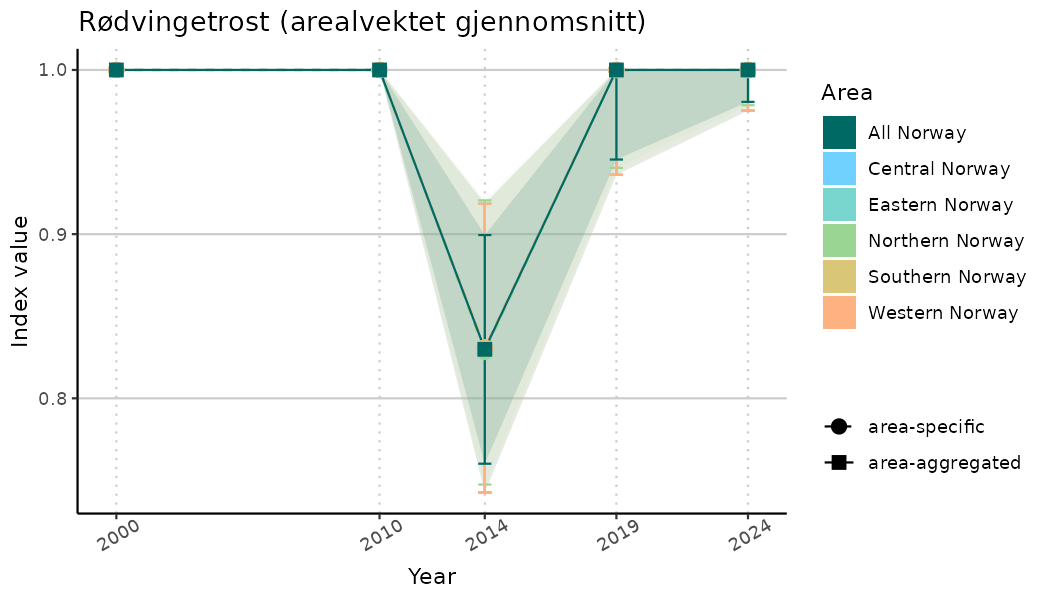
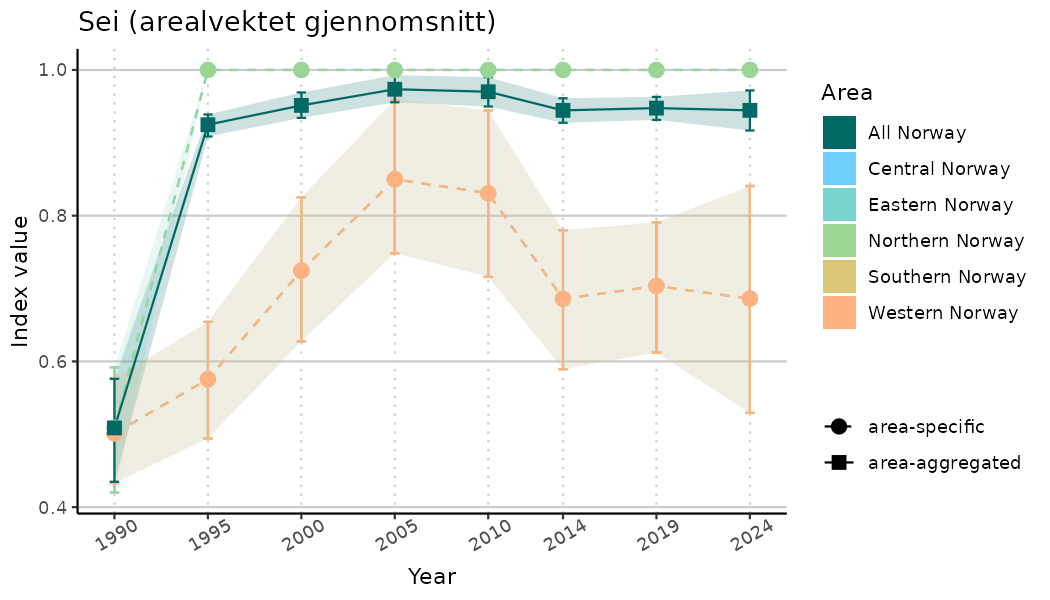
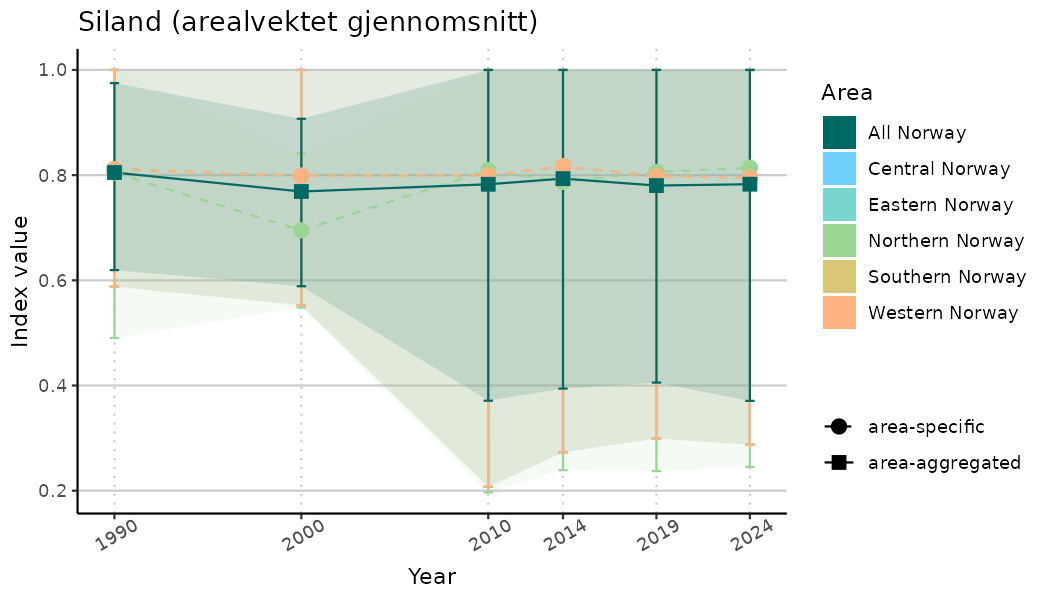
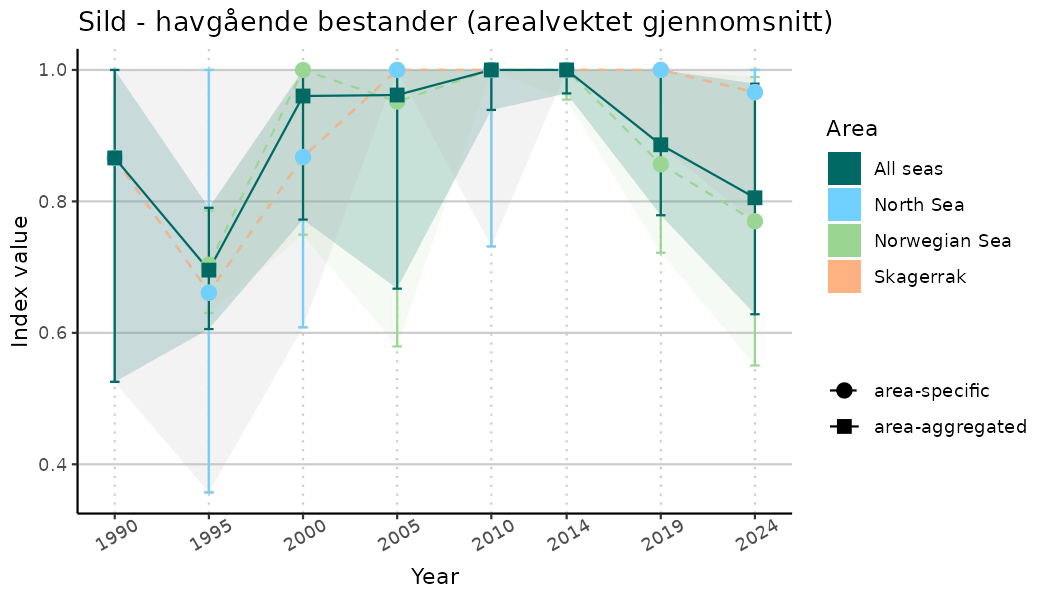
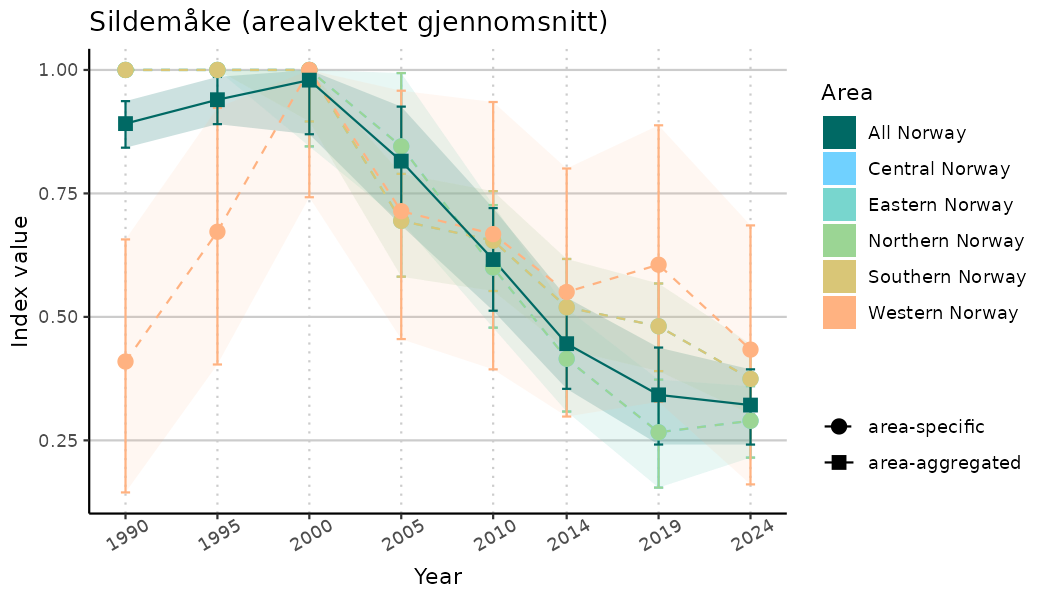
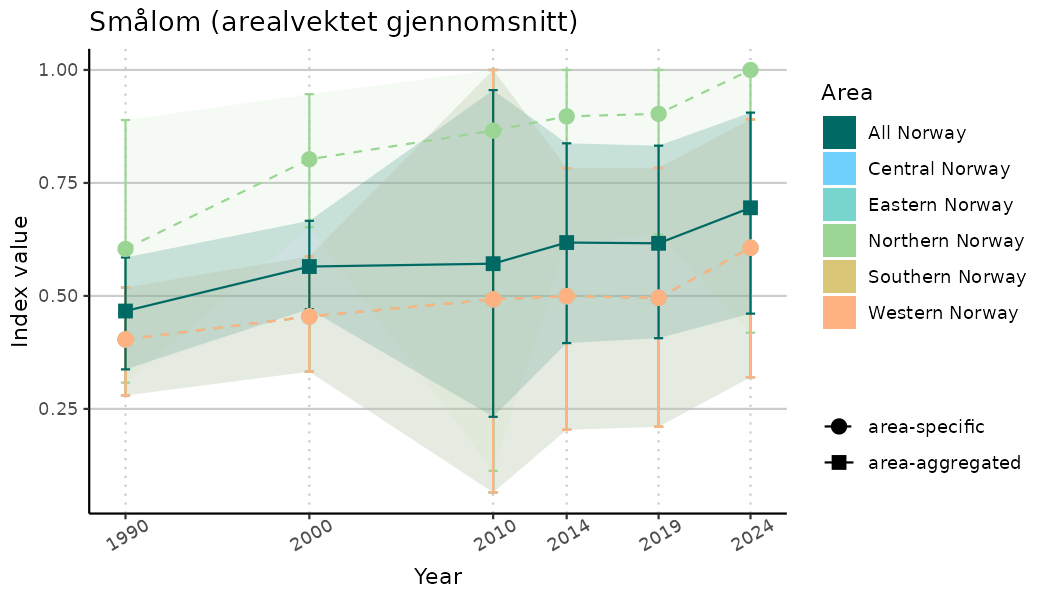
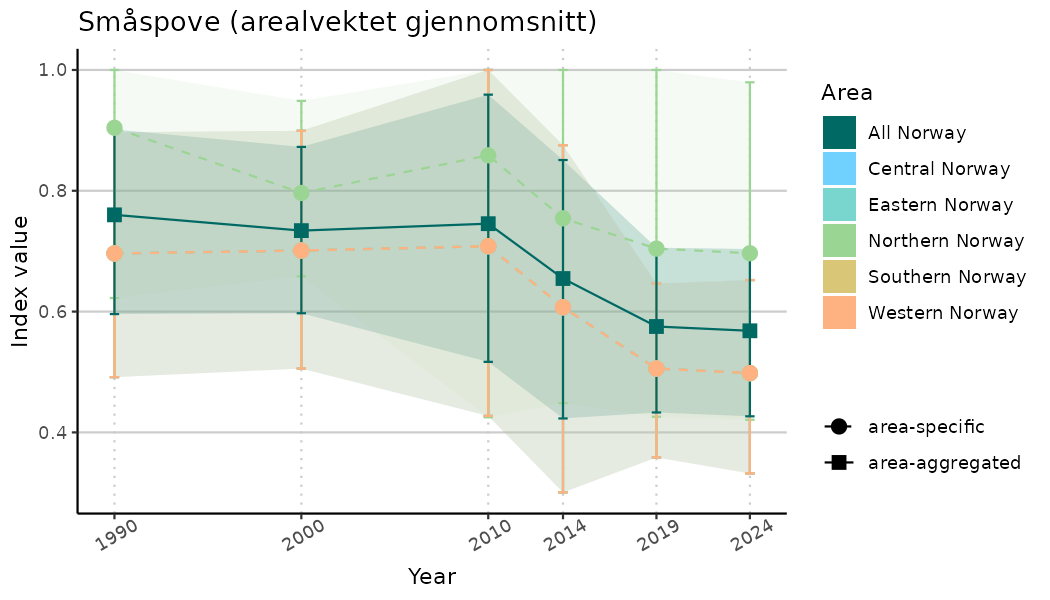
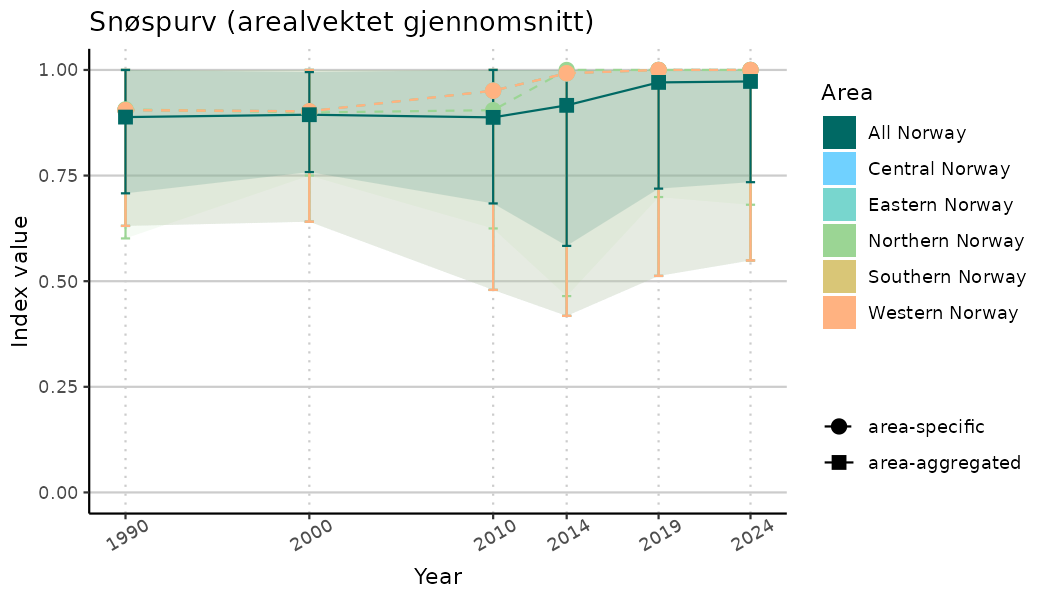
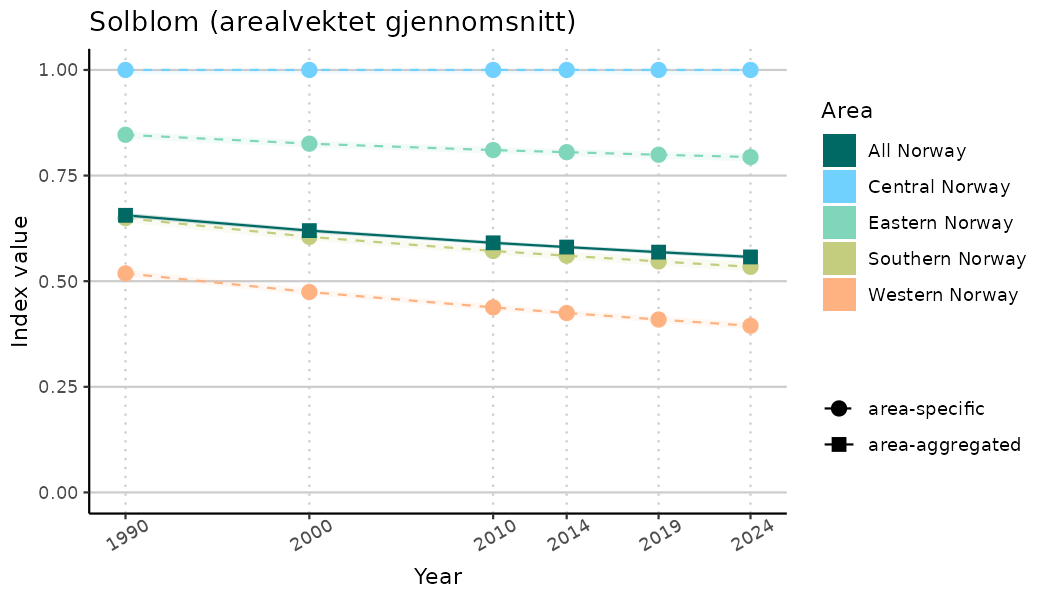
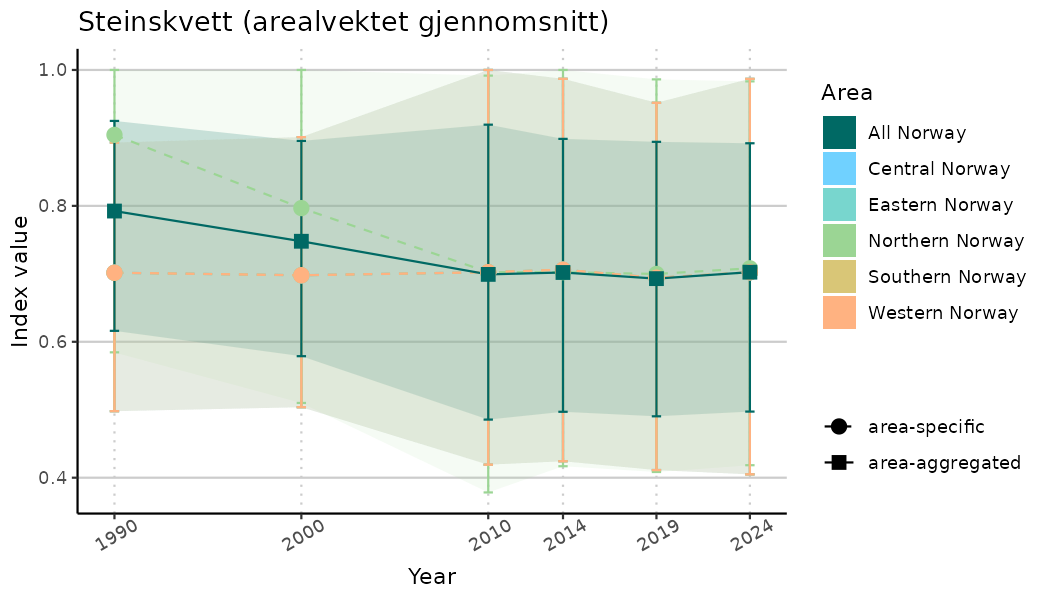
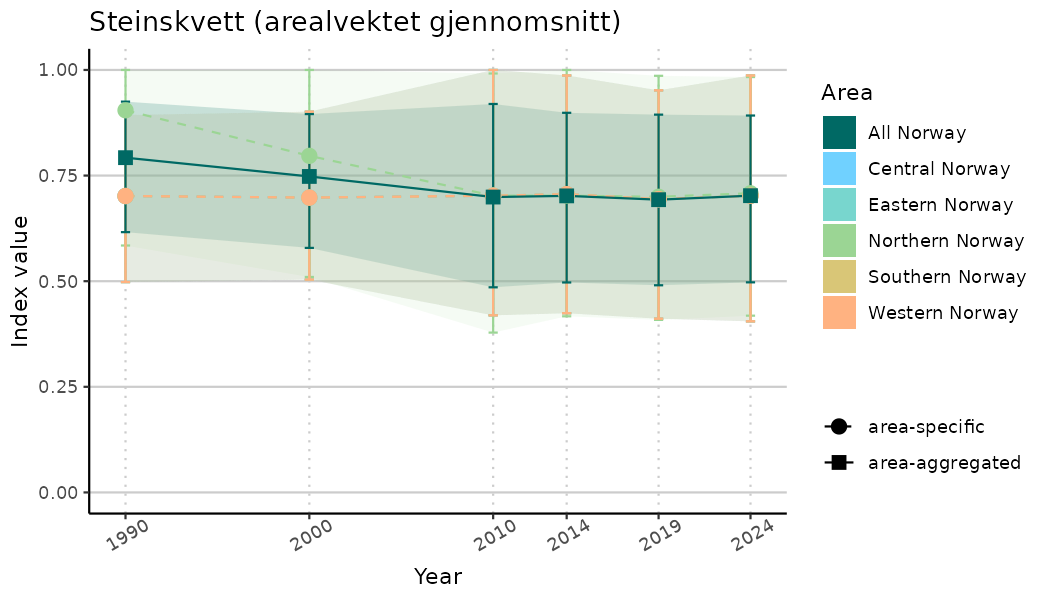
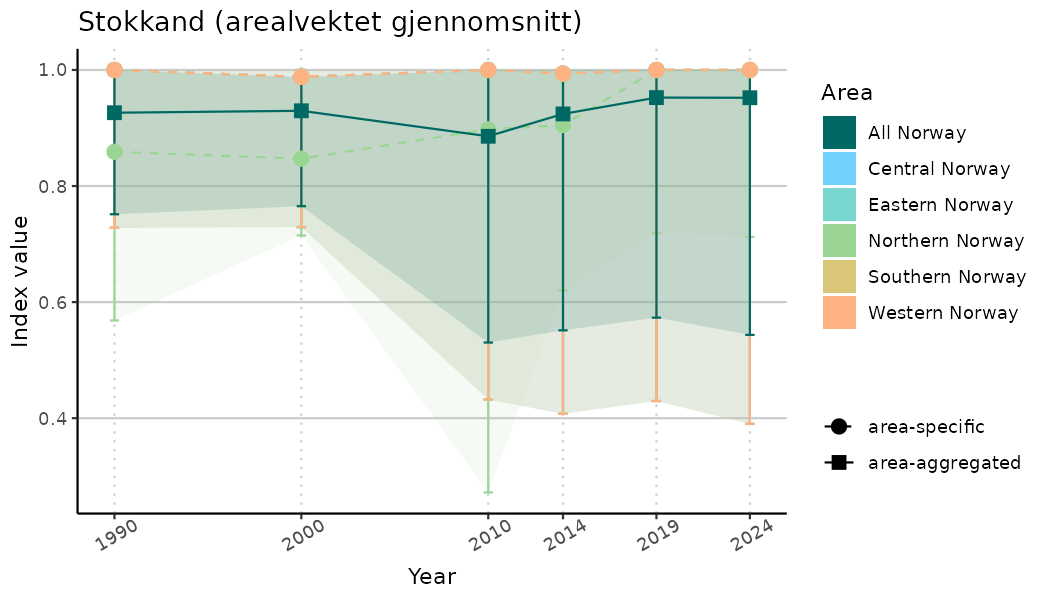
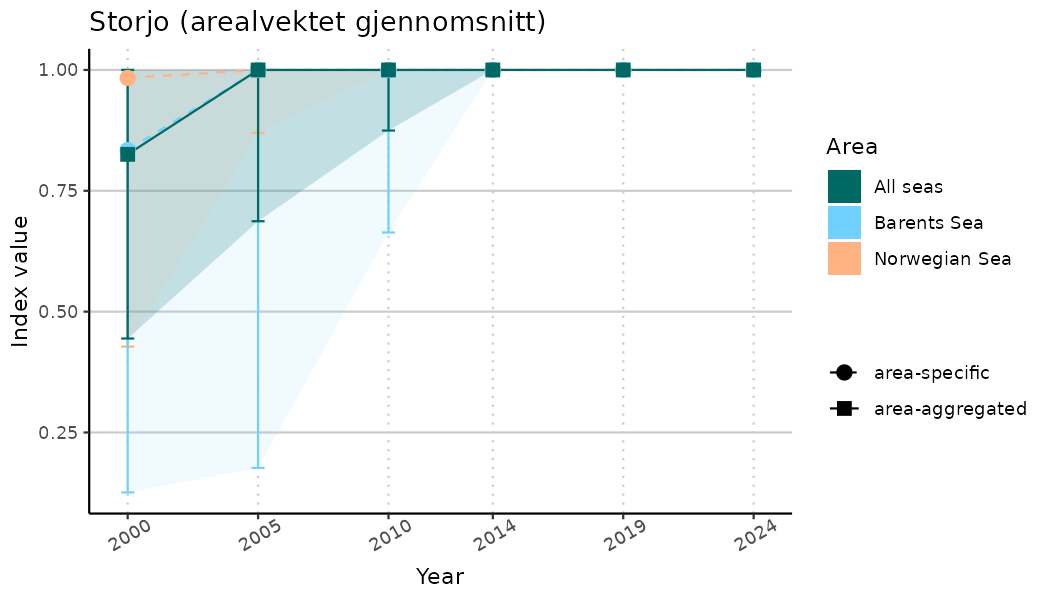
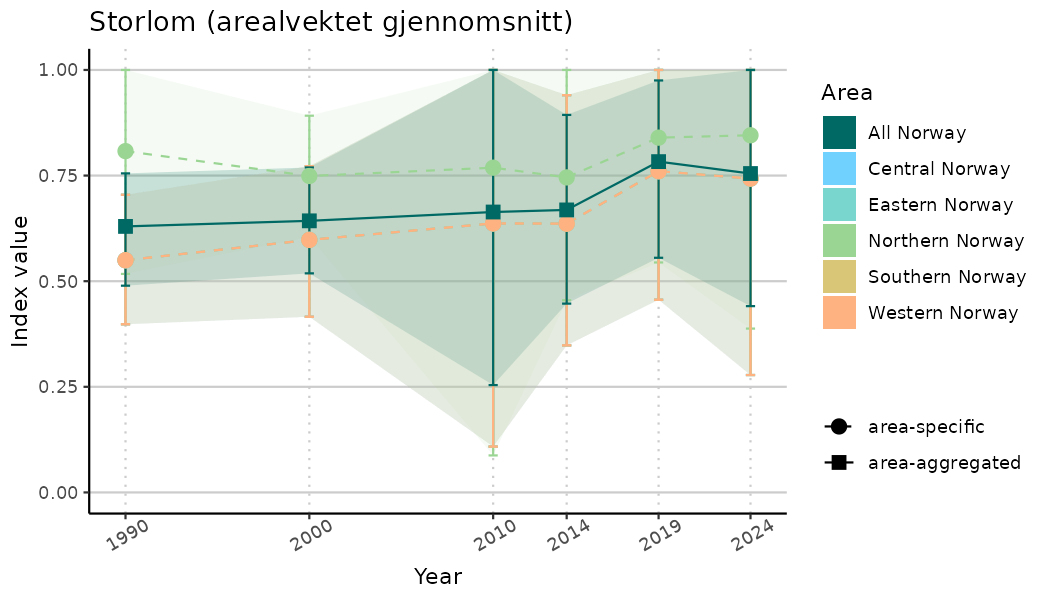
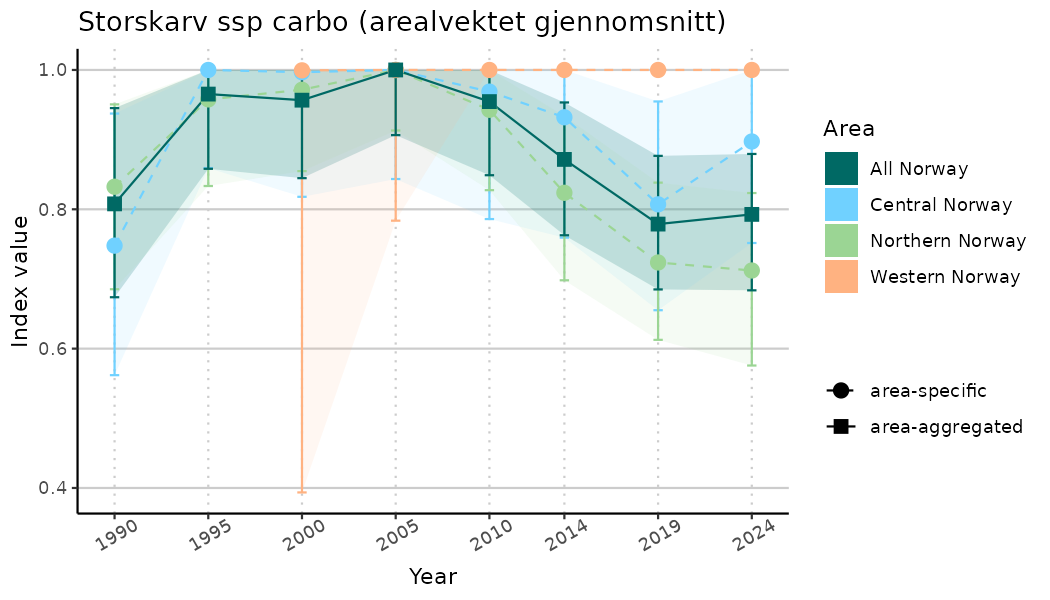
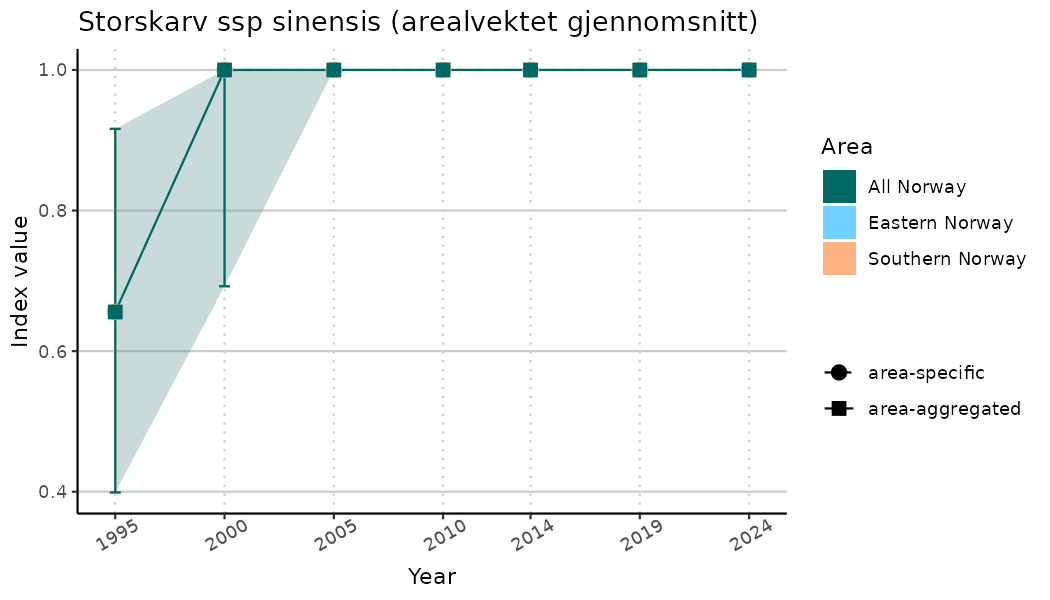
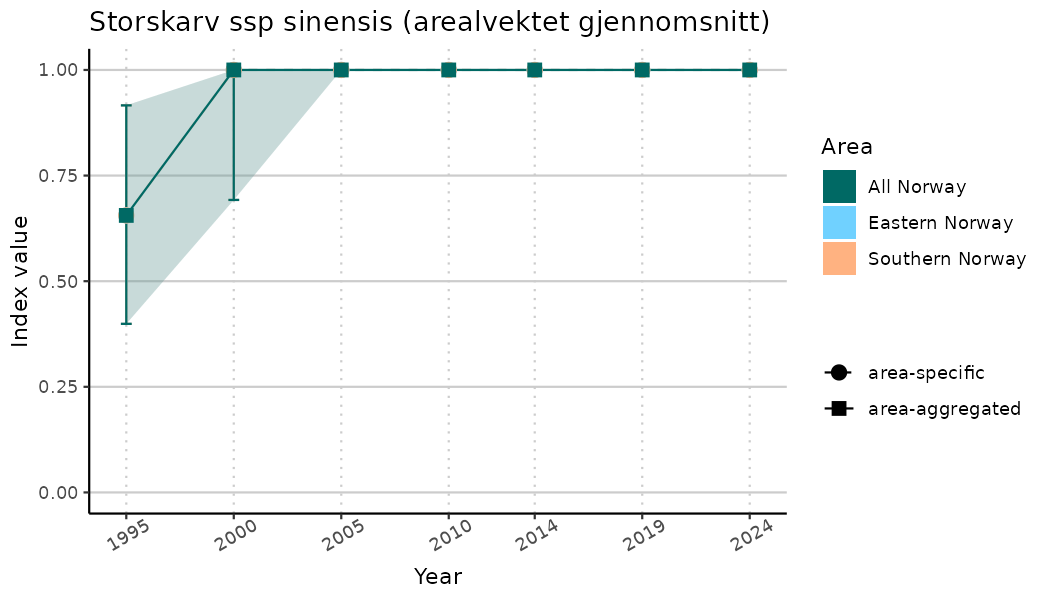
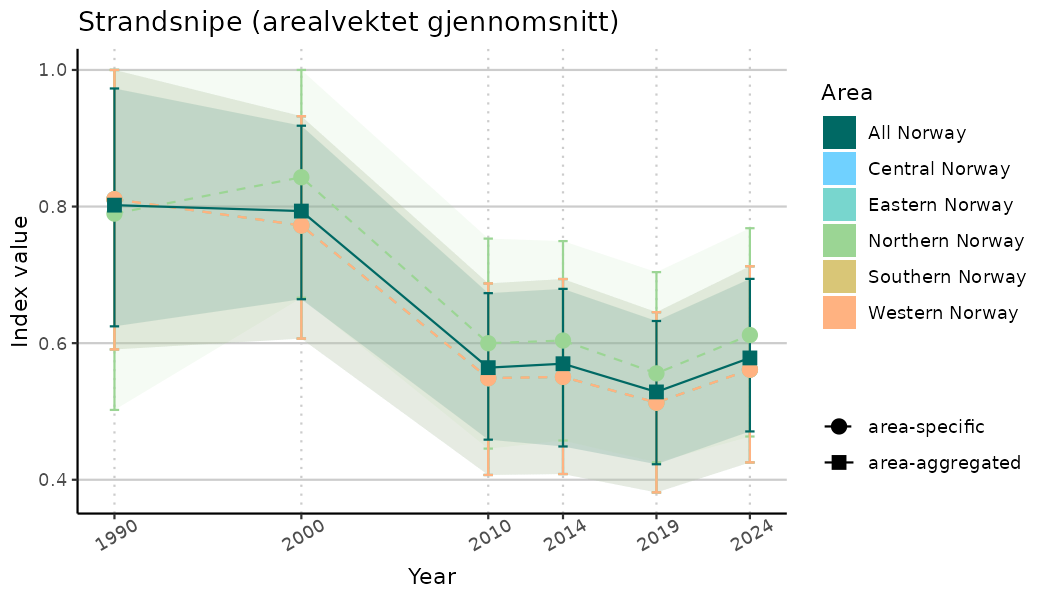
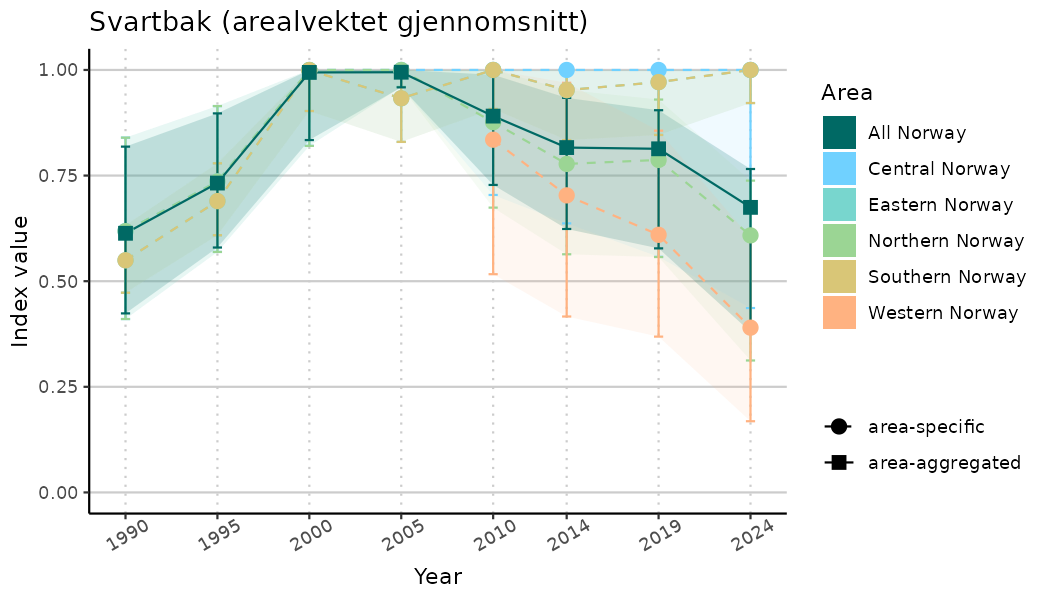
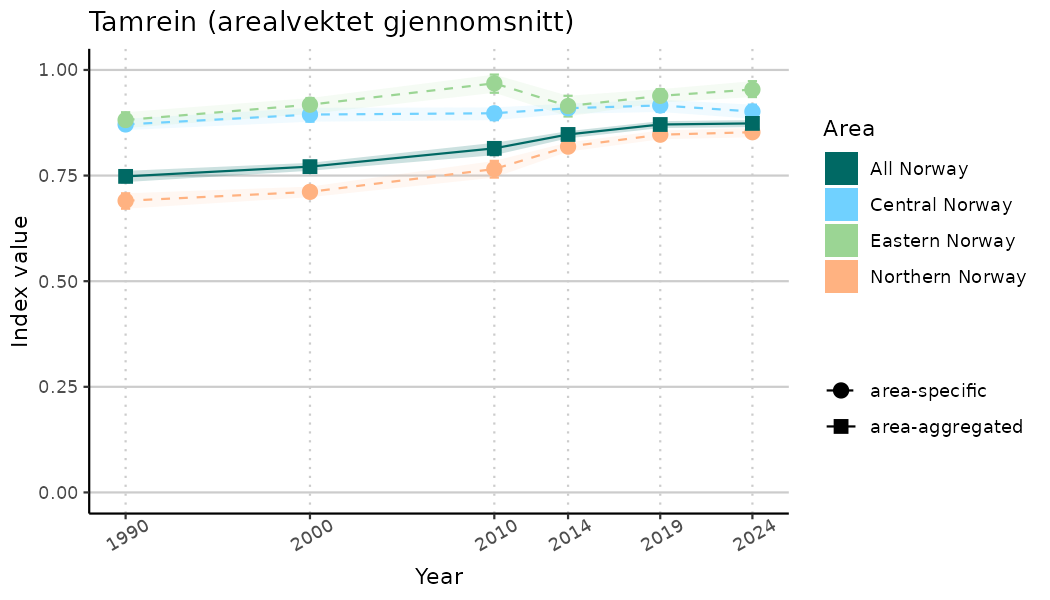
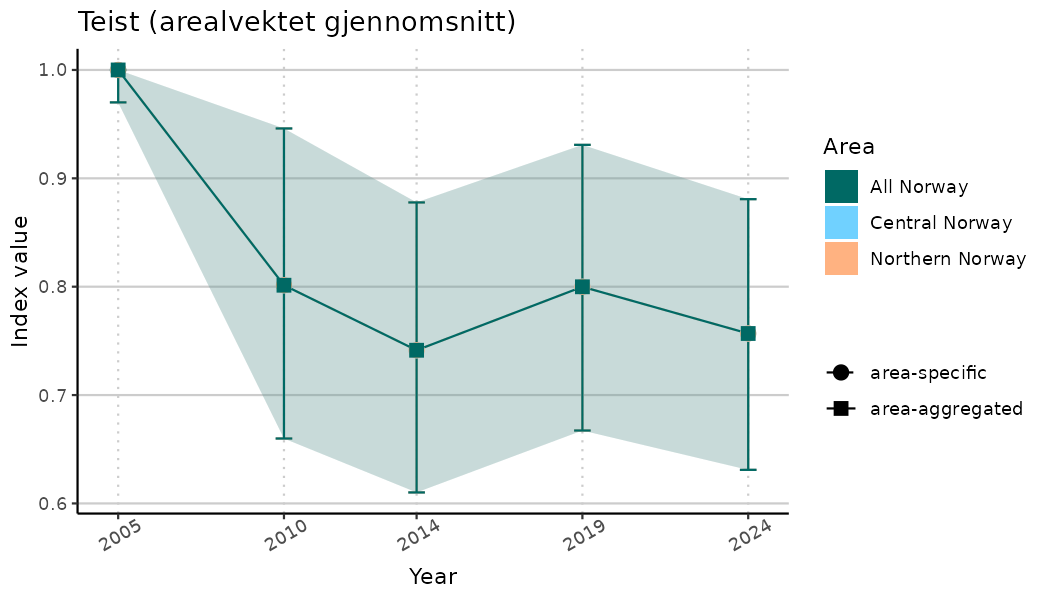
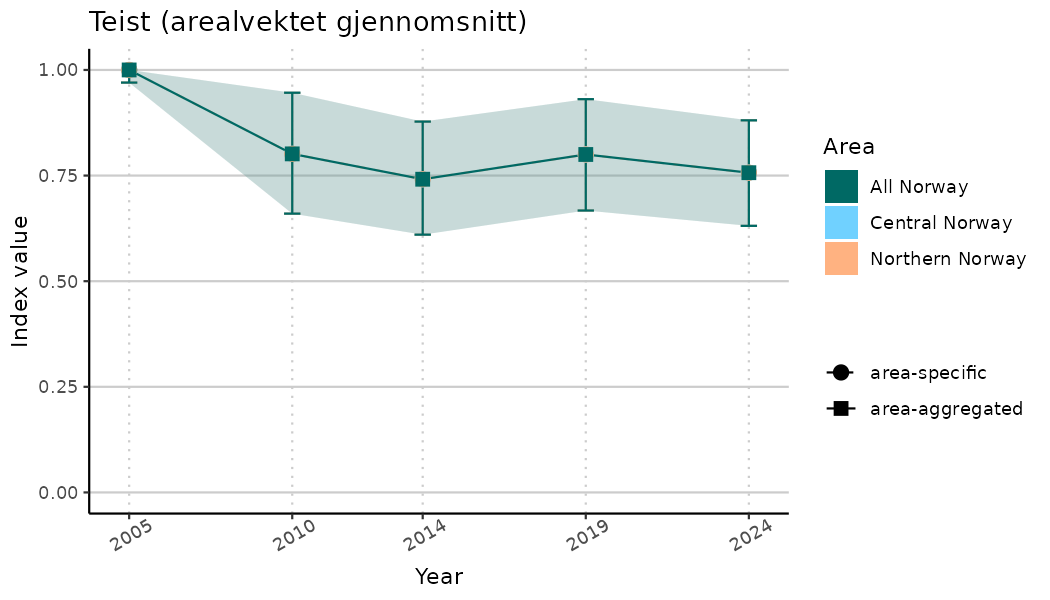
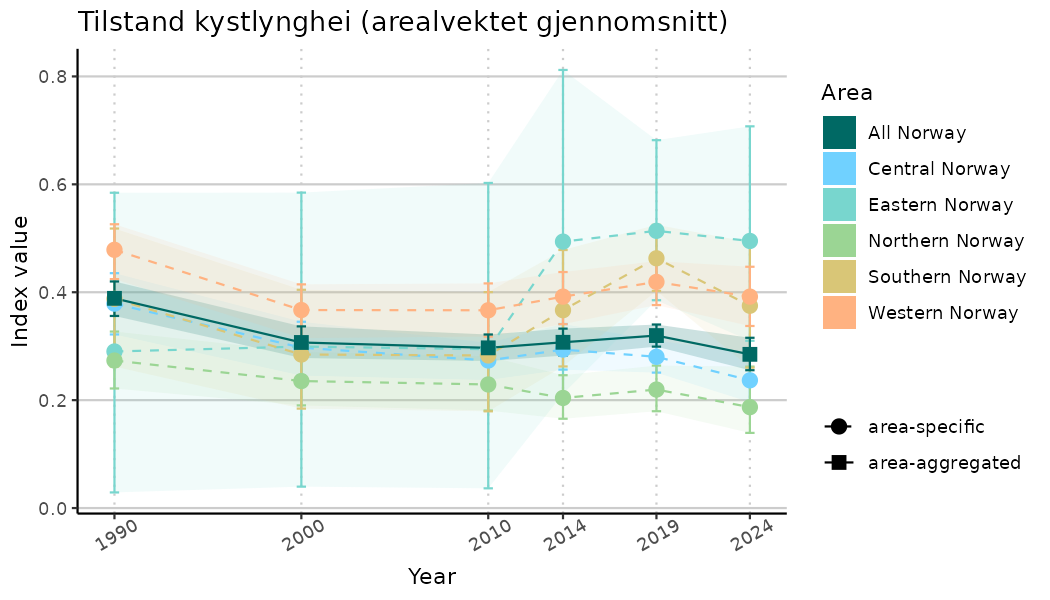
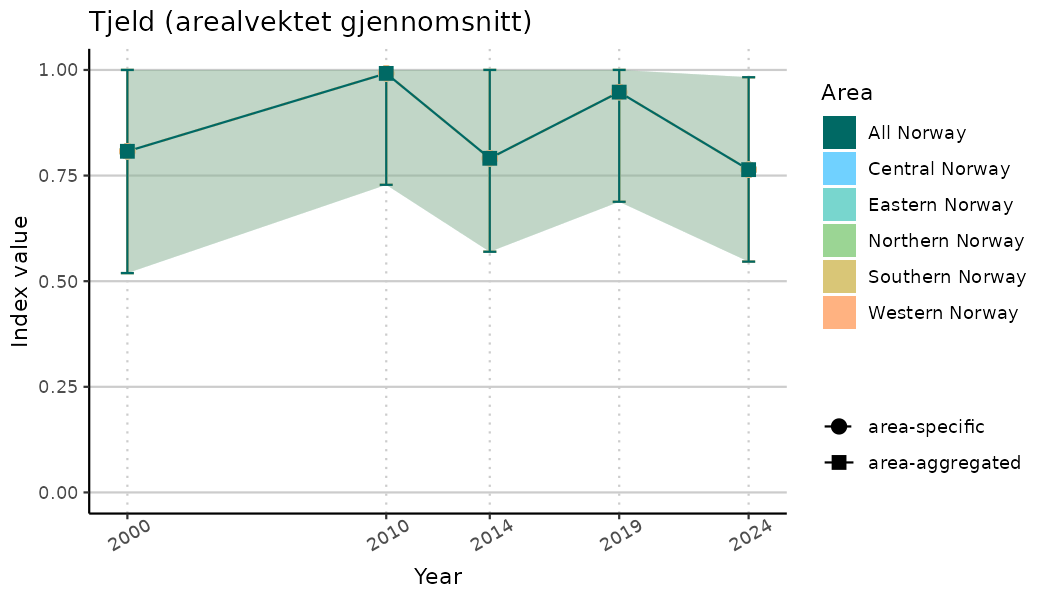
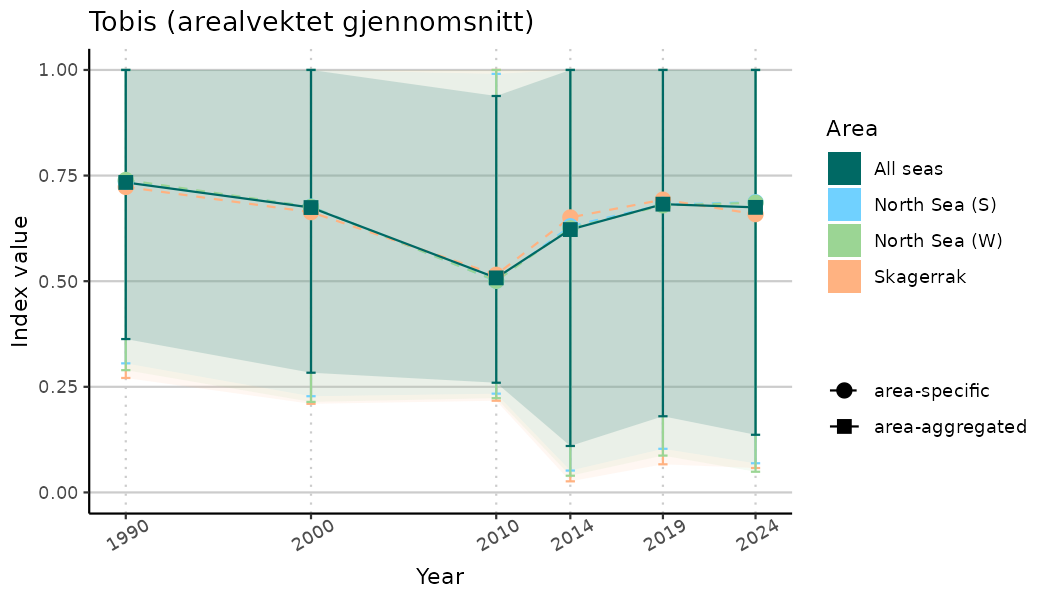
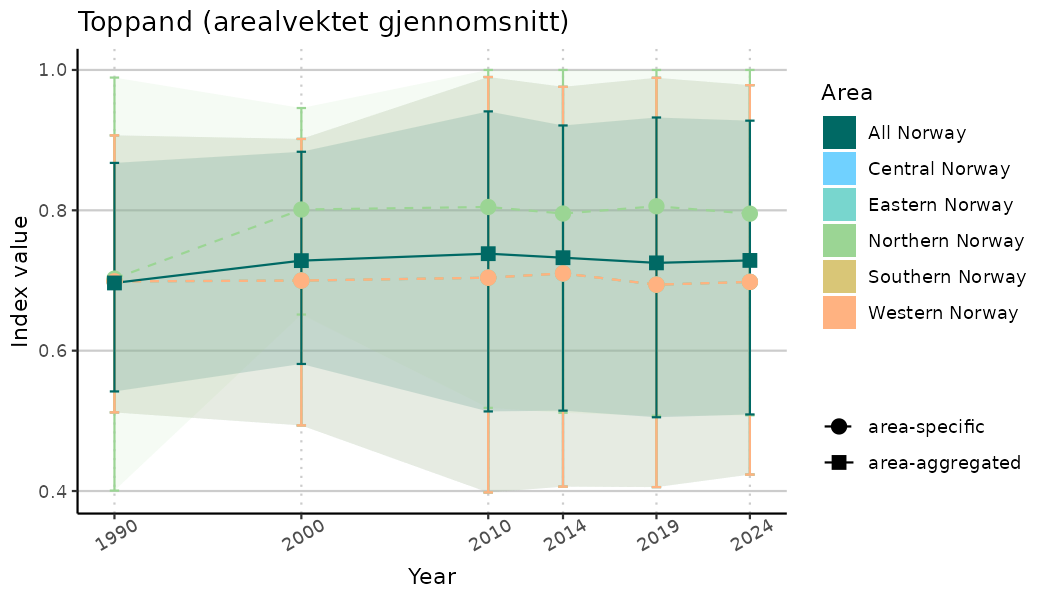
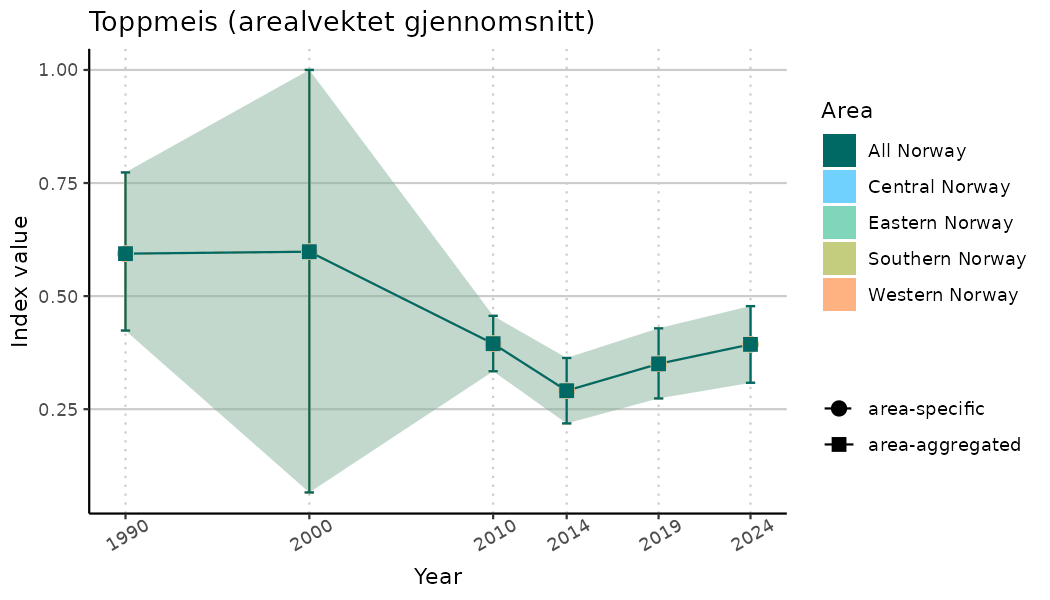
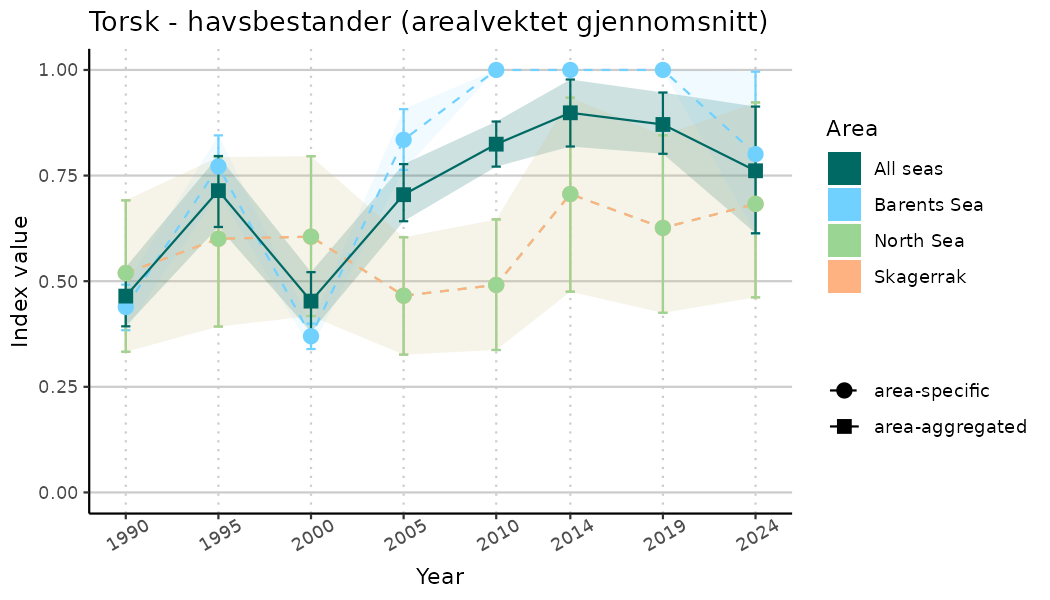
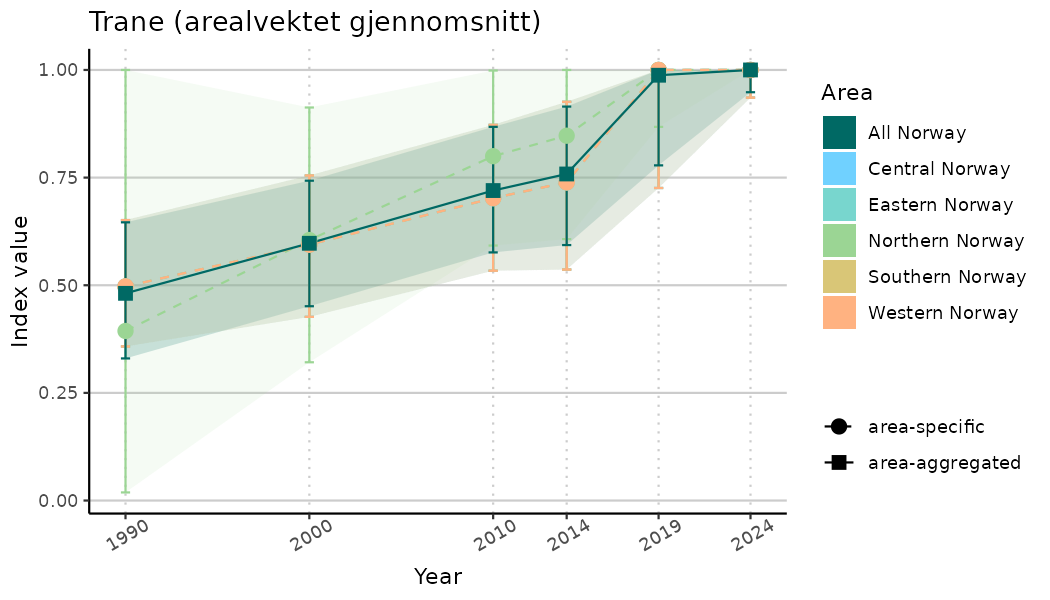
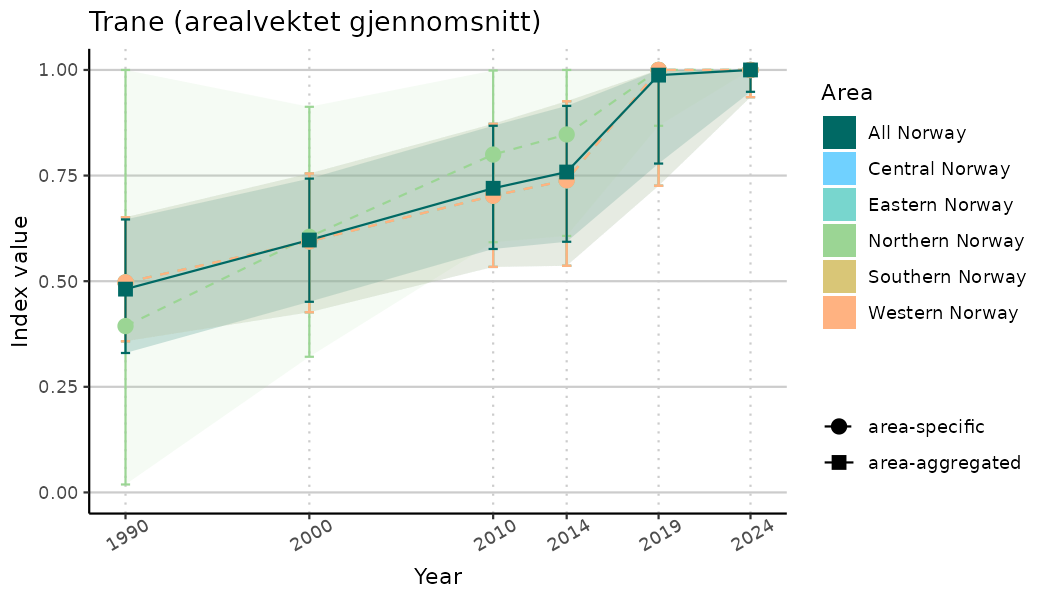
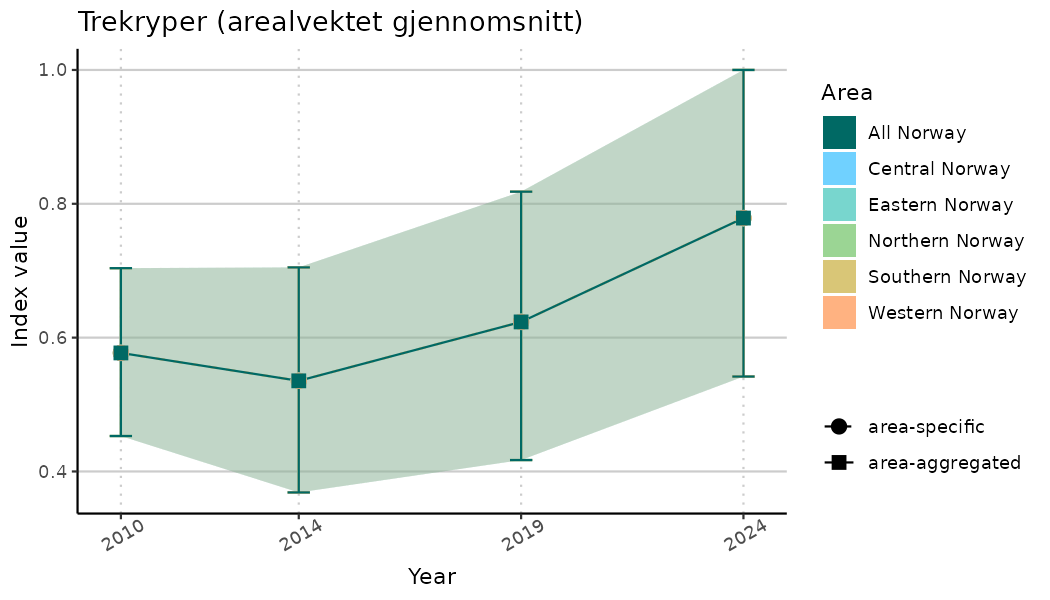
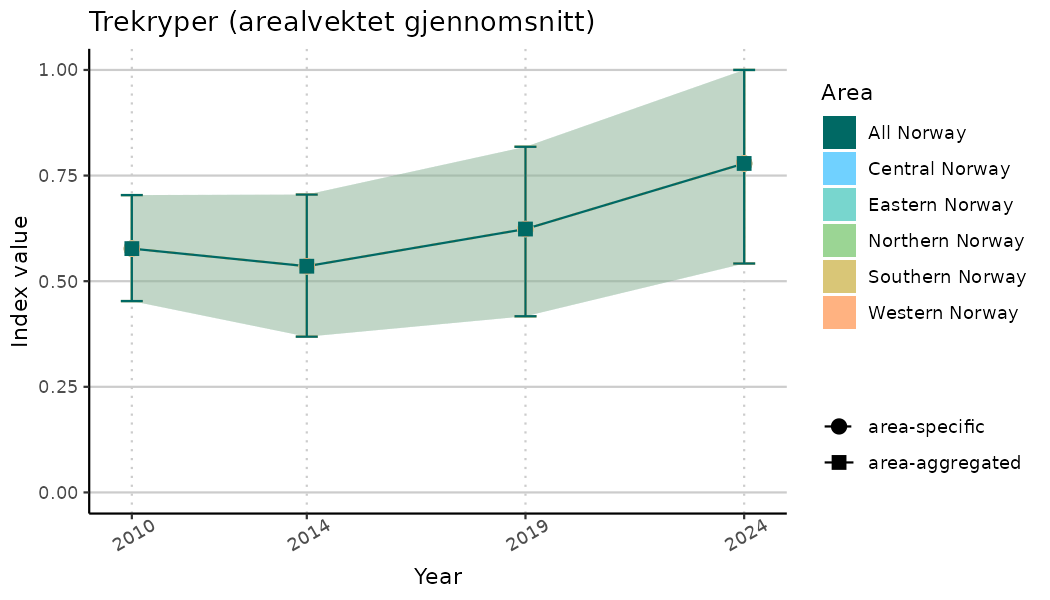
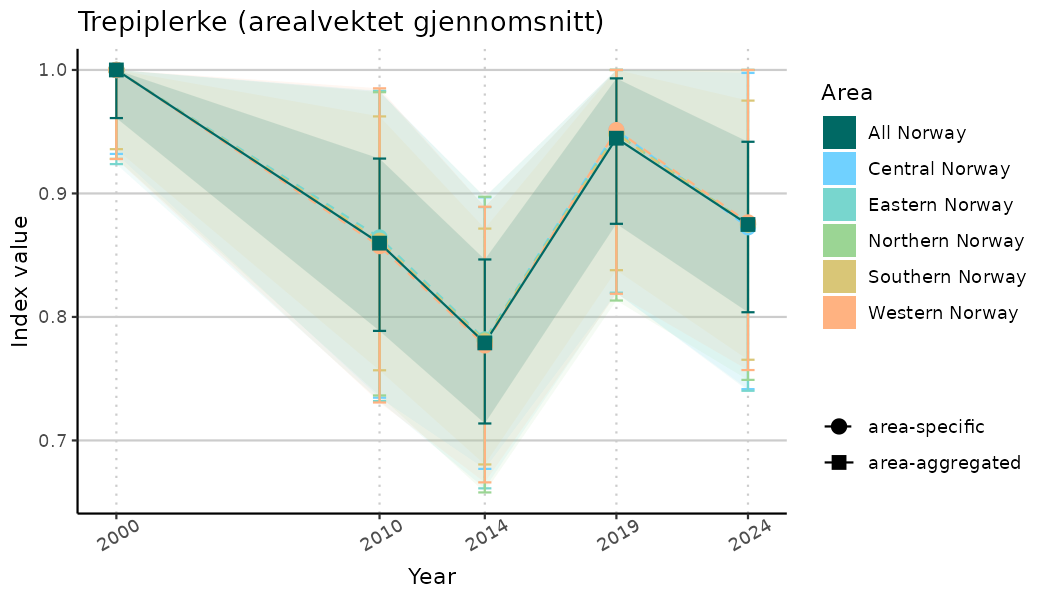
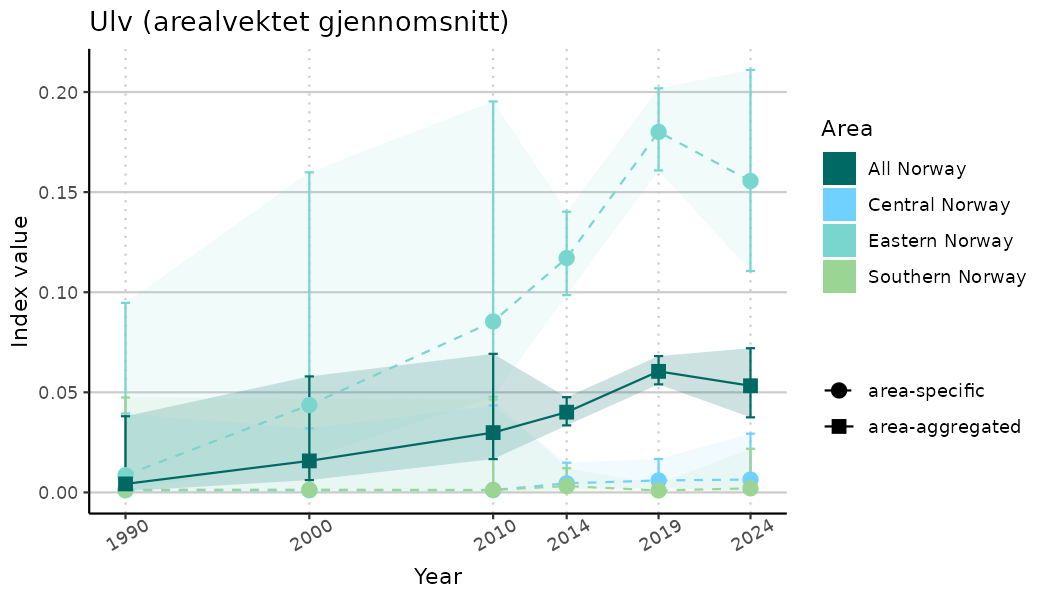
# Indicator indices (average and sub-areas)
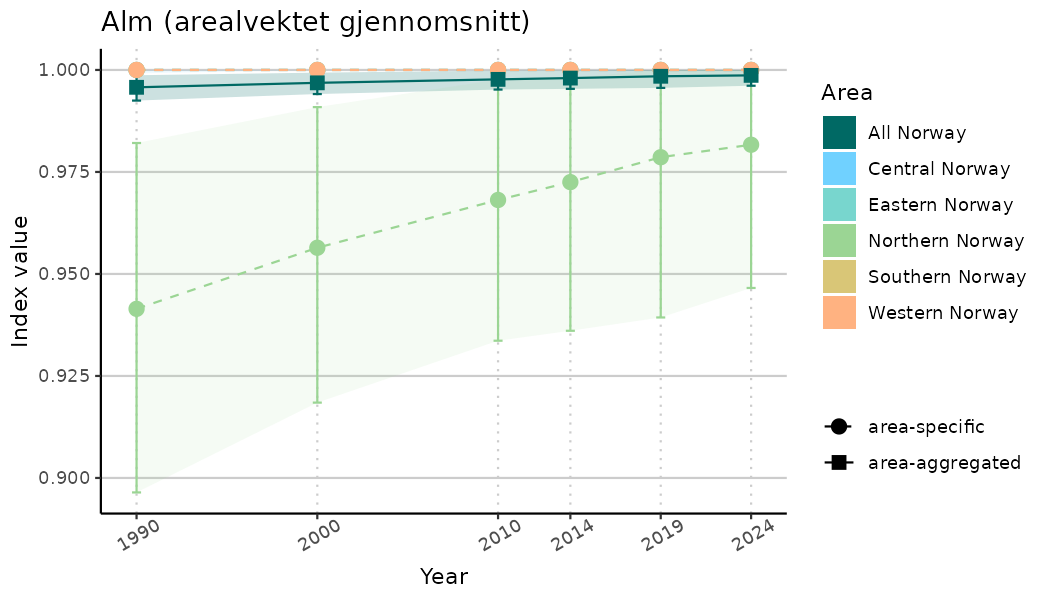
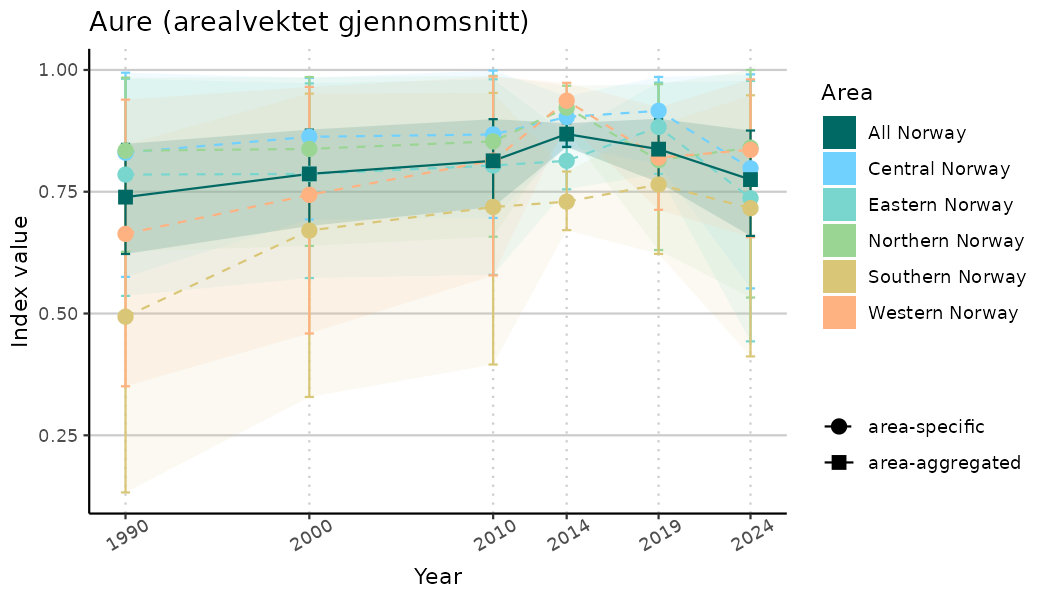
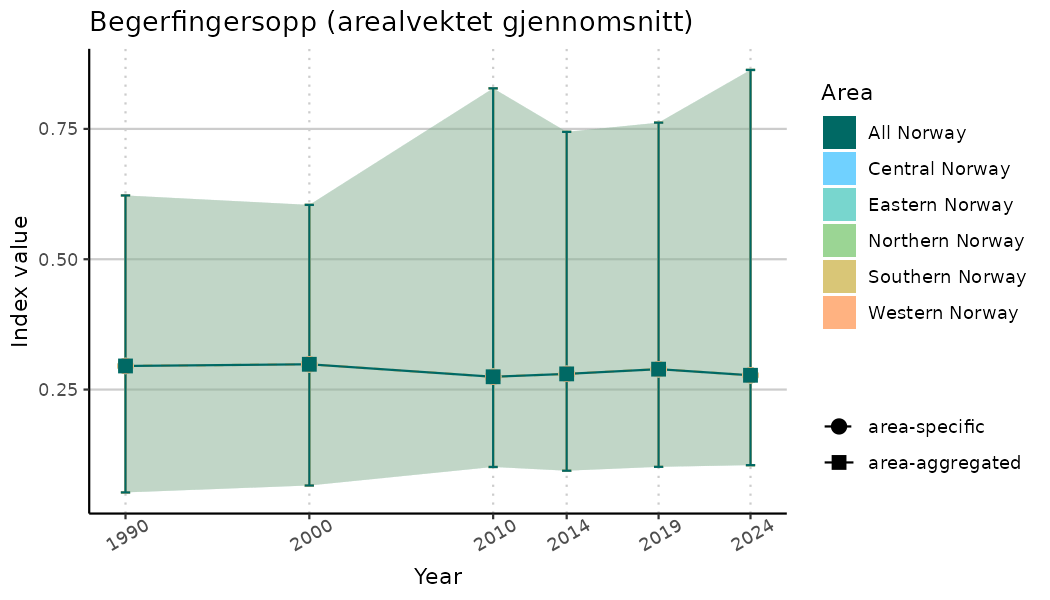
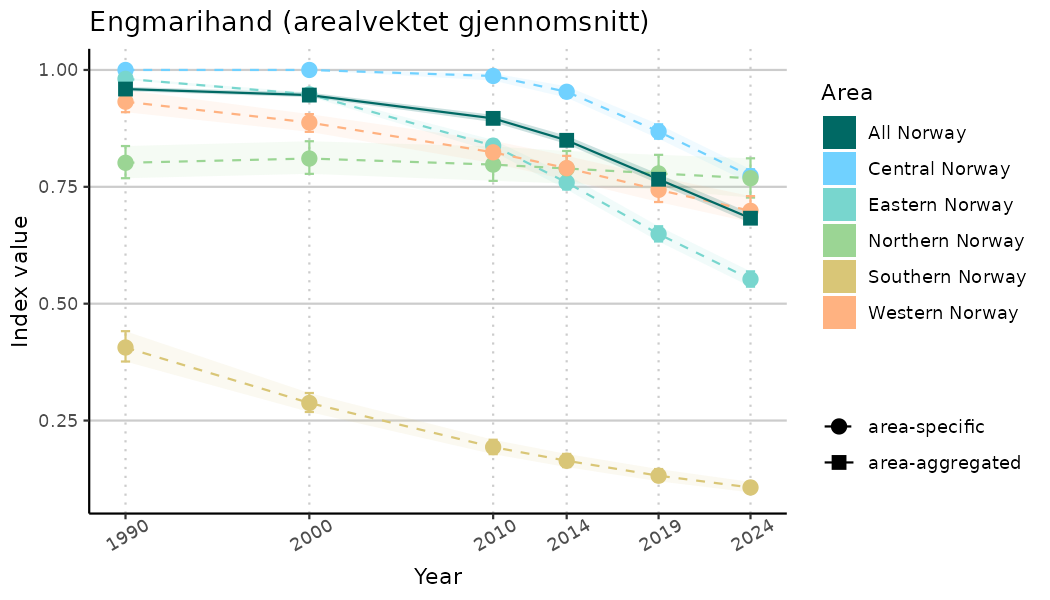
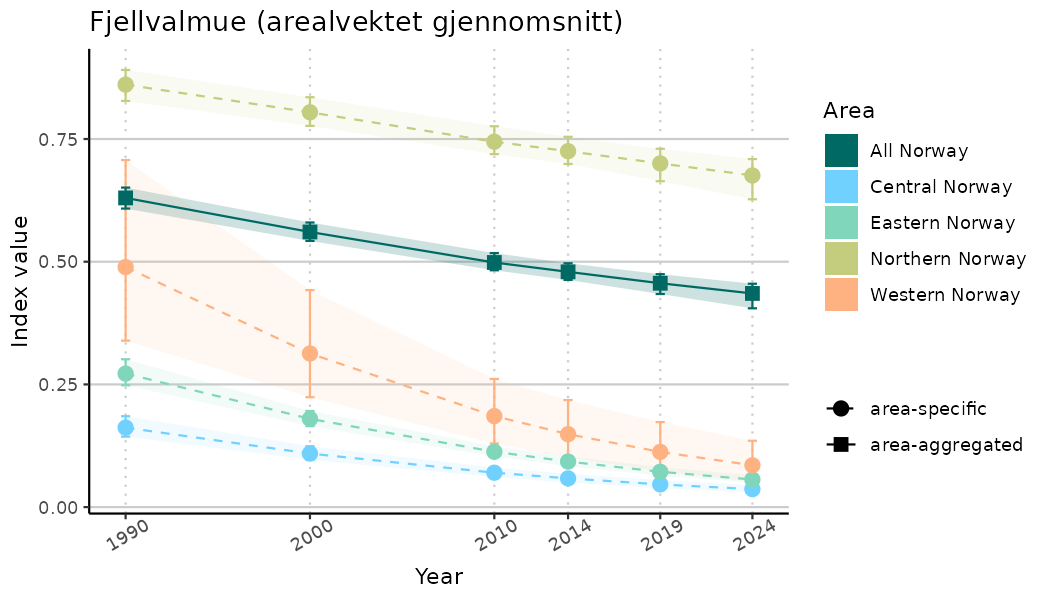
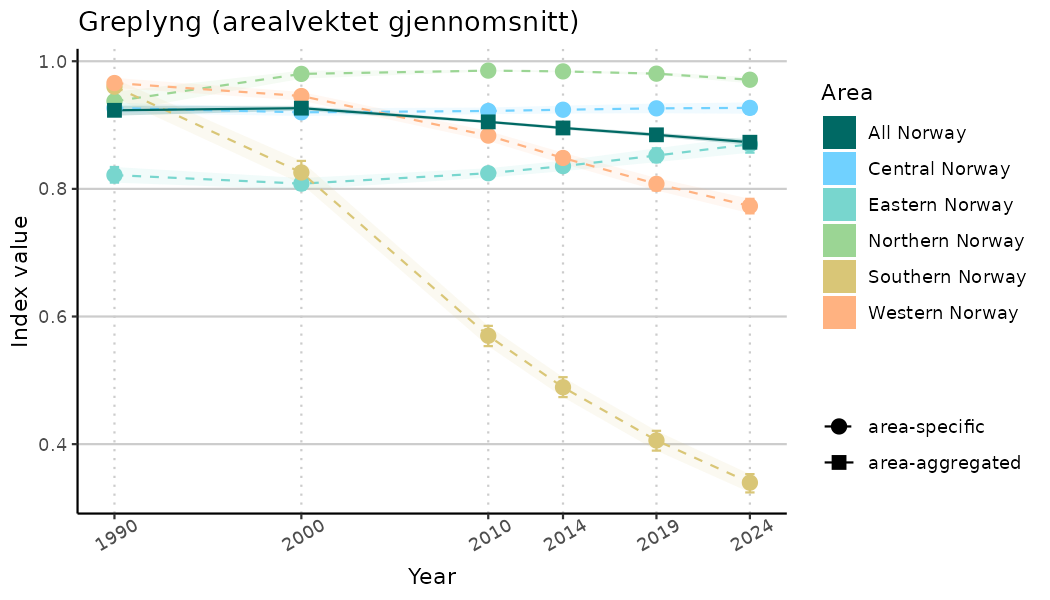
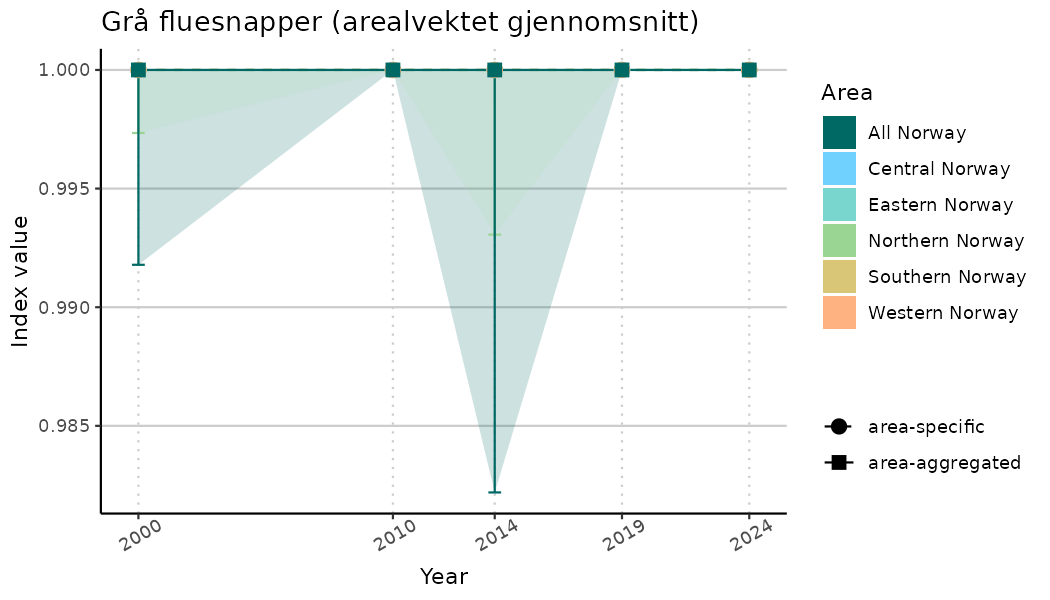
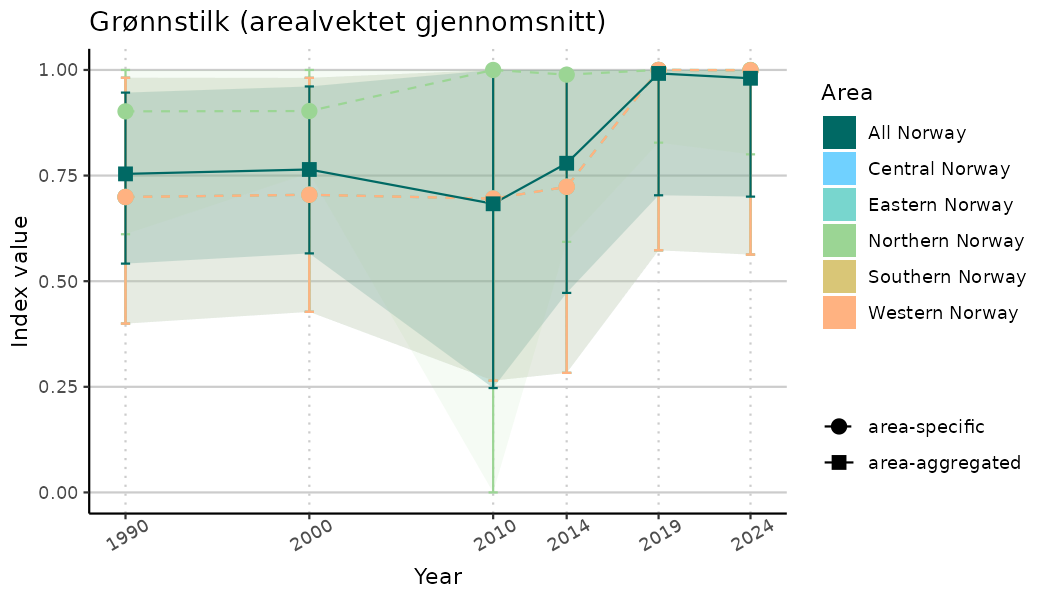
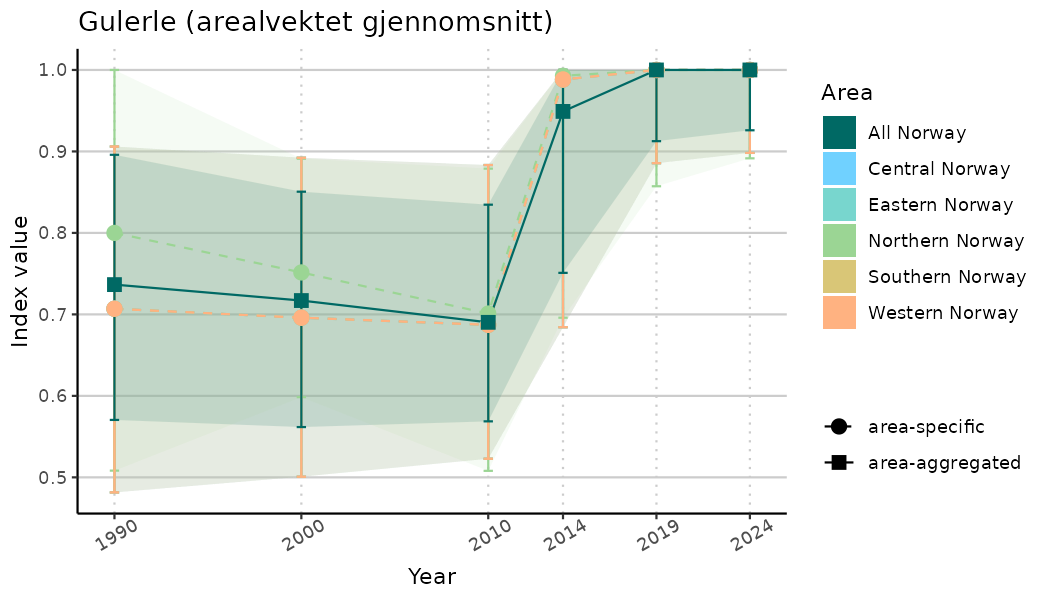
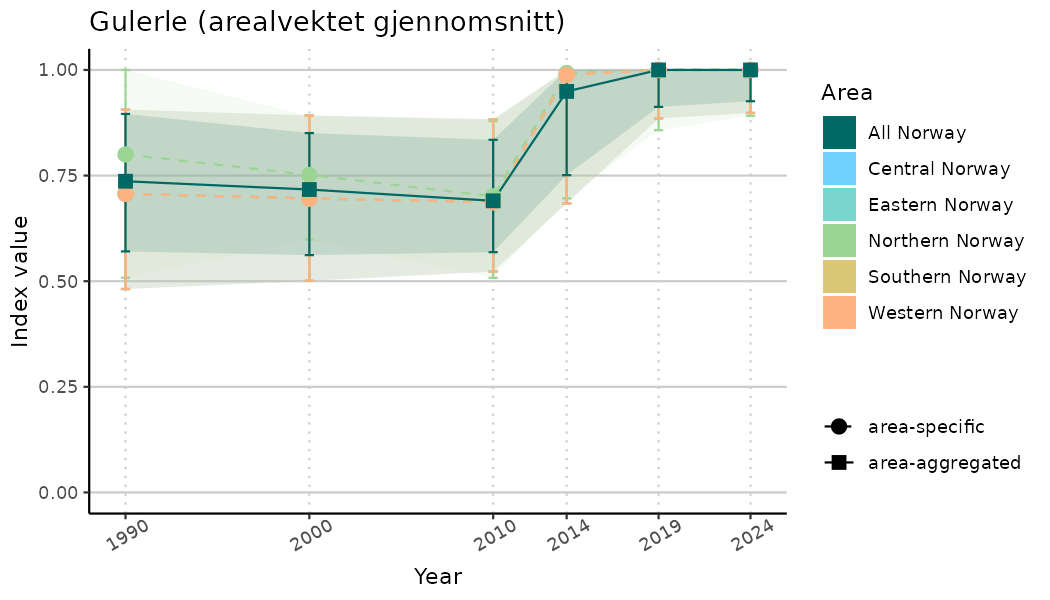
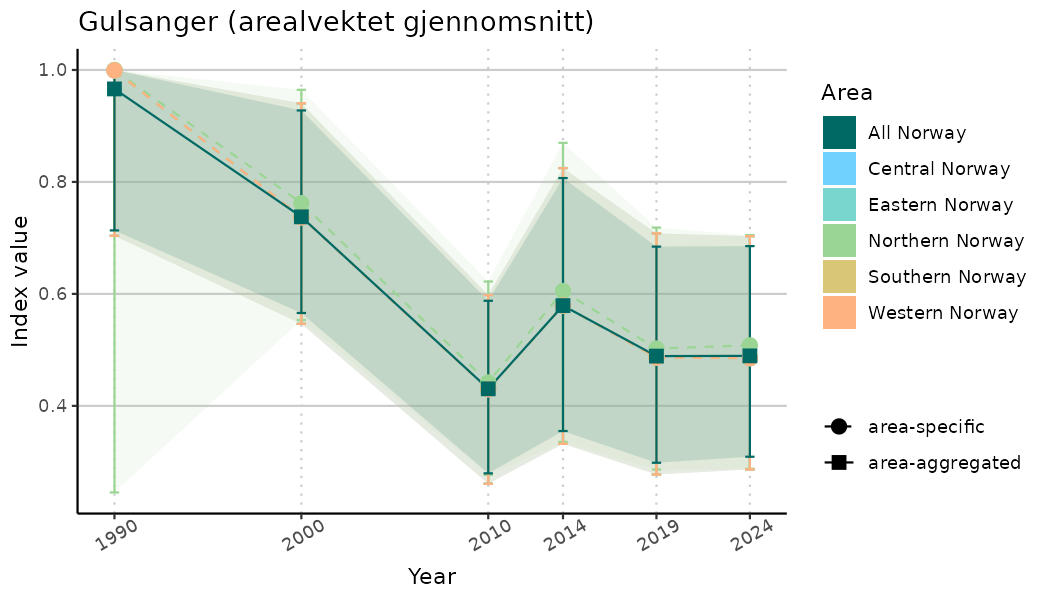
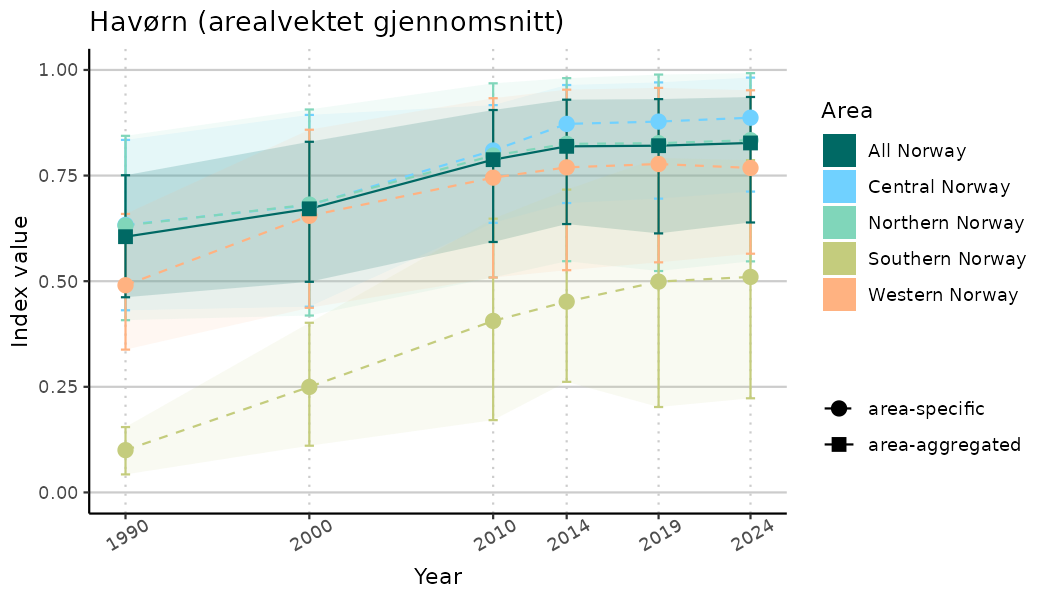
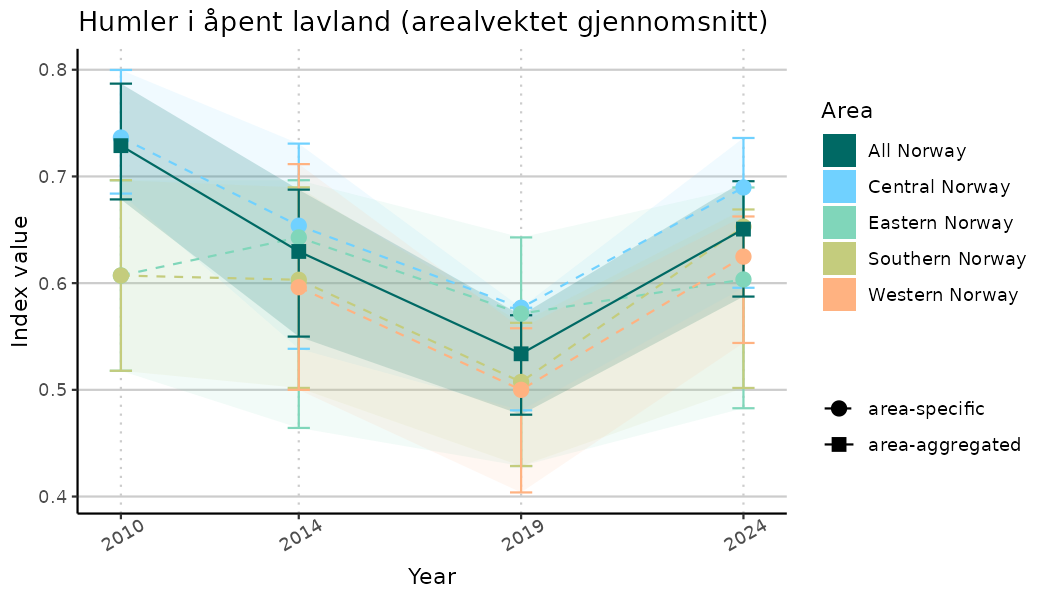
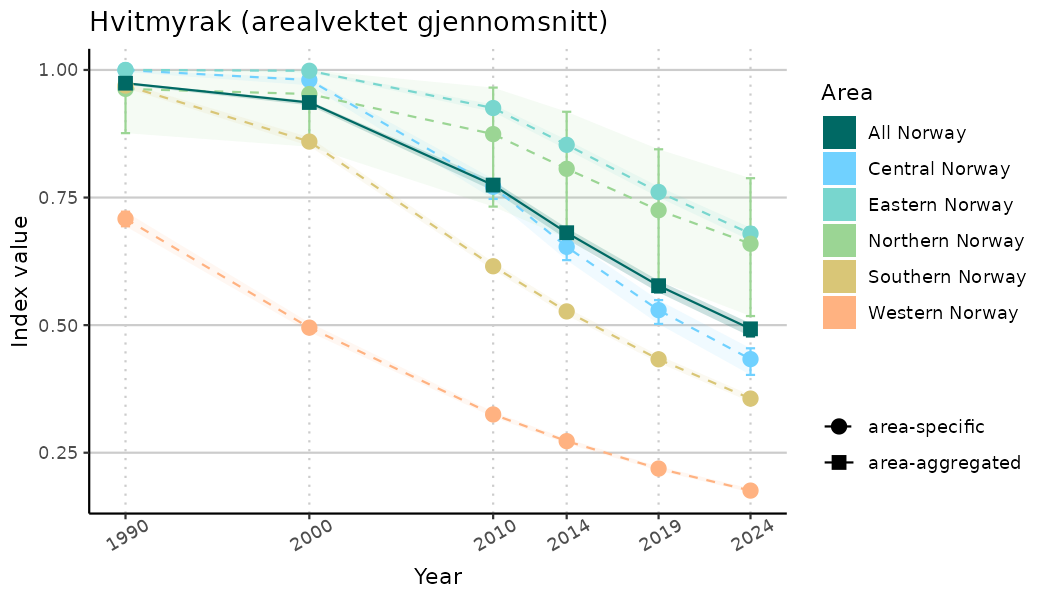
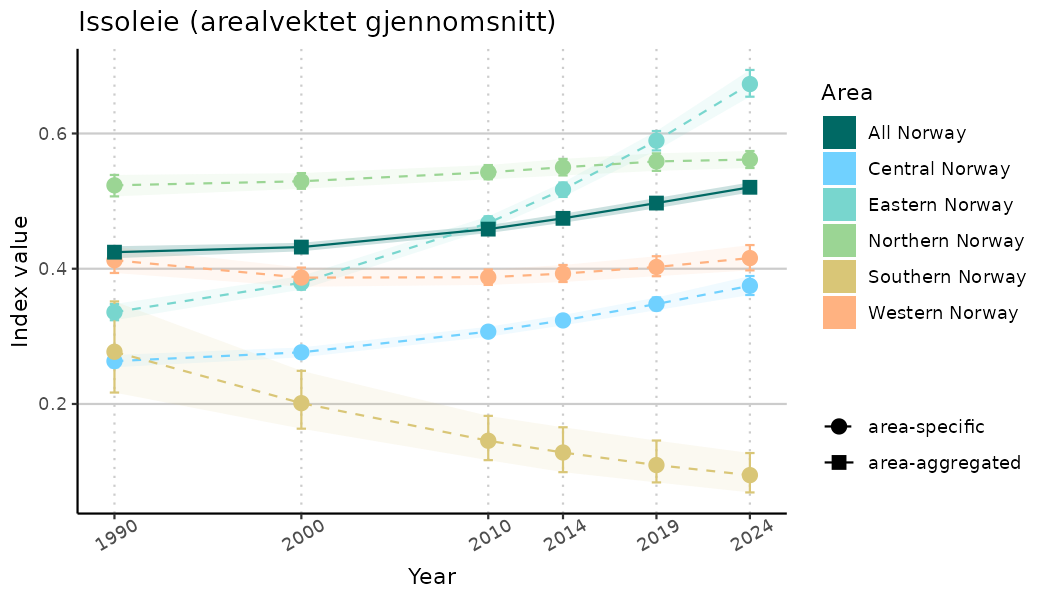
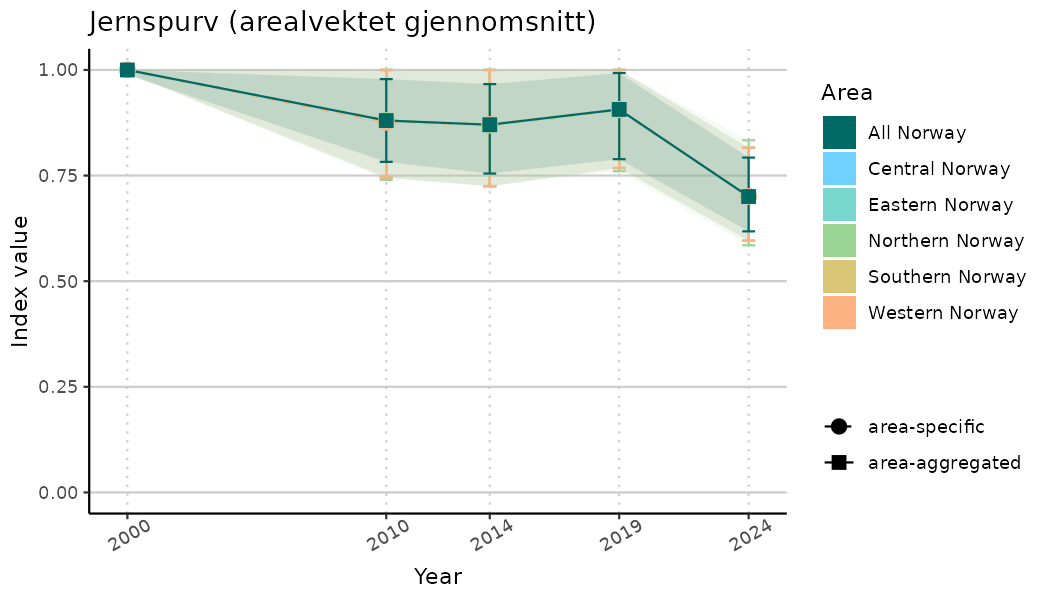
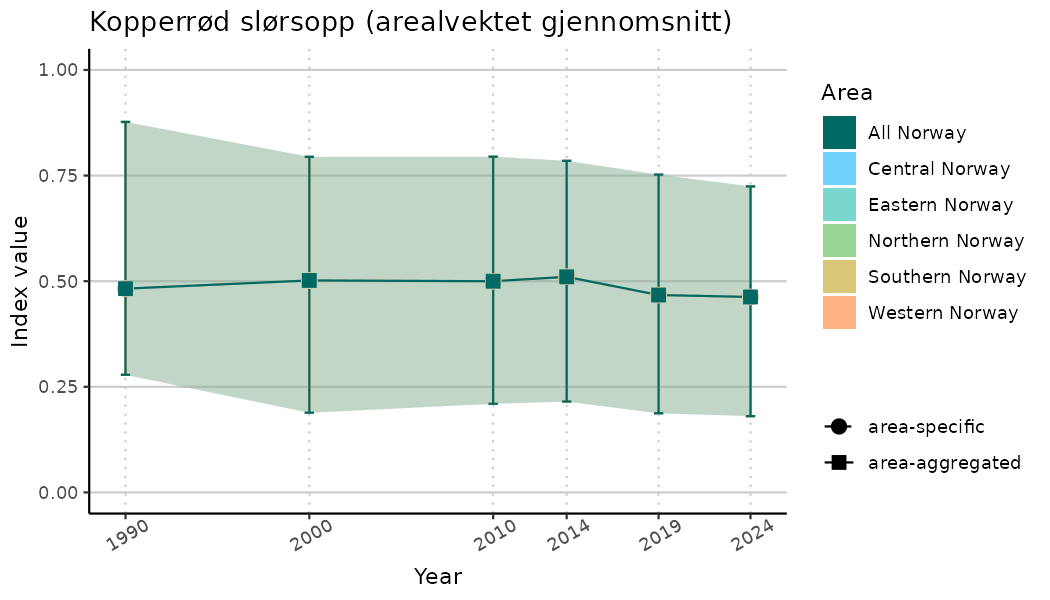
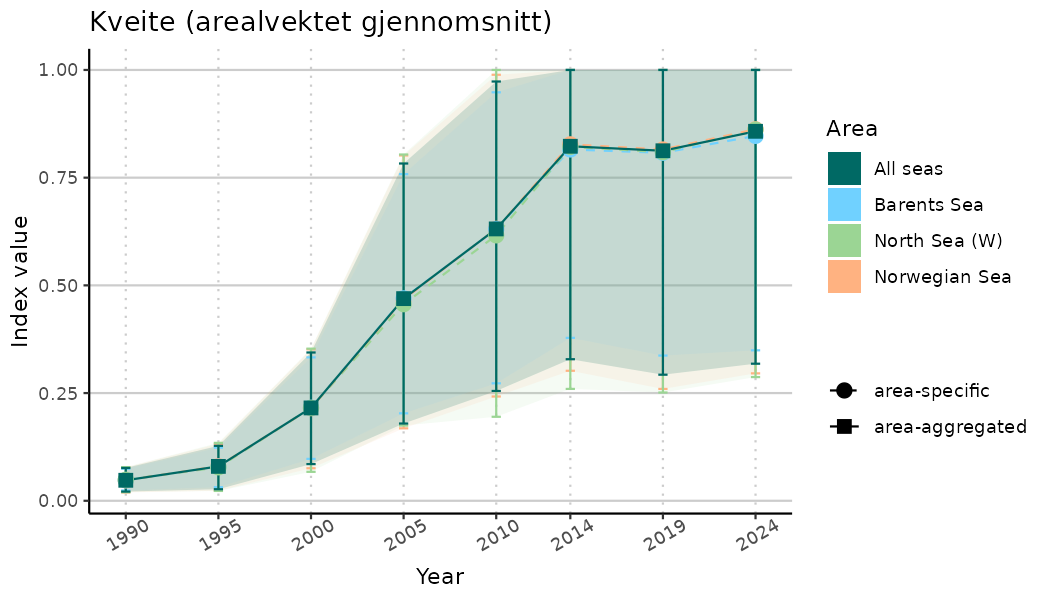
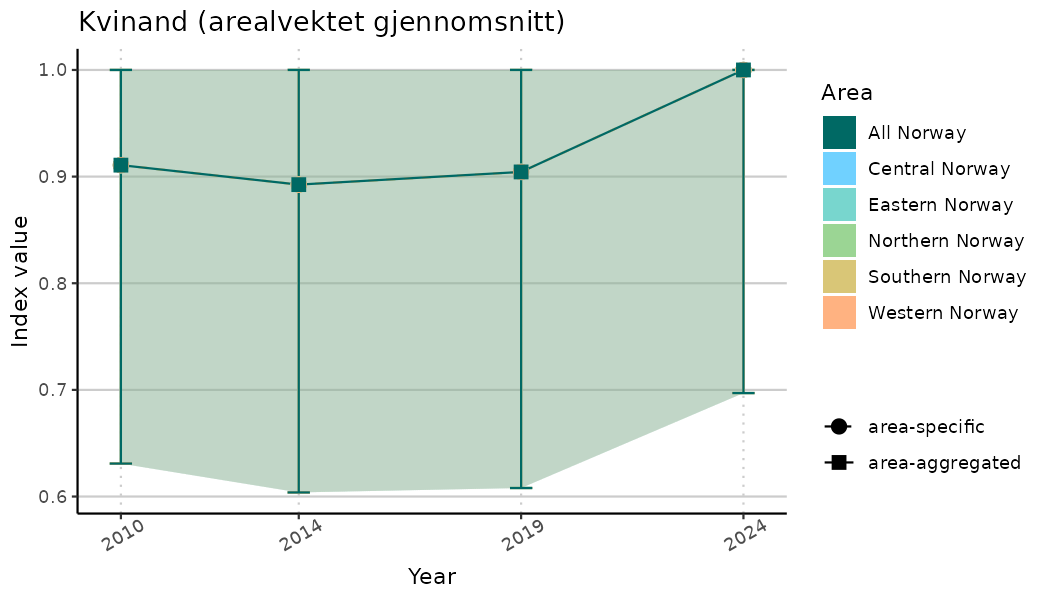
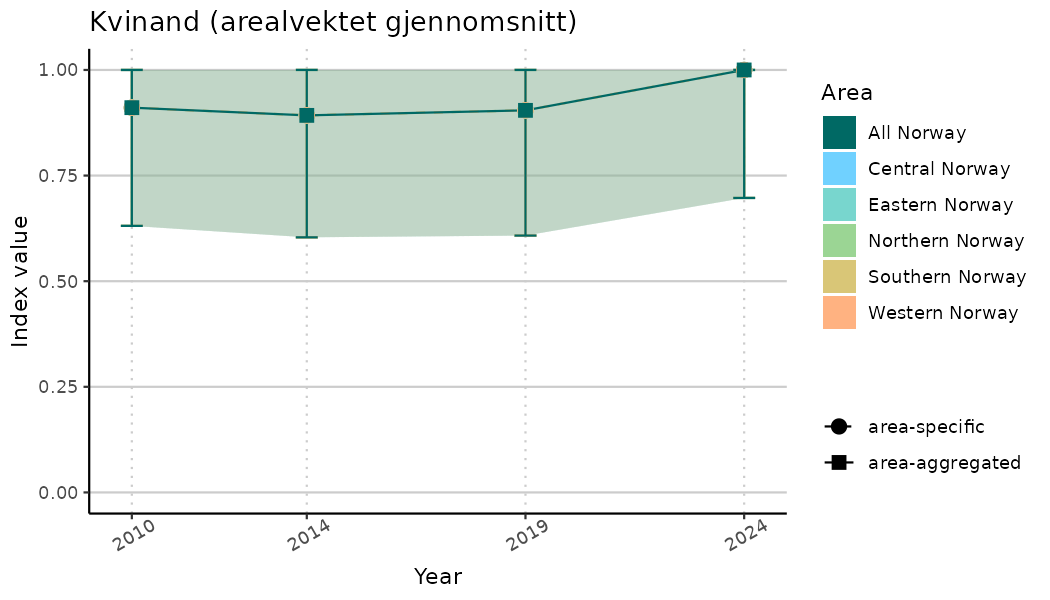
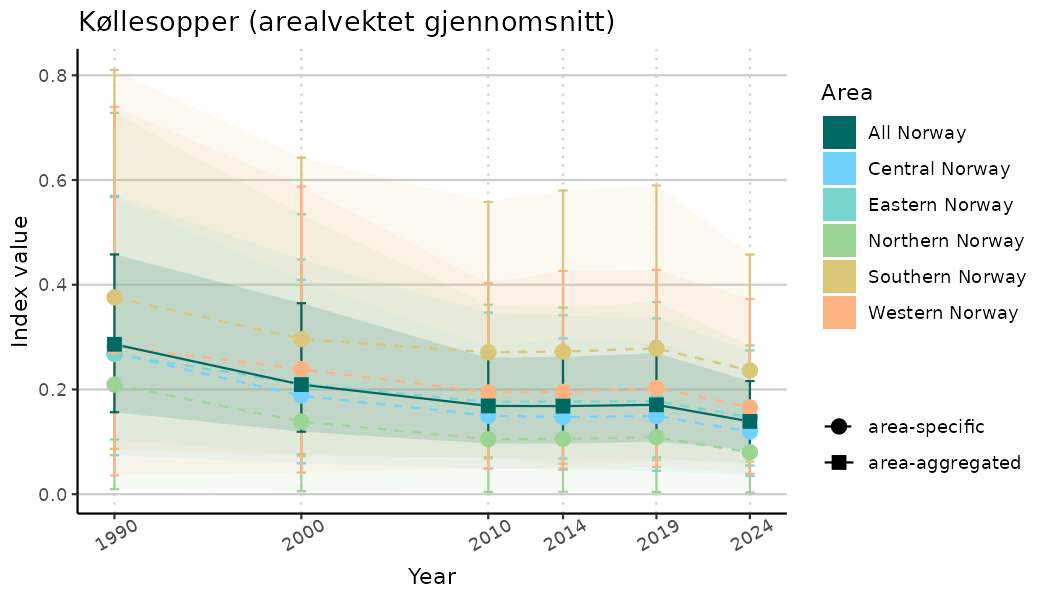
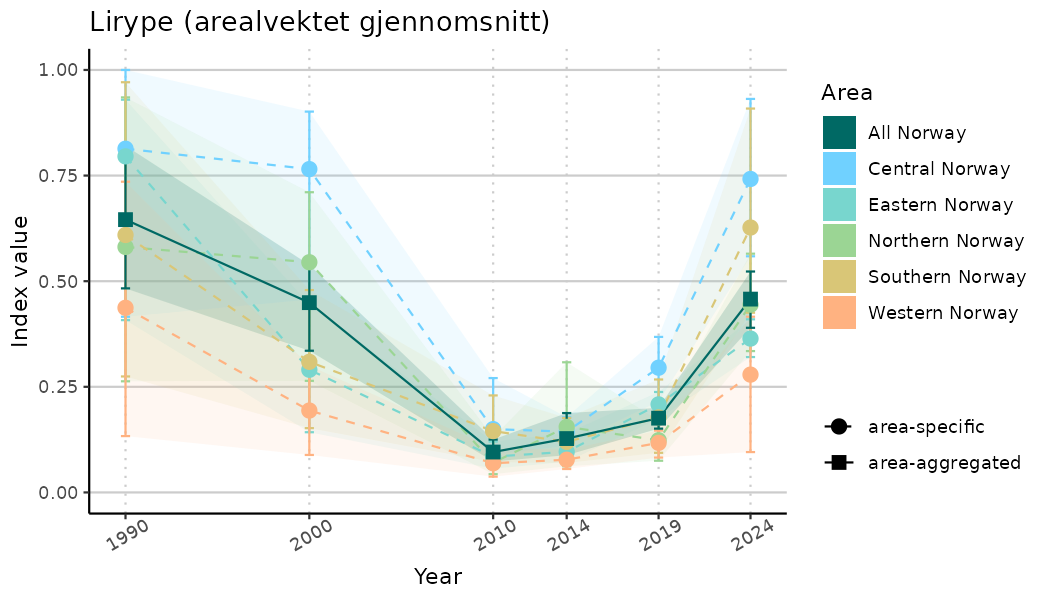
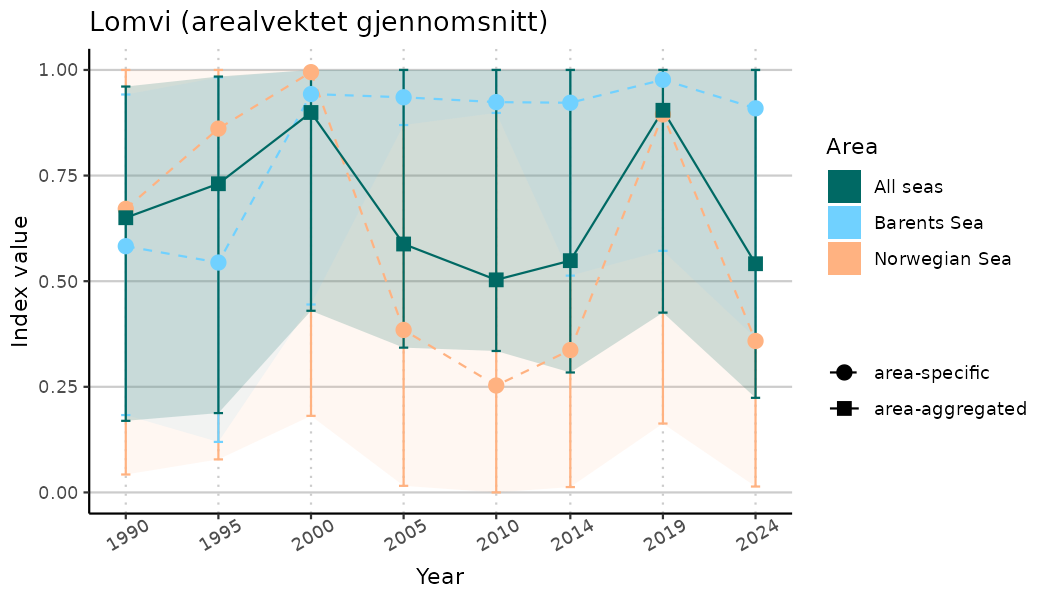
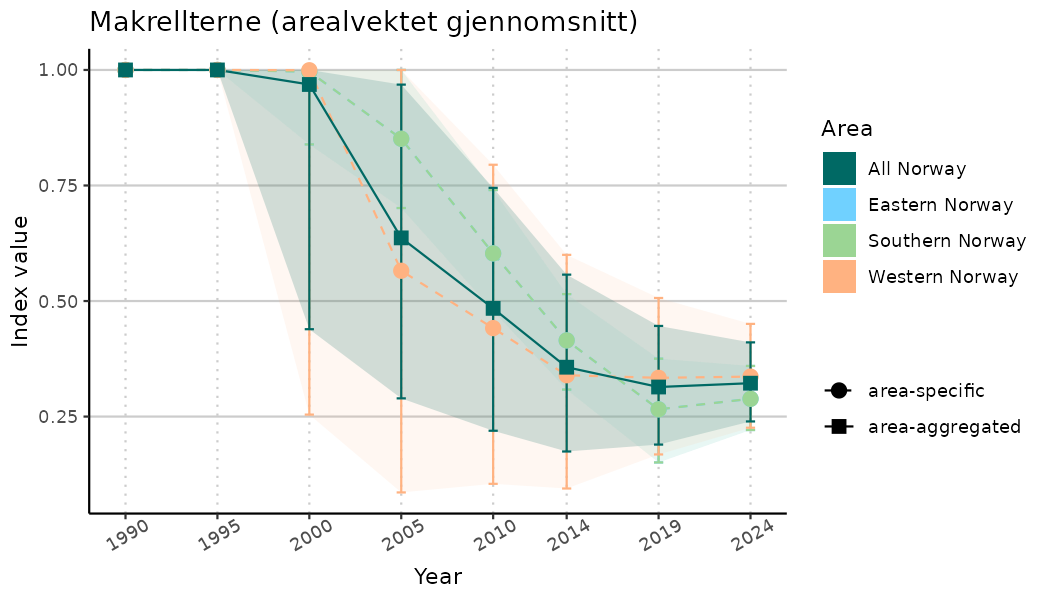
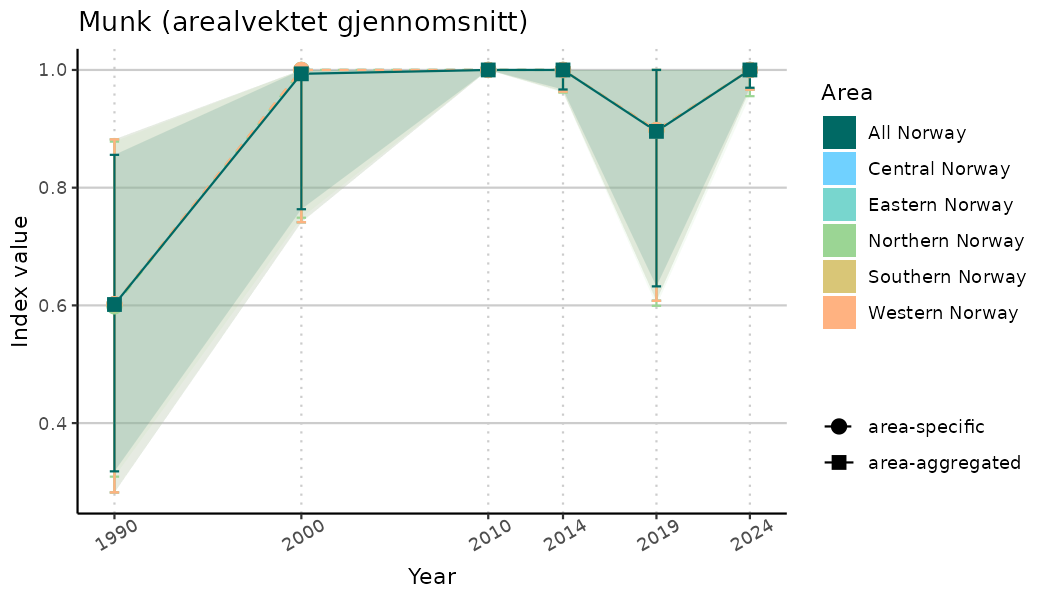
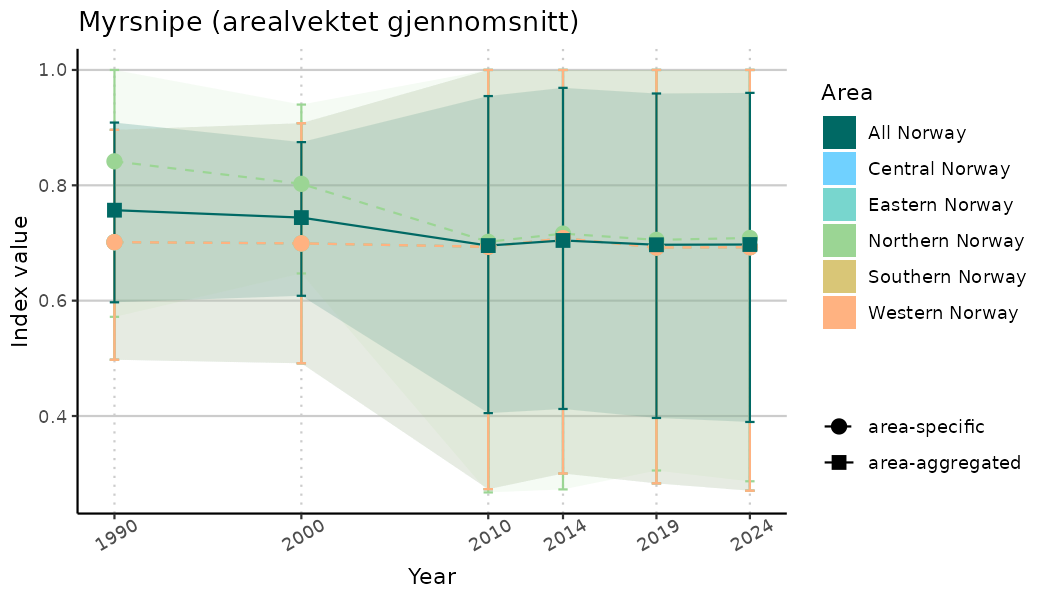
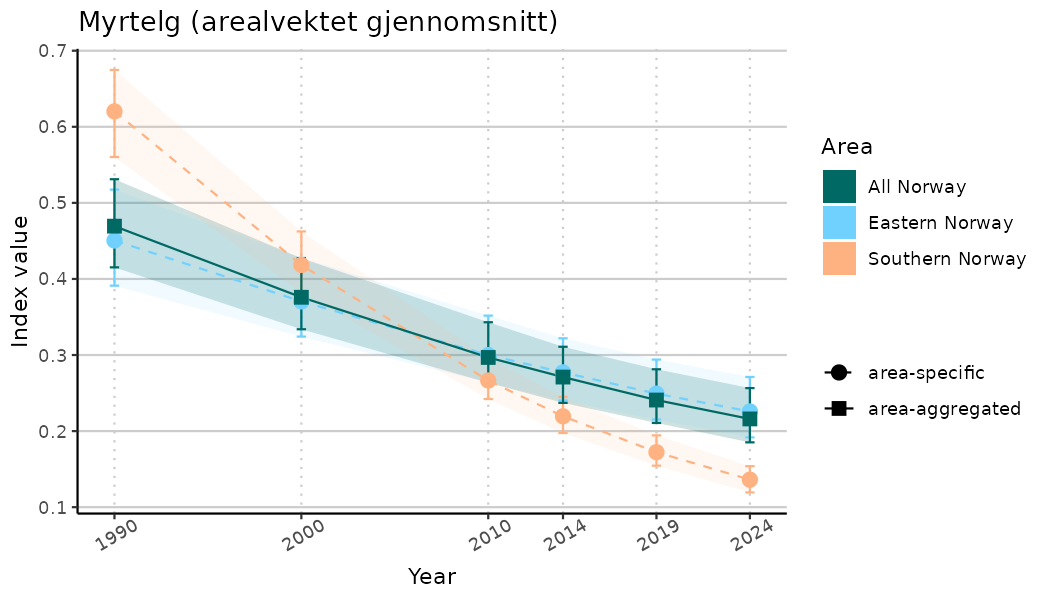
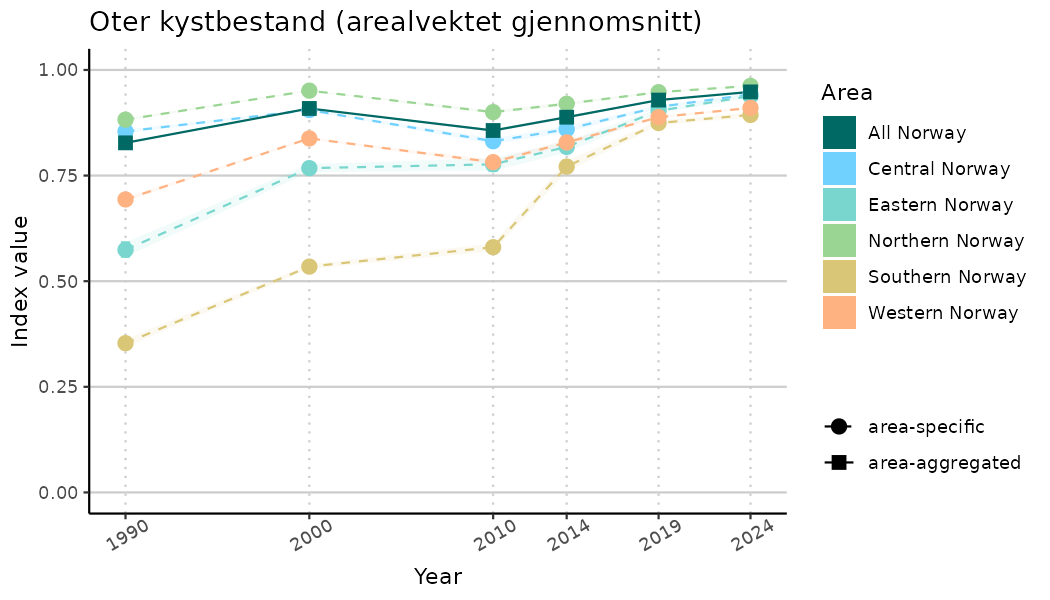
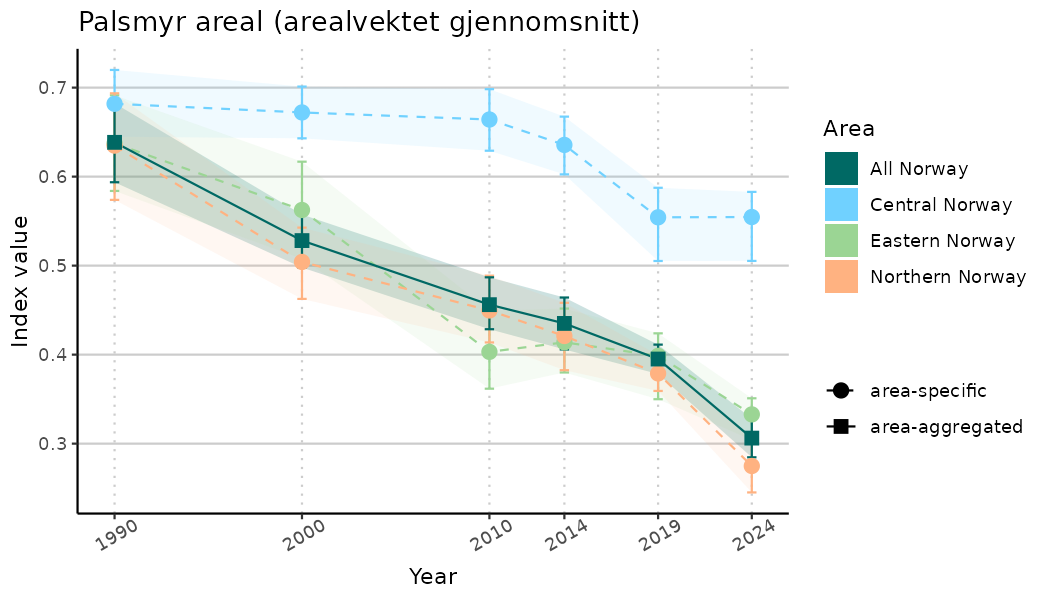
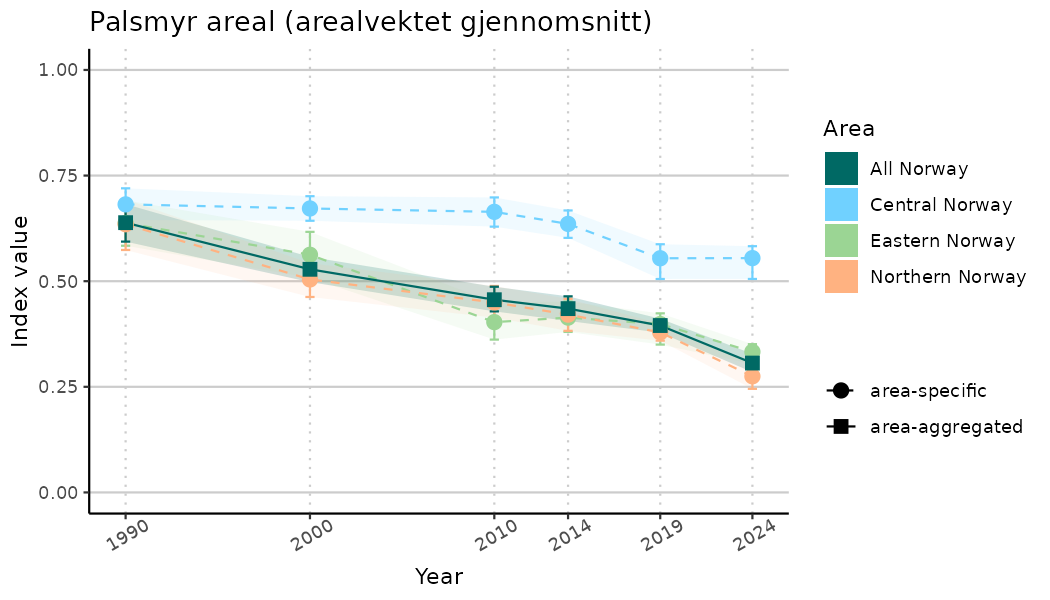
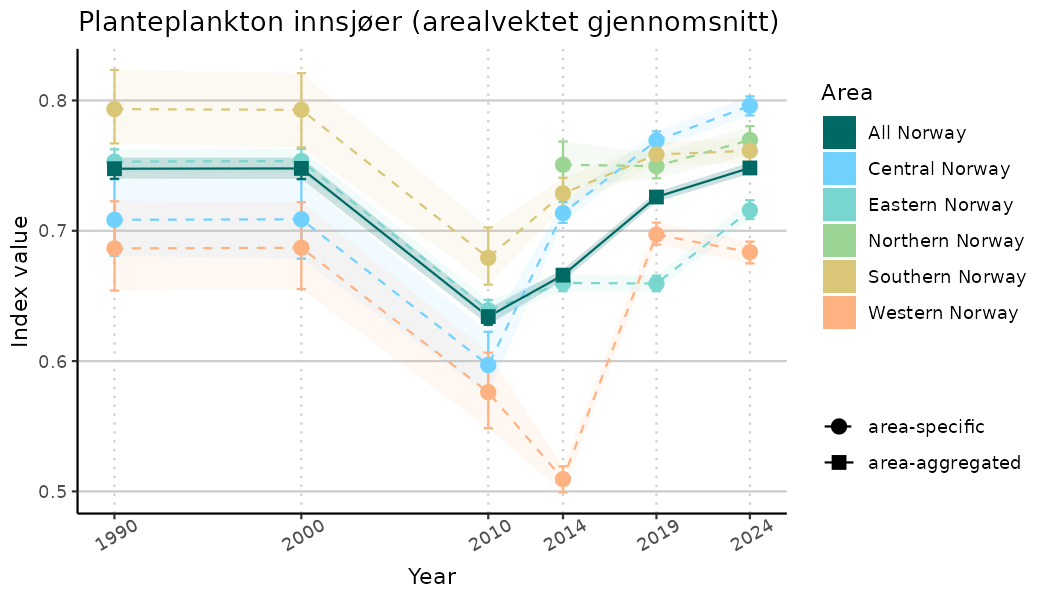
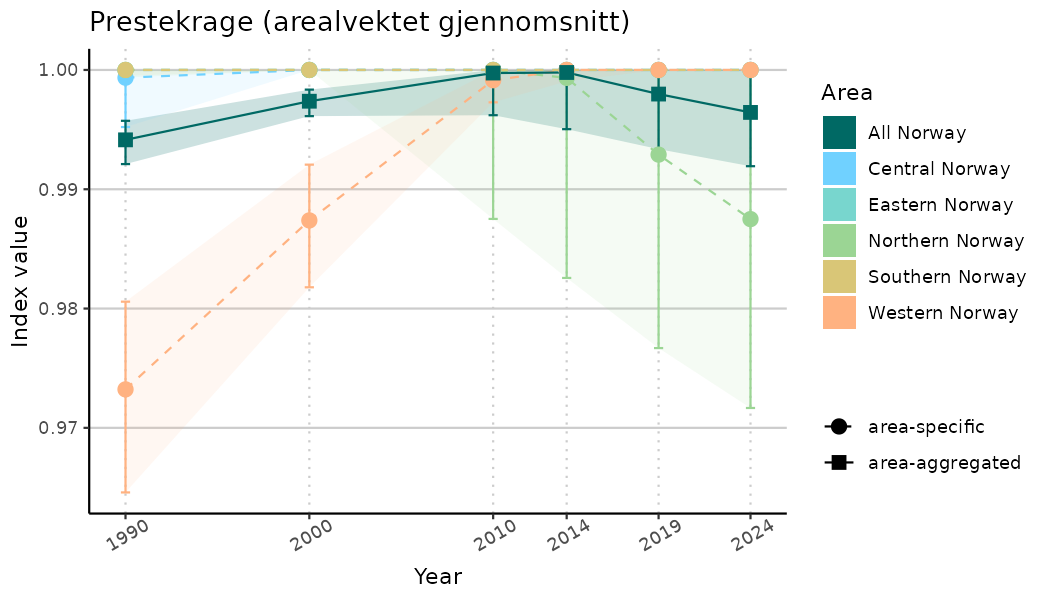
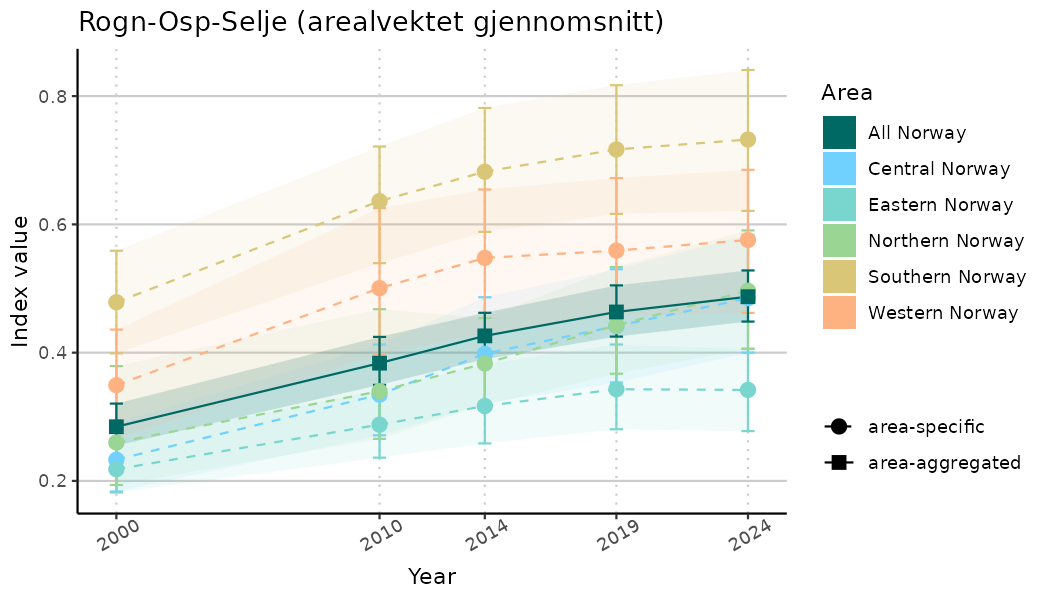
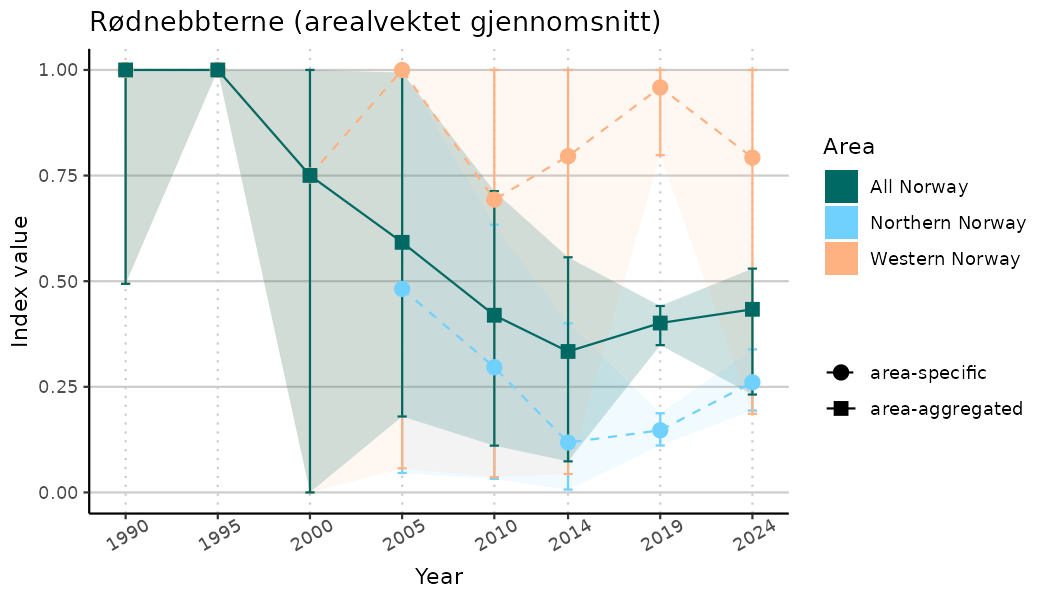
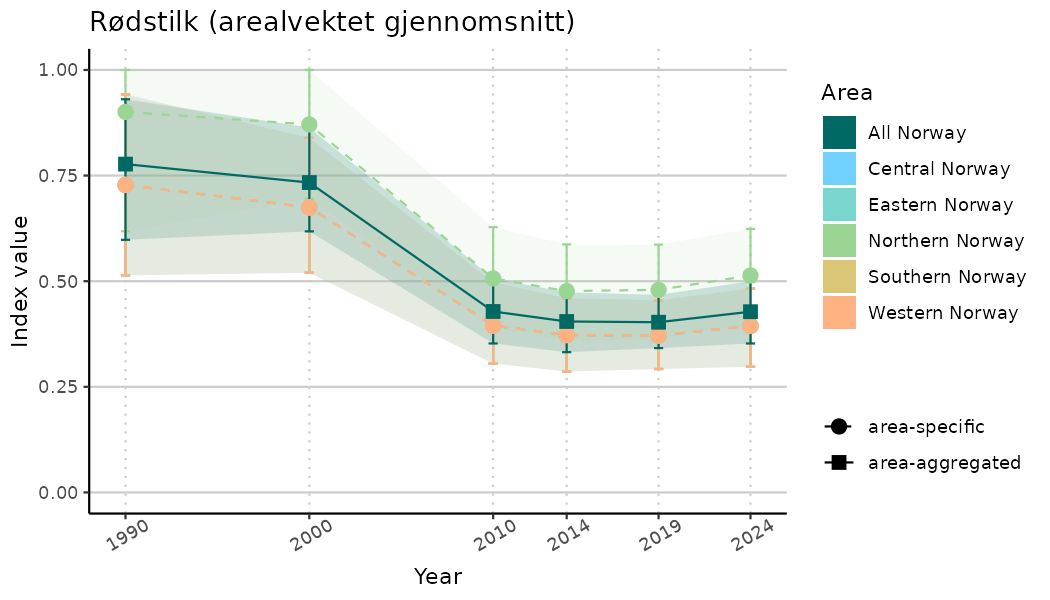
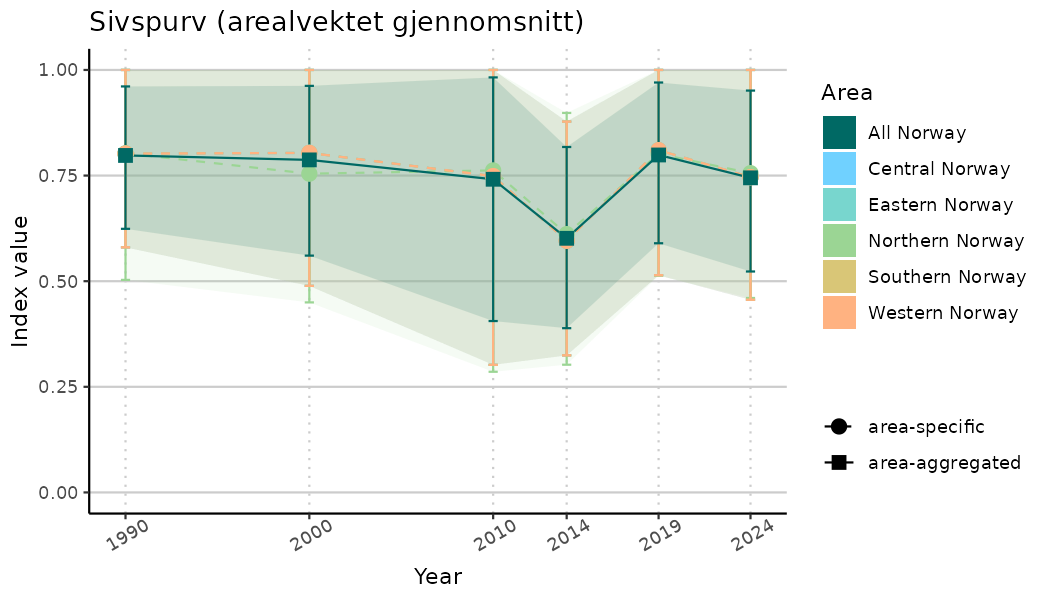
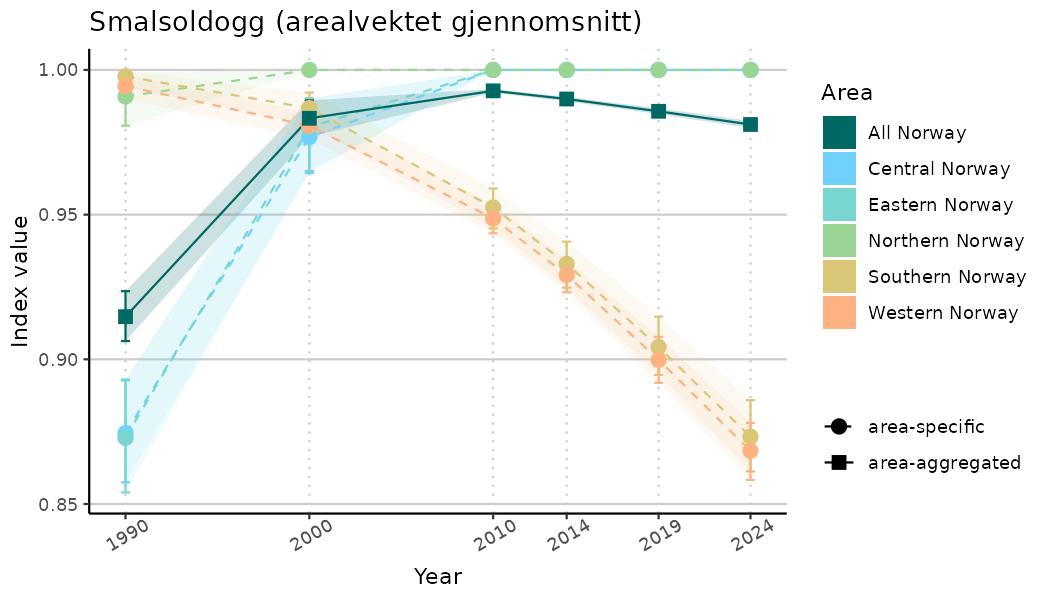
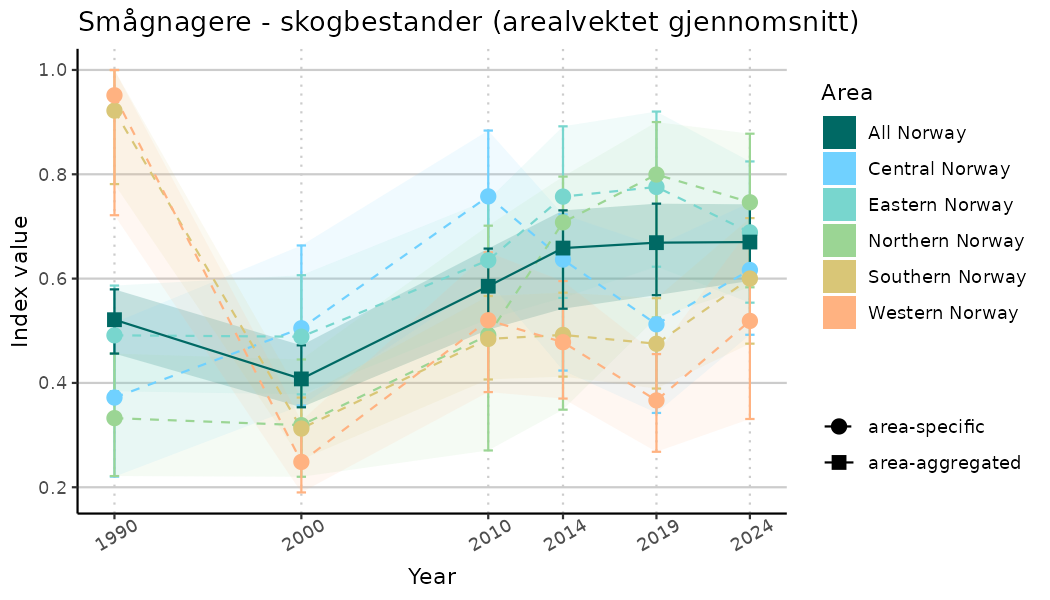
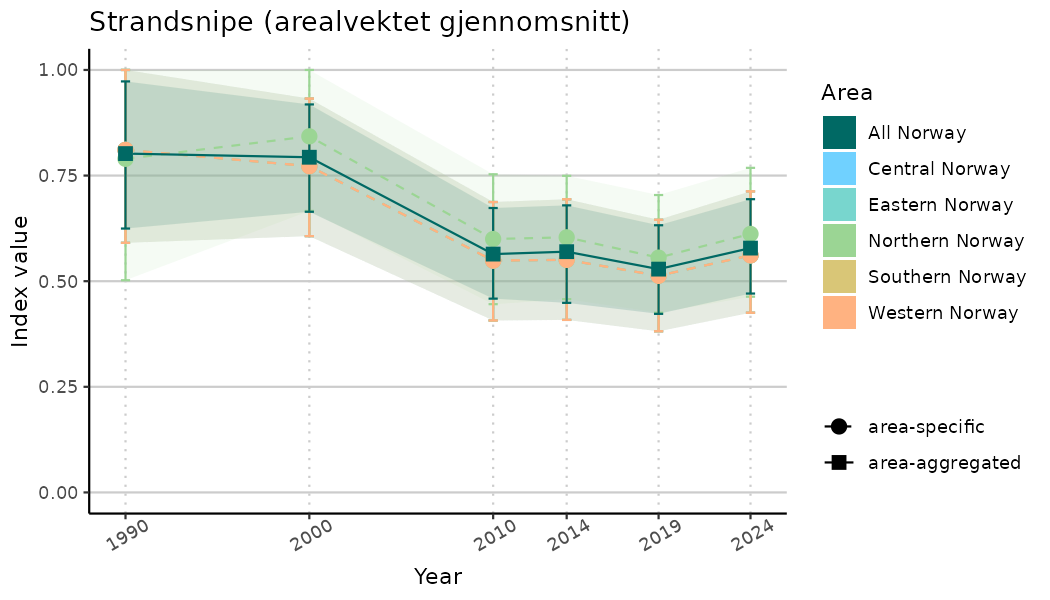
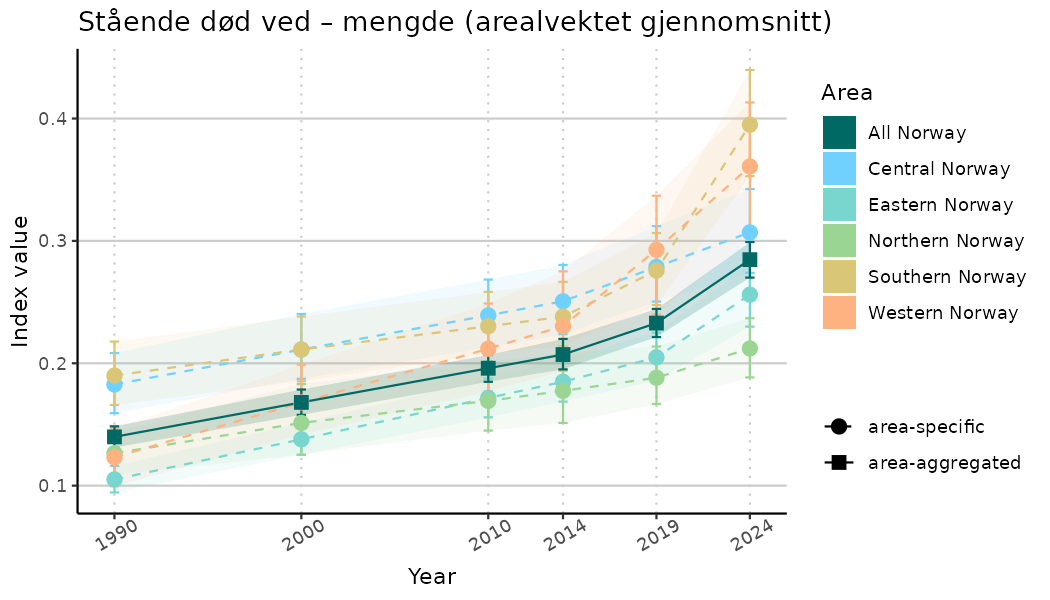
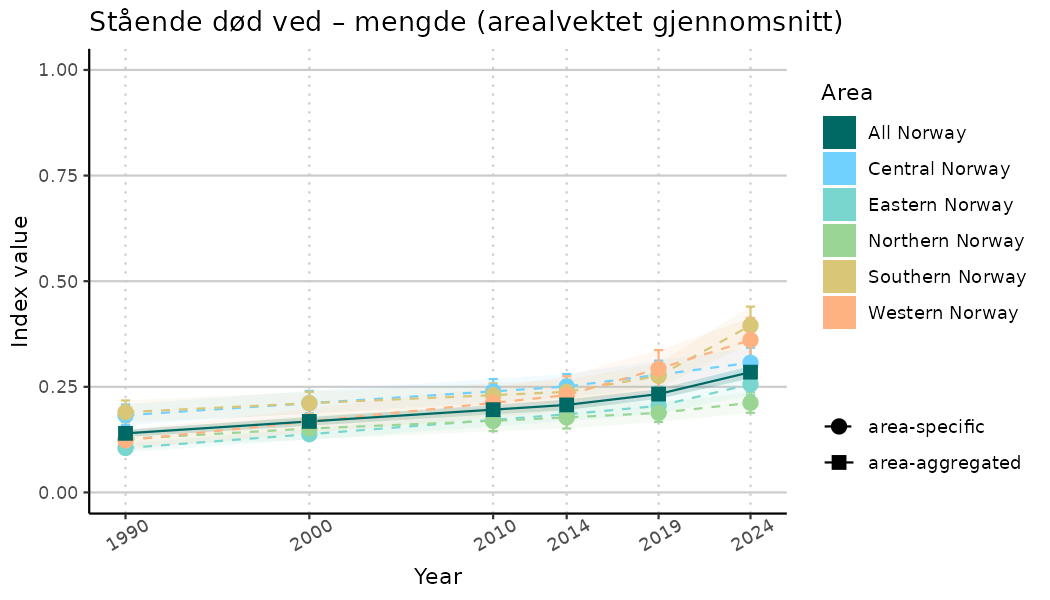
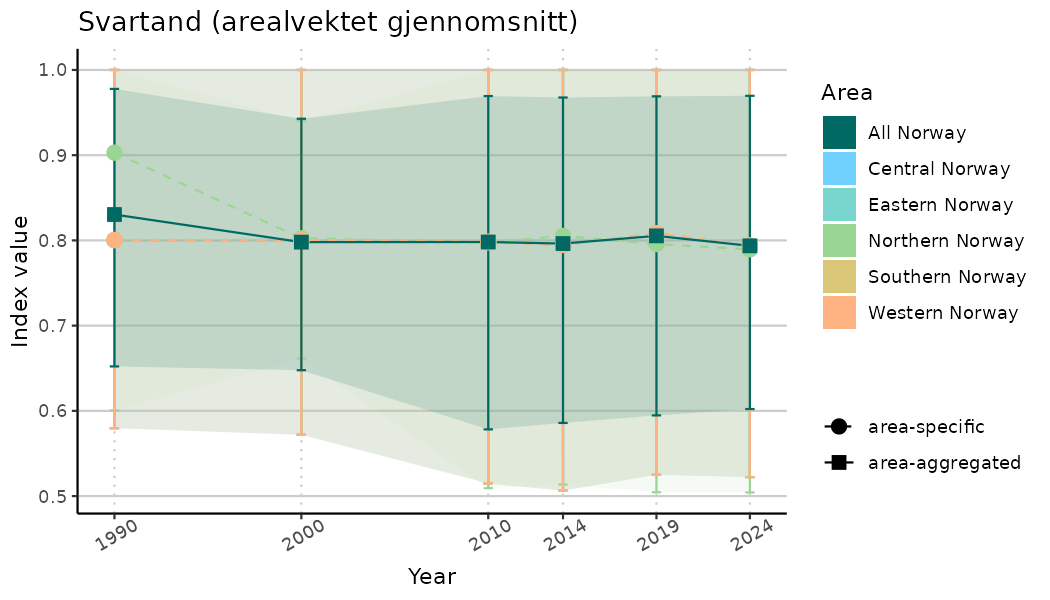
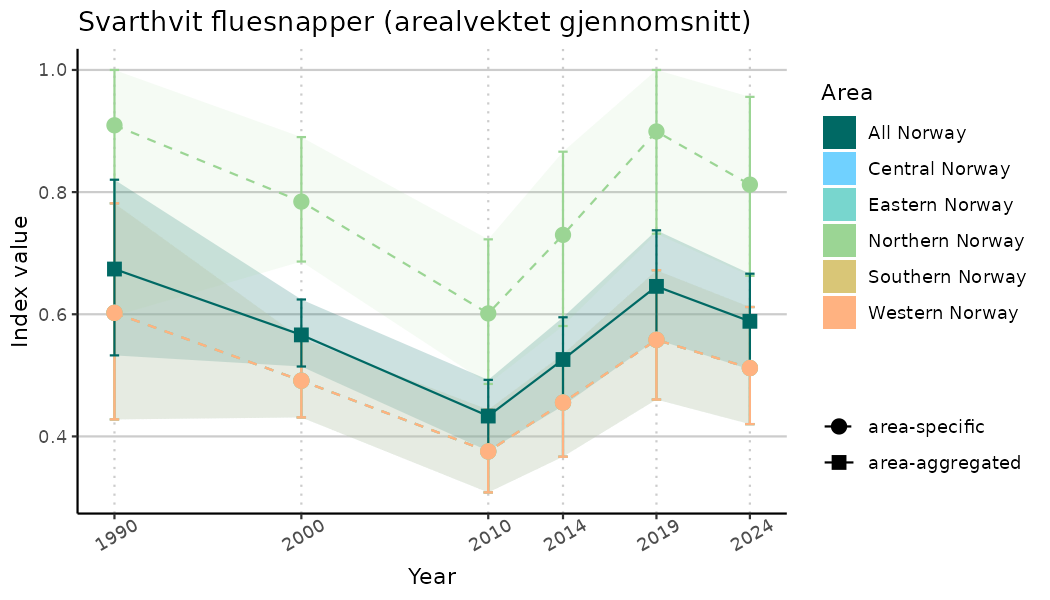
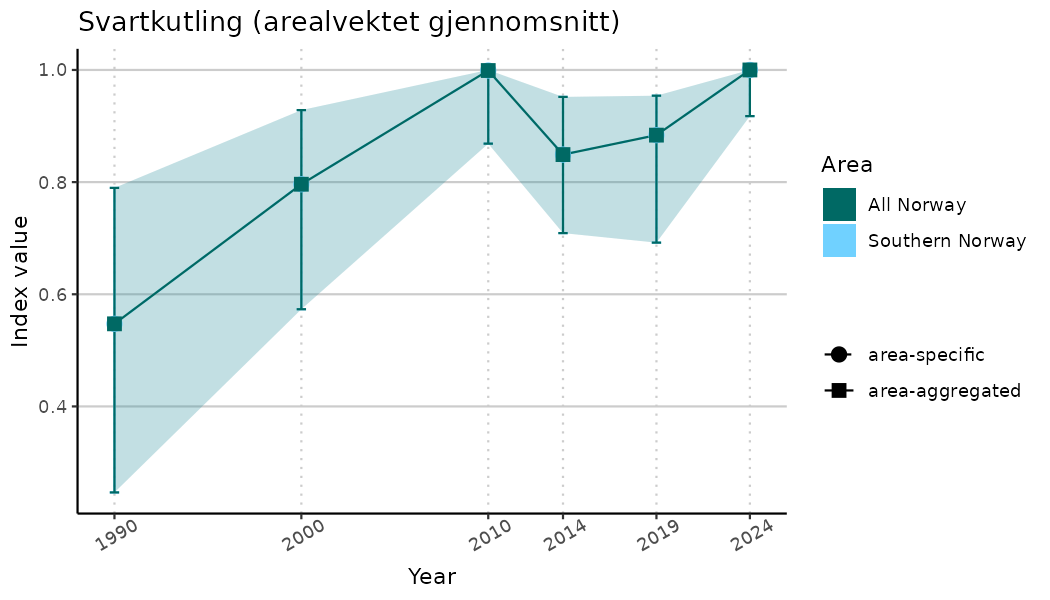
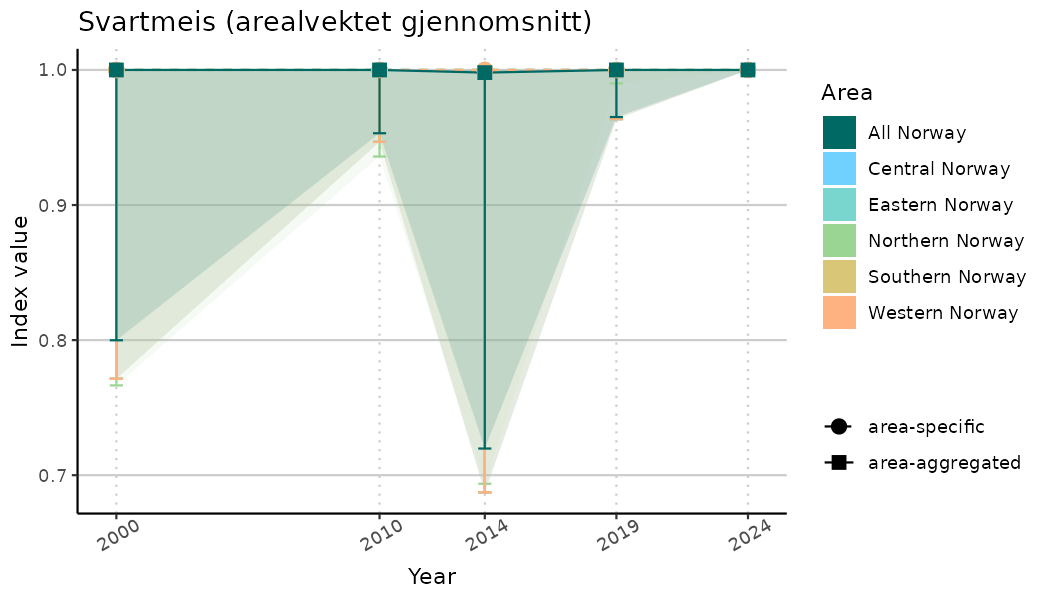
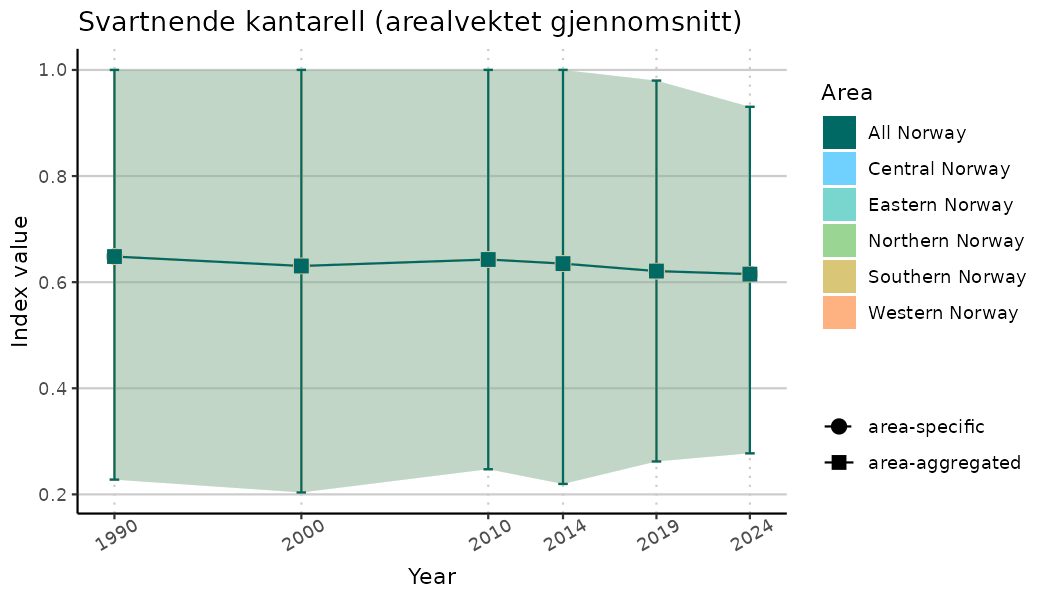
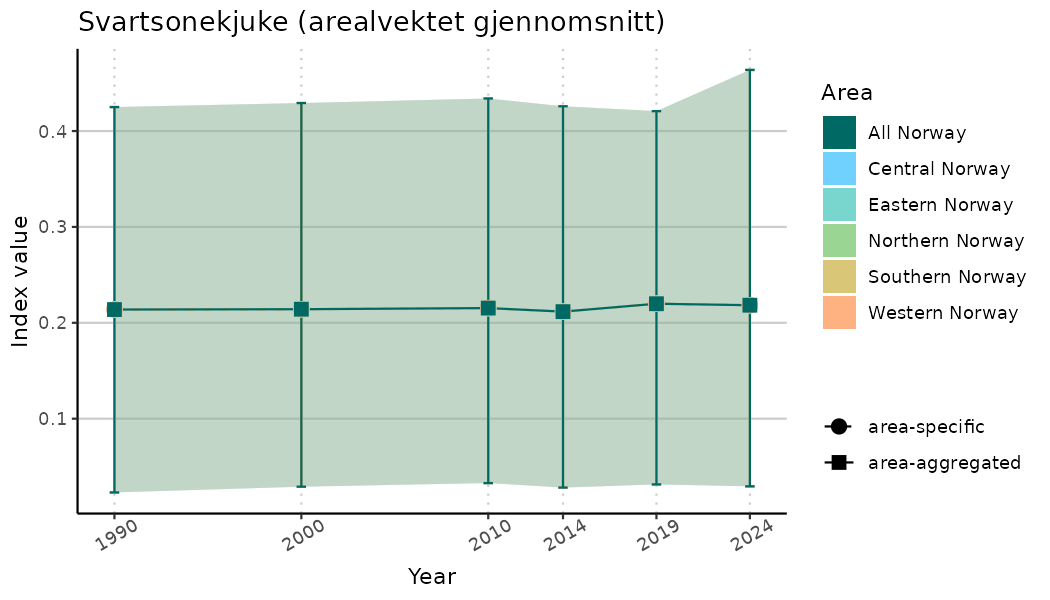
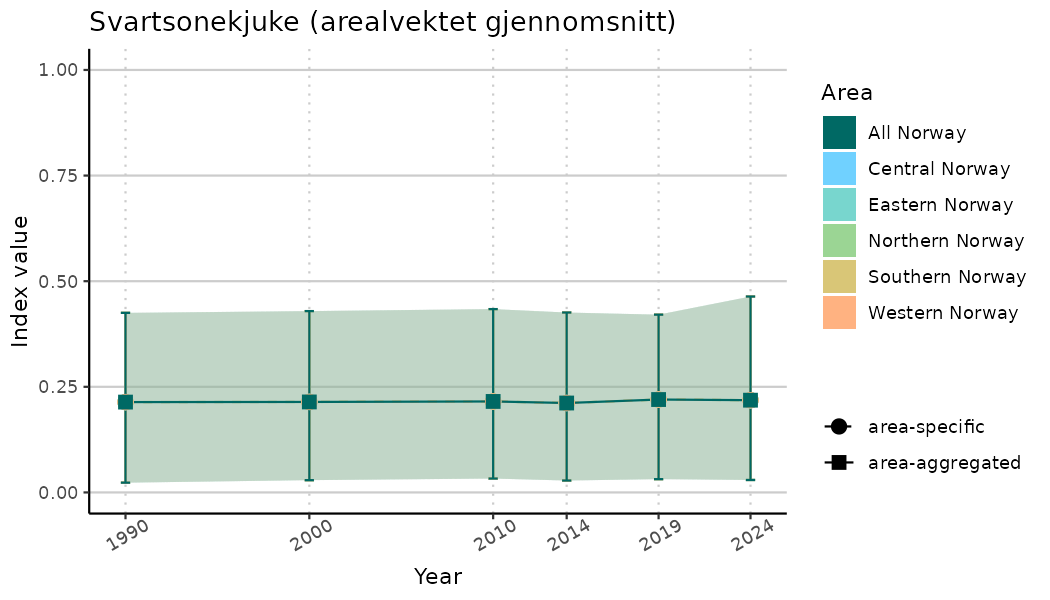
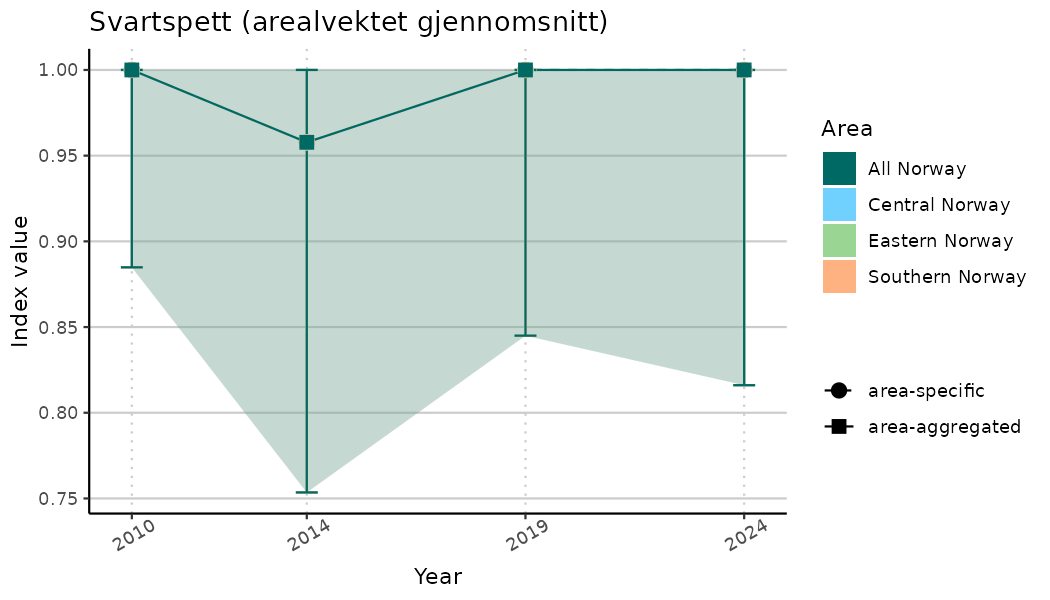
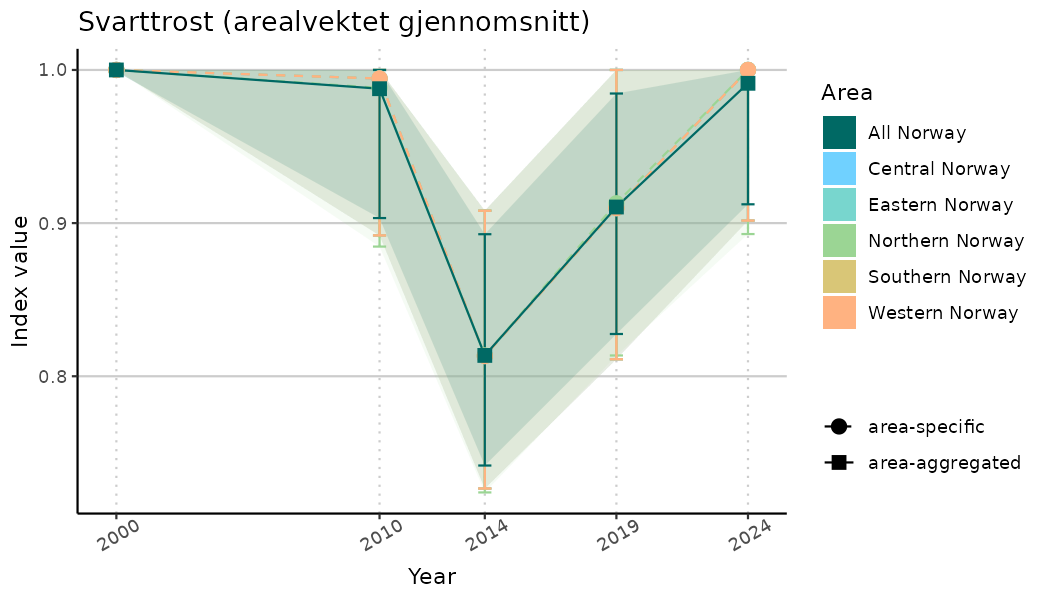
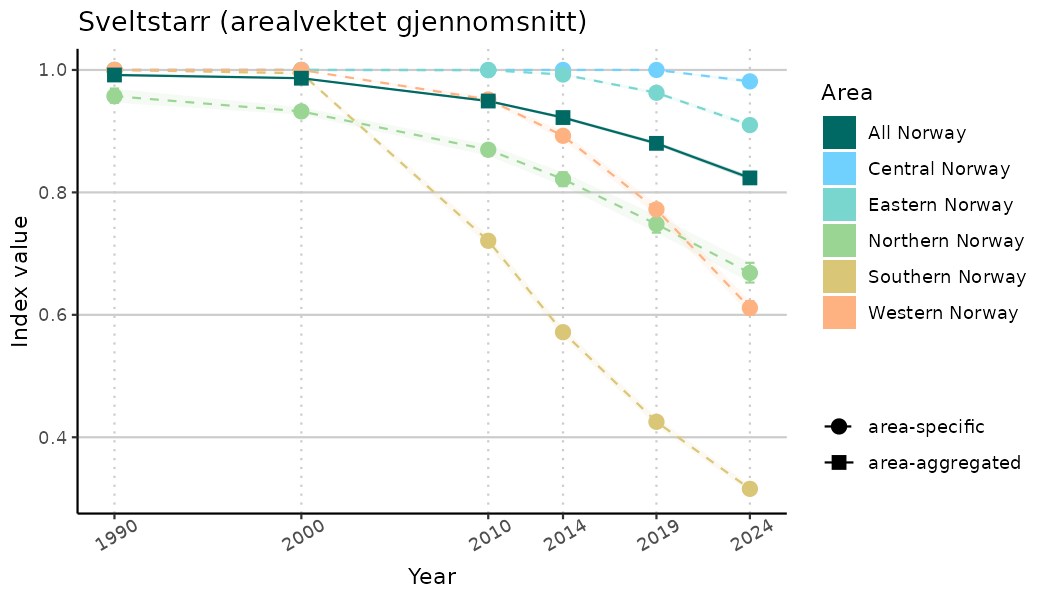
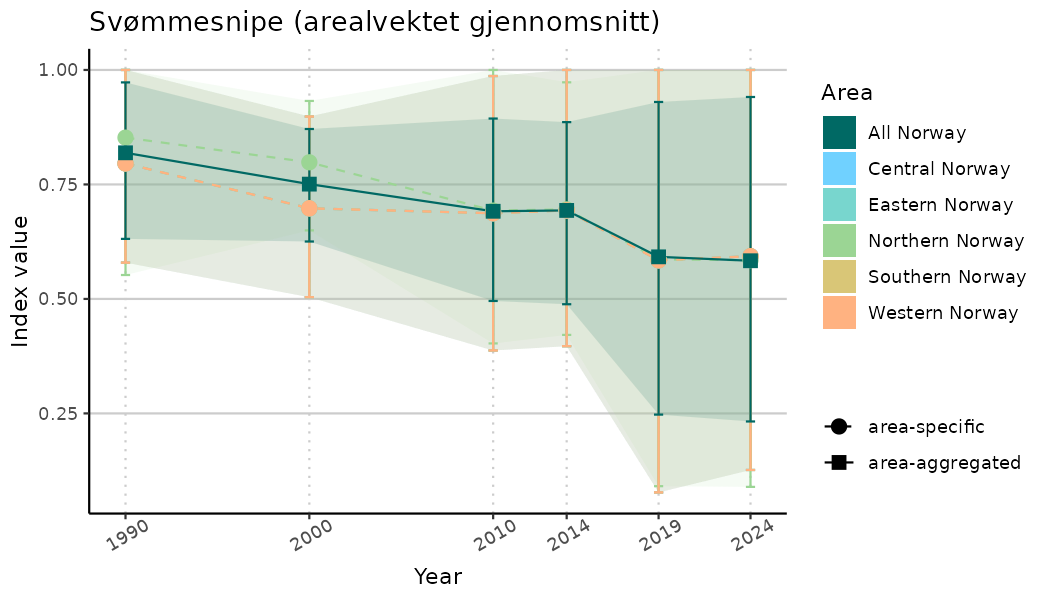
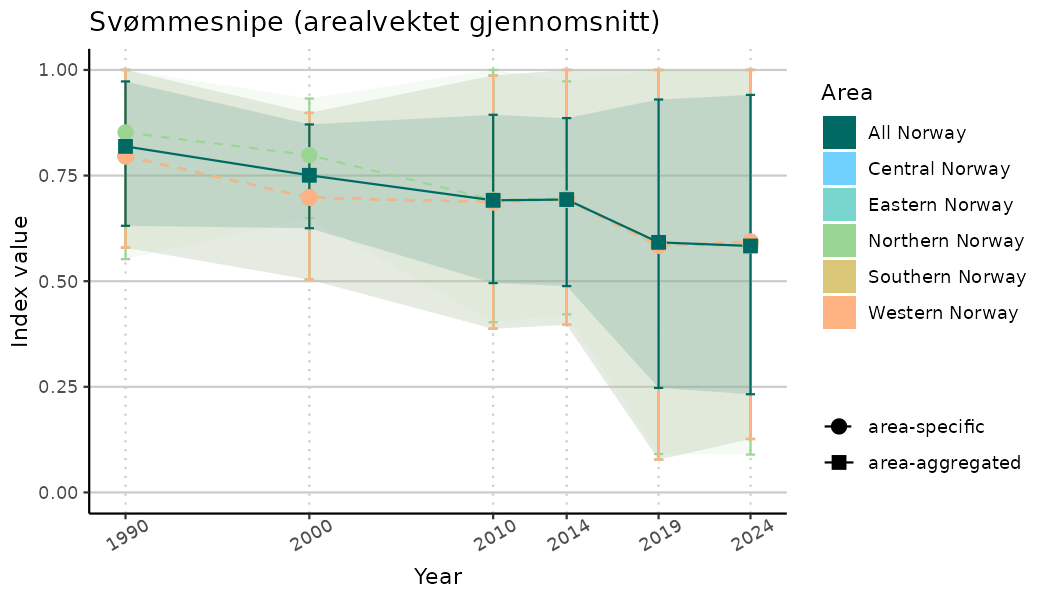
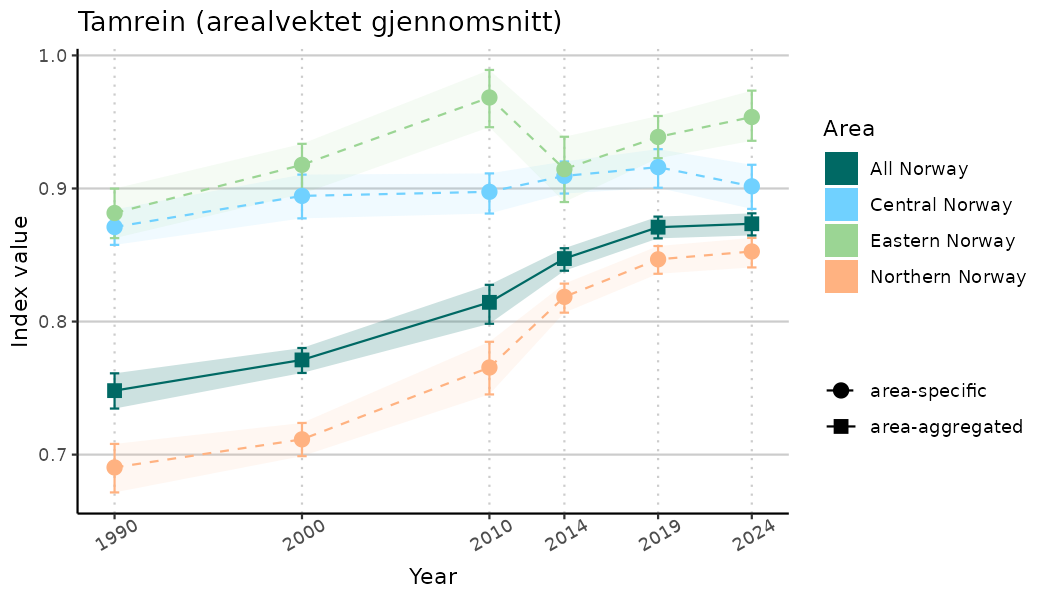
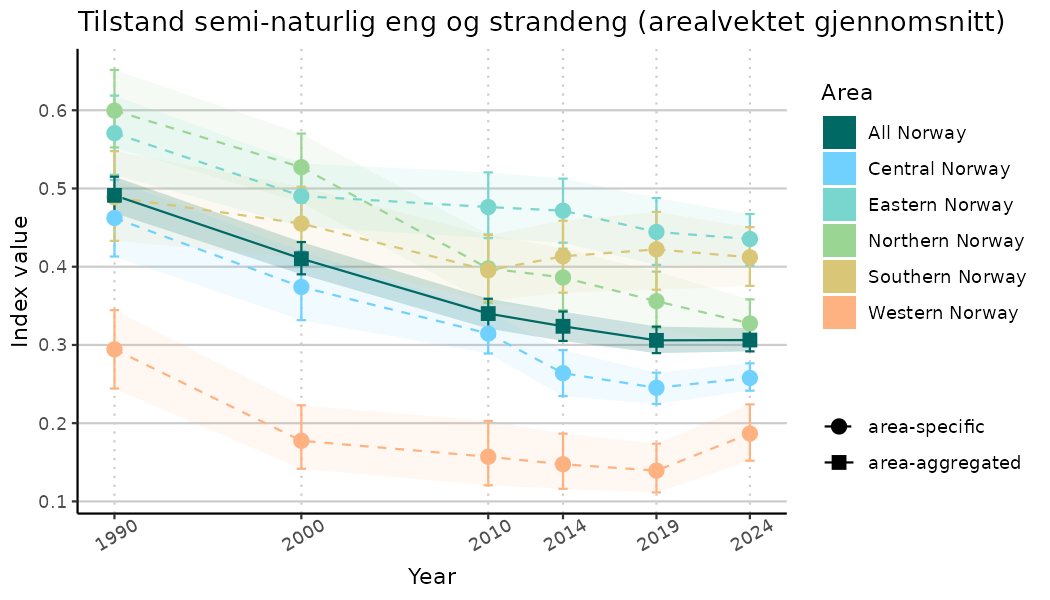
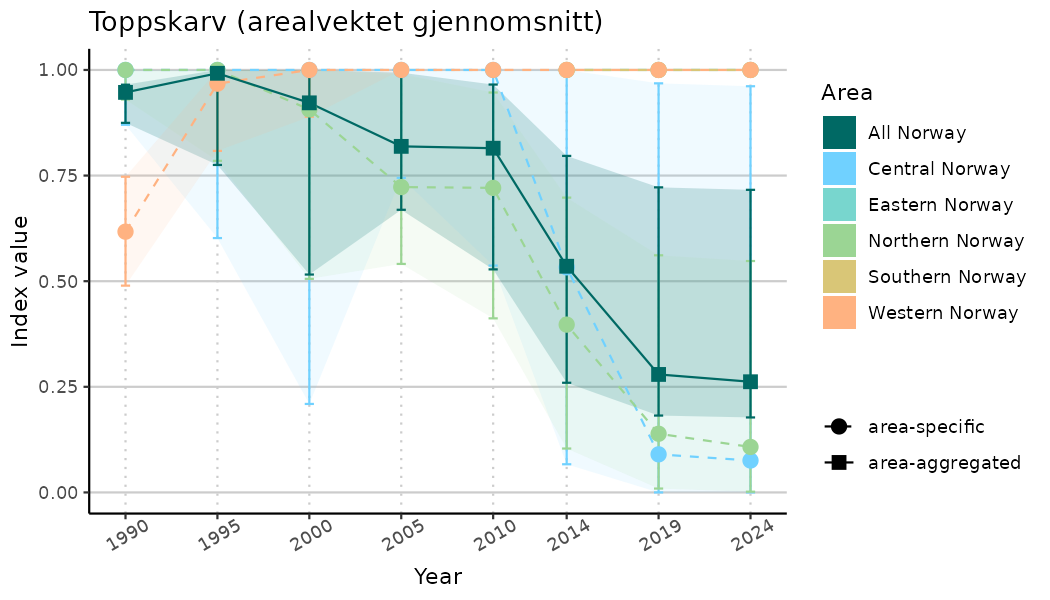
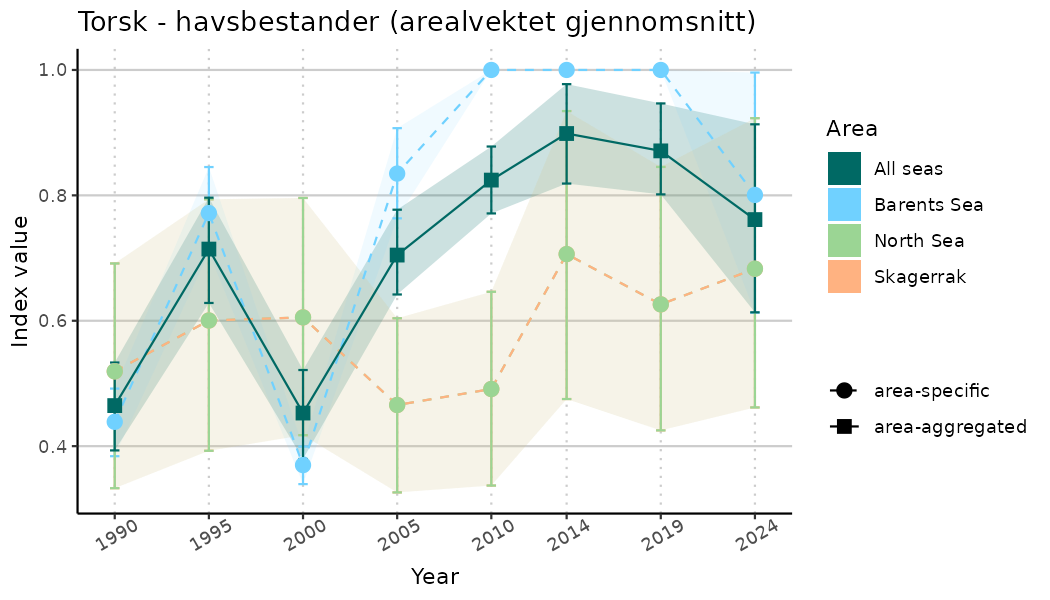
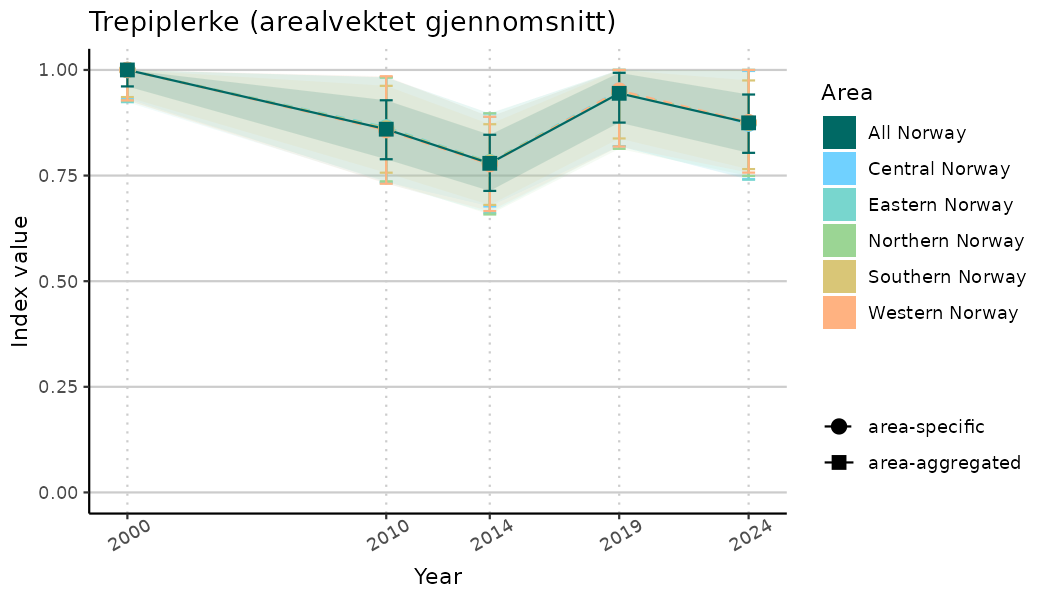
ts_indIdx <- NIflex::plotNI_StandardTS(Index = output_indIdx$indexOutput,
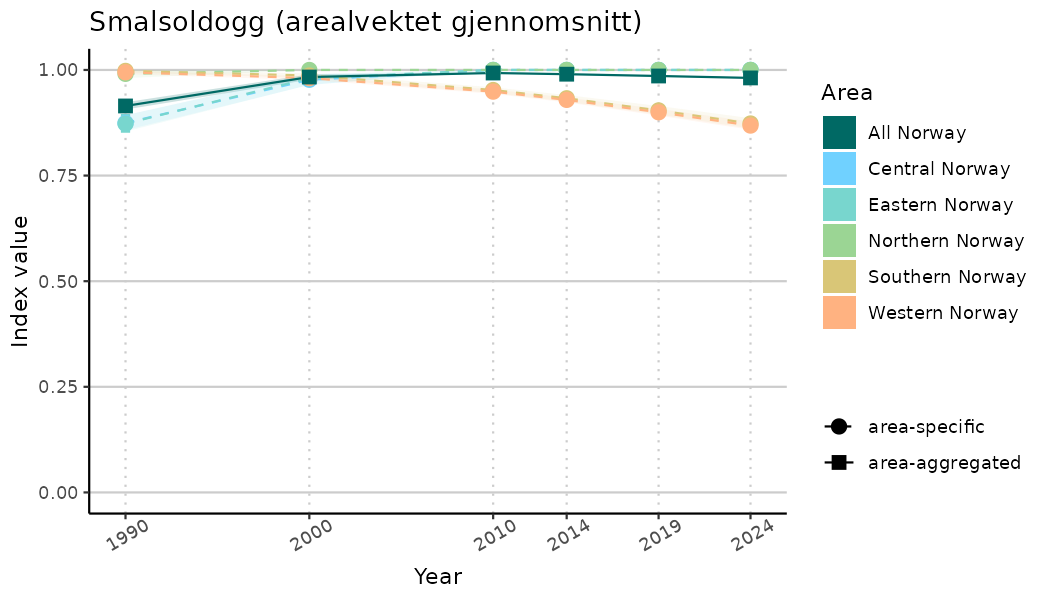
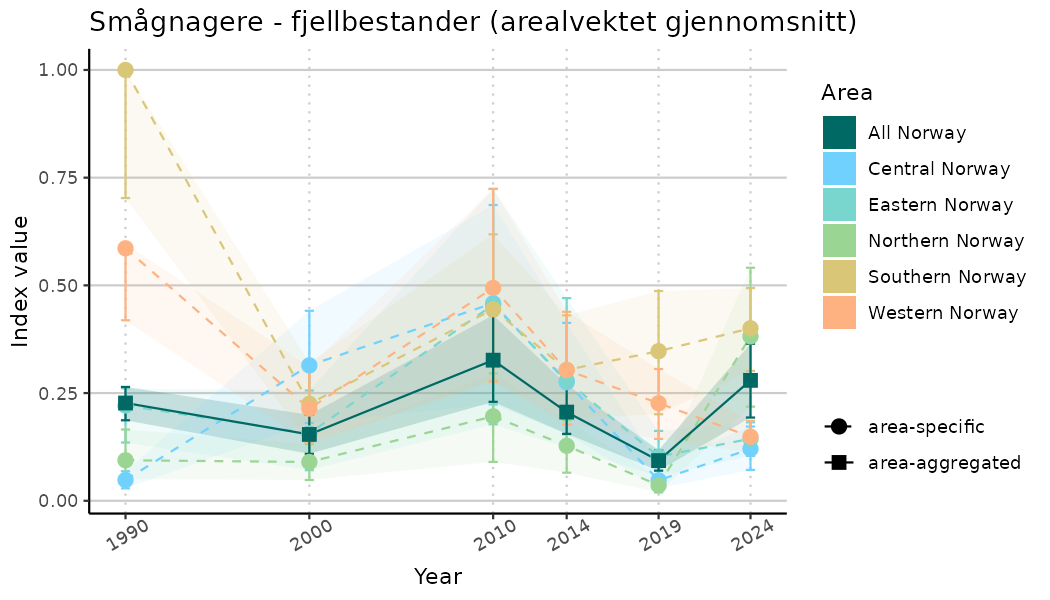
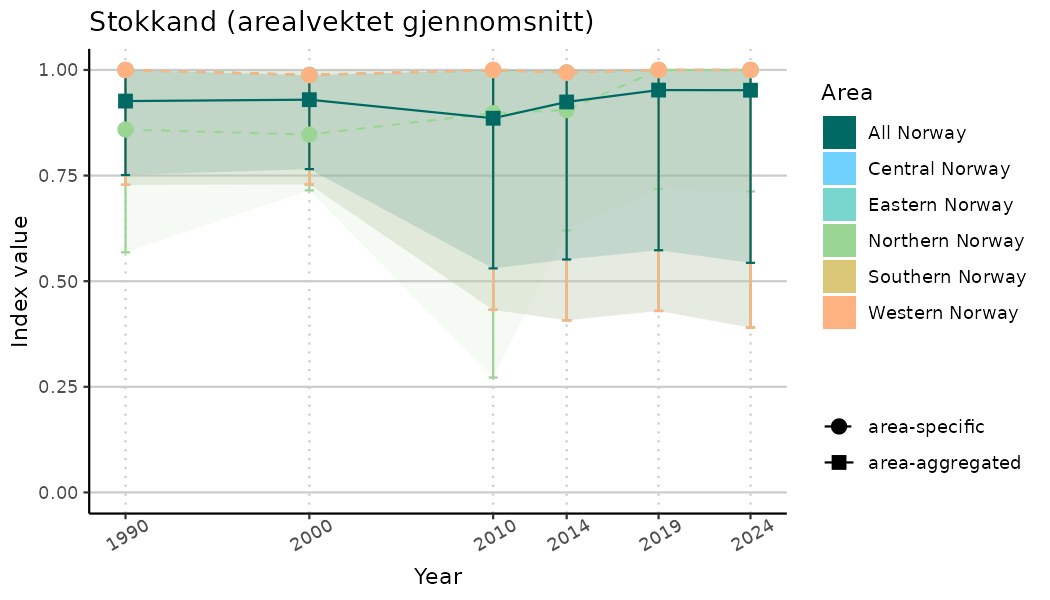
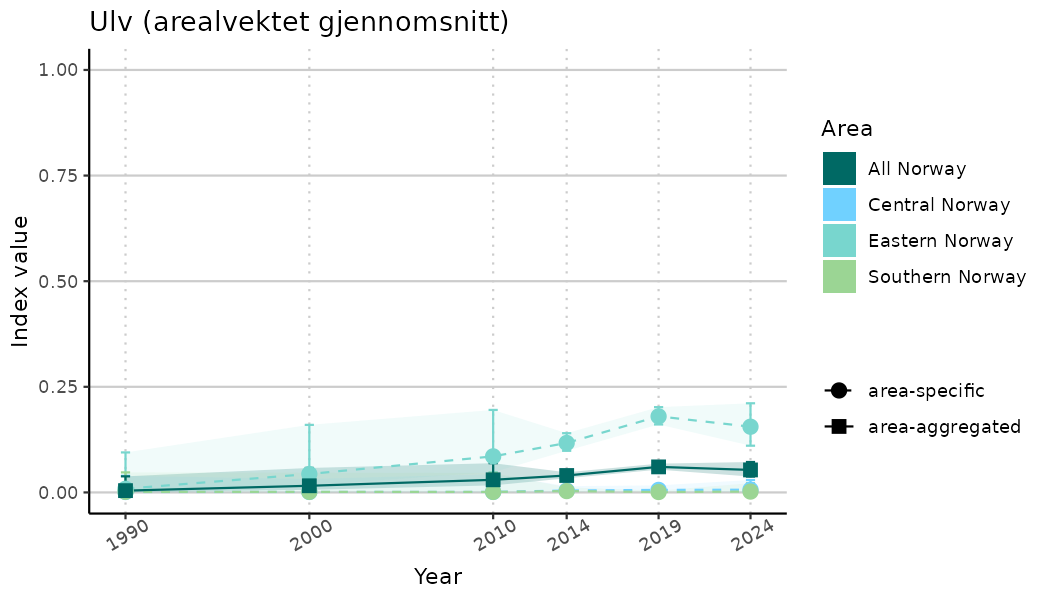
plotTitle = paste0(ind_name, " (arealvektet gjennomsnitt)"),
plotYears = ind_focalYears,
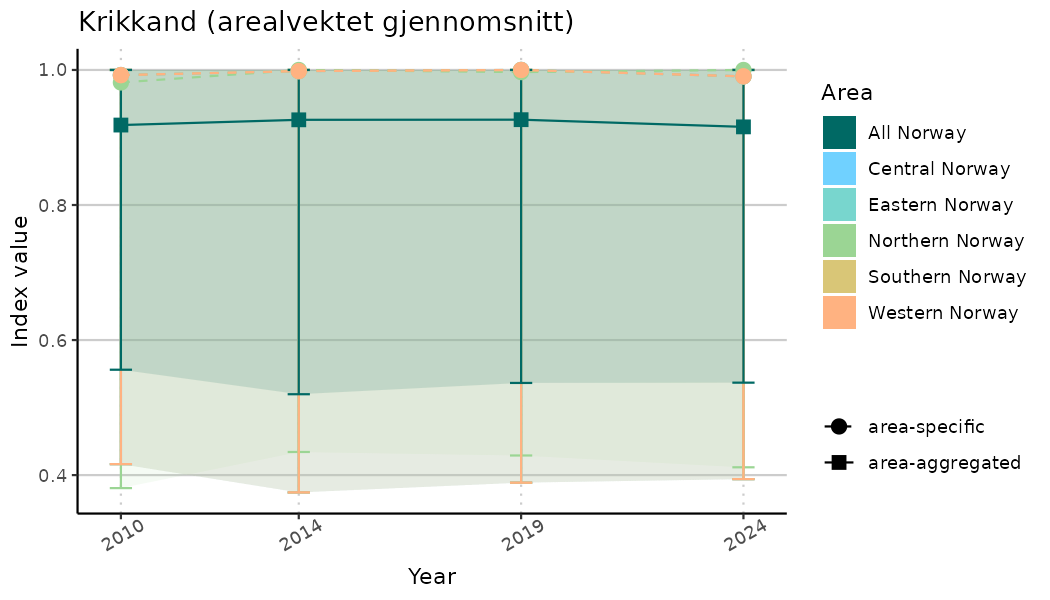
addAverage = TRUE,
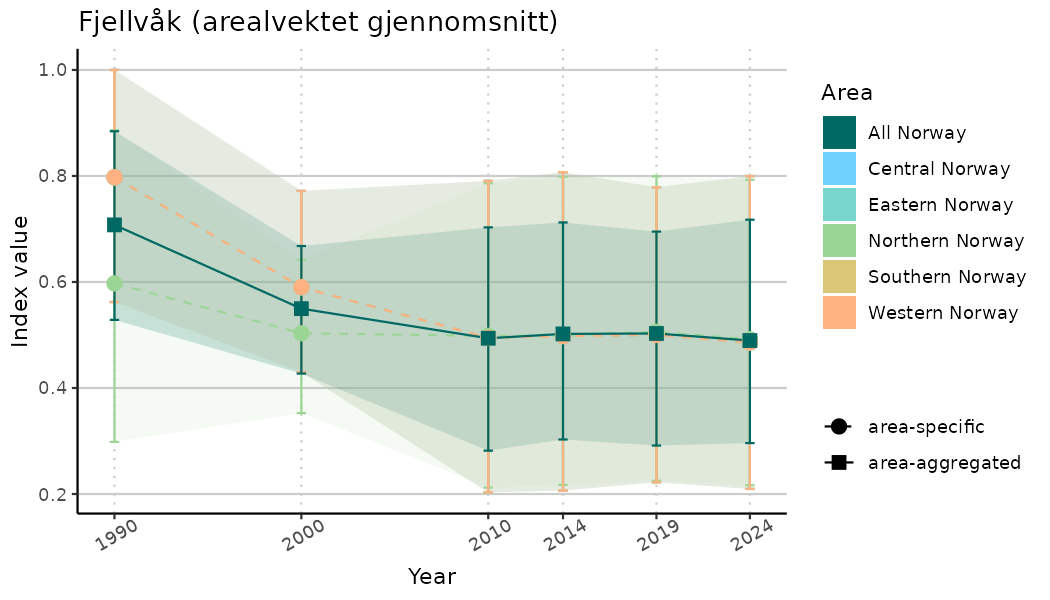
truncateY = TRUE) +
theme(axis.text.x = element_text(angle = 30, vjust = 0.8))
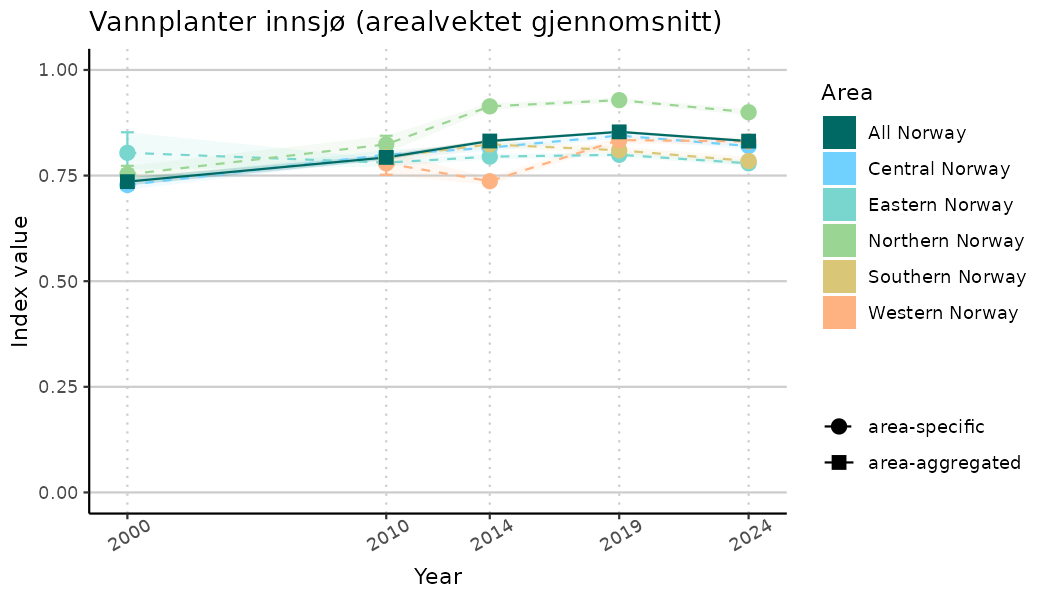
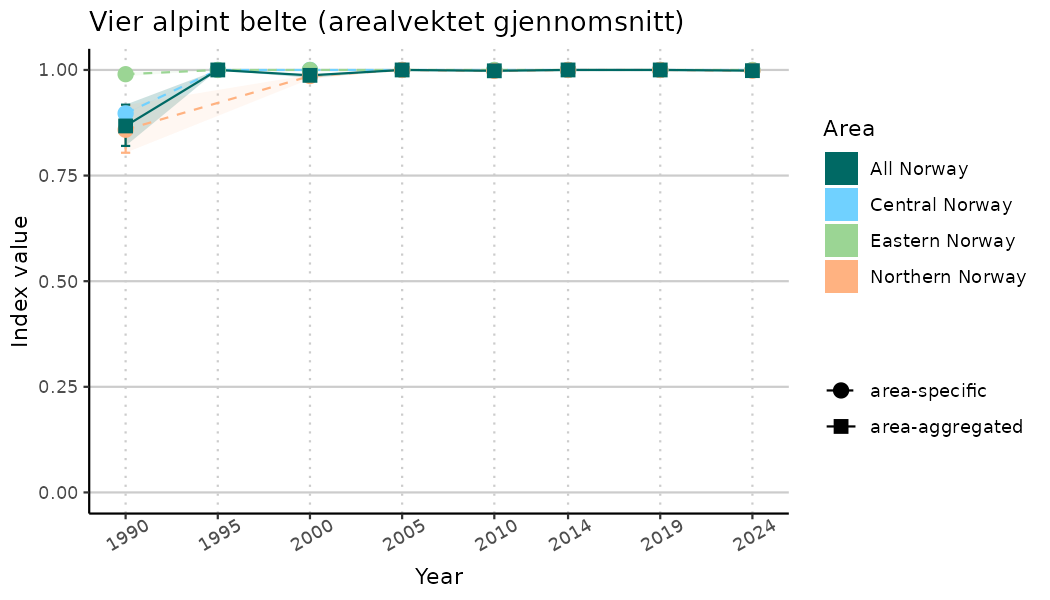
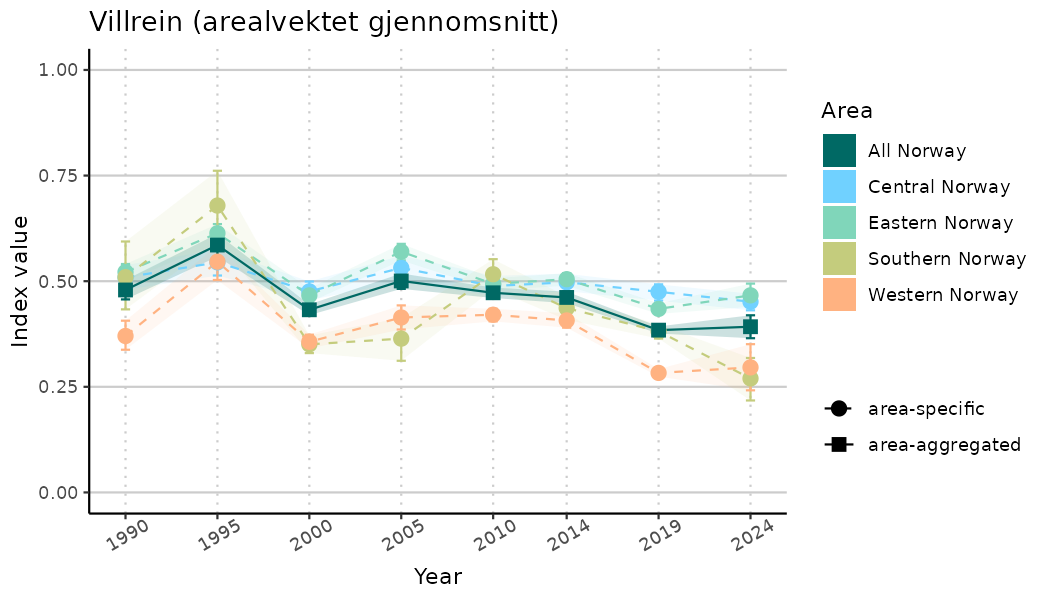
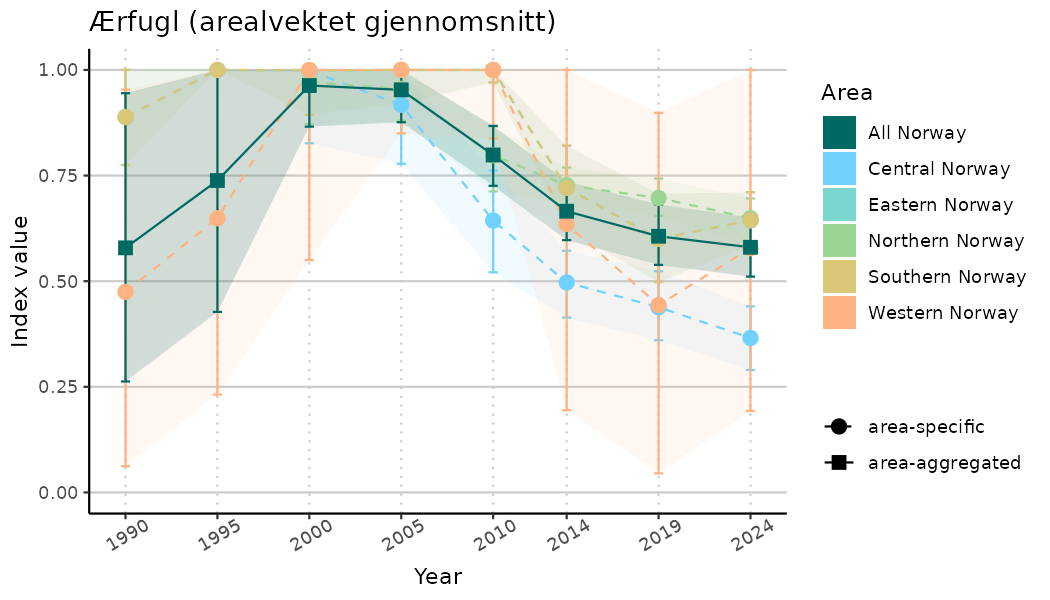
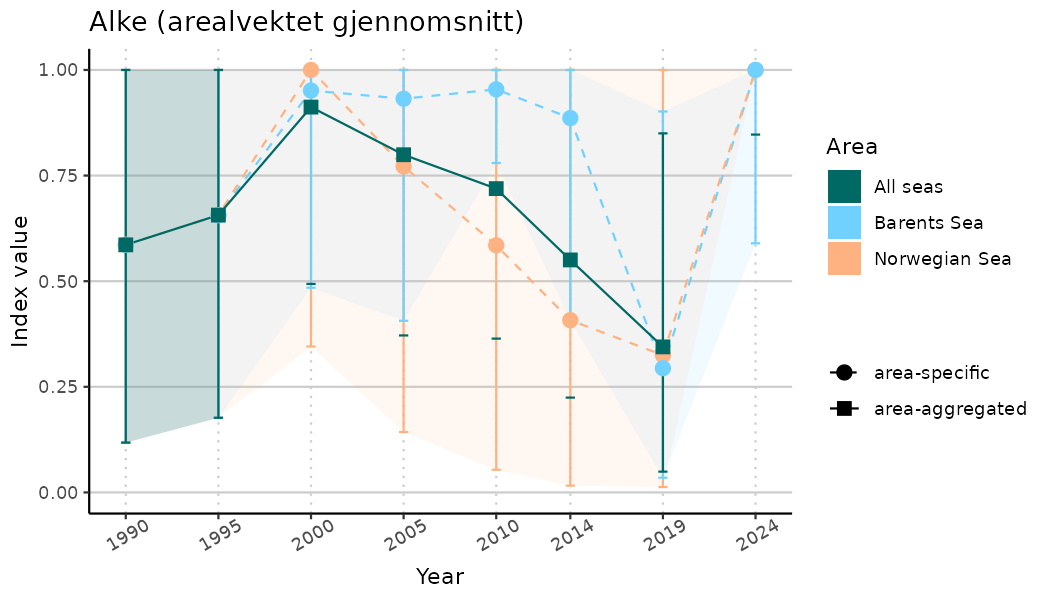
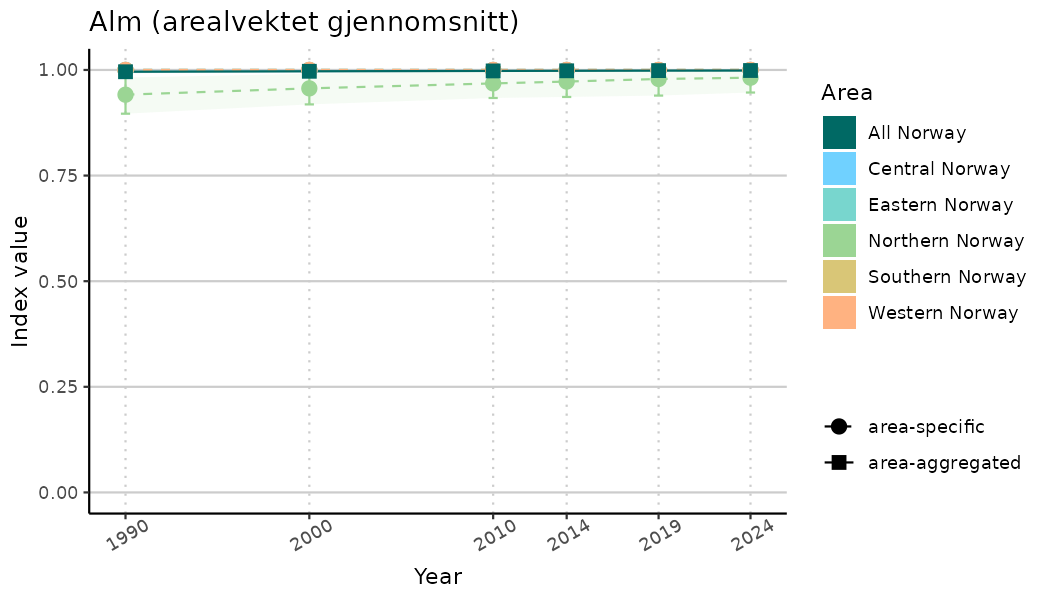
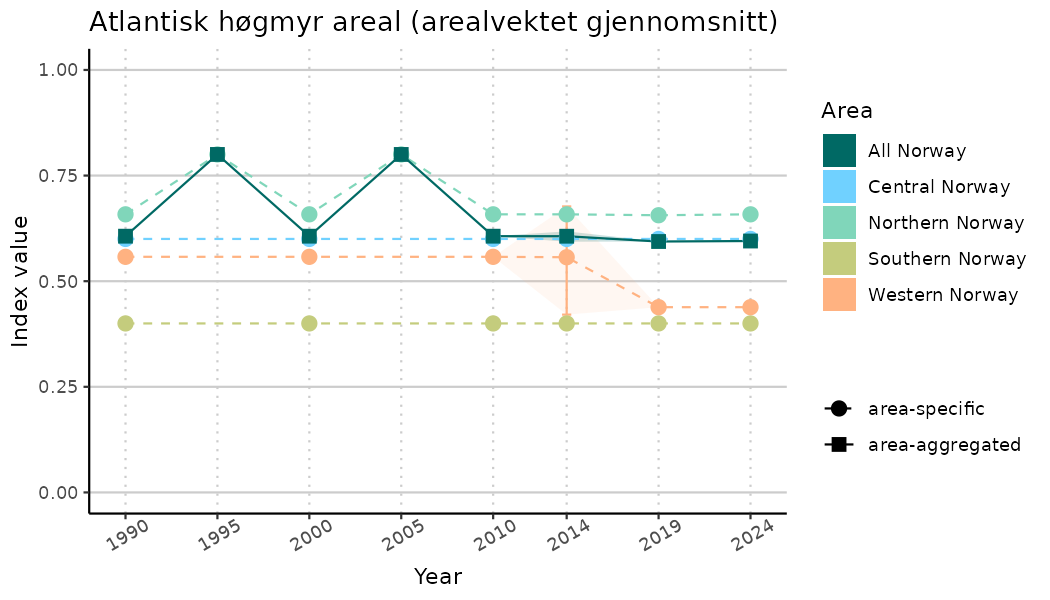
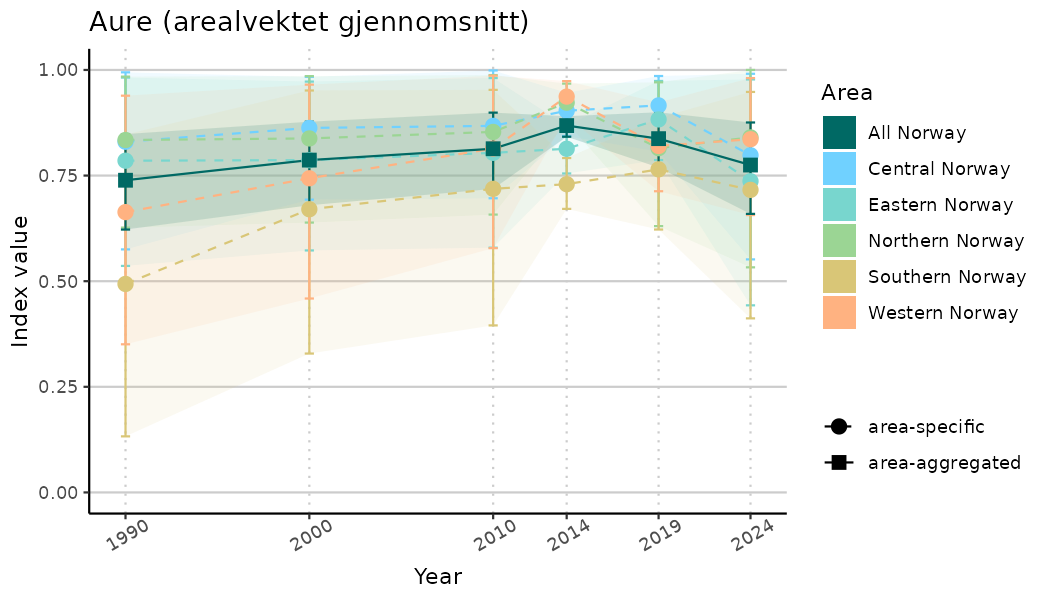
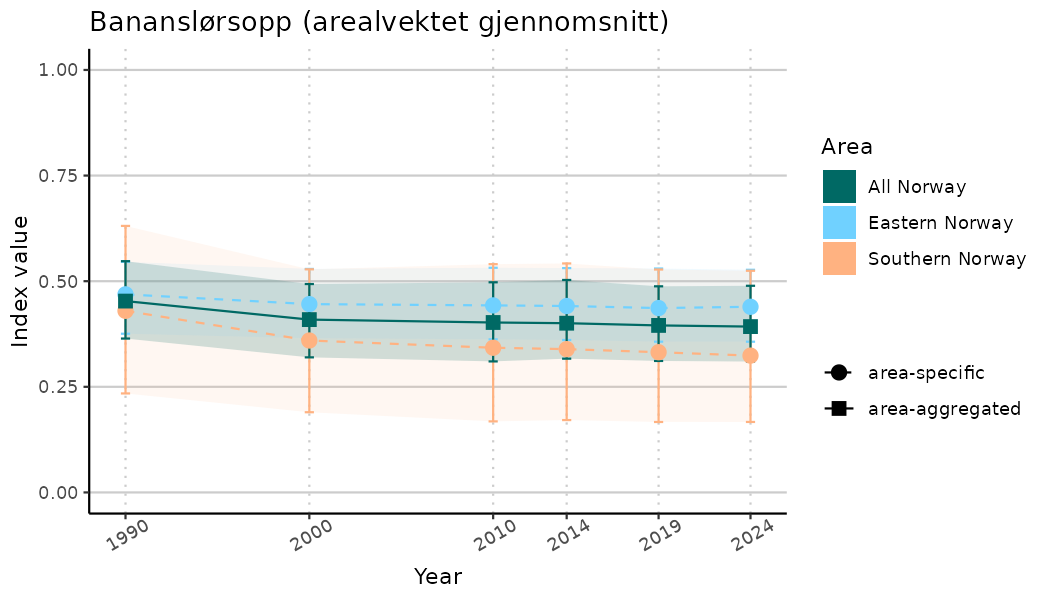
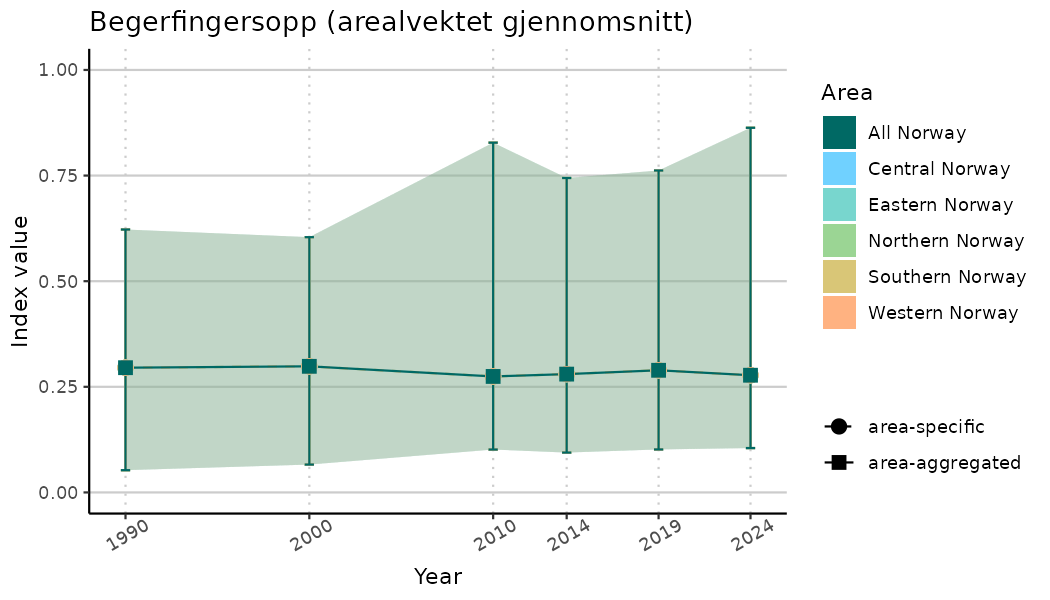
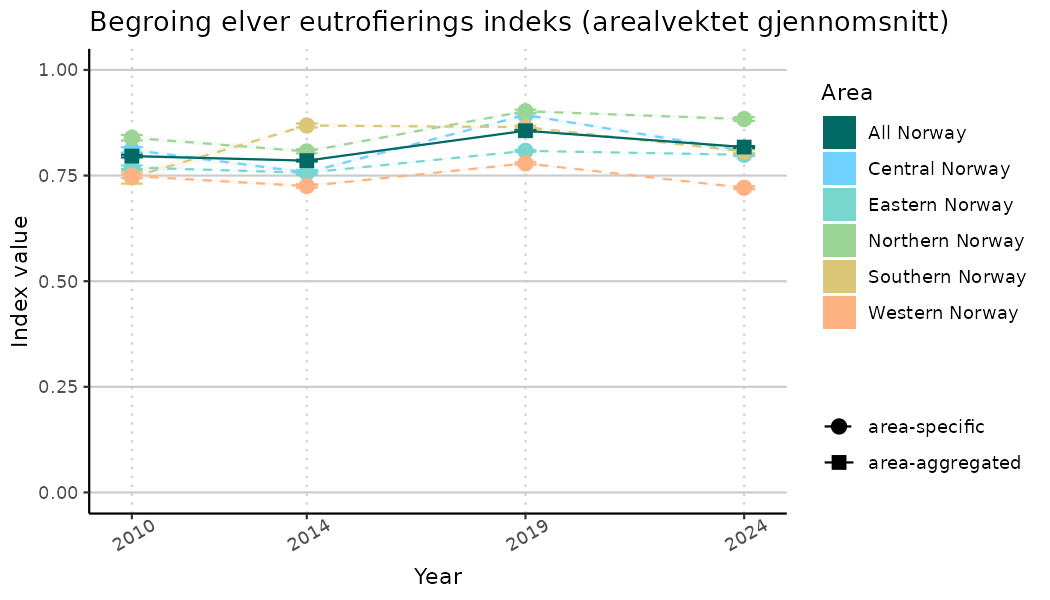
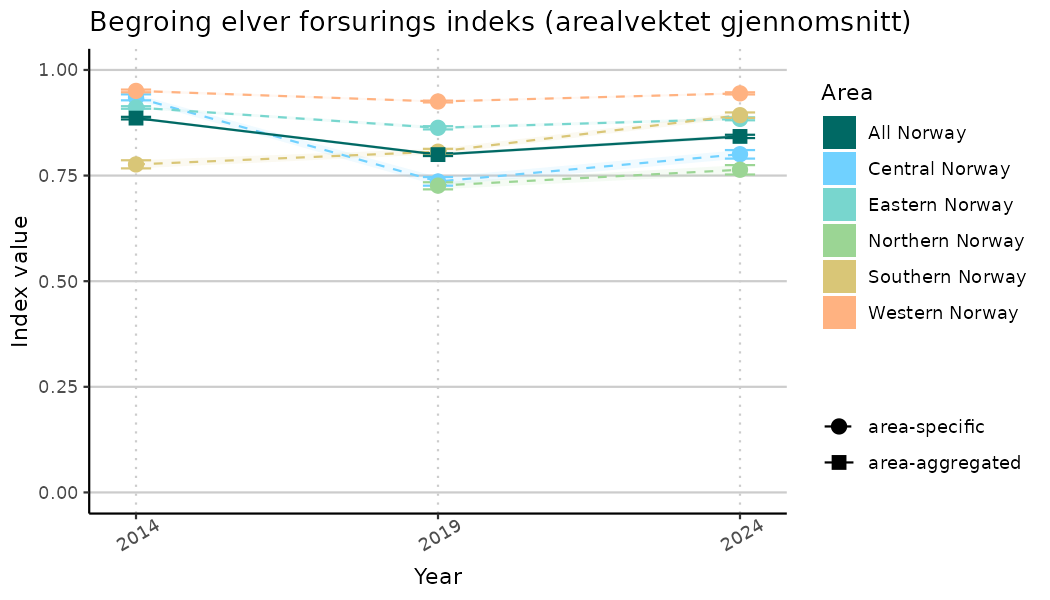
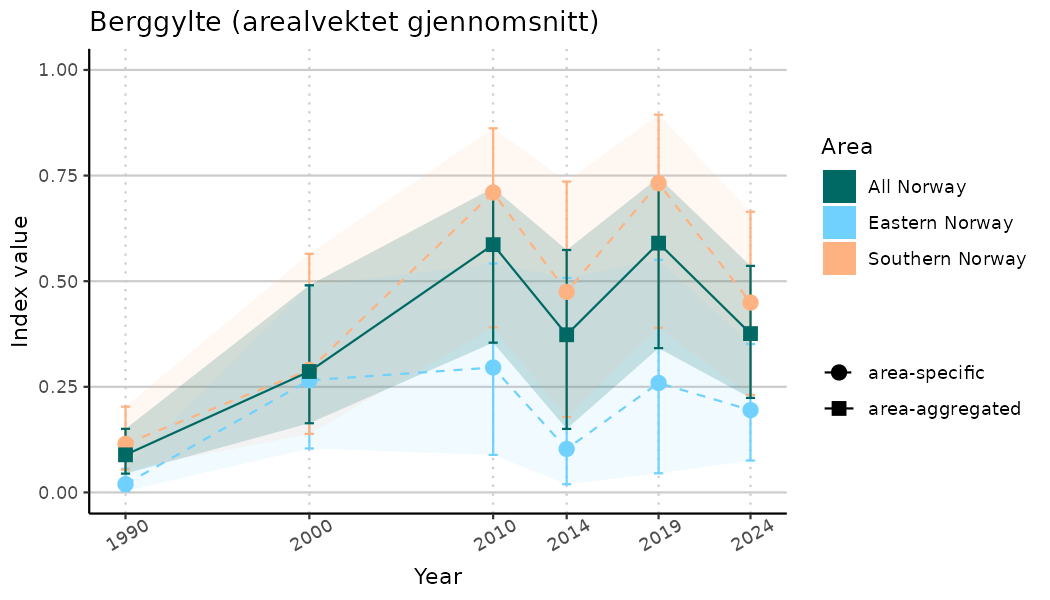
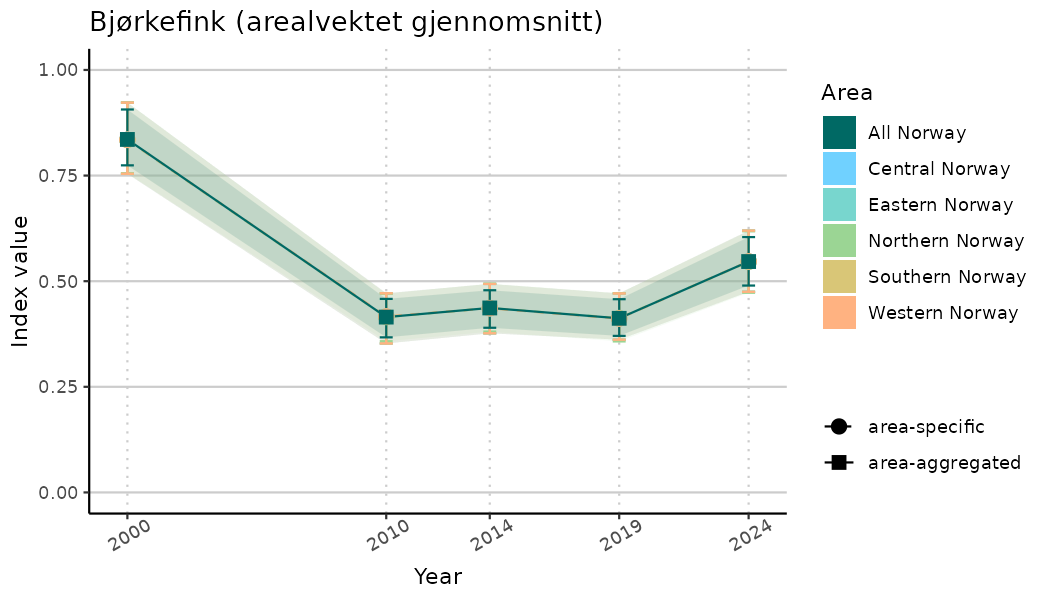
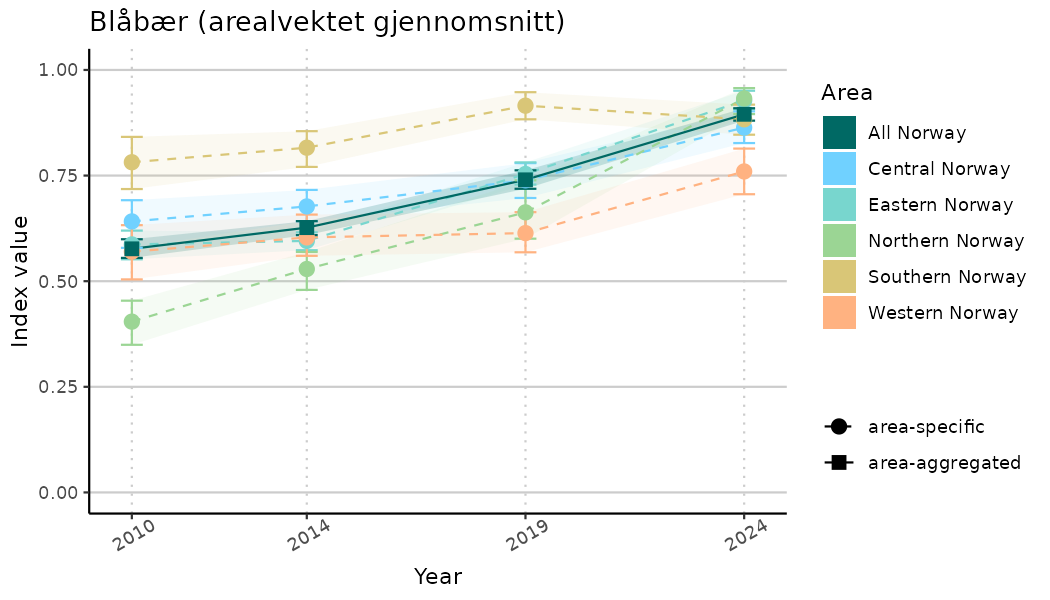
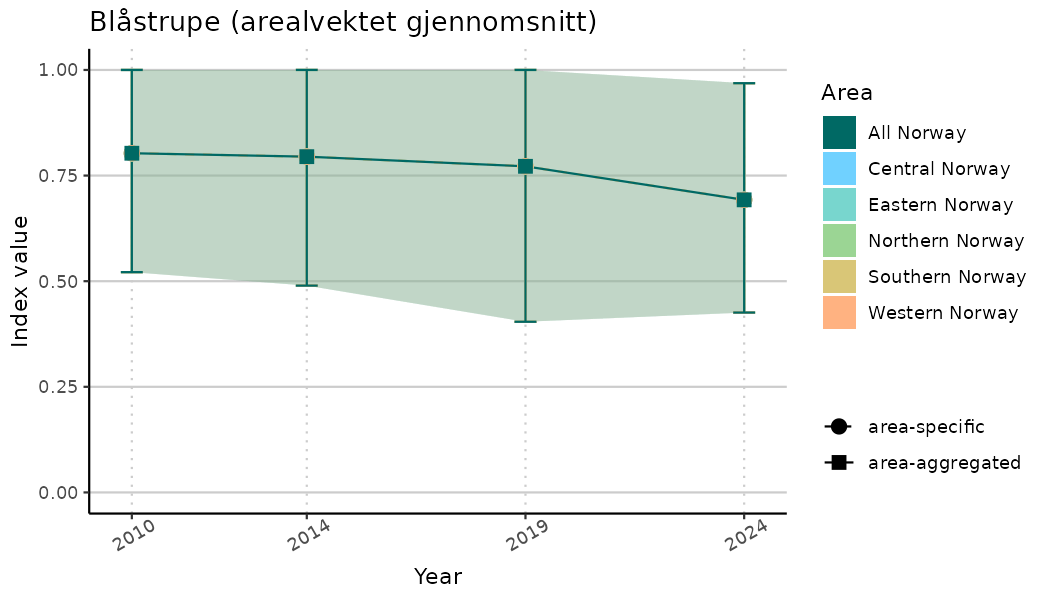
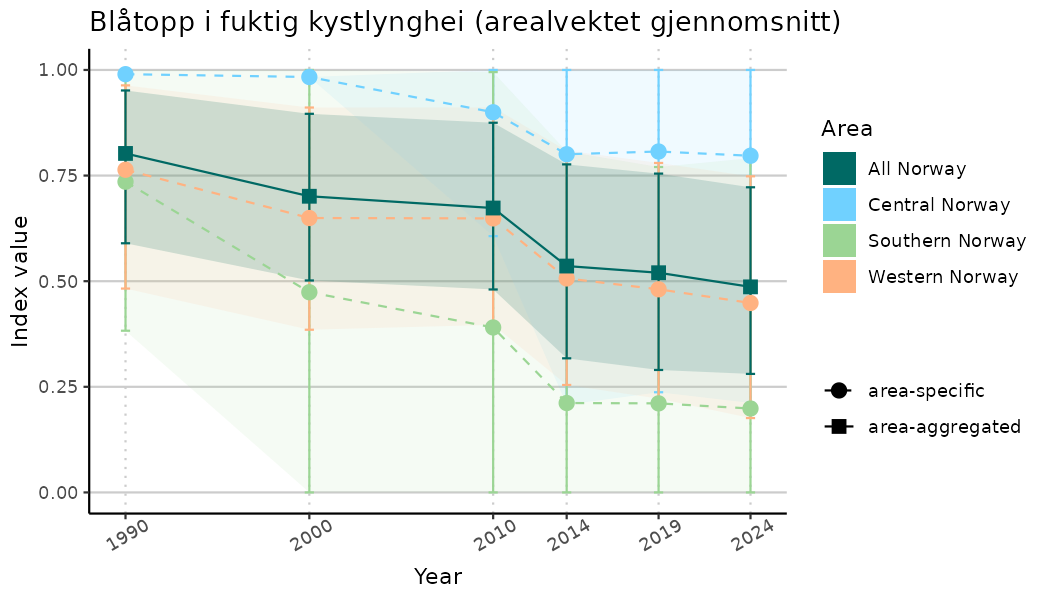
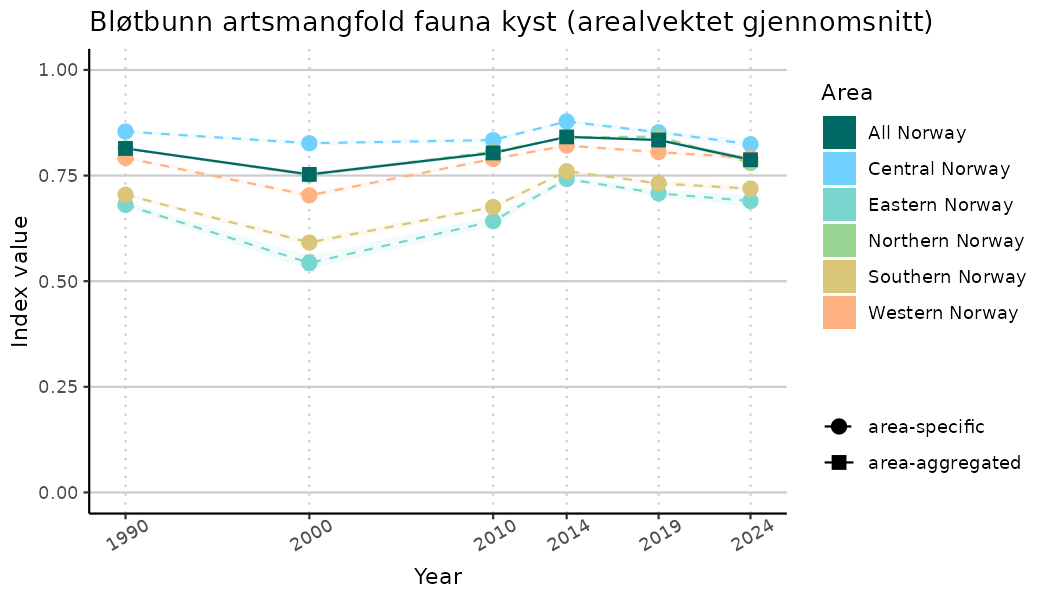
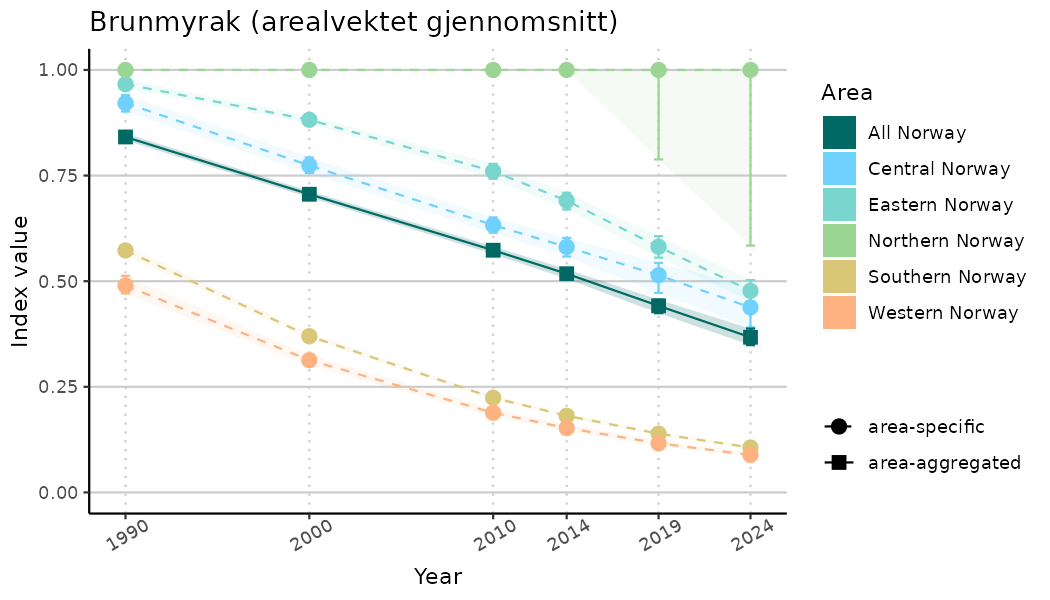
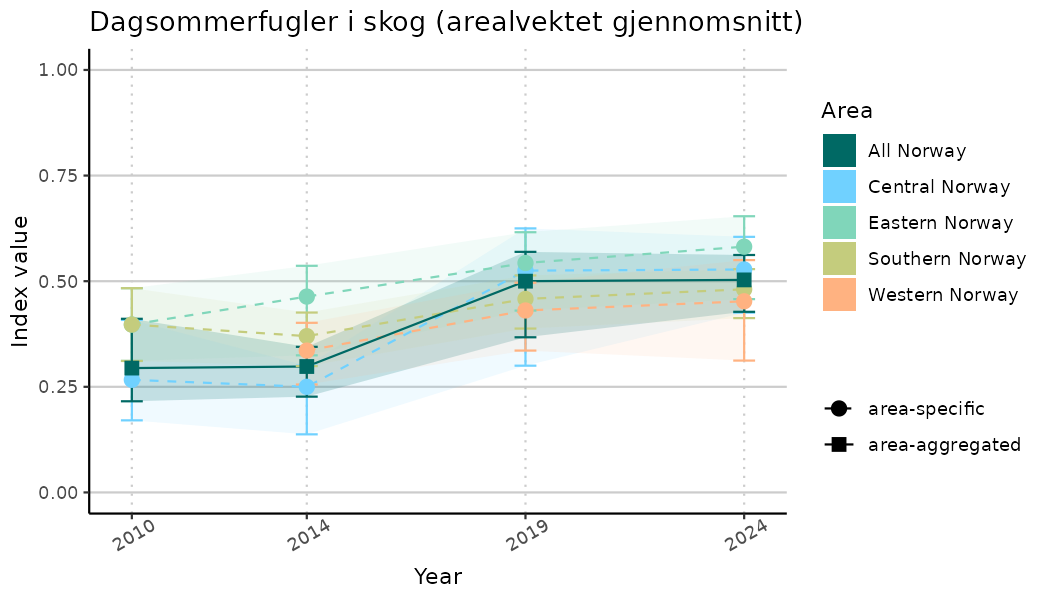
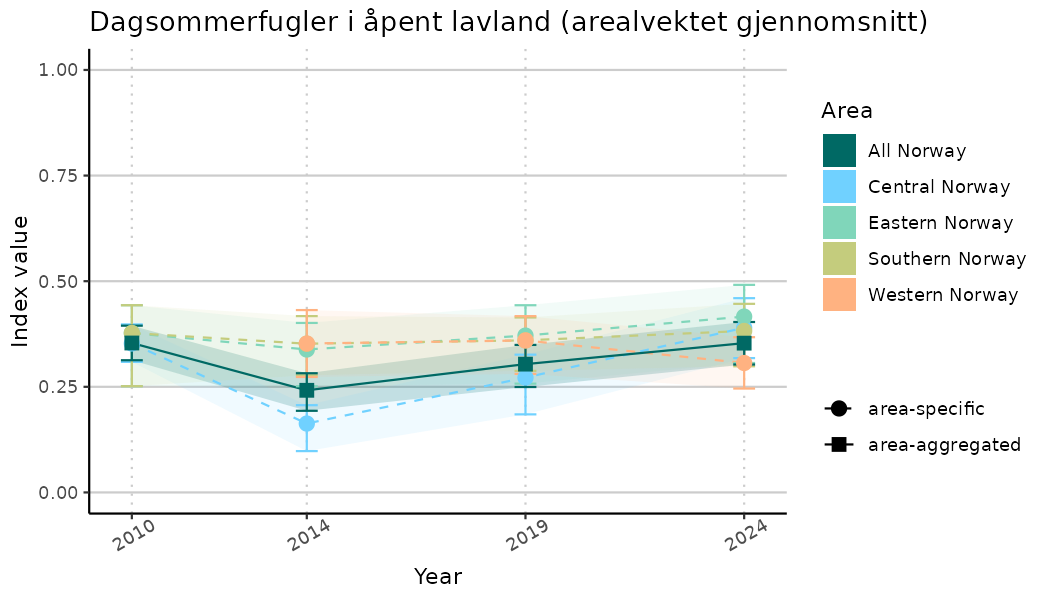
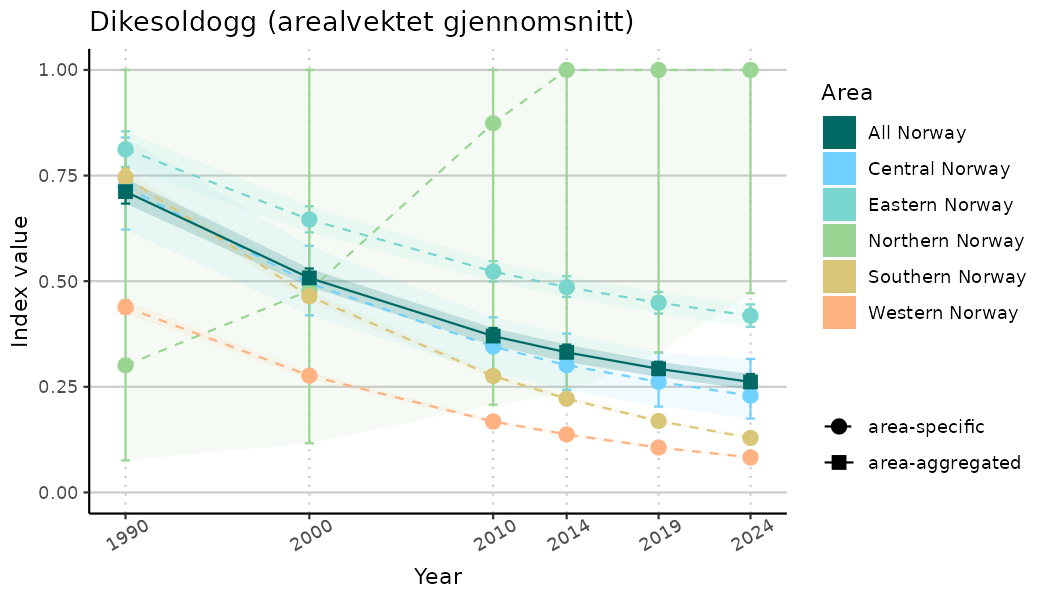
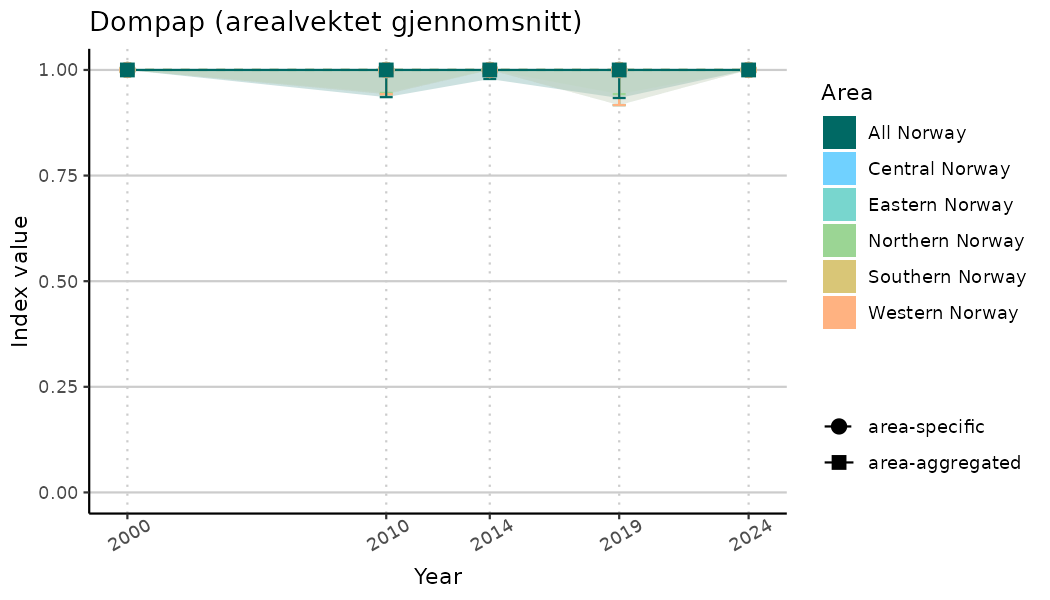
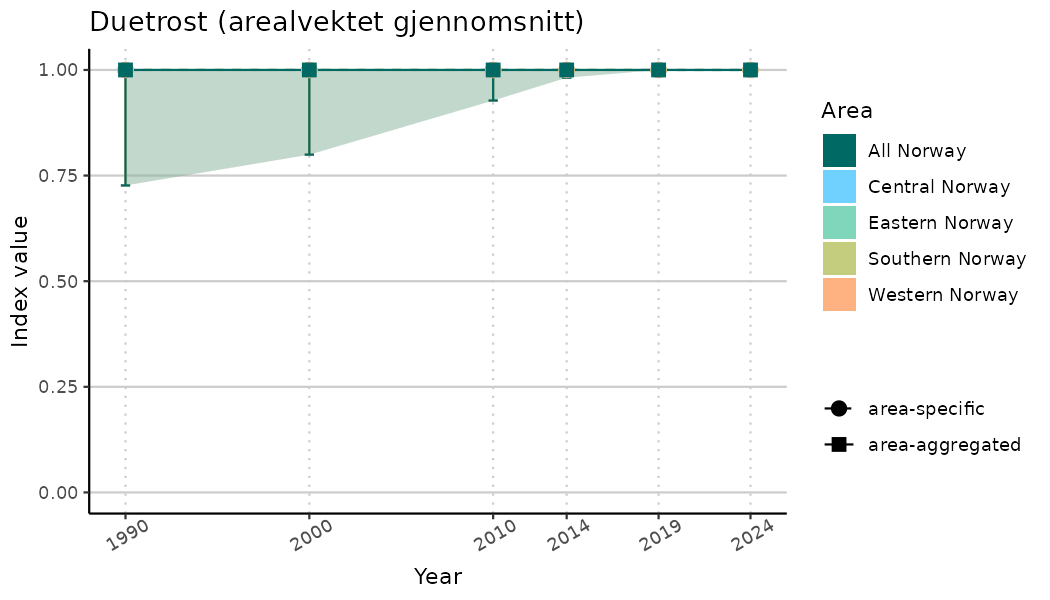
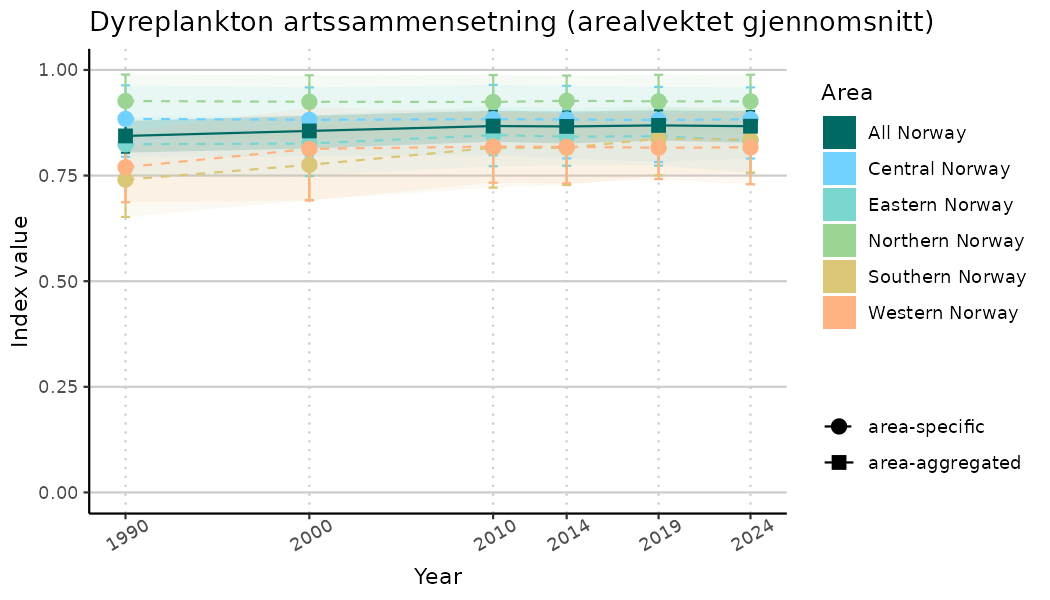
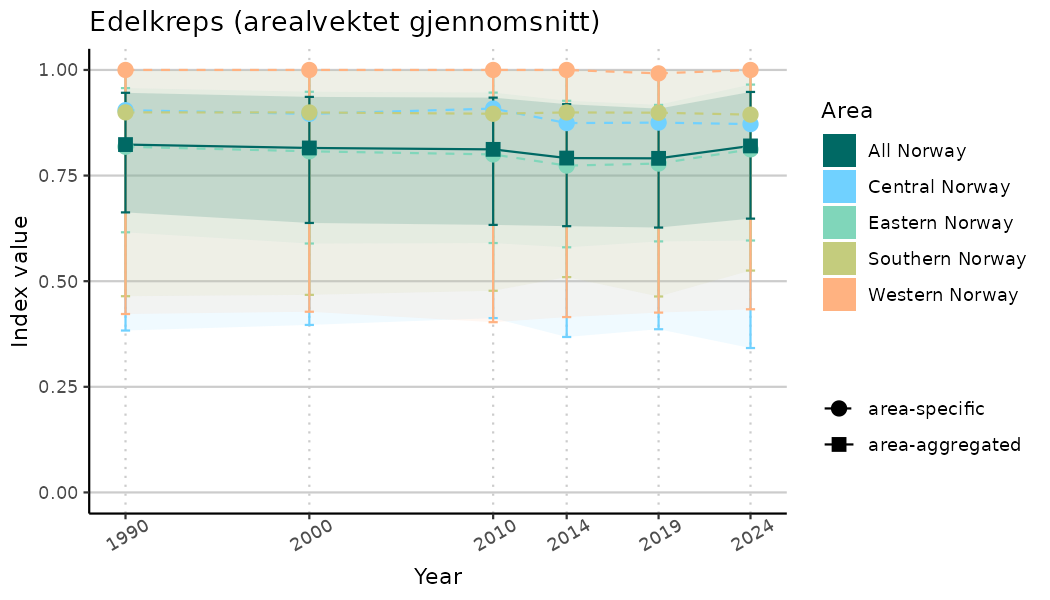
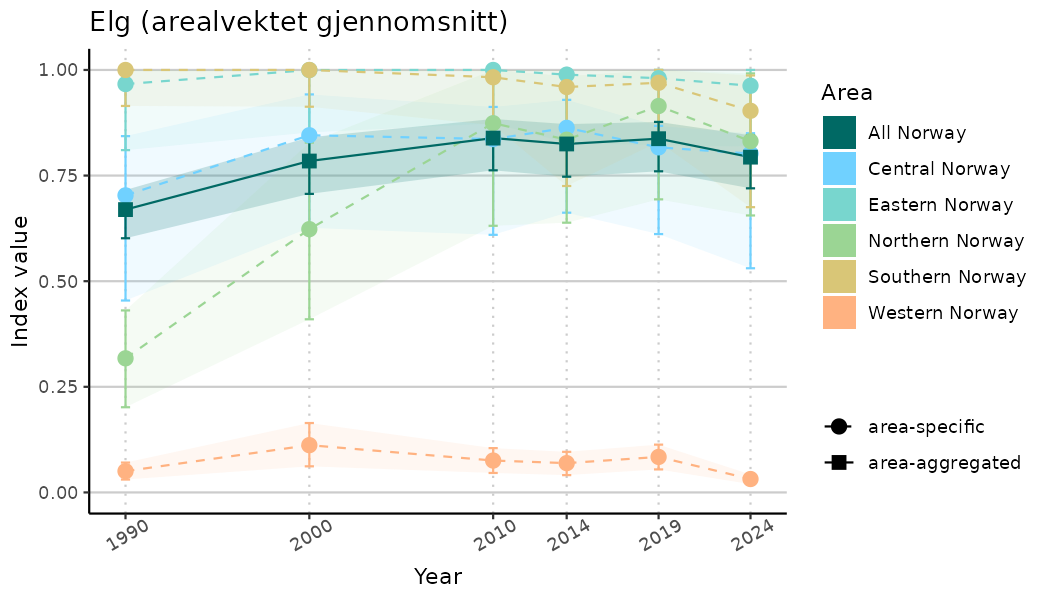
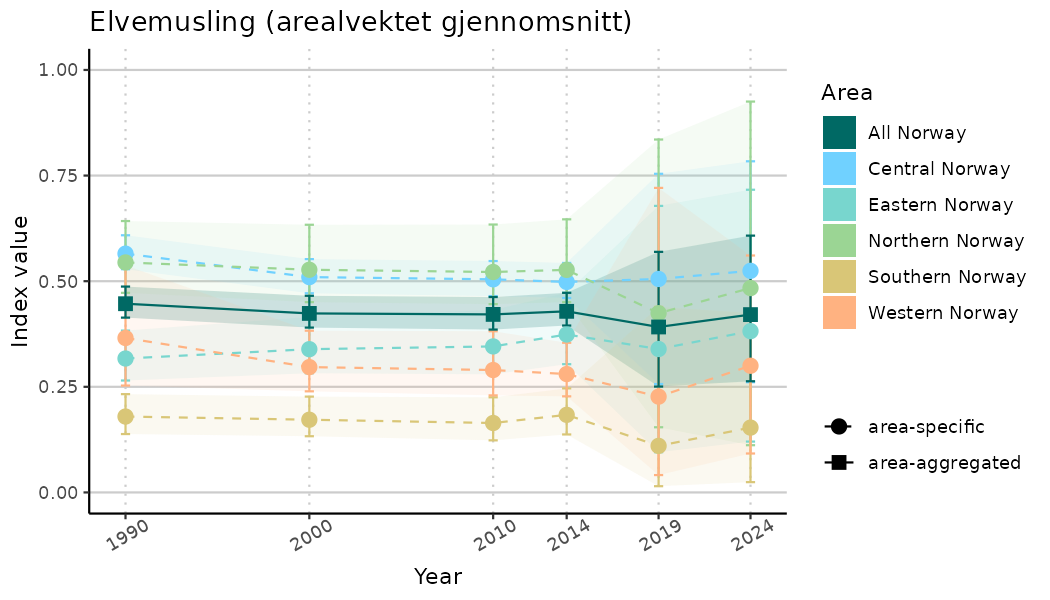
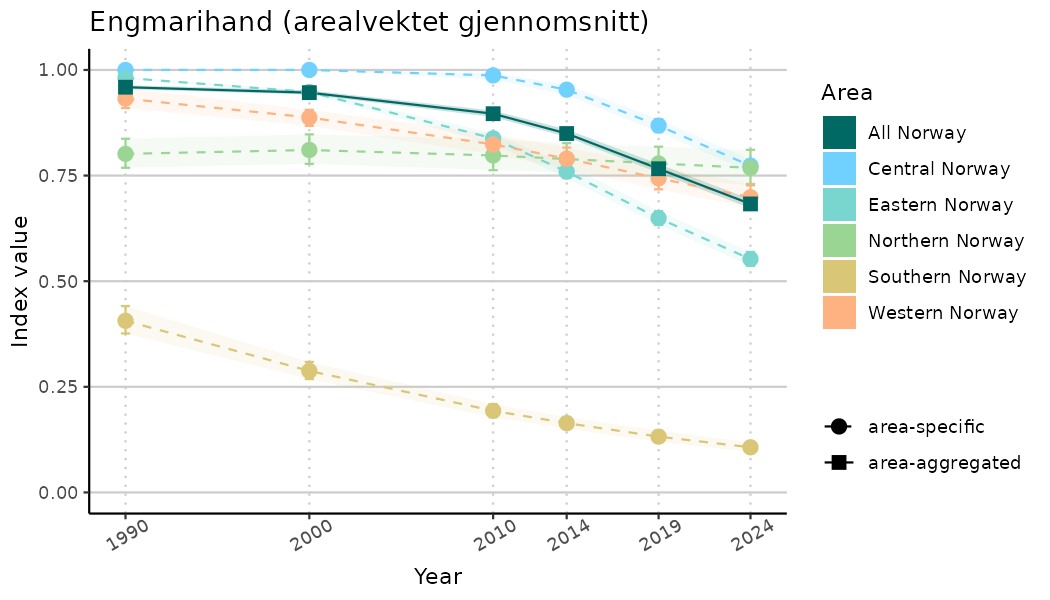
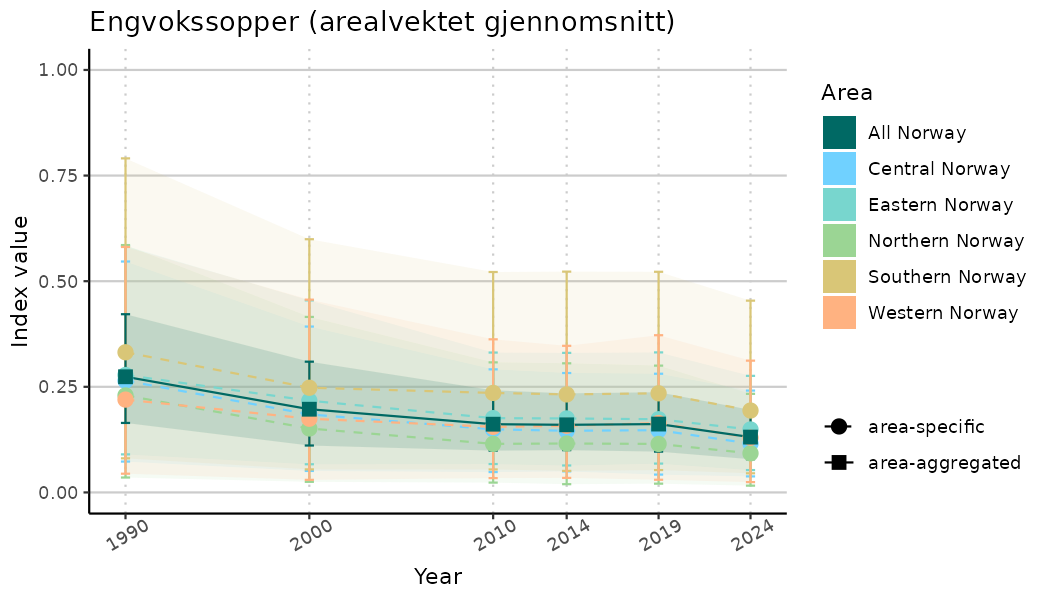
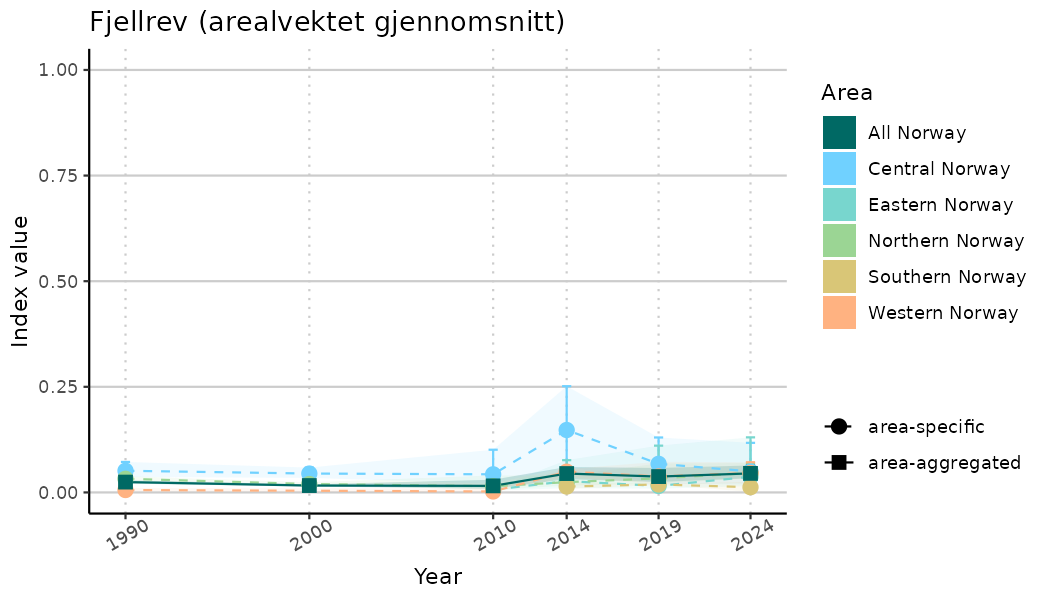
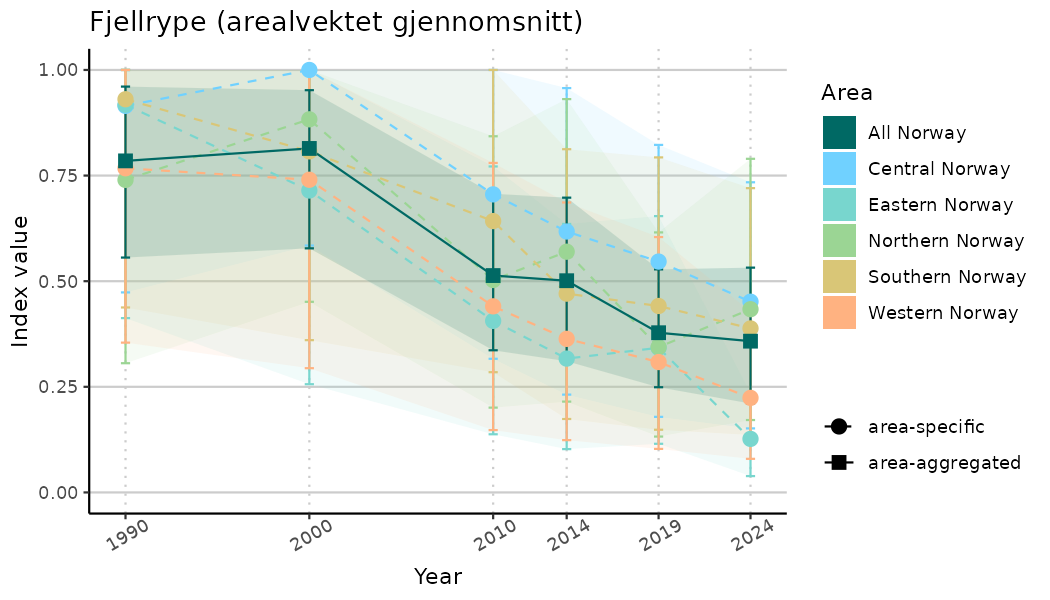
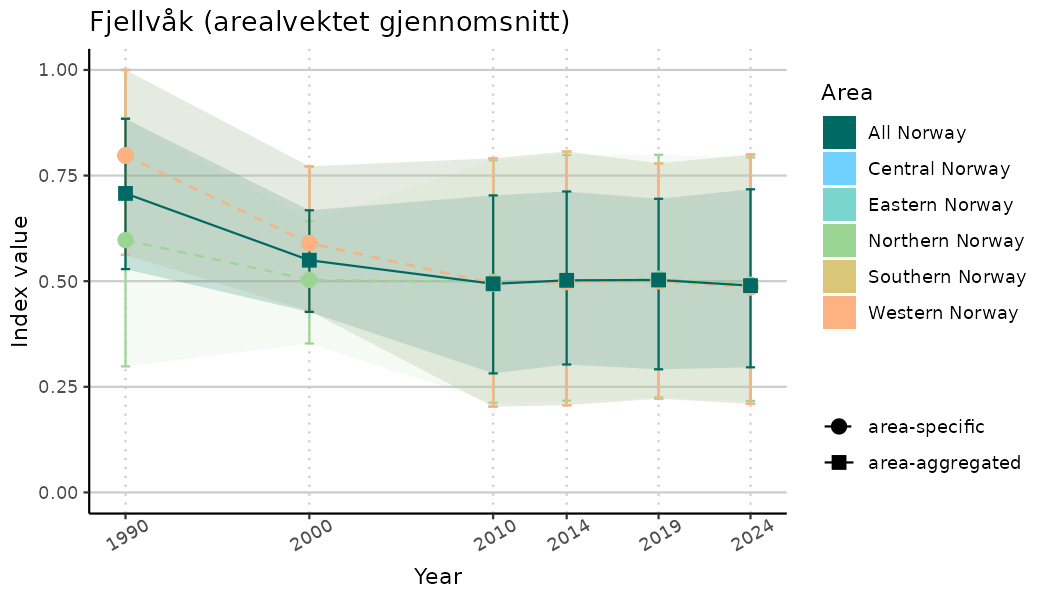
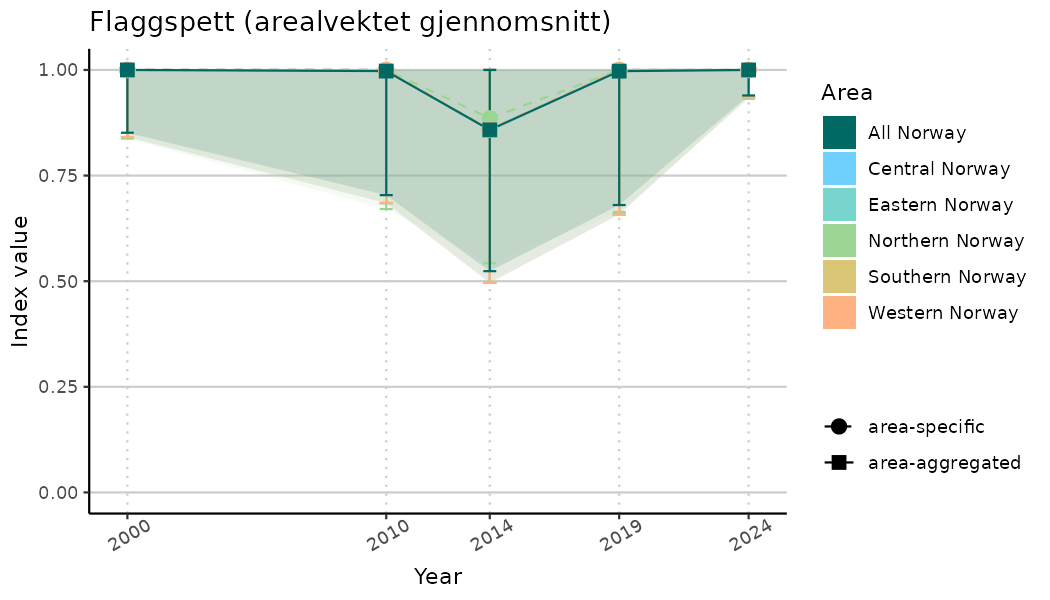
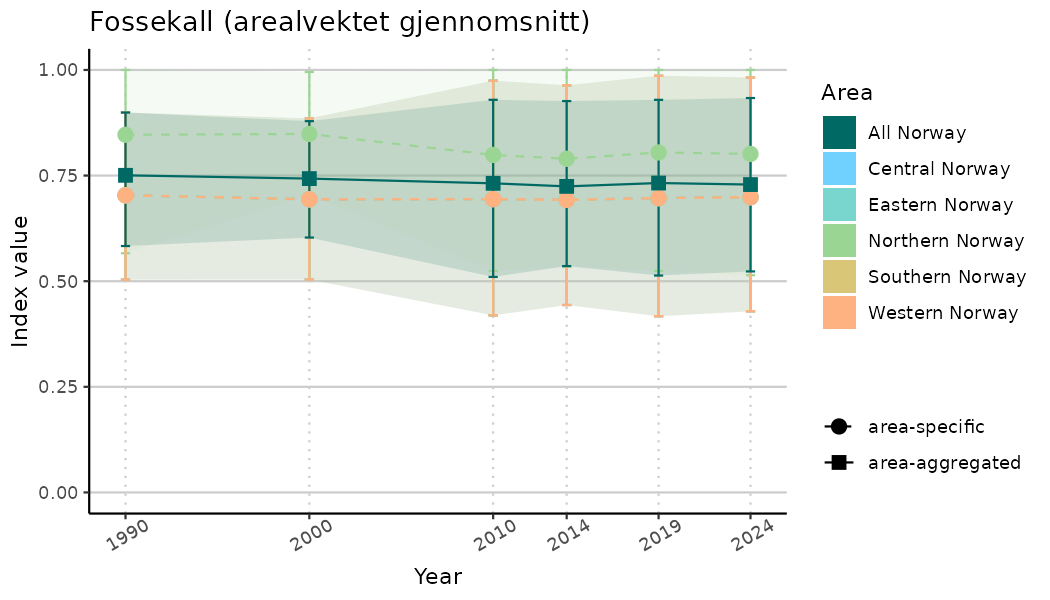
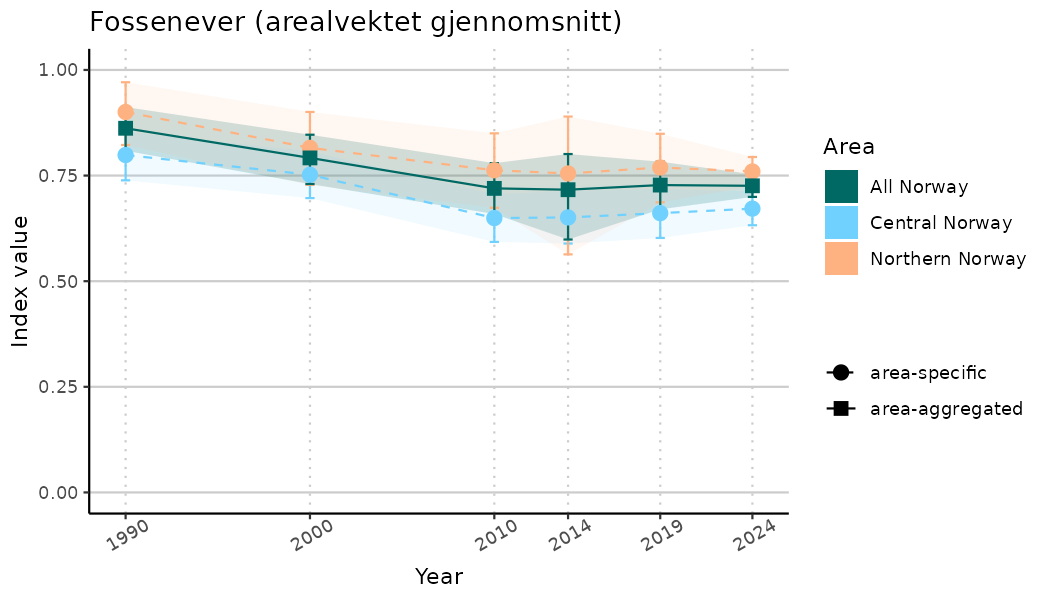
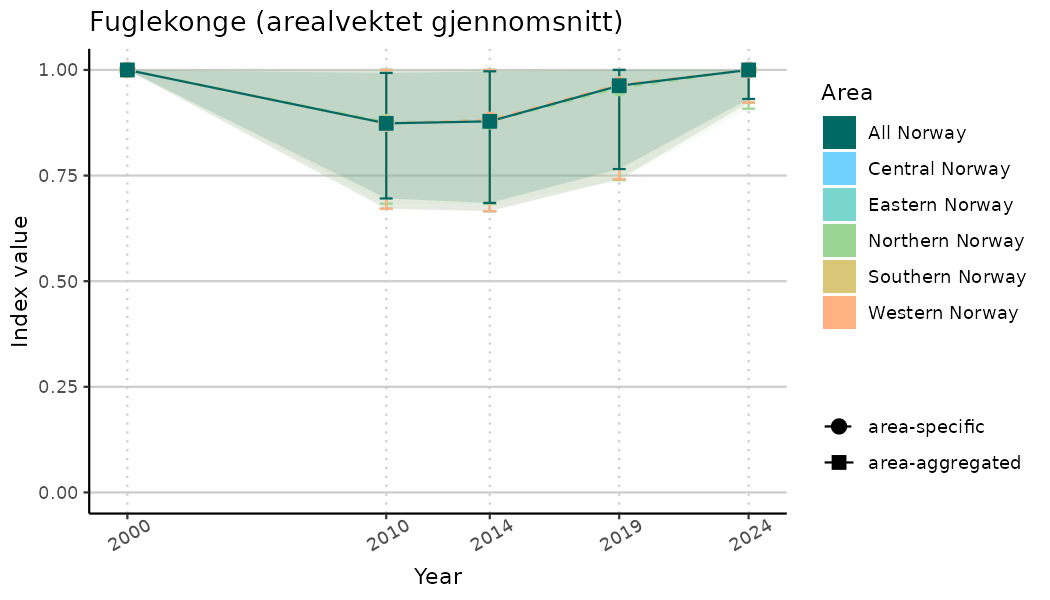
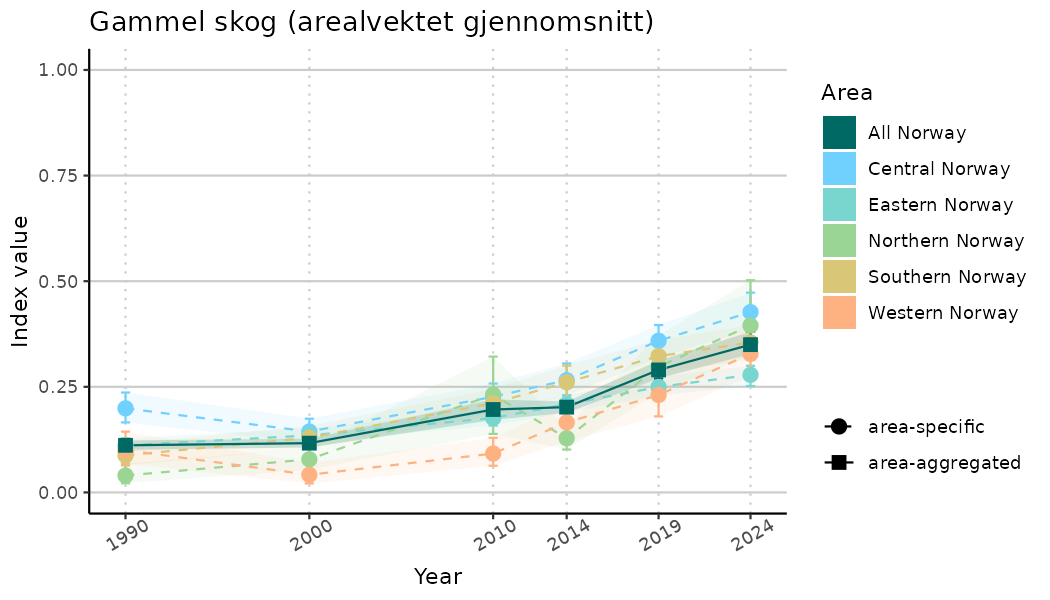
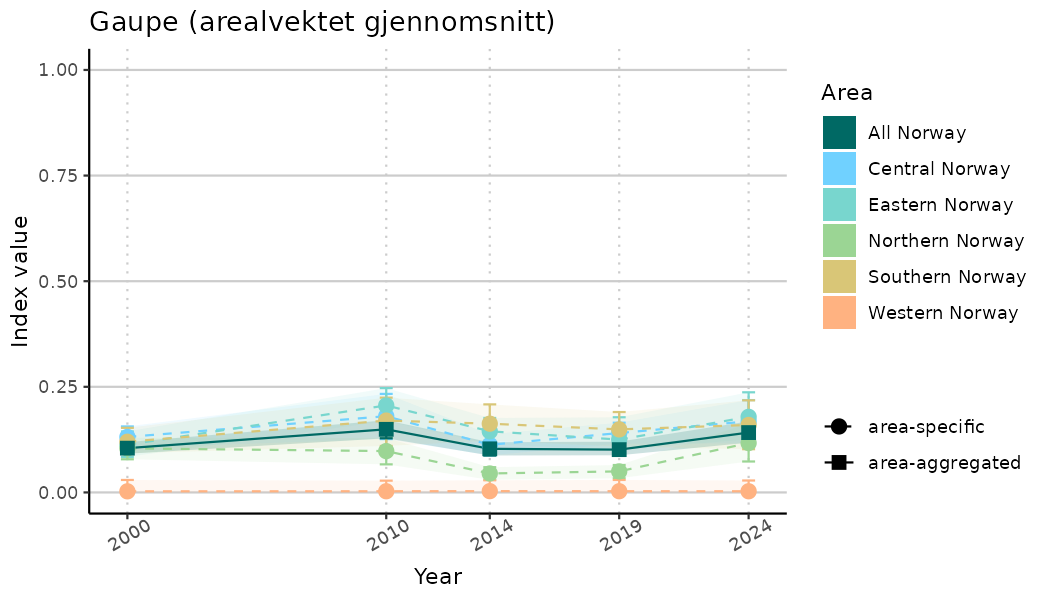
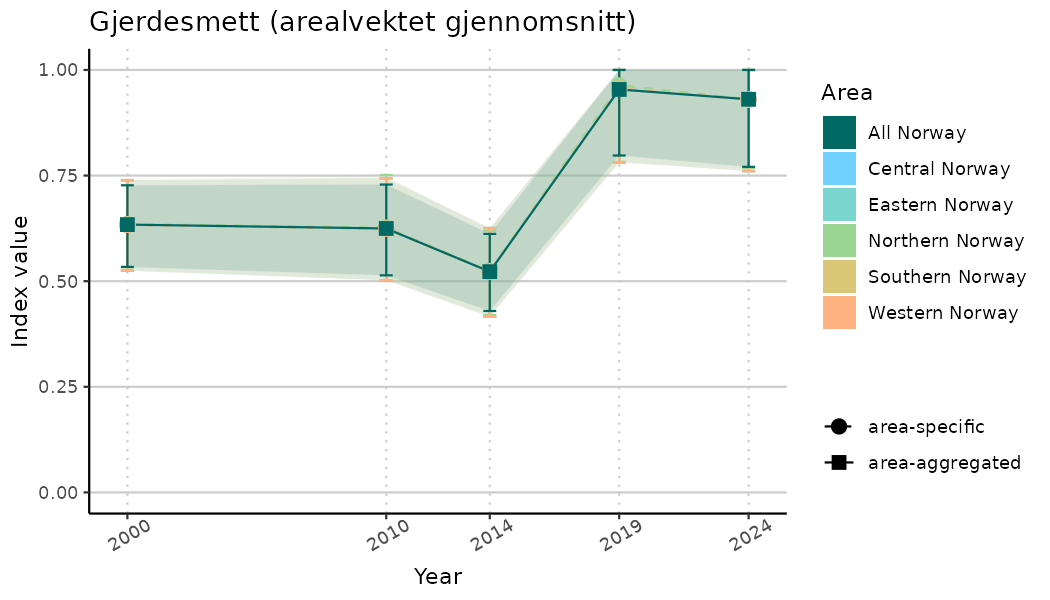
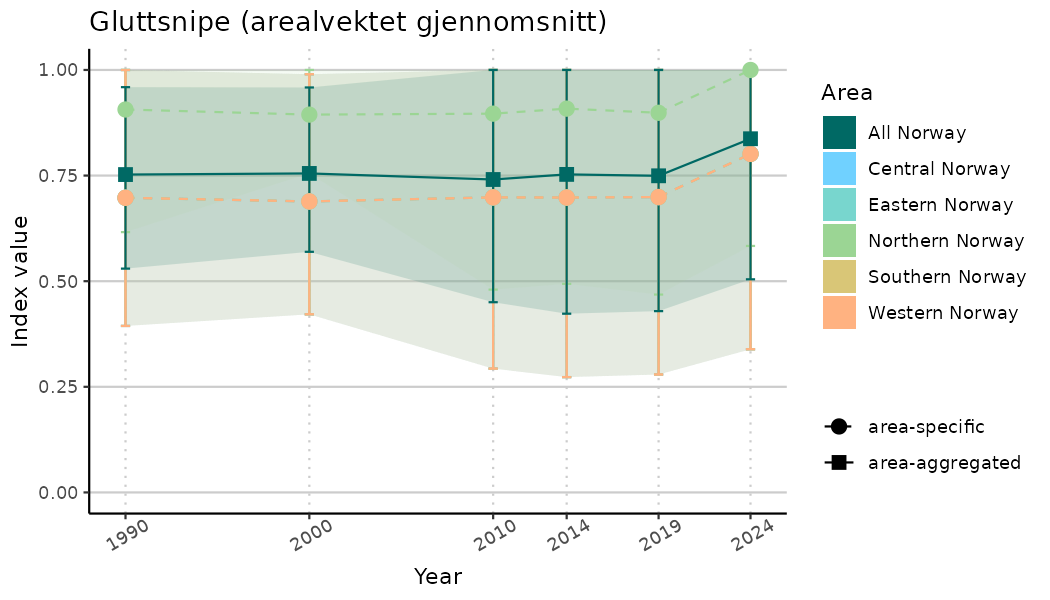
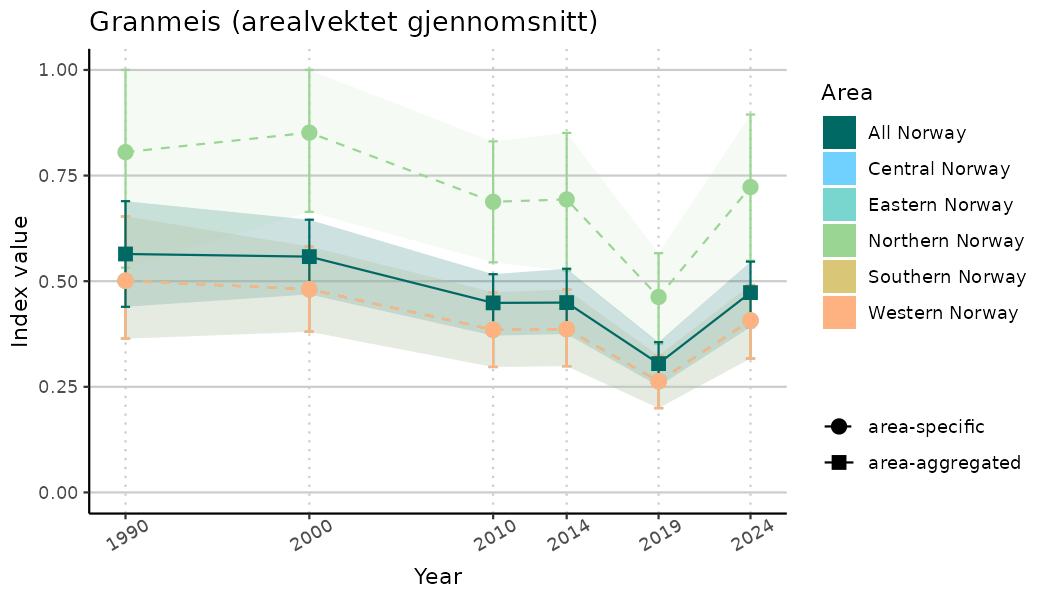
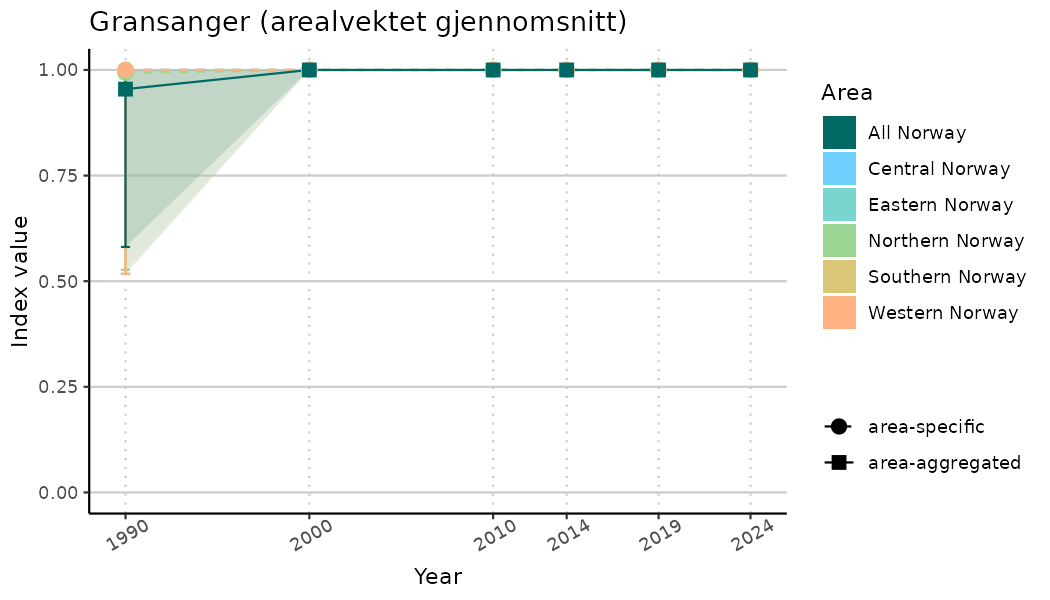
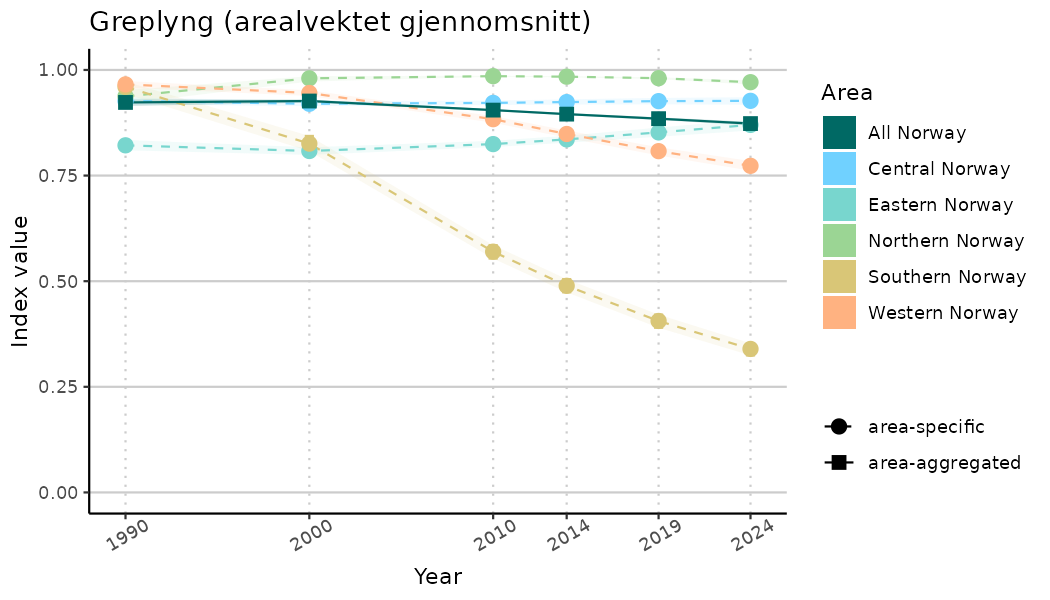
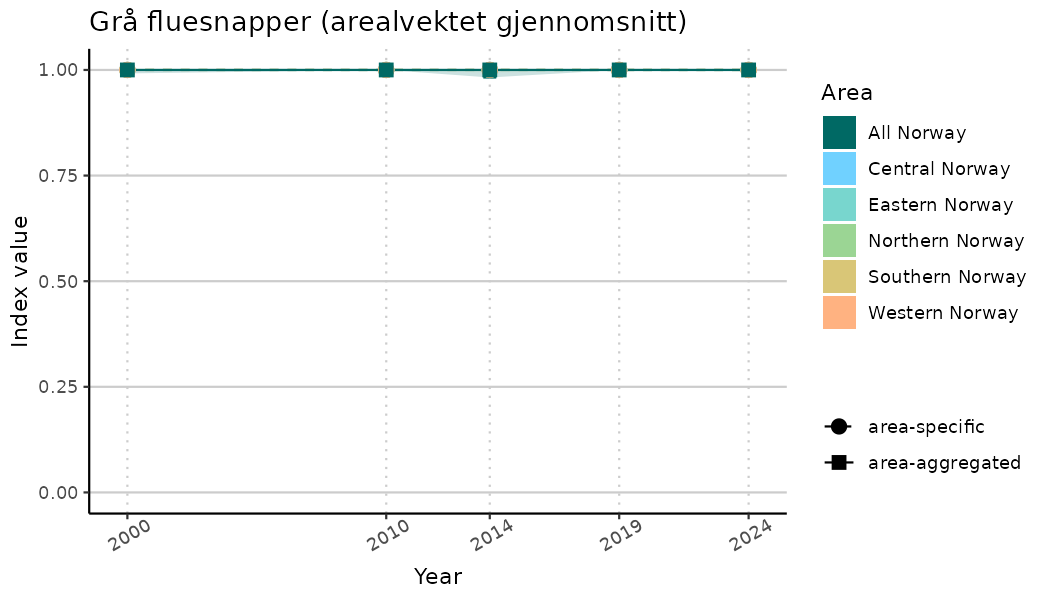
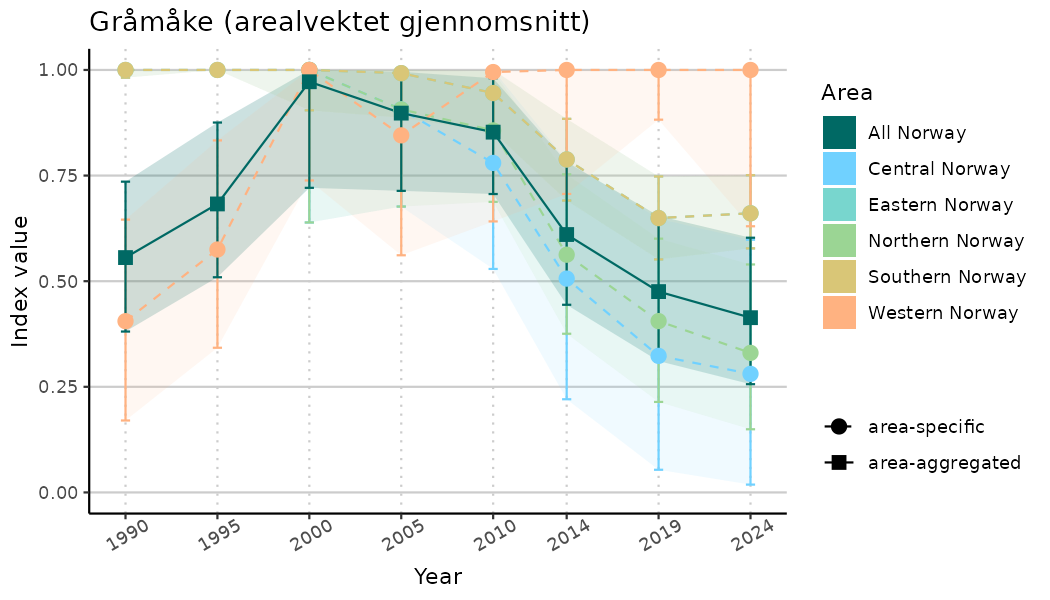
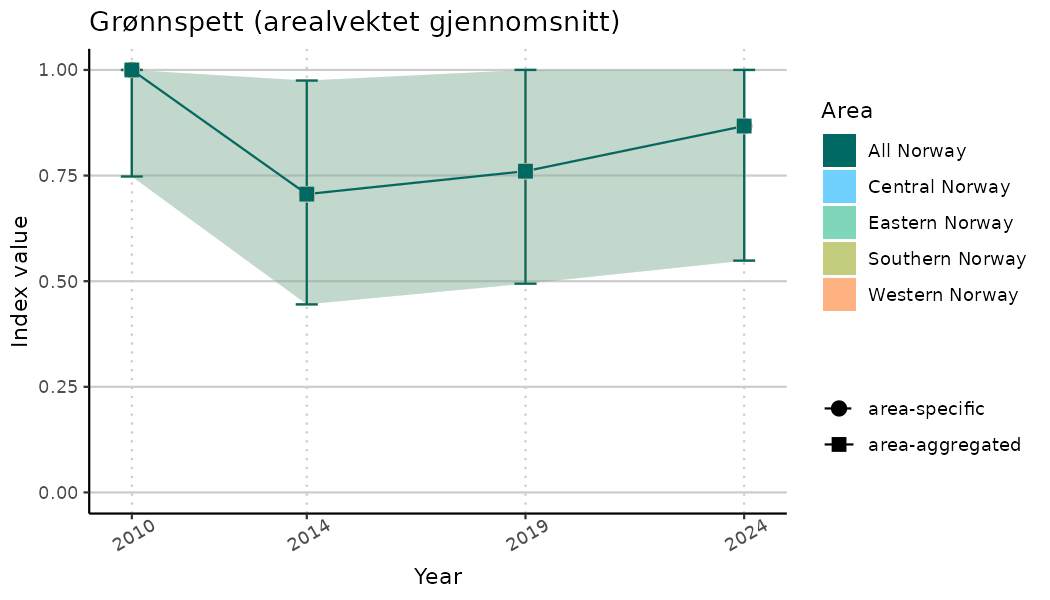
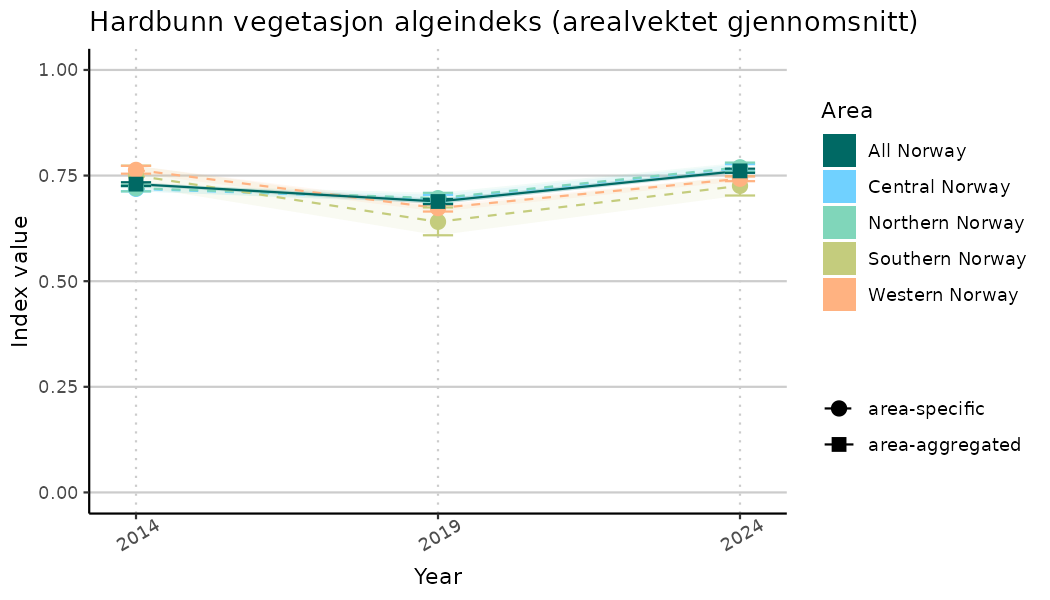
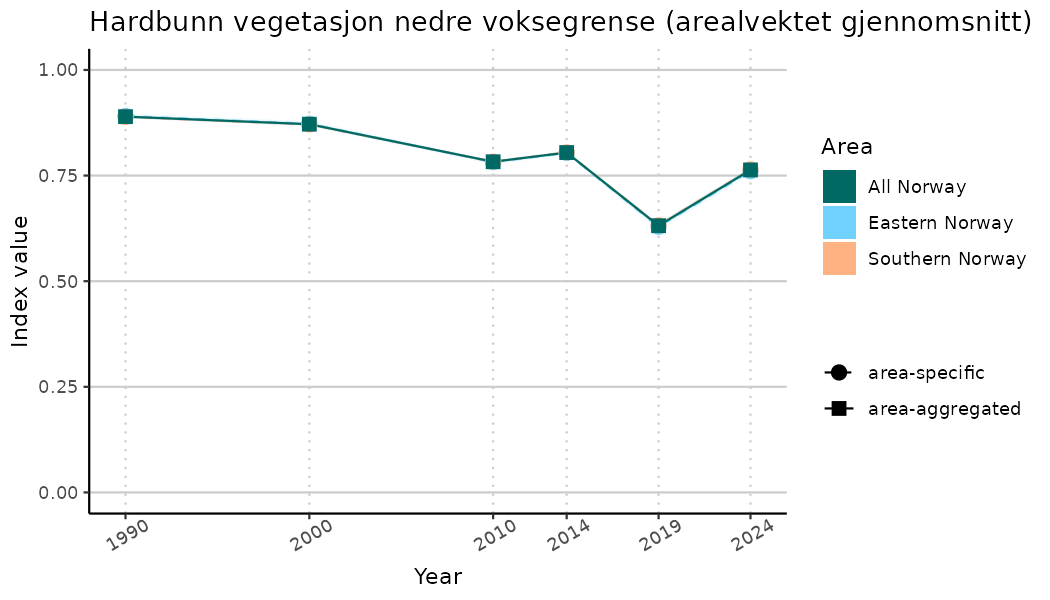
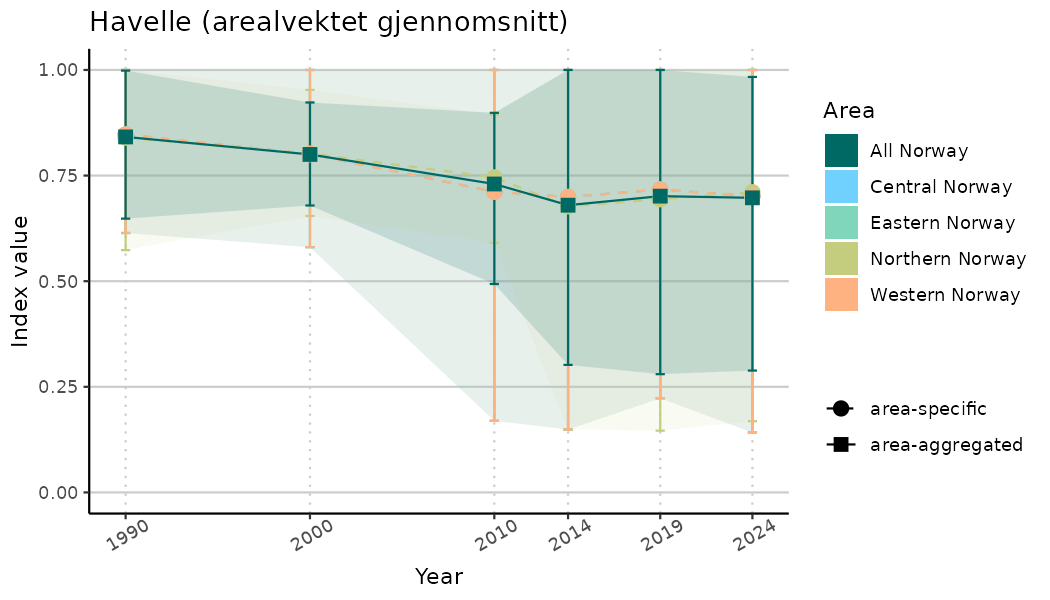
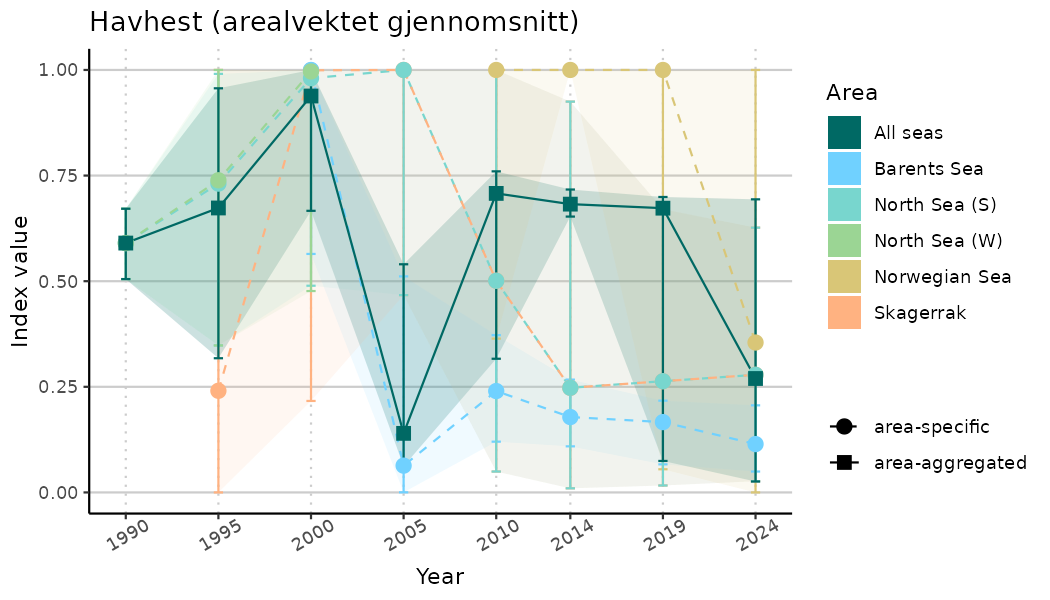
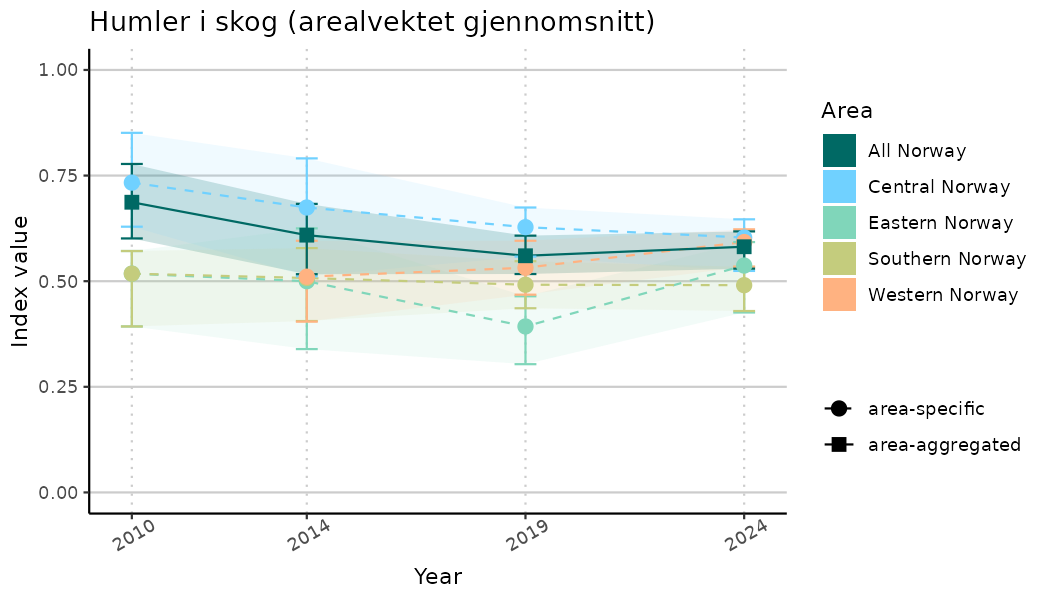
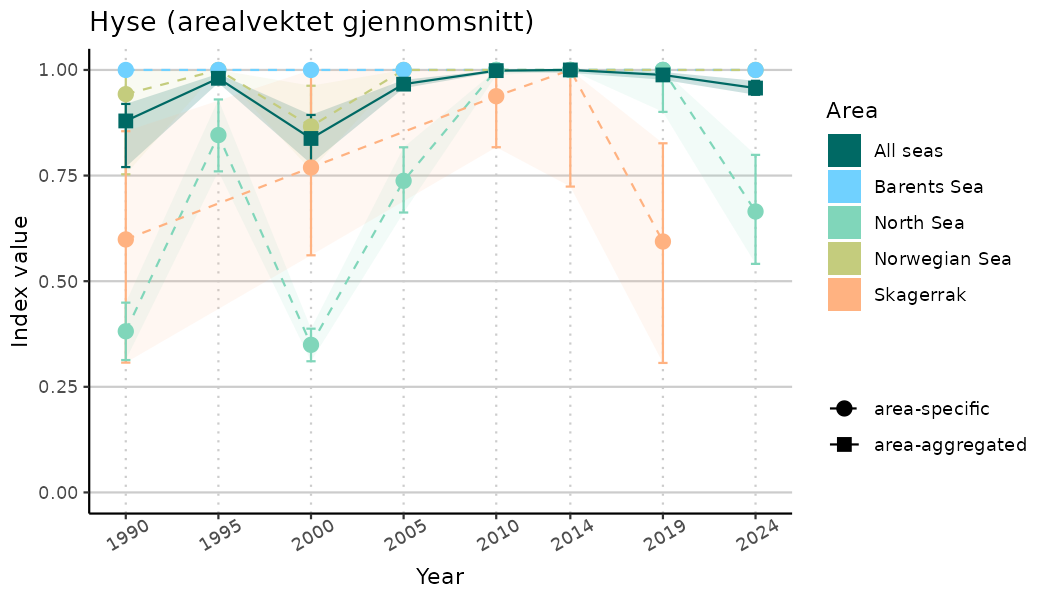
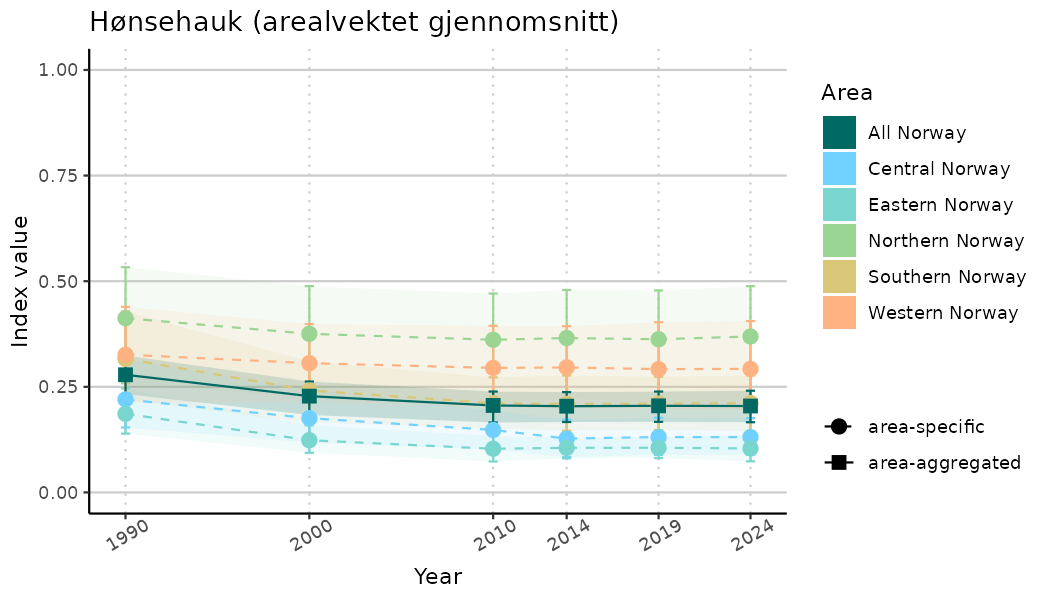
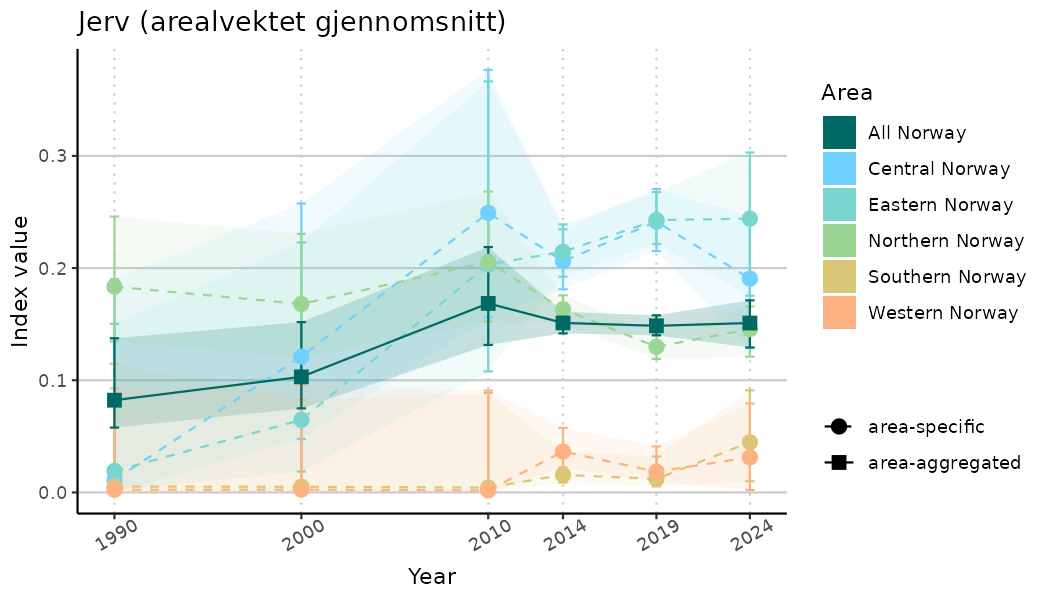
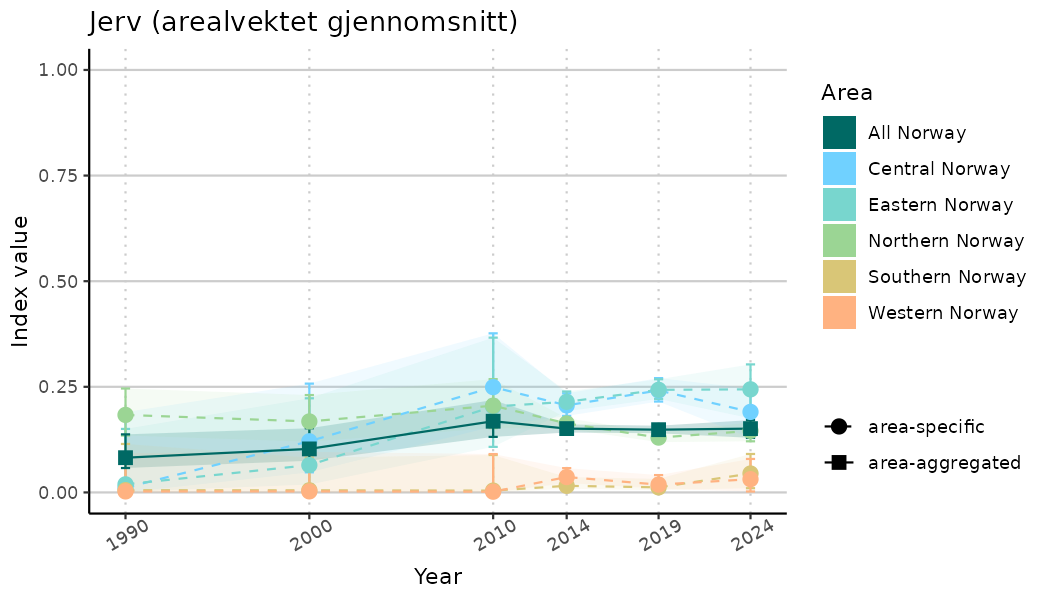
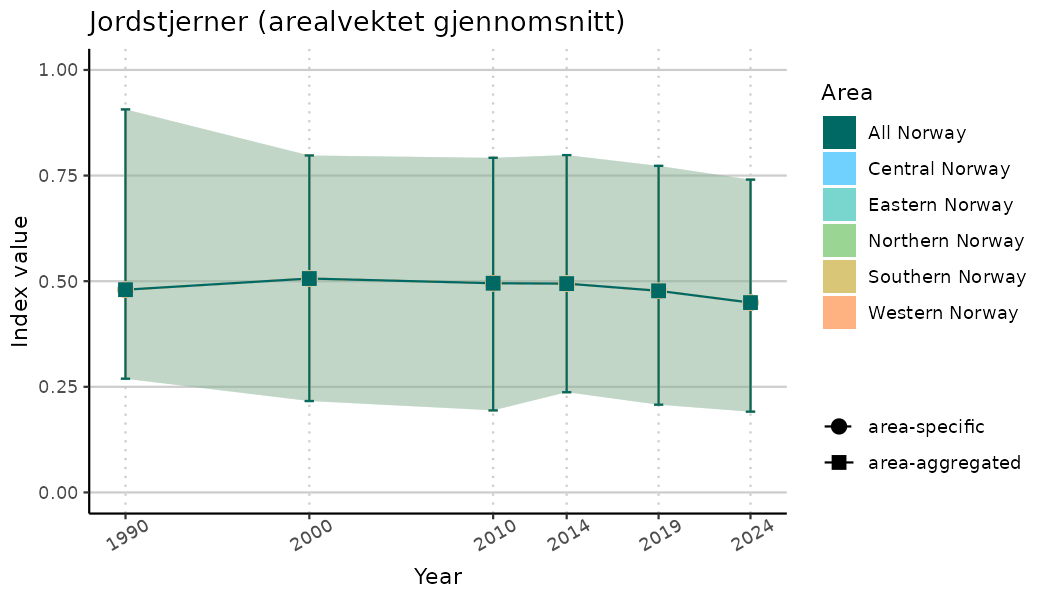
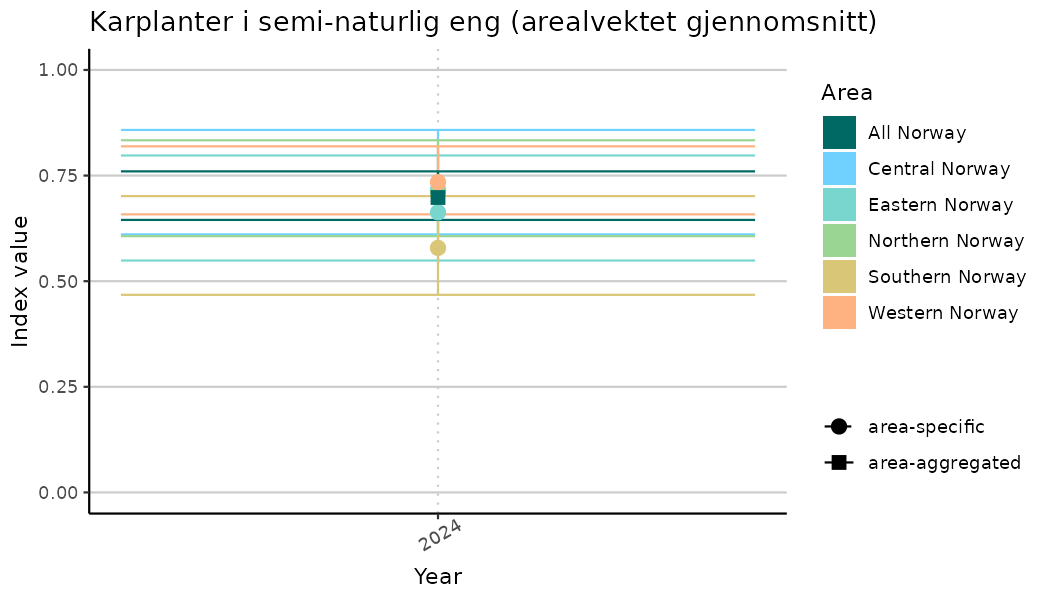
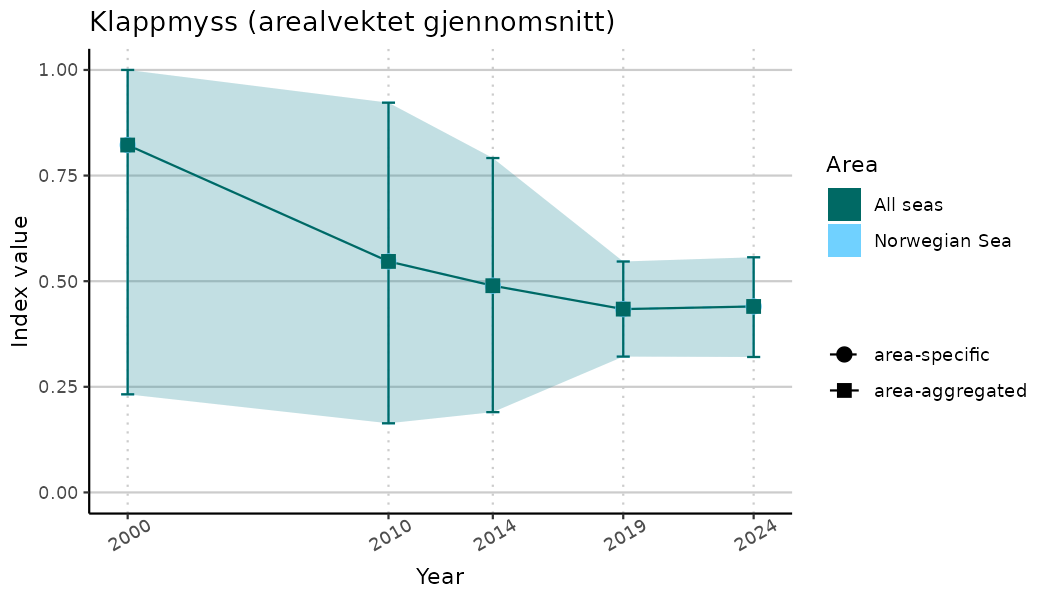
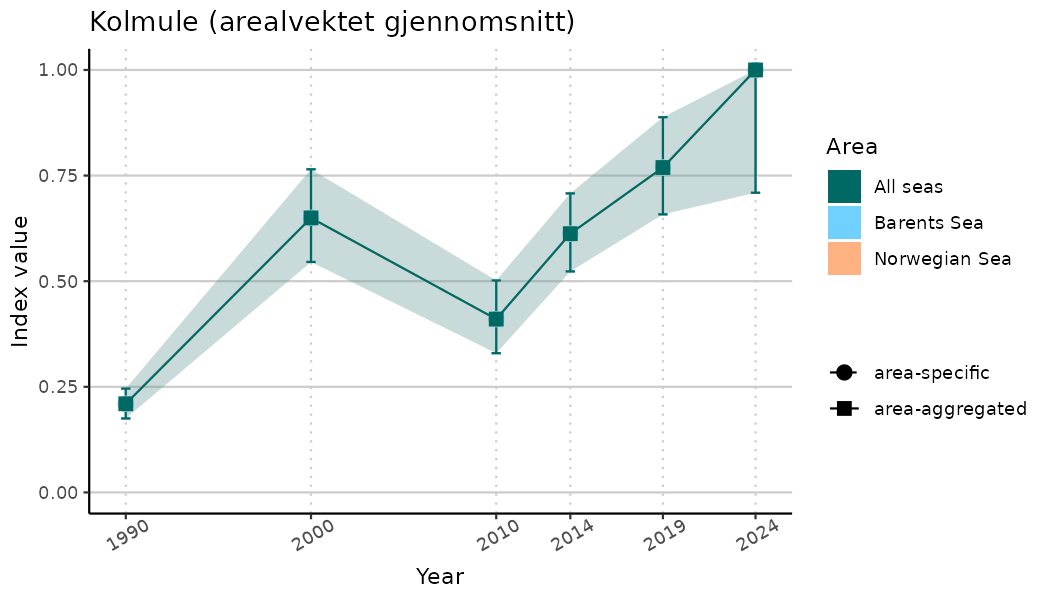
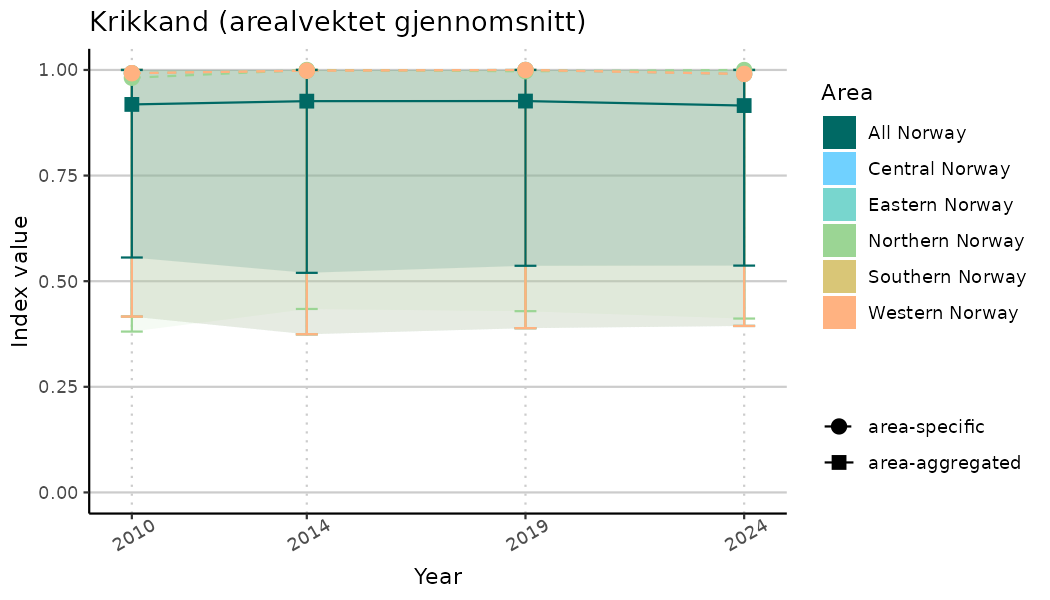
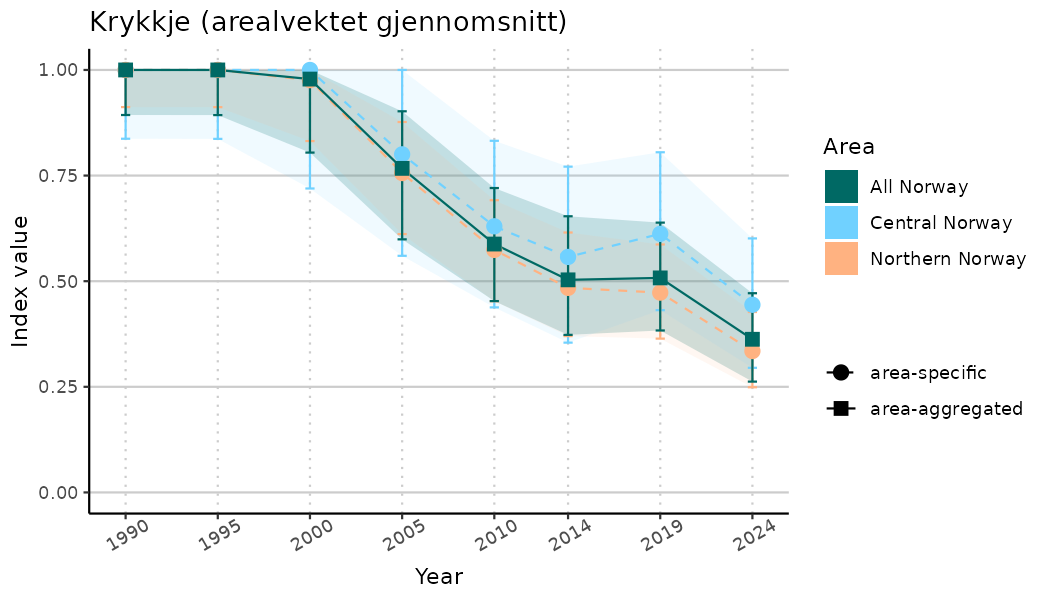
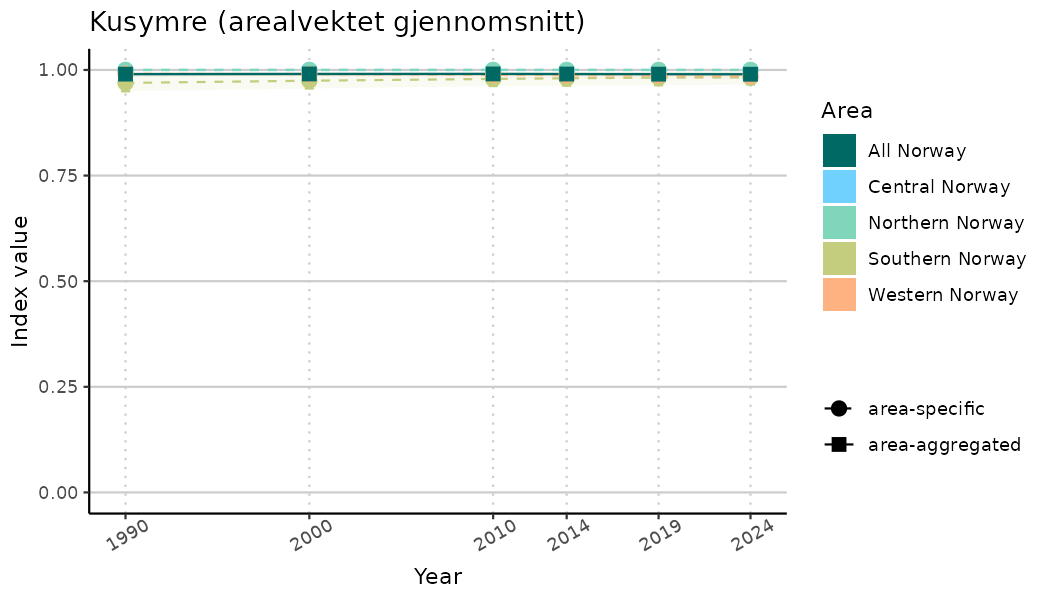
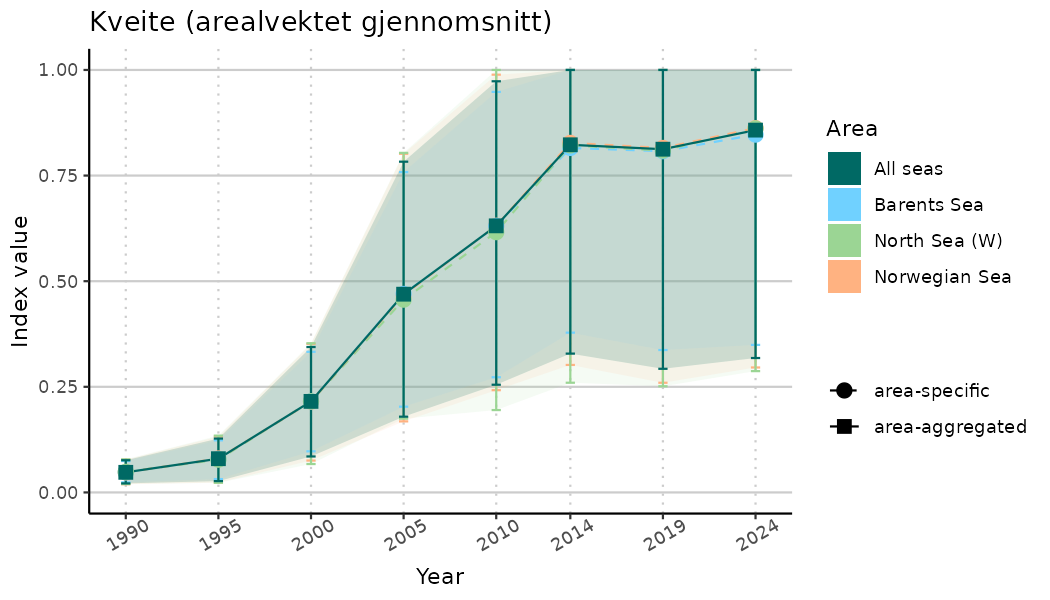
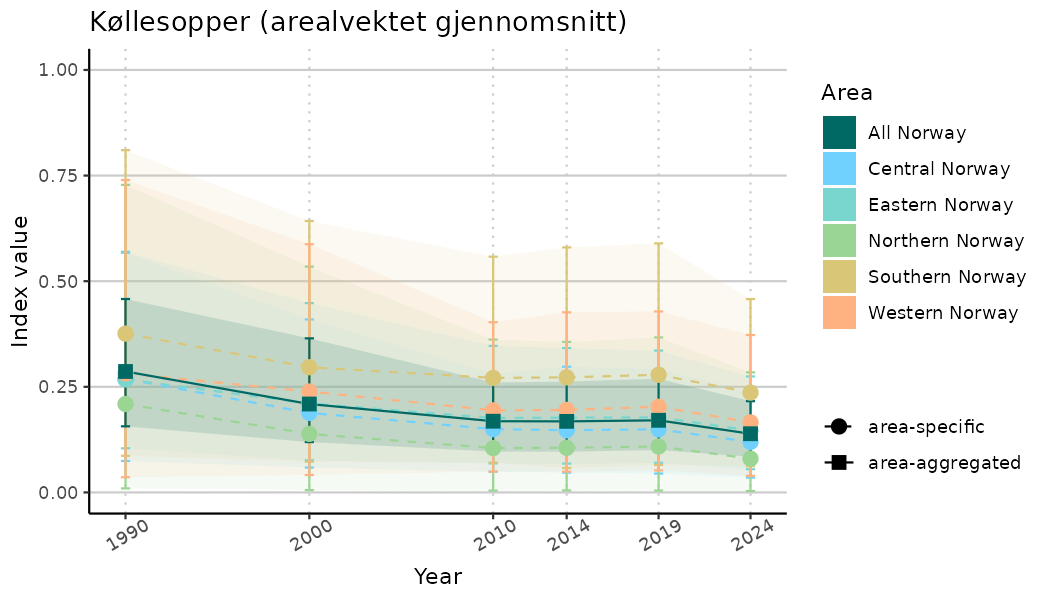
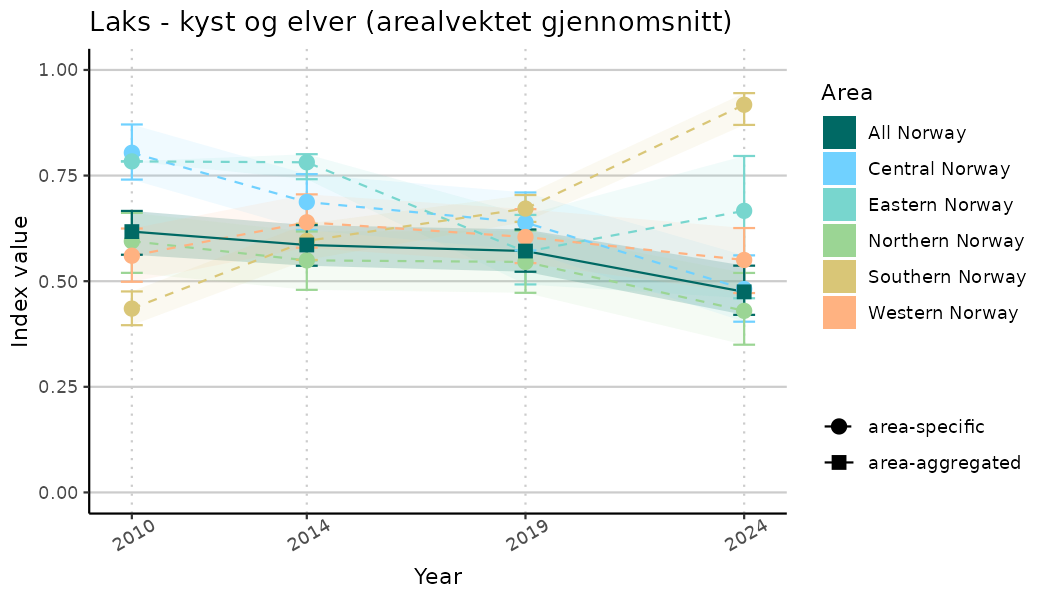
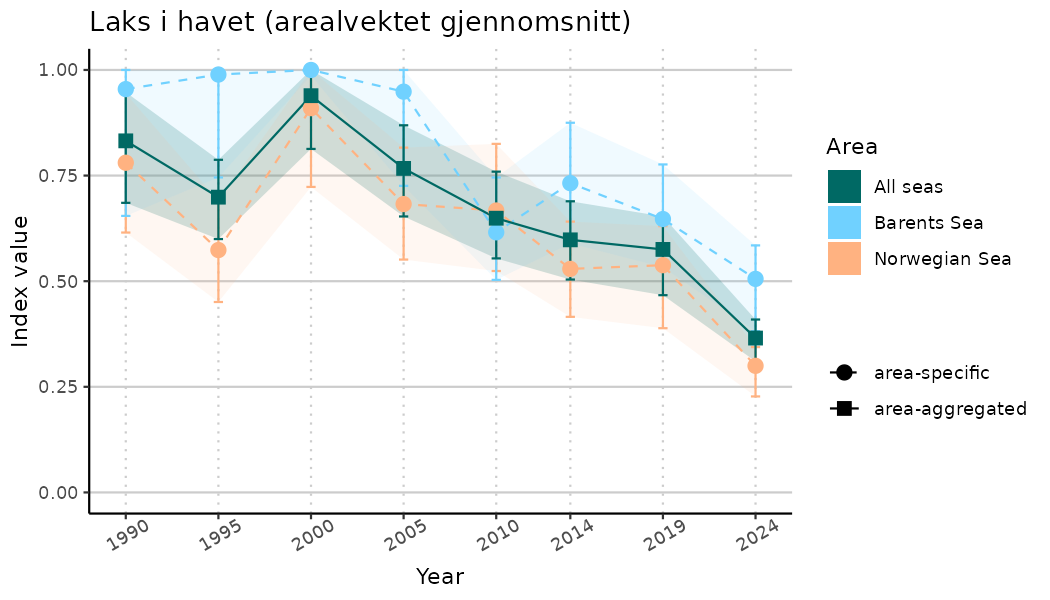
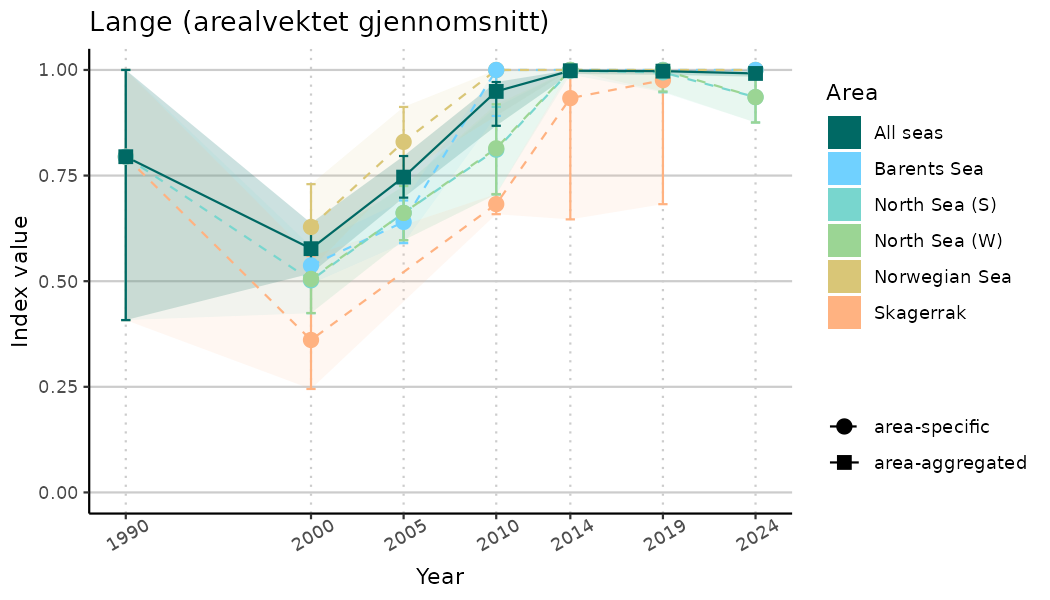
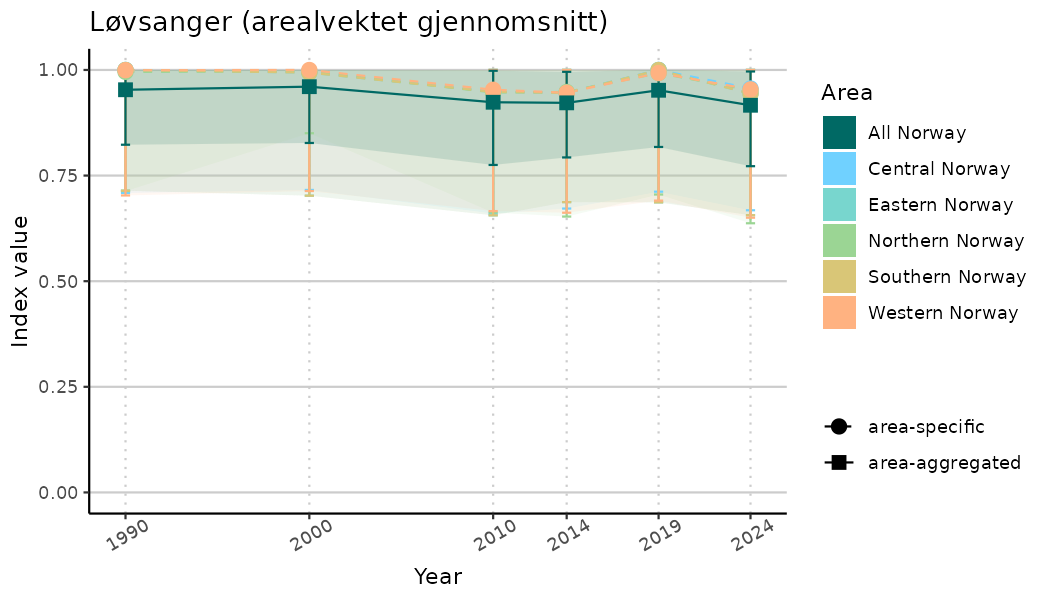
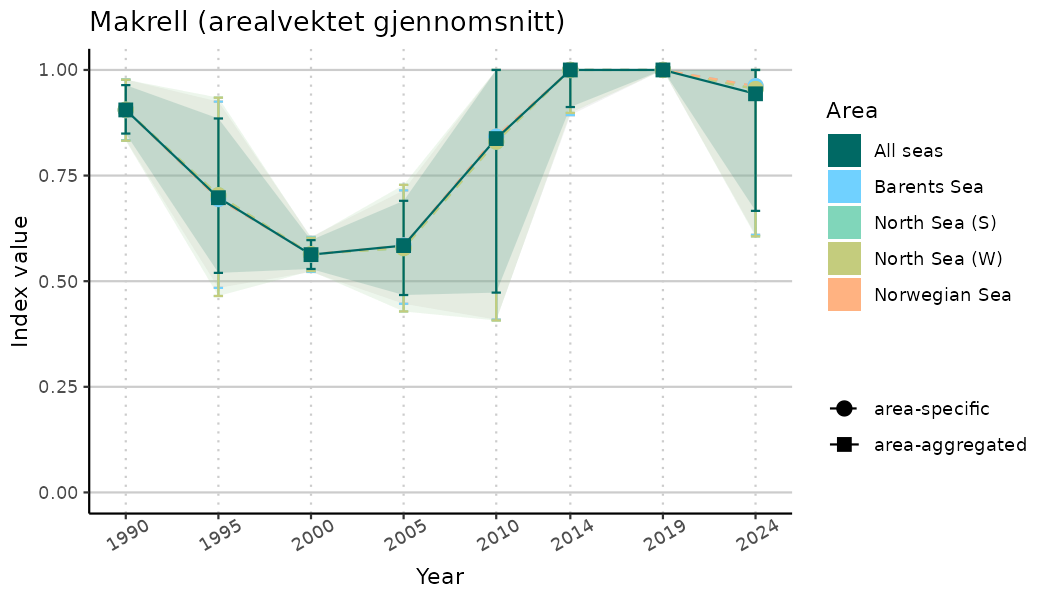
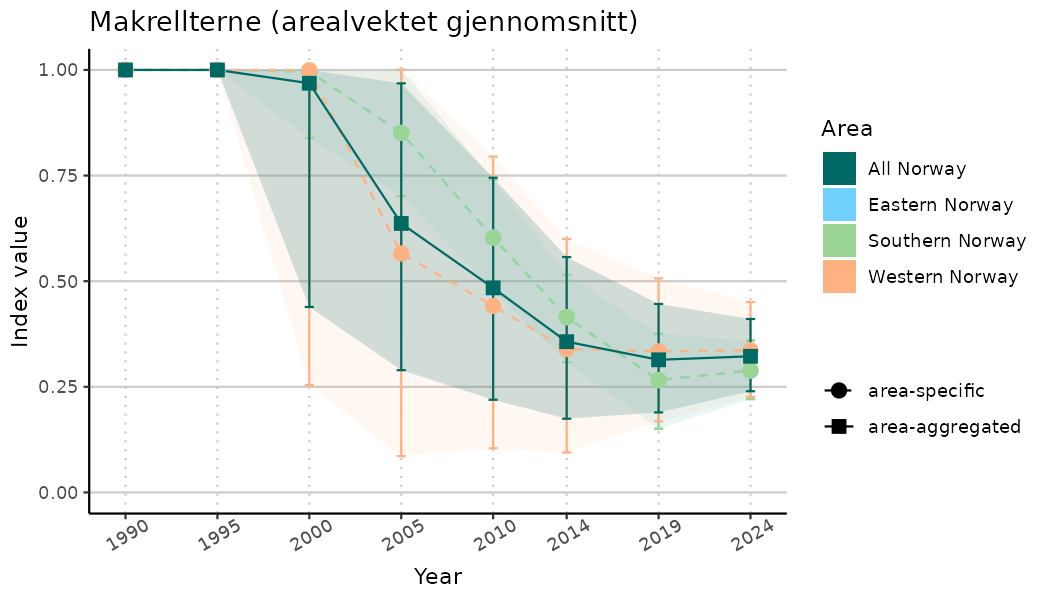
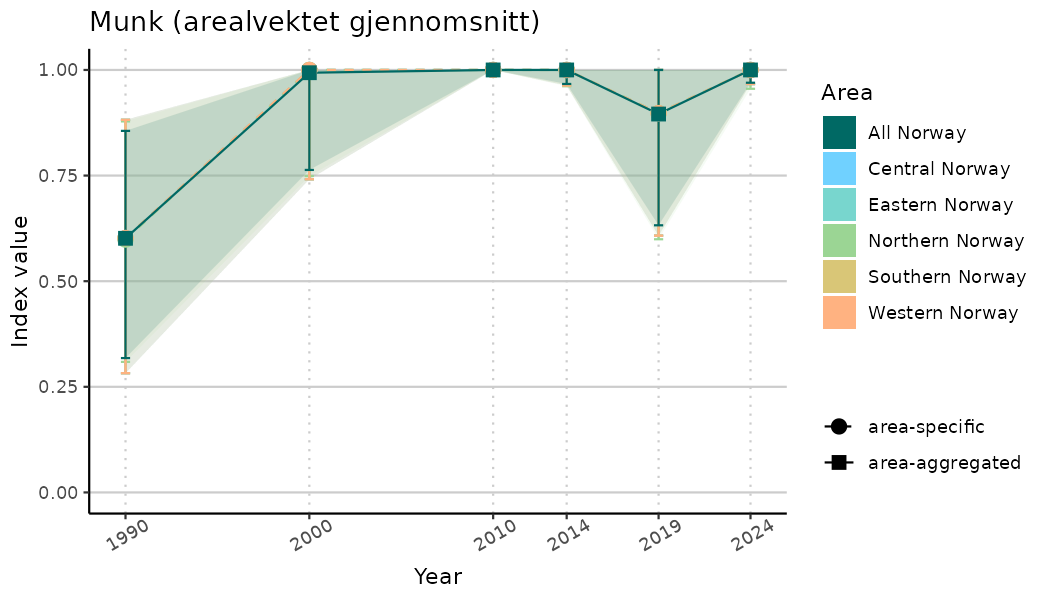
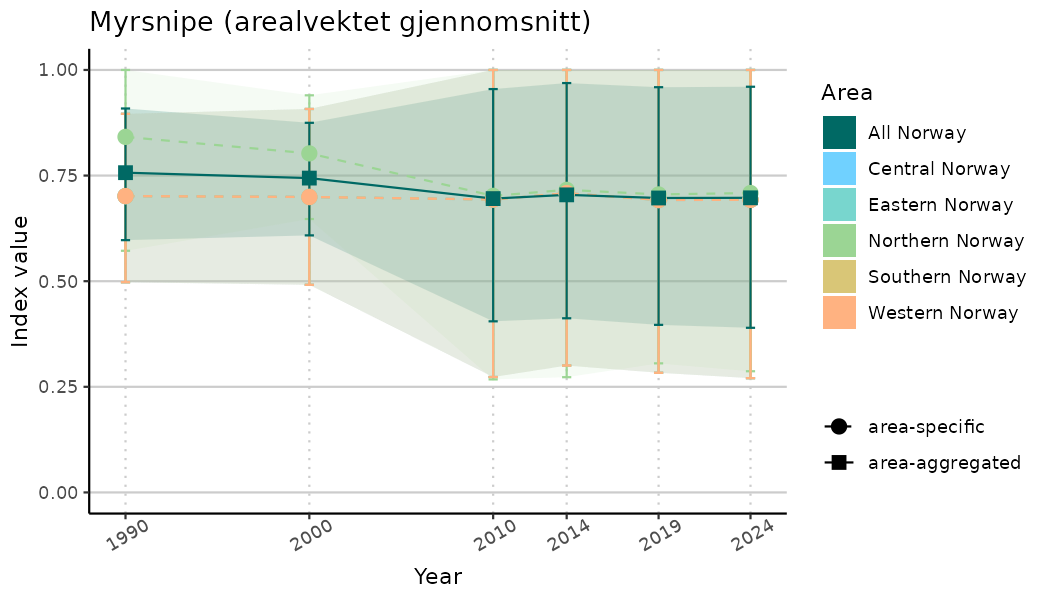
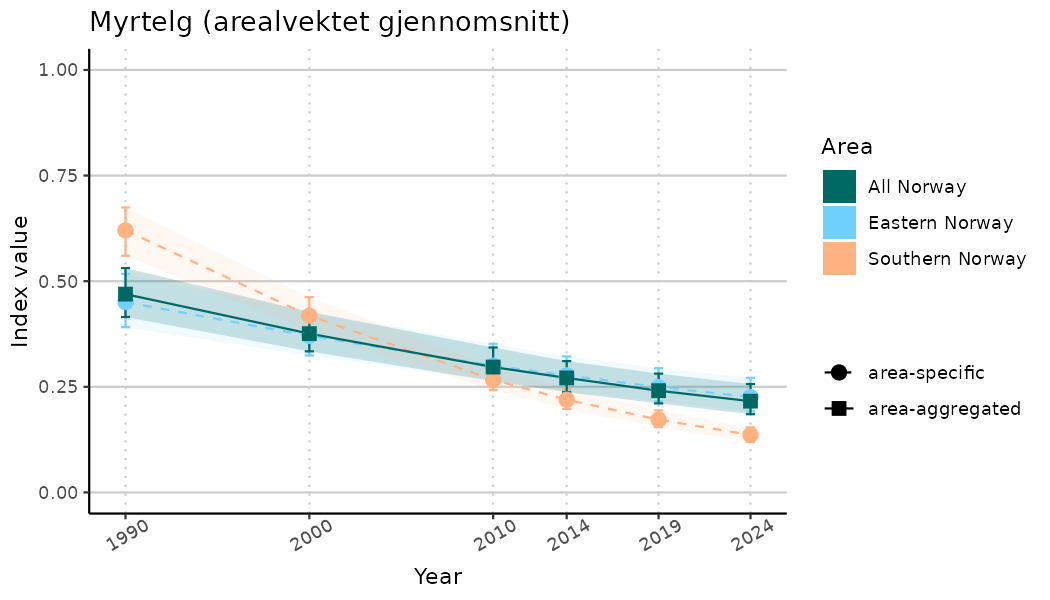
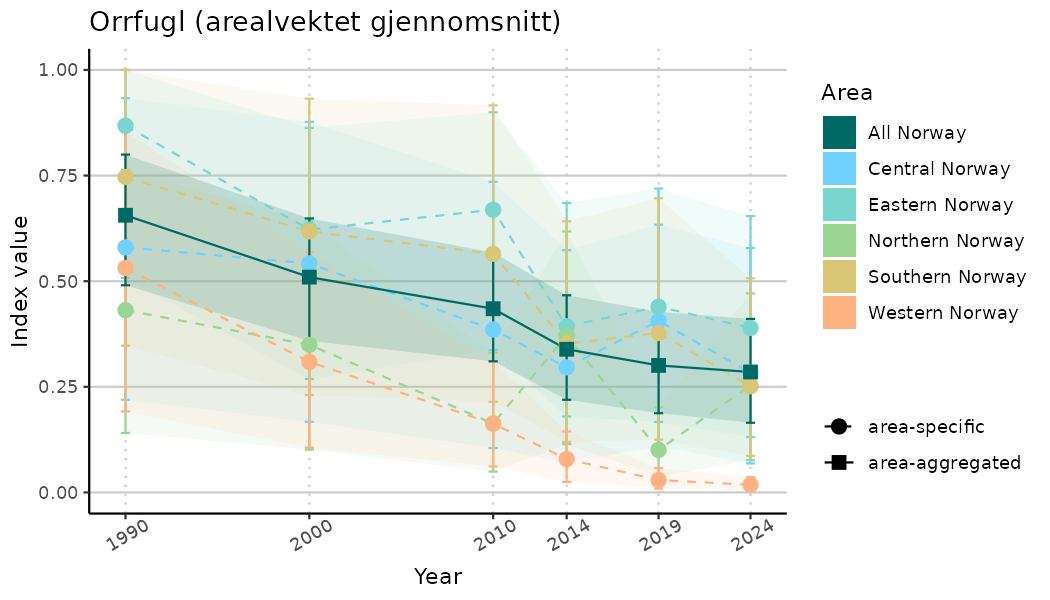
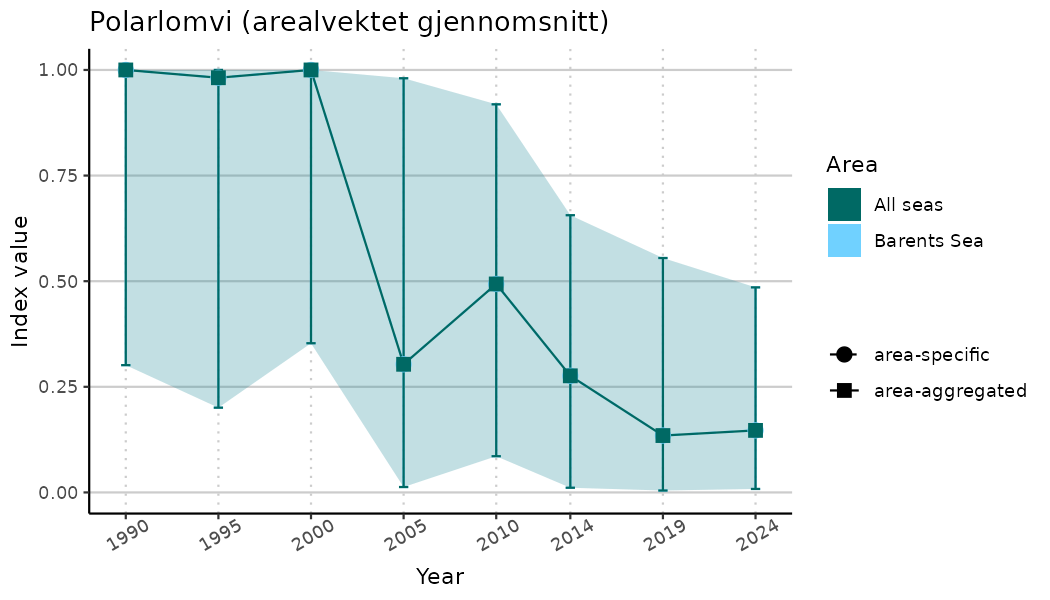
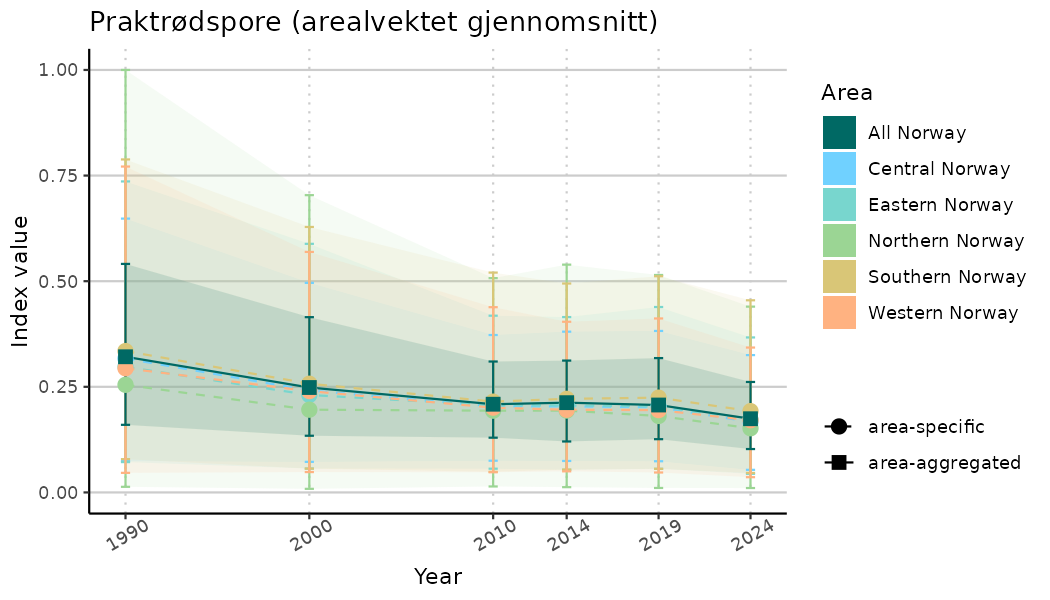
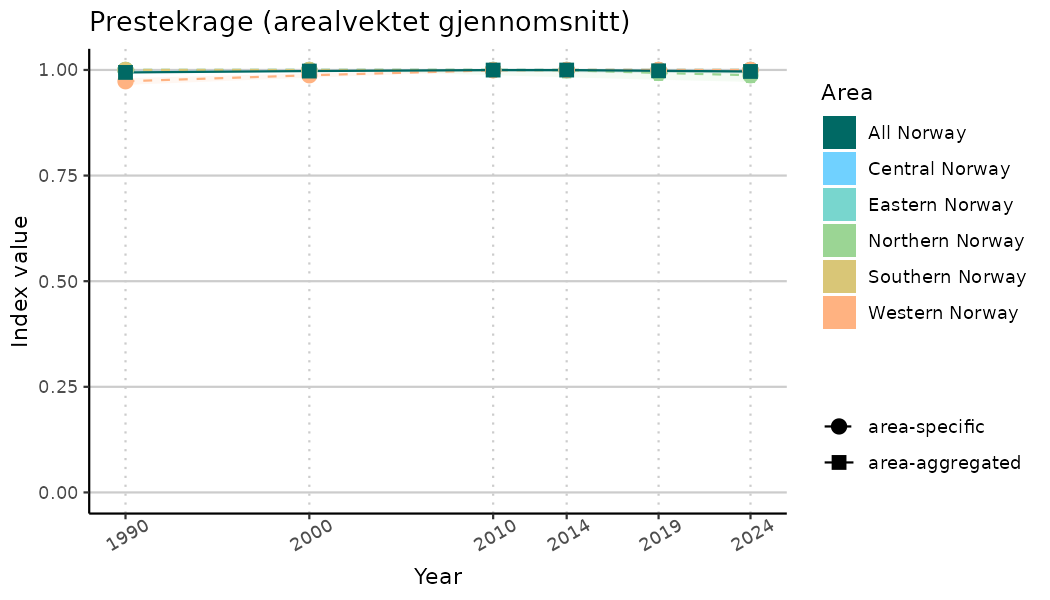
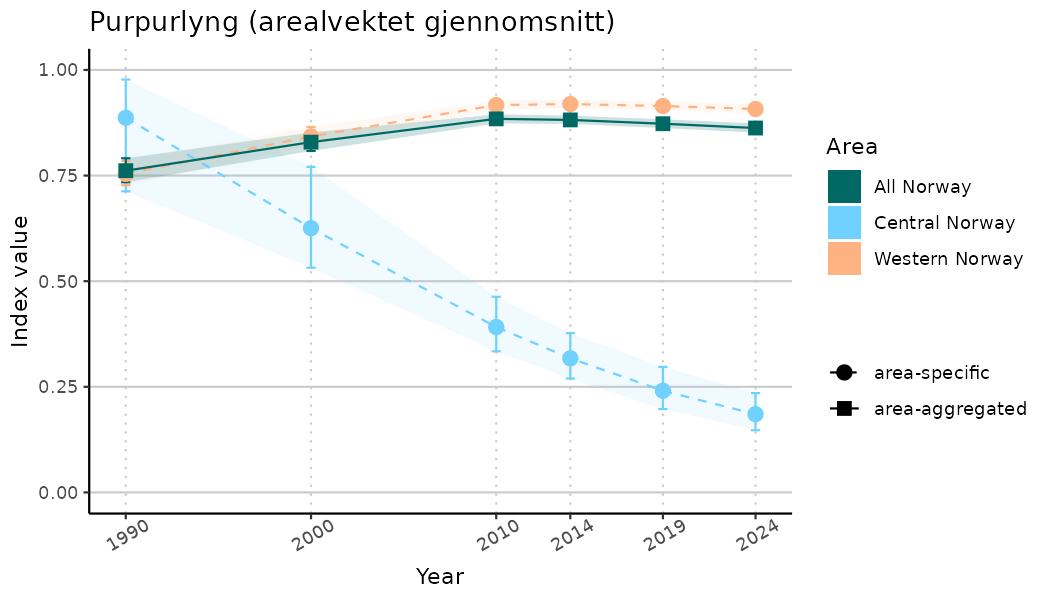
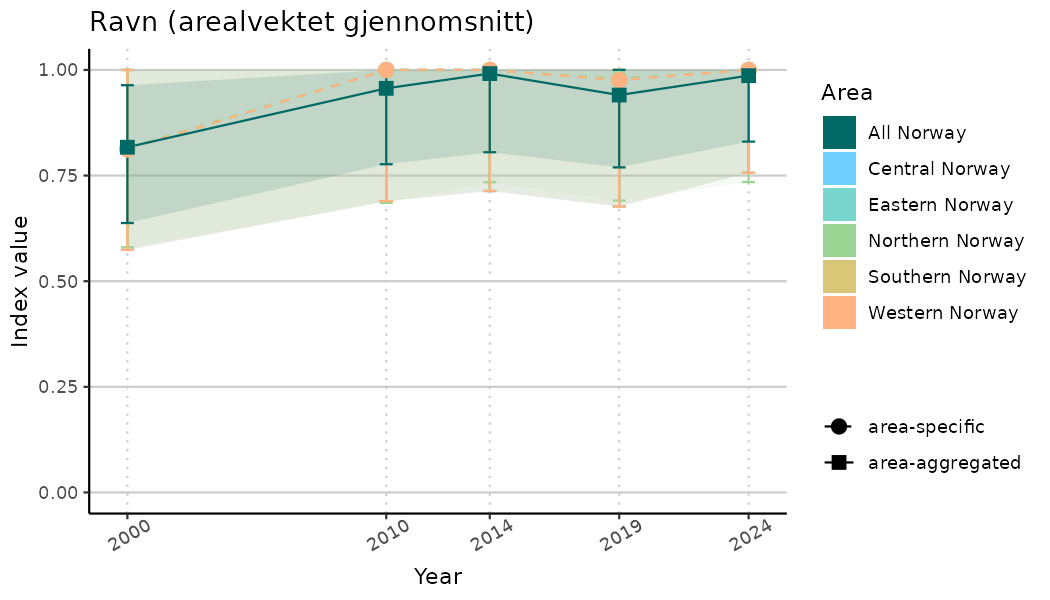
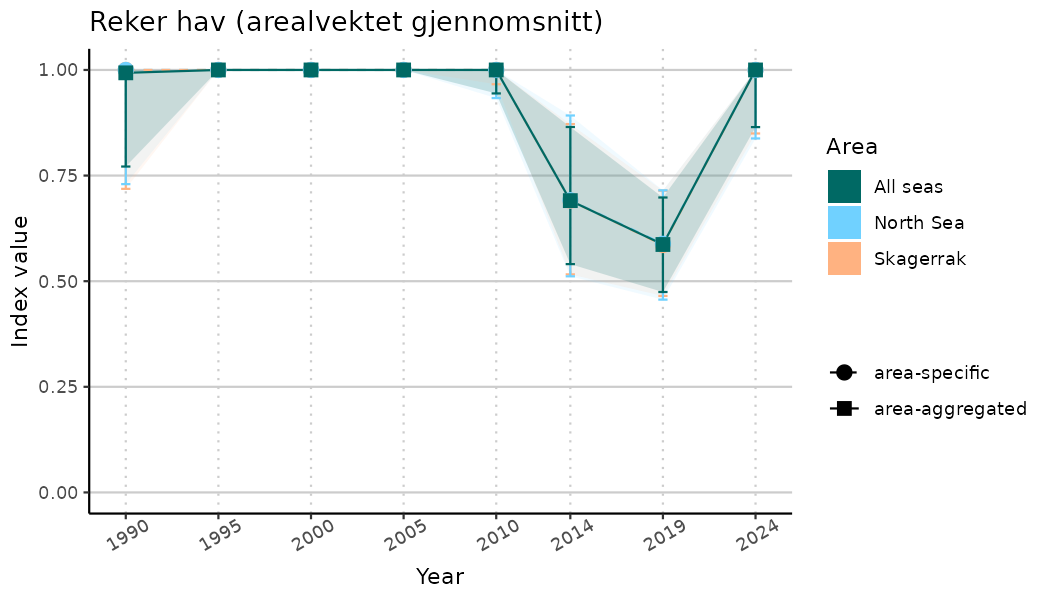
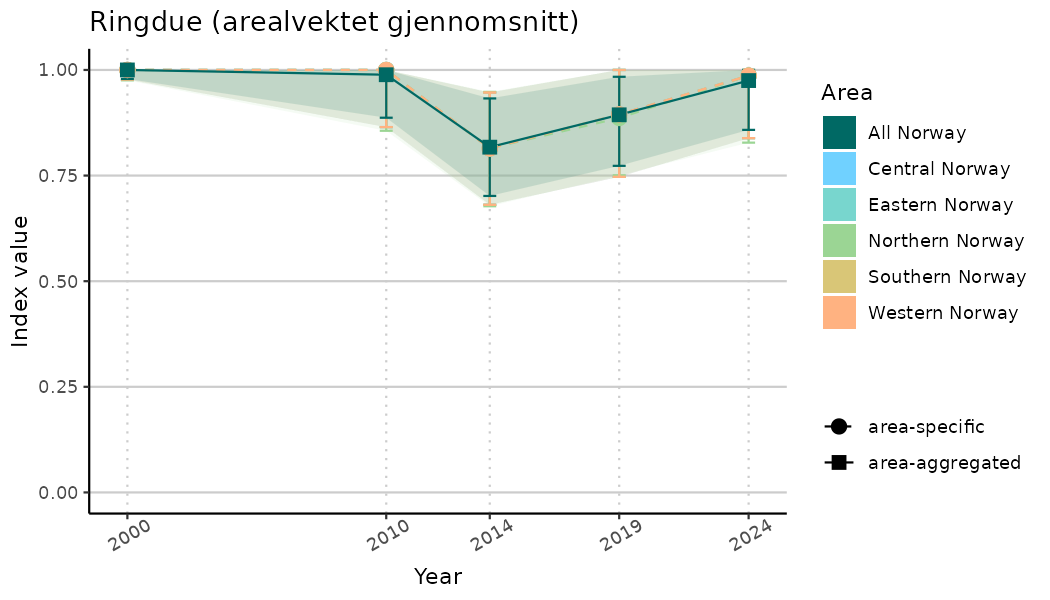
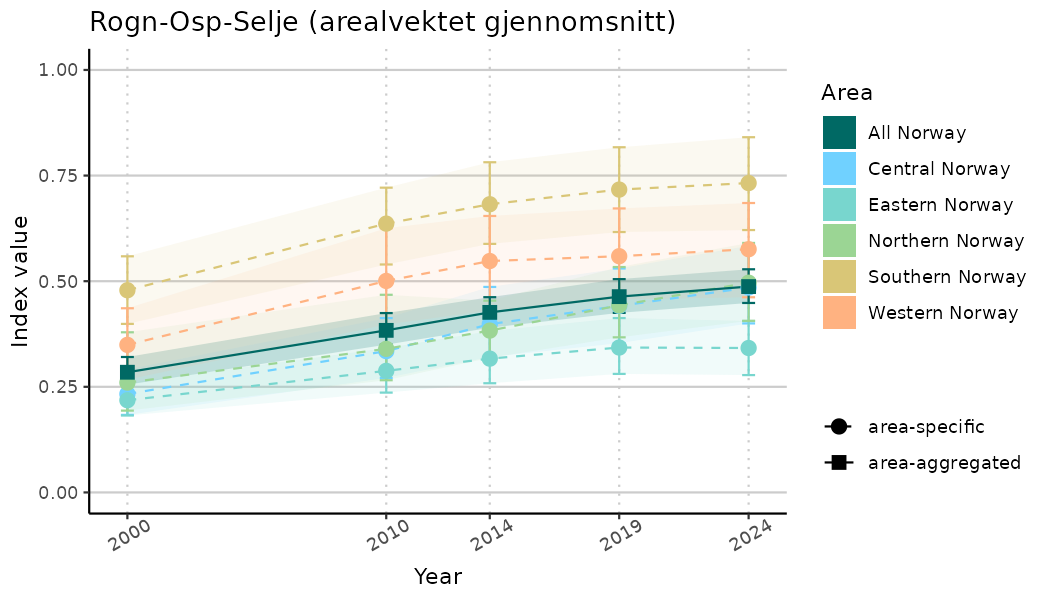
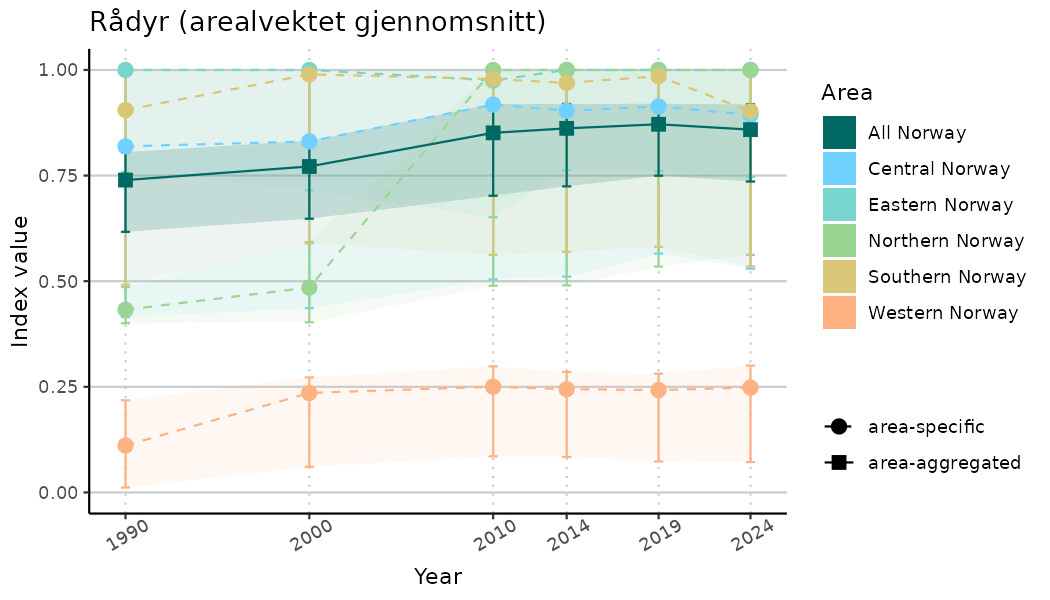
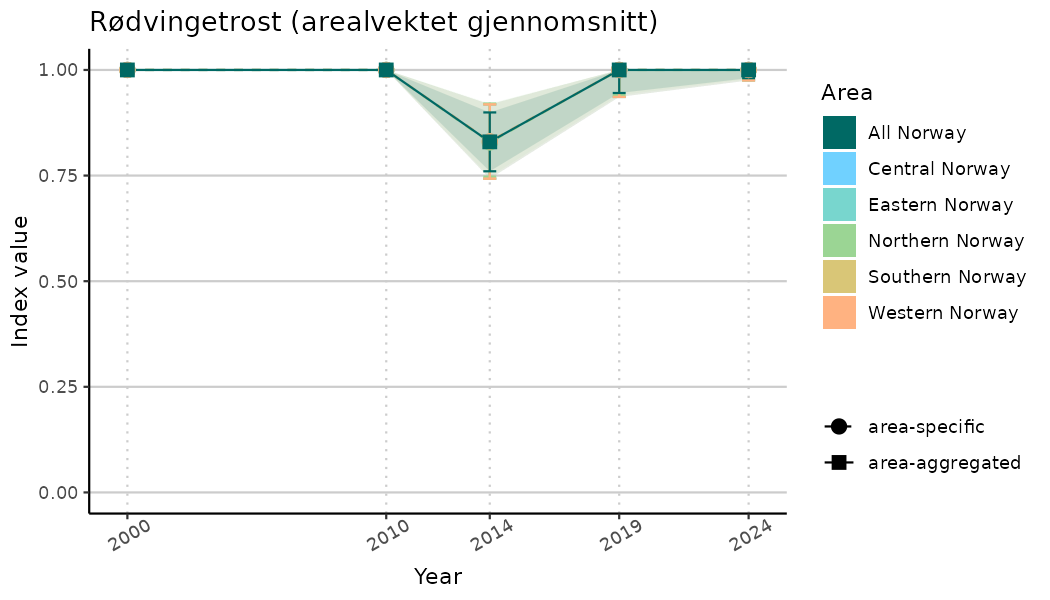
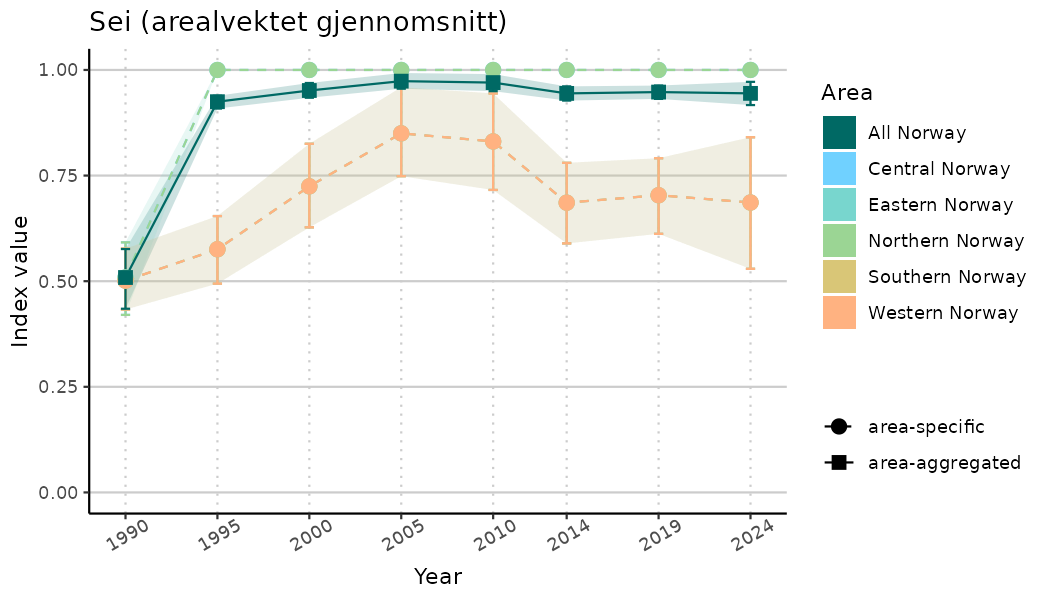
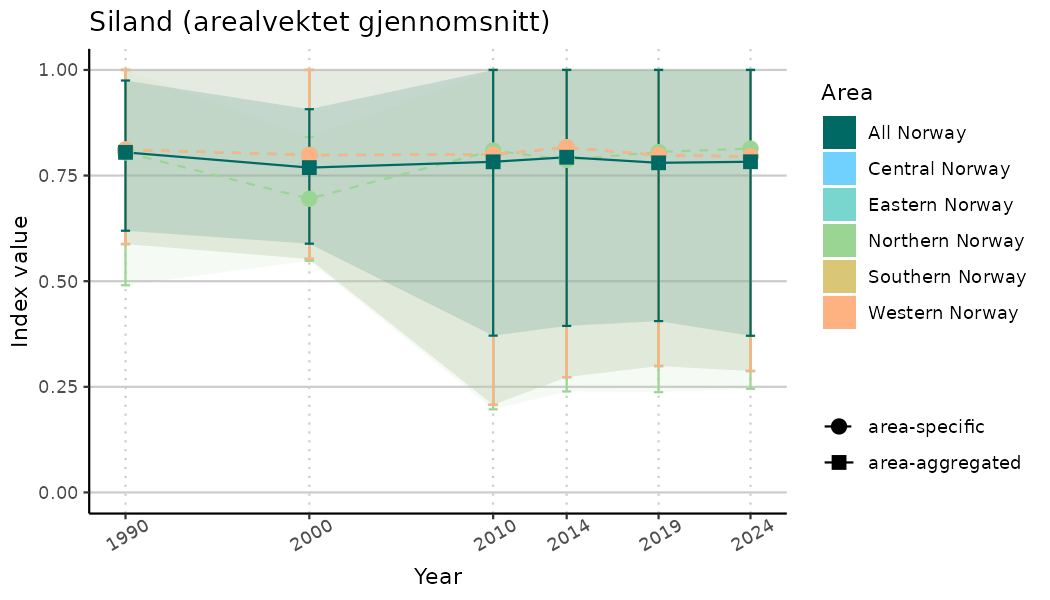
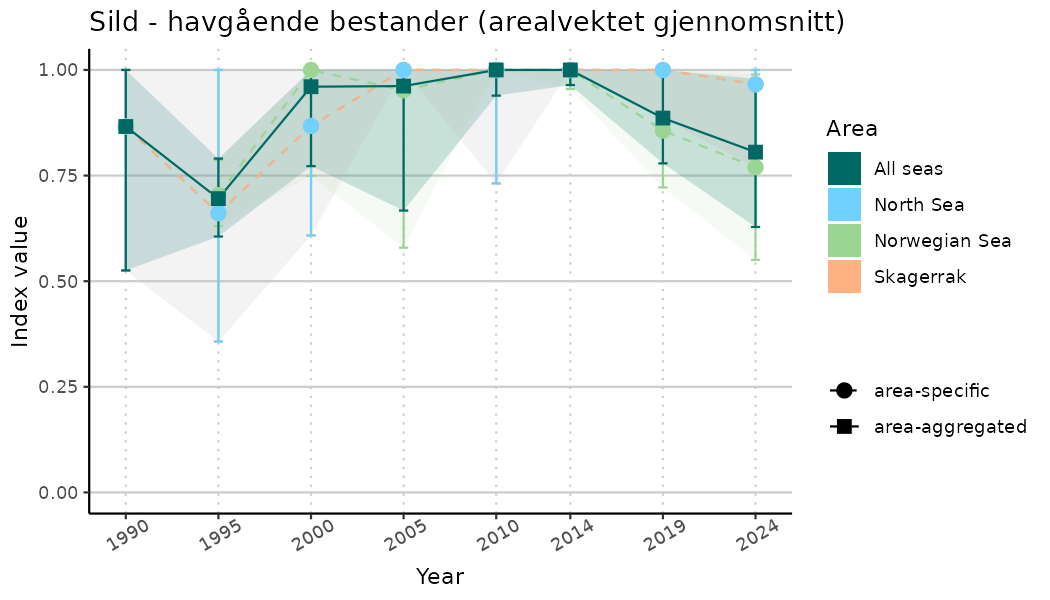
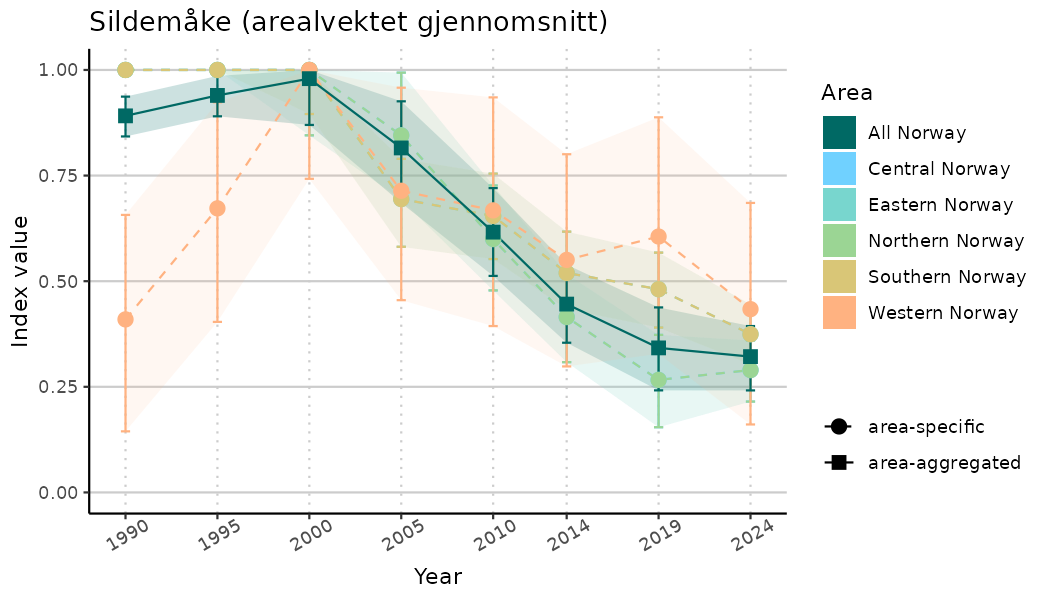
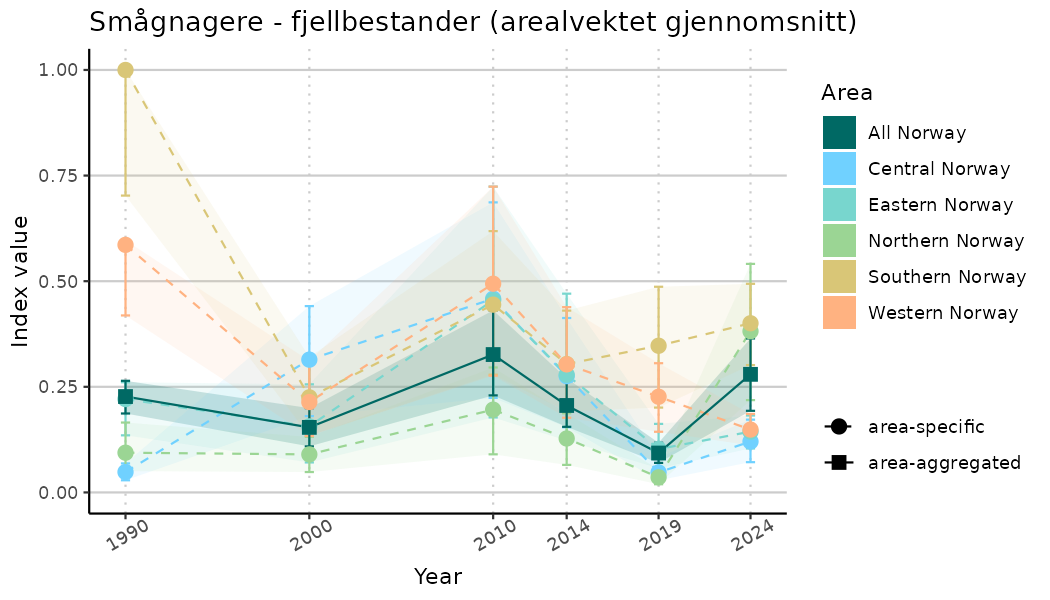
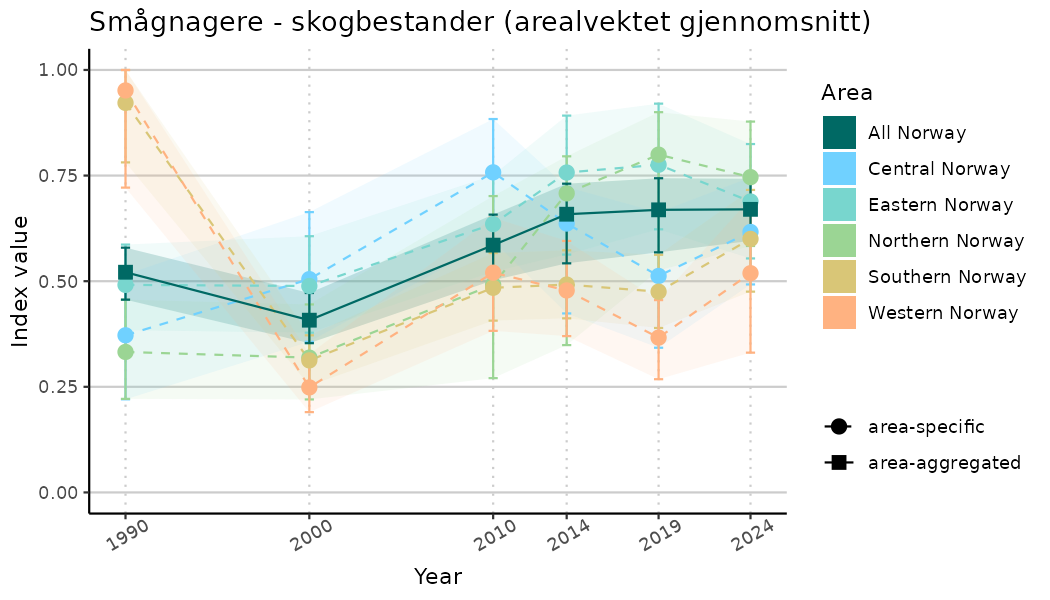
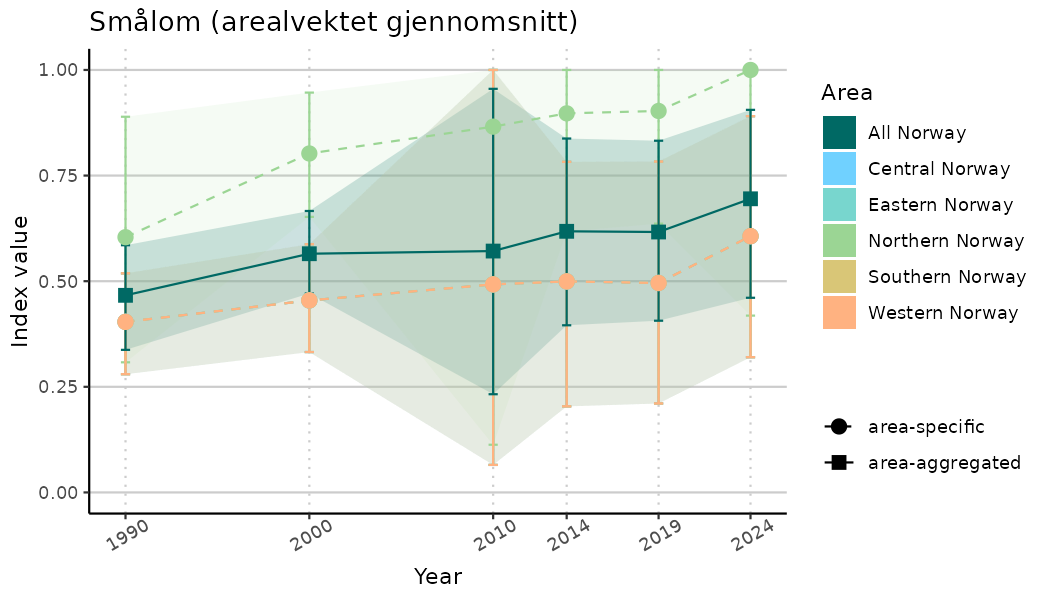
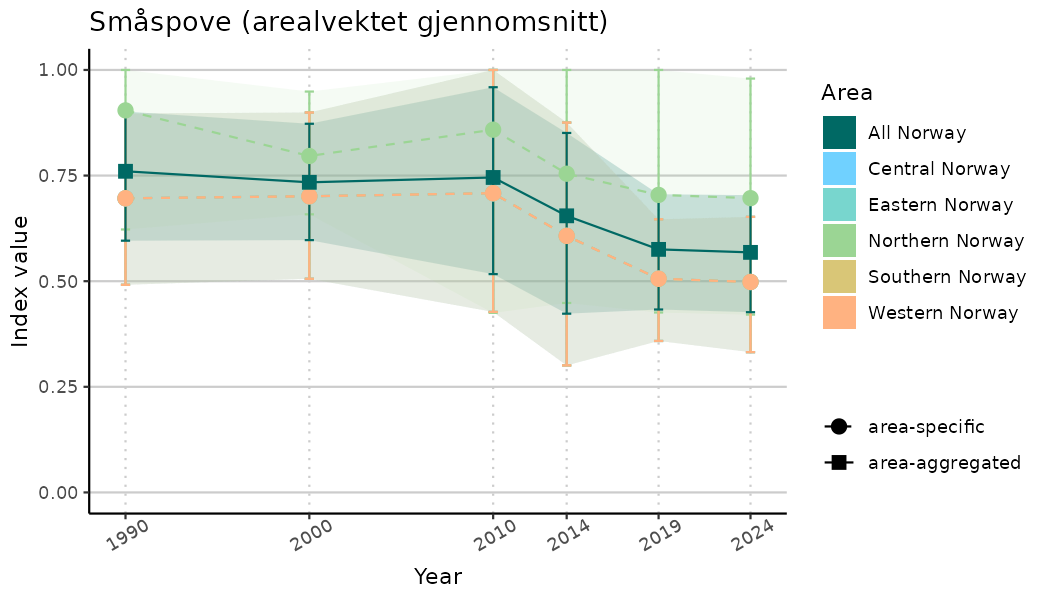
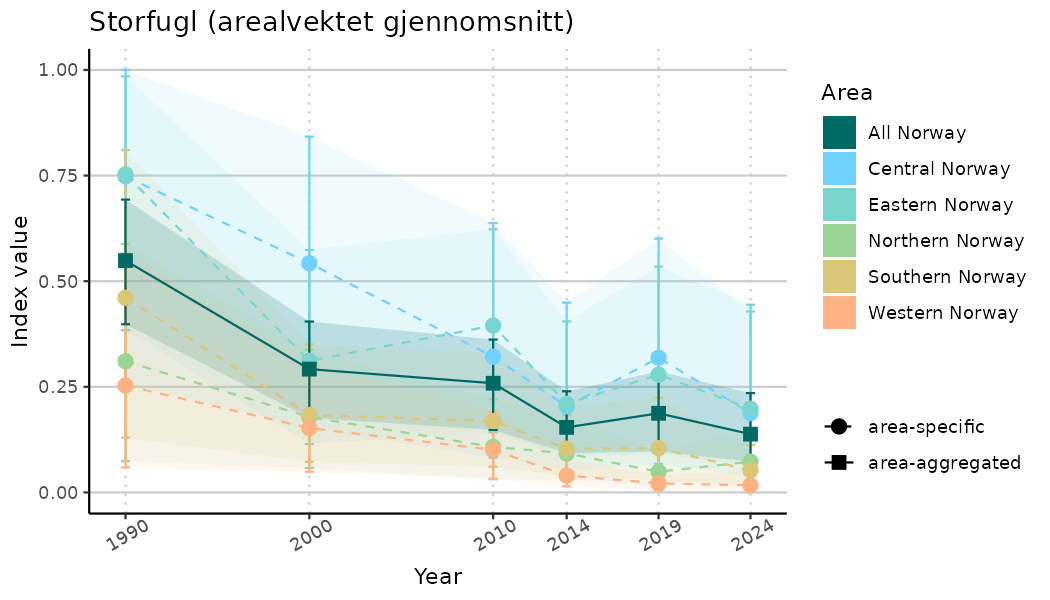
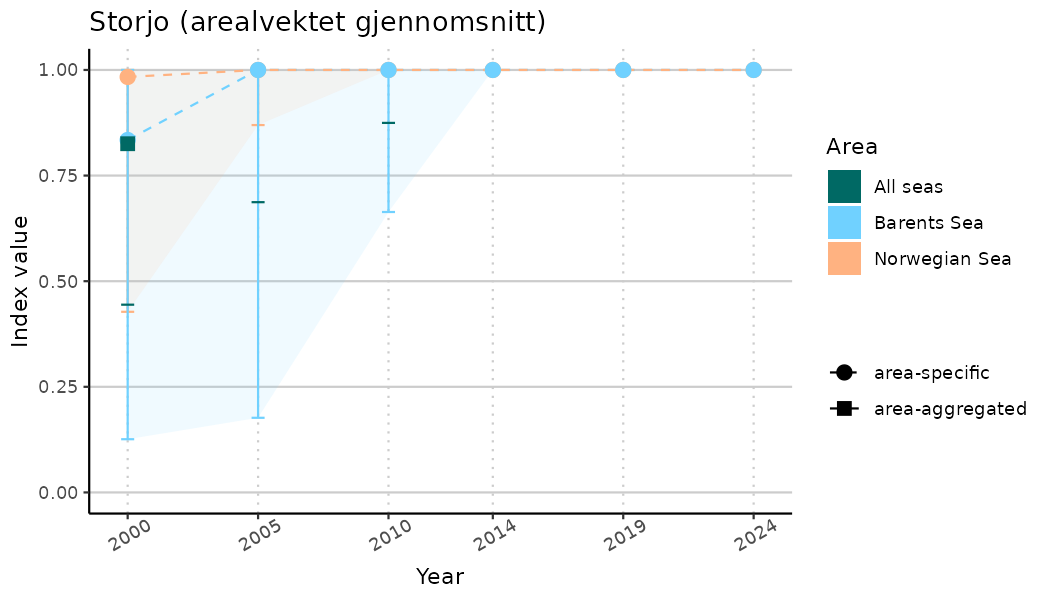
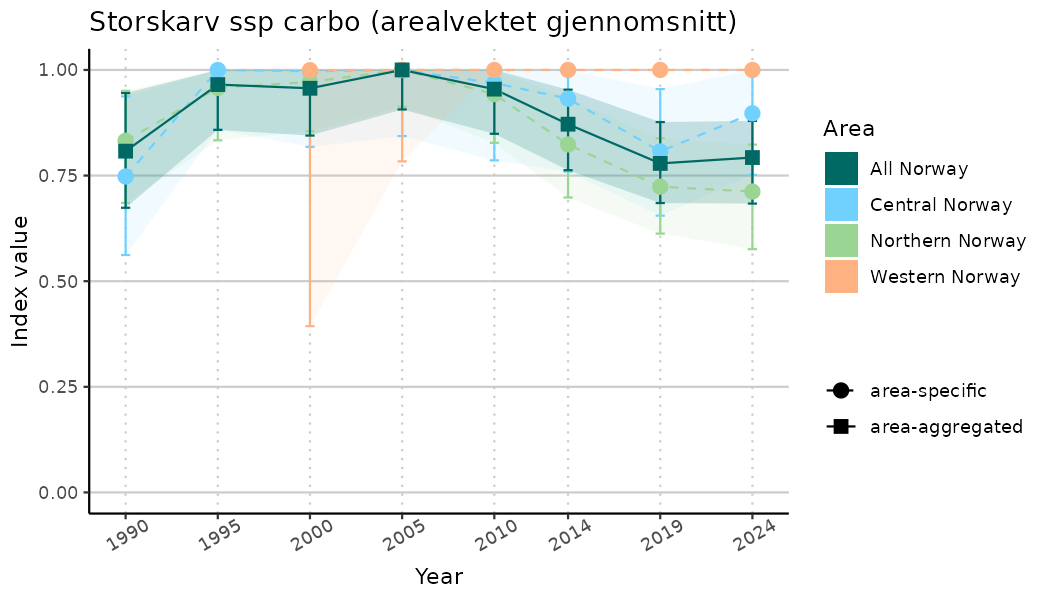
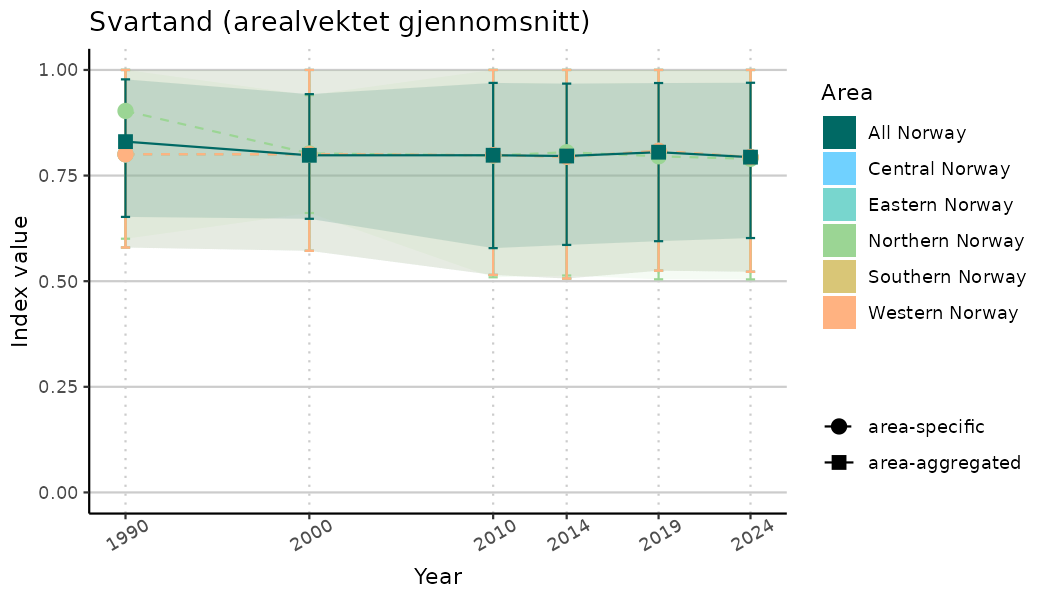
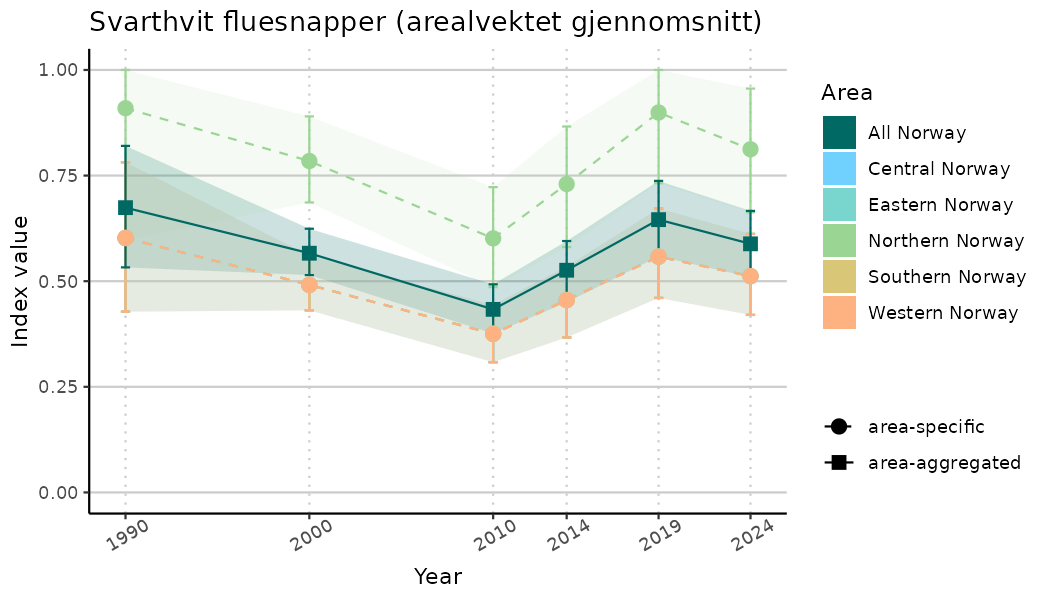
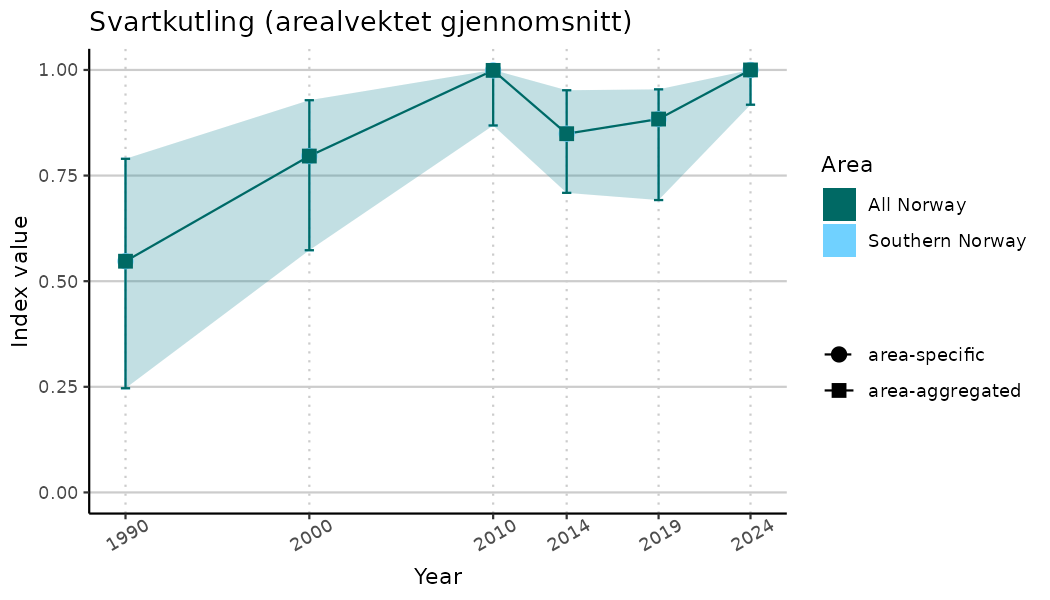
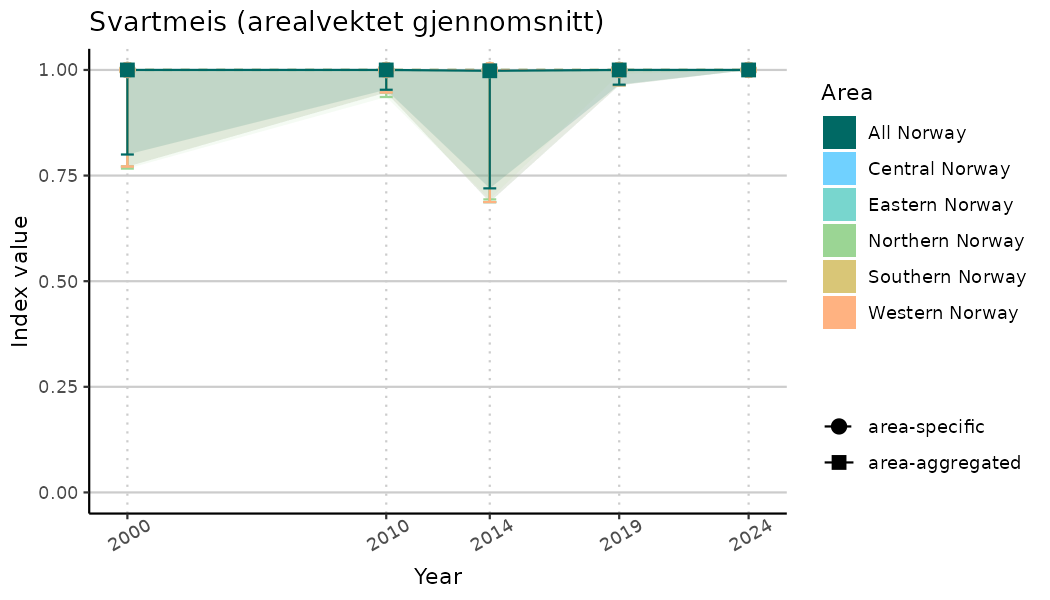
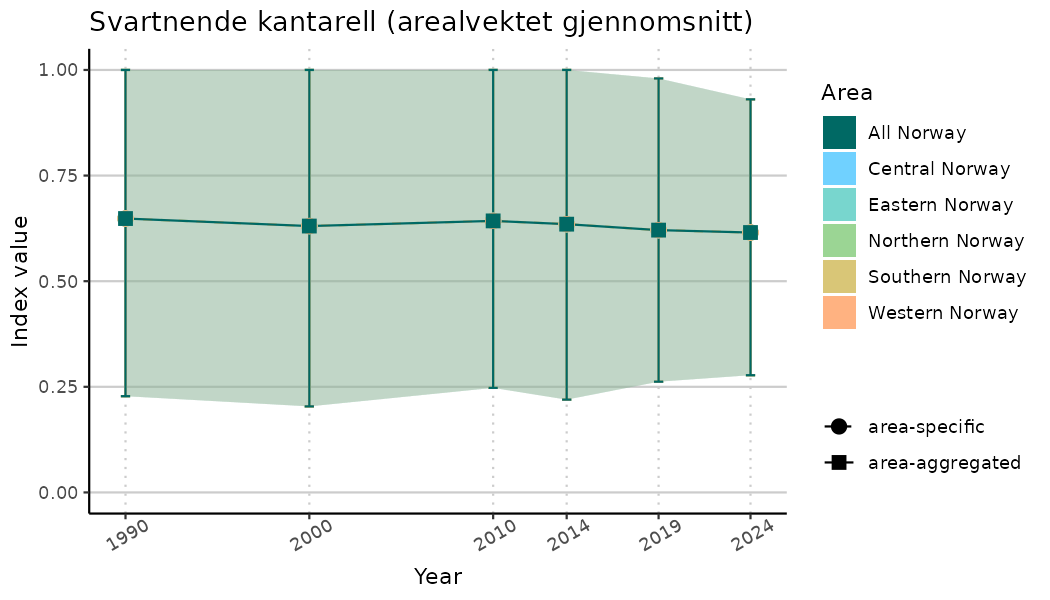
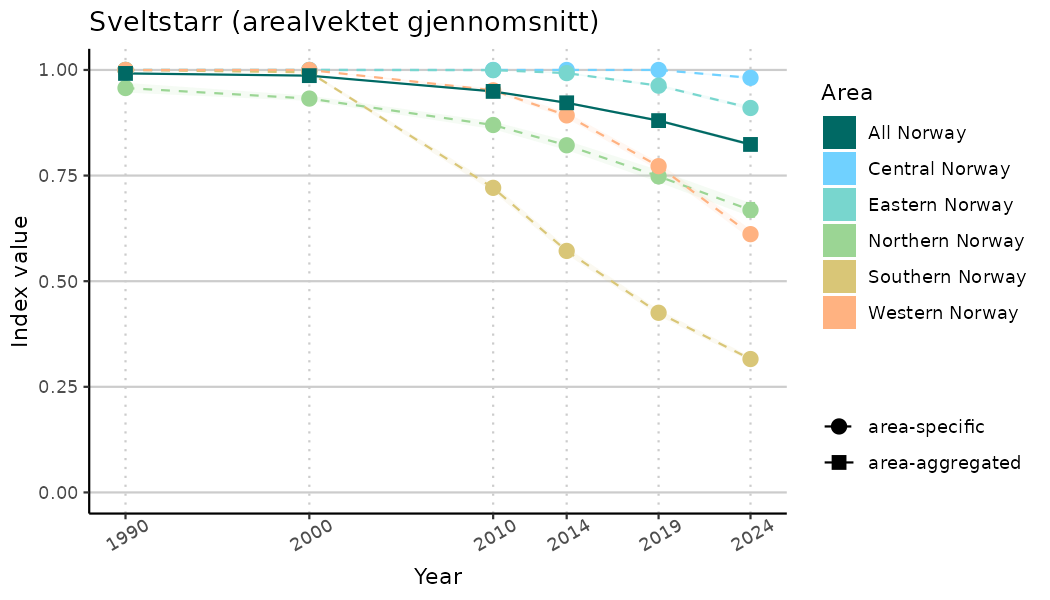
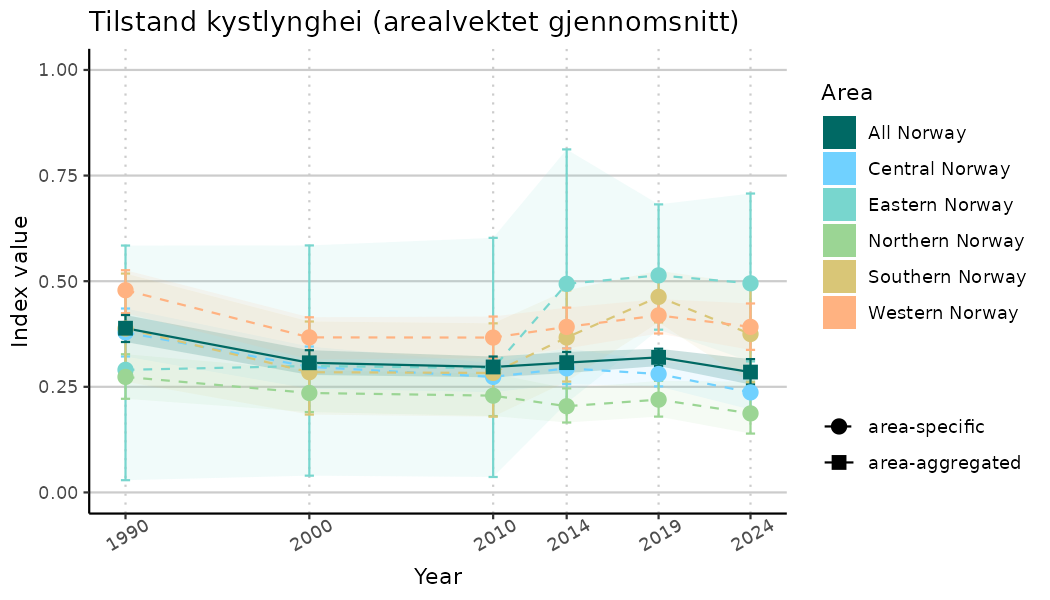
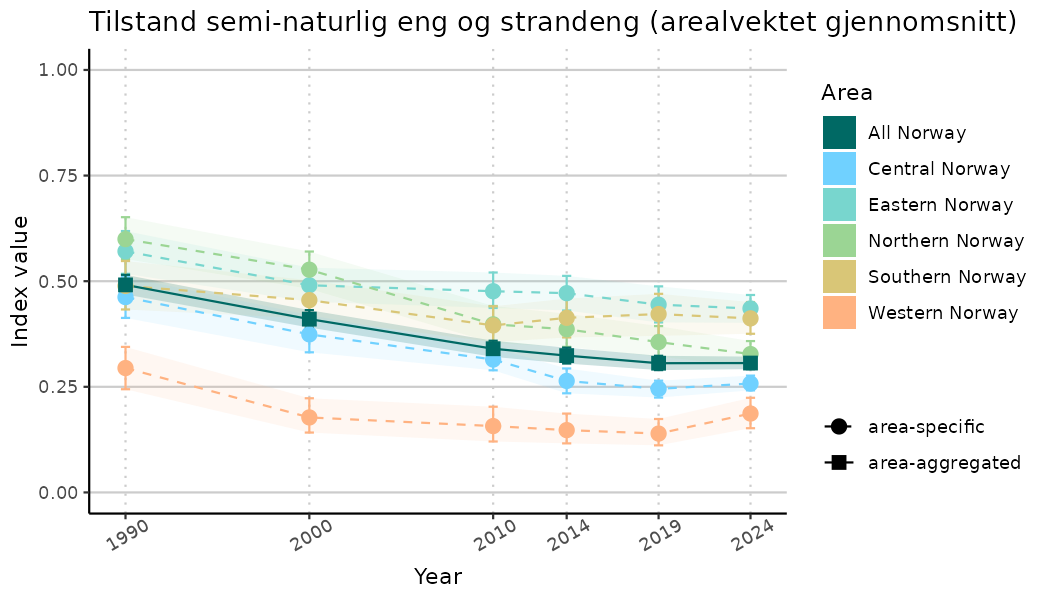
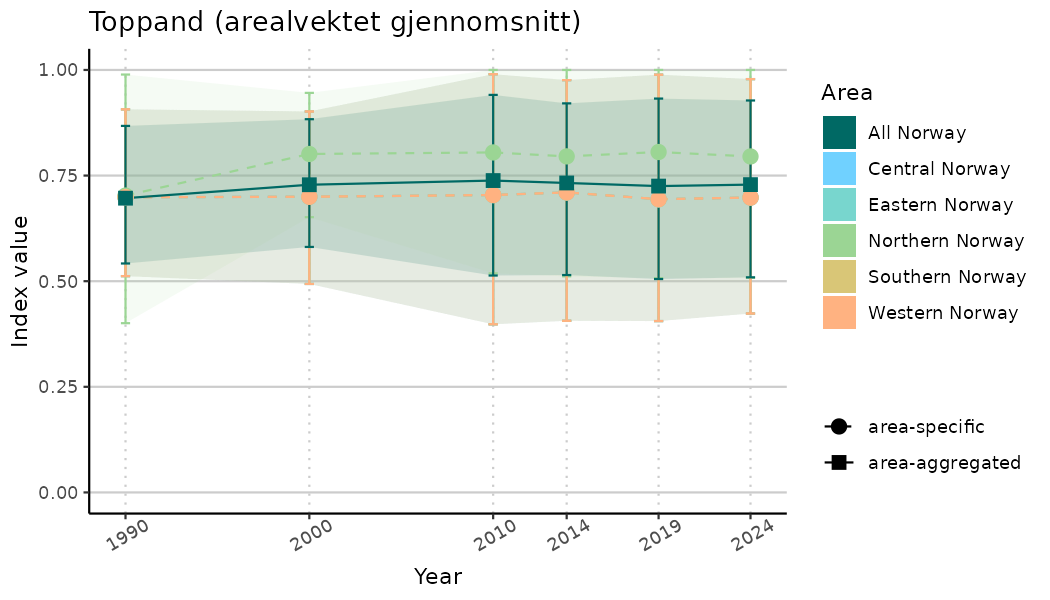
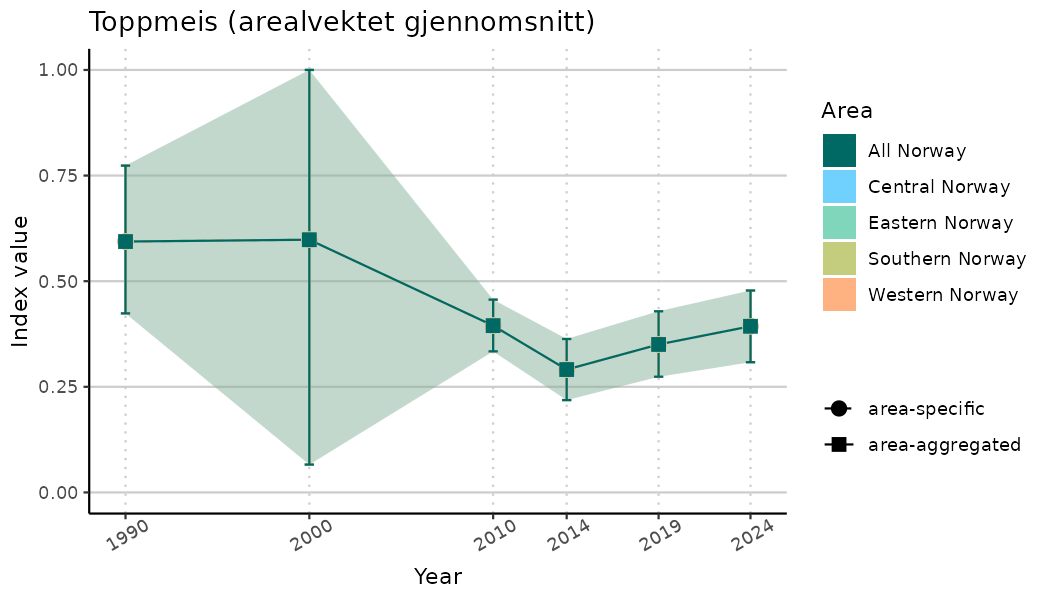
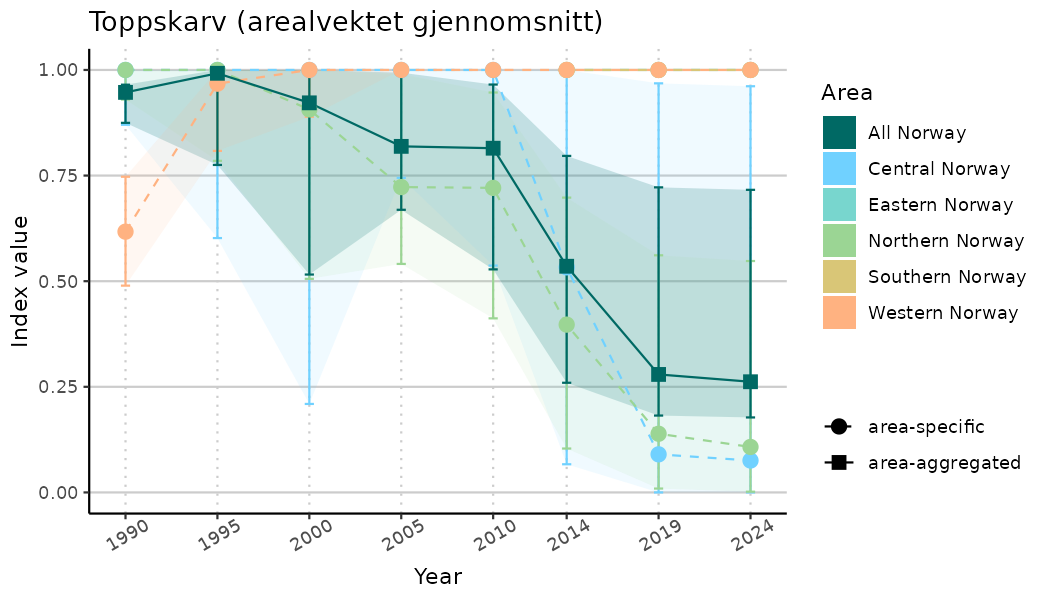
ts_indIdx2 <- NIflex::plotNI_StandardTS(Index = output_indIdx$indexOutput,
plotTitle = paste0(ind_name, " (arealvektet gjennomsnitt)"),
plotYears = ind_focalYears,
addAverage = TRUE,
truncateY = FALSE) +
theme(axis.text.x = element_text(angle = 30, vjust = 0.8))
TS_indIdx[[i]] <- ts_indIdx
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx.svg"), plot = ts_indIdx, width = 7, height = 4)
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx.pdf"), plot = ts_indIdx, width = 7, height = 4)
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx.png"), plot = ts_indIdx, width = 7, height = 4, dpi = 150)
TS_indIdx_std[[i]] <- ts_indIdx2
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx_std.svg"), plot = ts_indIdx2, width = 7, height = 4)
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx_std.pdf"), plot = ts_indIdx2, width = 7, height = 4)
ggsave(file = paste0(savePath_i, "/TimeSeries_indIdx_std.png"), plot = ts_indIdx2, width = 7, height = 4, dpi = 150)
message("")
}
## Save entire plot lists for re-use later
saveRDS(TS_raw, file = "results_Indicators/PlotList_TimeSeries_raw.rds")
saveRDS(TS_scaled, file = "results_Indicators/PlotList_TimeSeries_scaled.rds")
saveRDS(TS_indIdx, file = "results_Indicators/PlotList_TimeSeries_indIdx.rds")
saveRDS(TS_indIdx_std, file = "results_Indicators/PlotList_TimeSeries_indIdx_std.rds")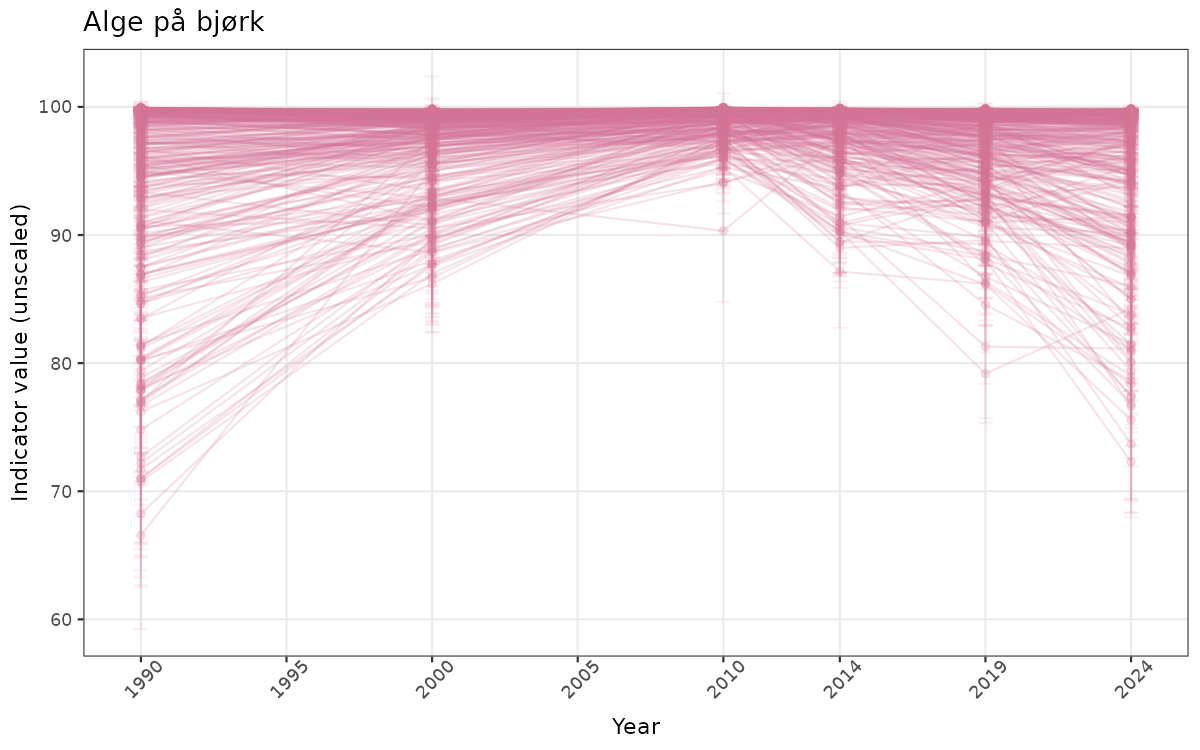










/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)
/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)
/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)











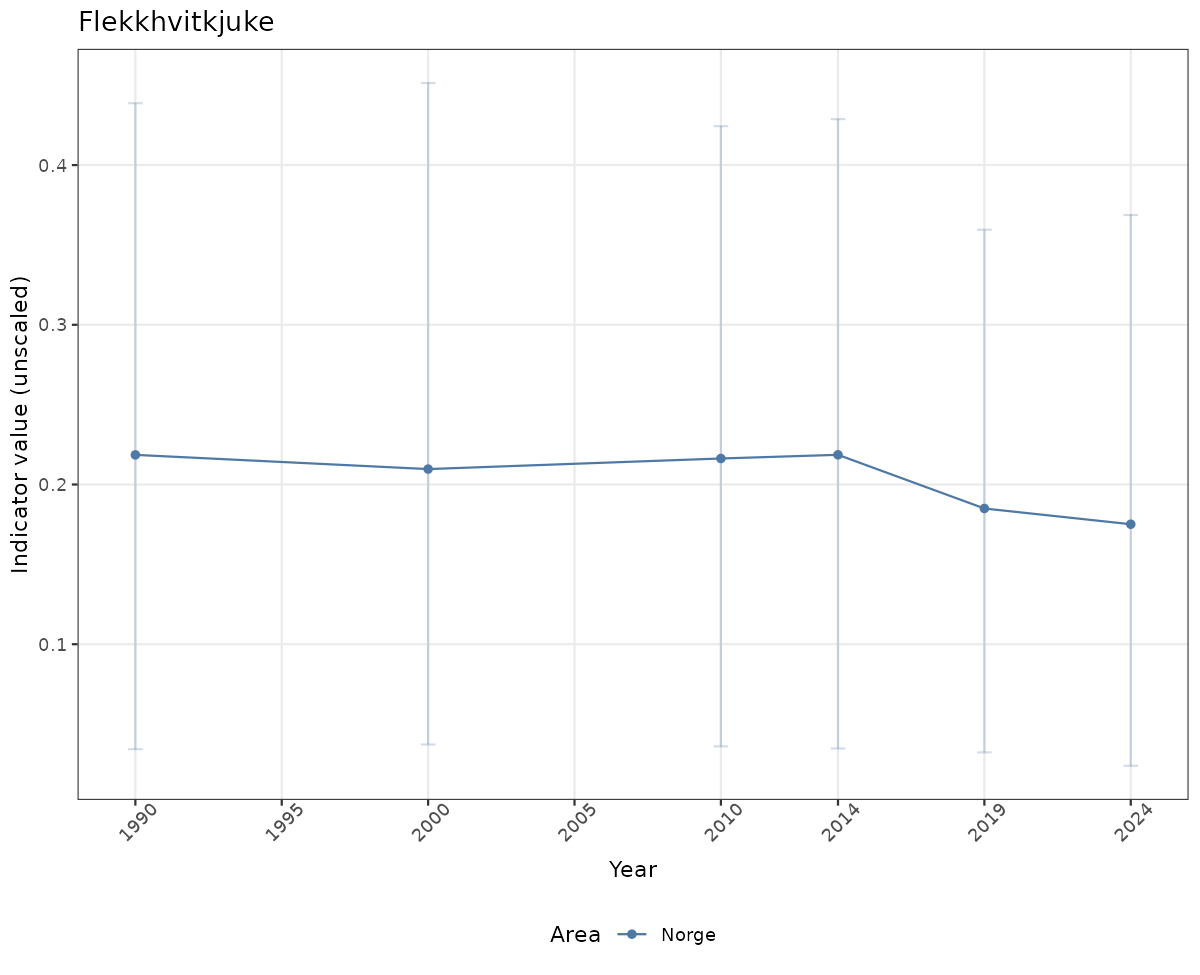
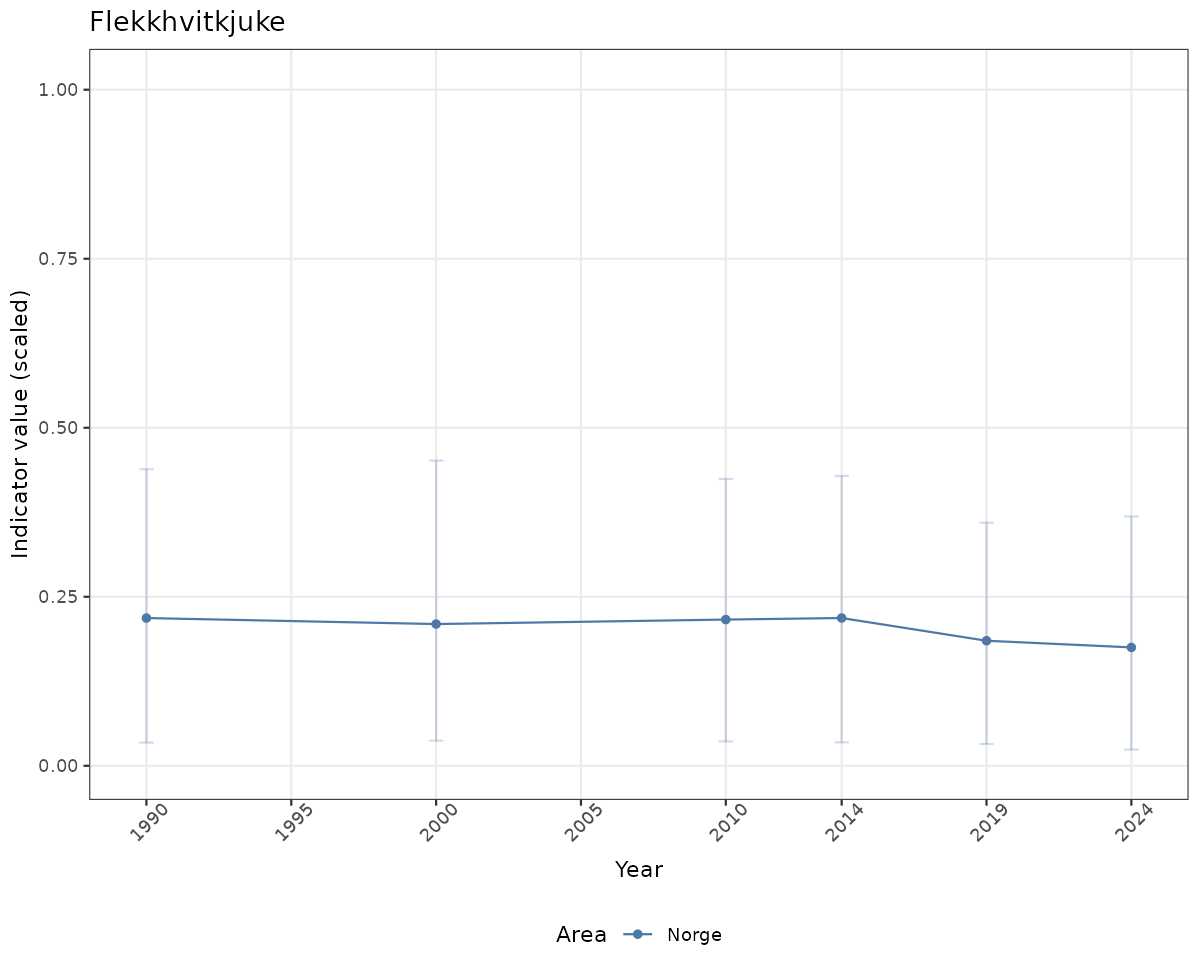
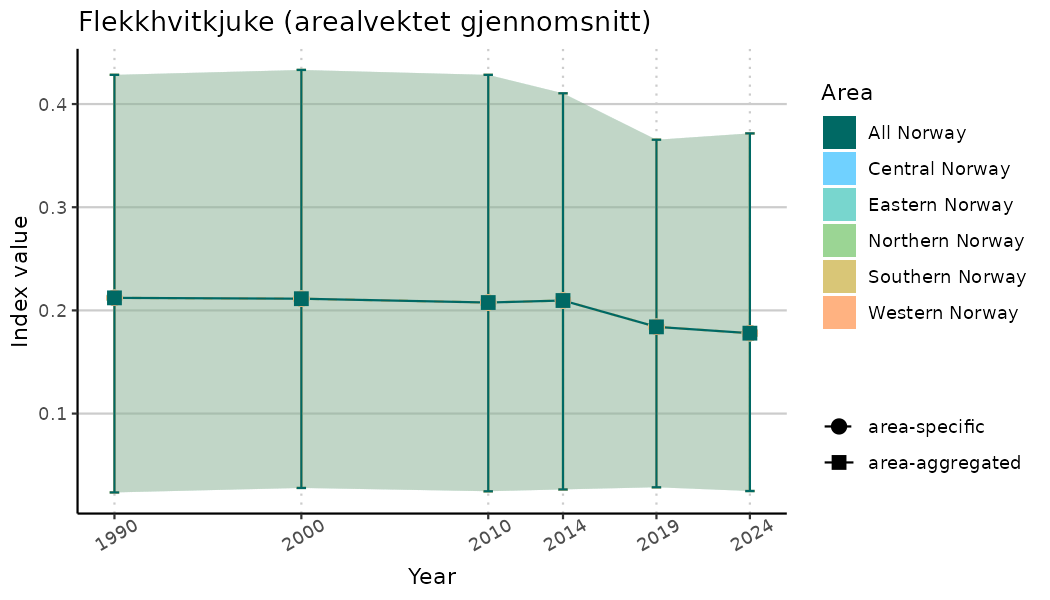
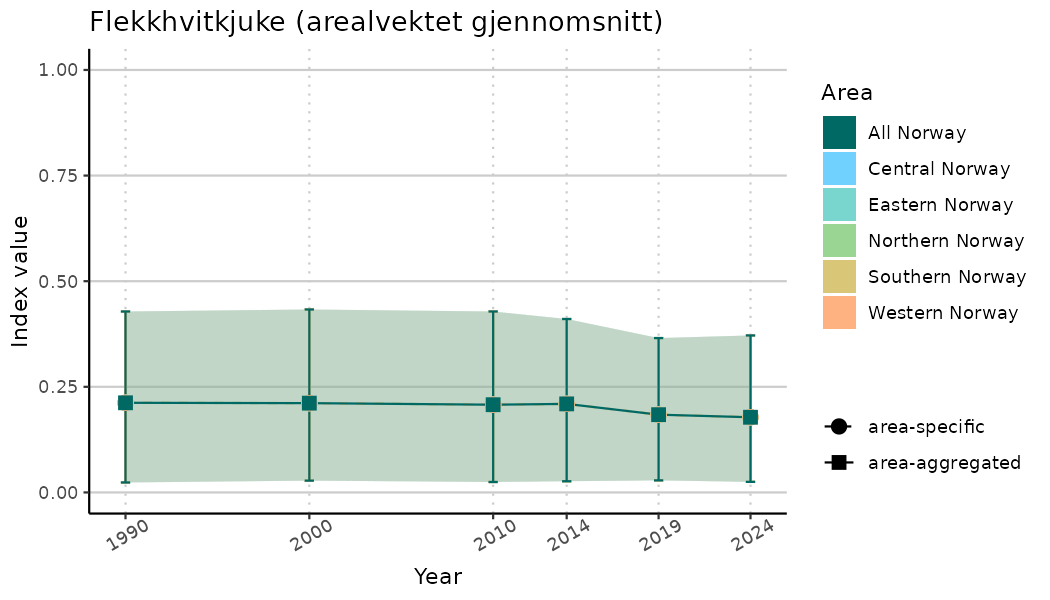
/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)































_%E2%80%93_arealandel/TimeSeries_raw.png)
_%E2%80%93_arealandel/TimeSeries_scaled.png)
_%E2%80%93_arealandel/TimeSeries_indIdx.png)
_%E2%80%93_arealandel/TimeSeries_indIdx_std.png)








/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)





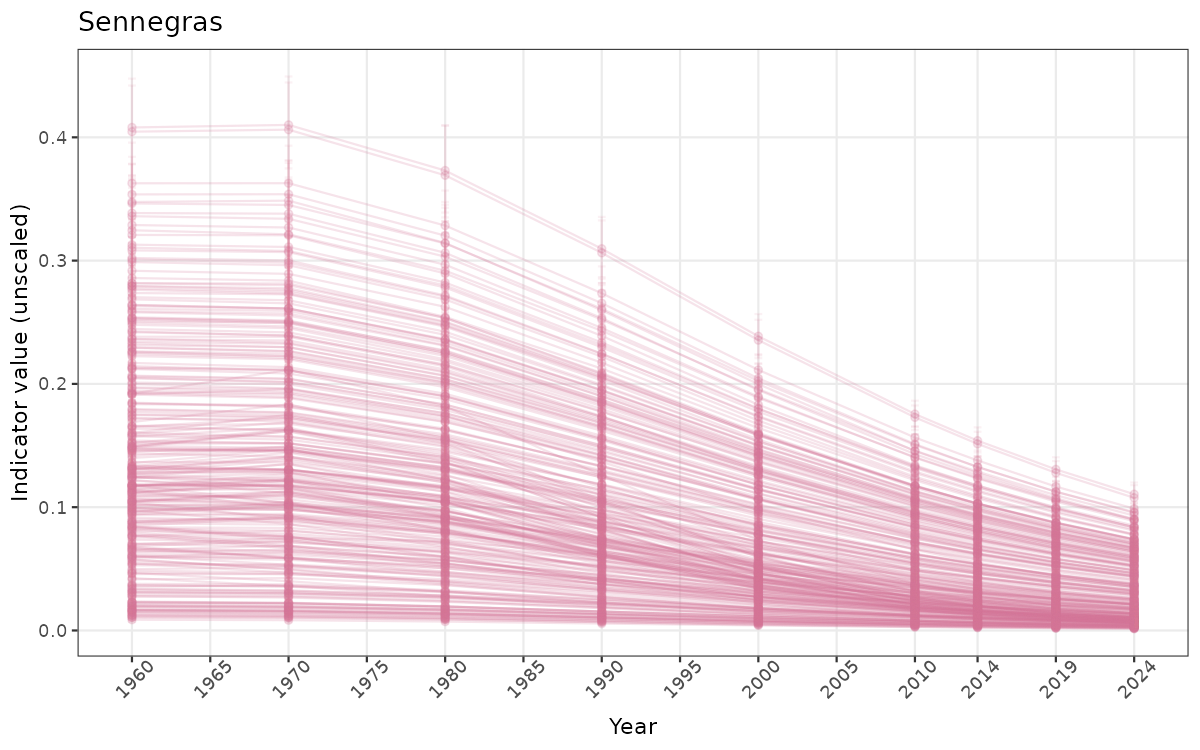
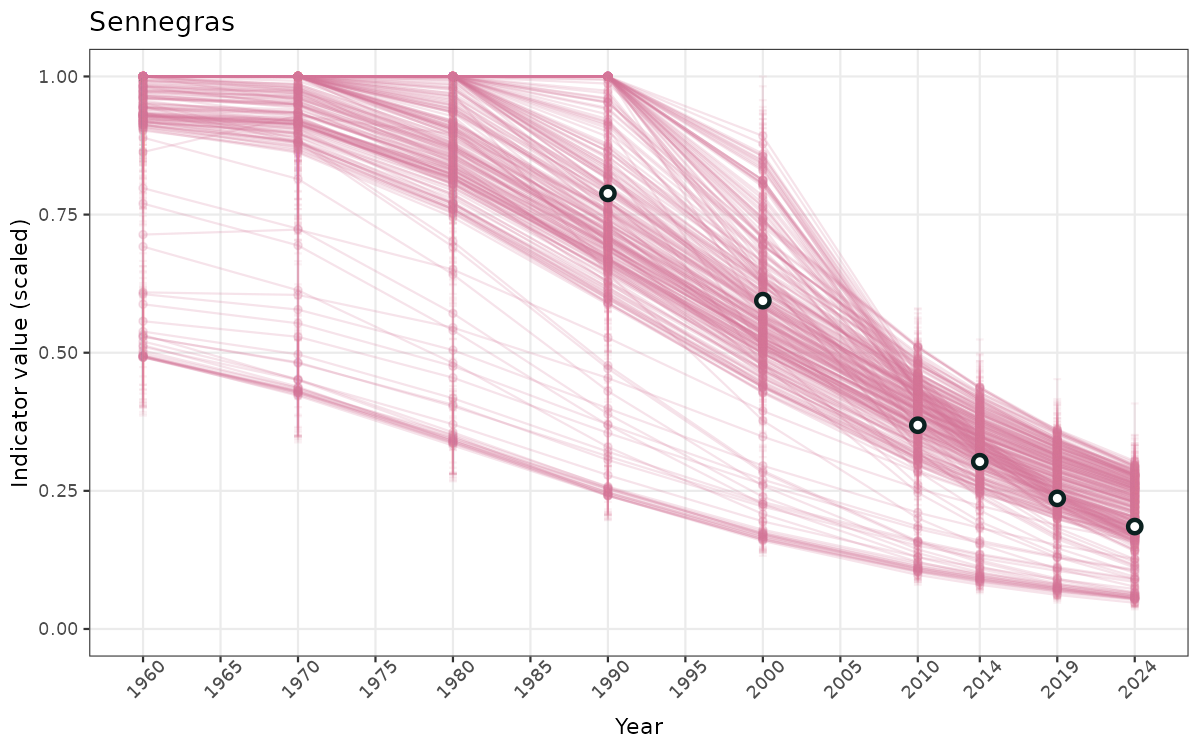
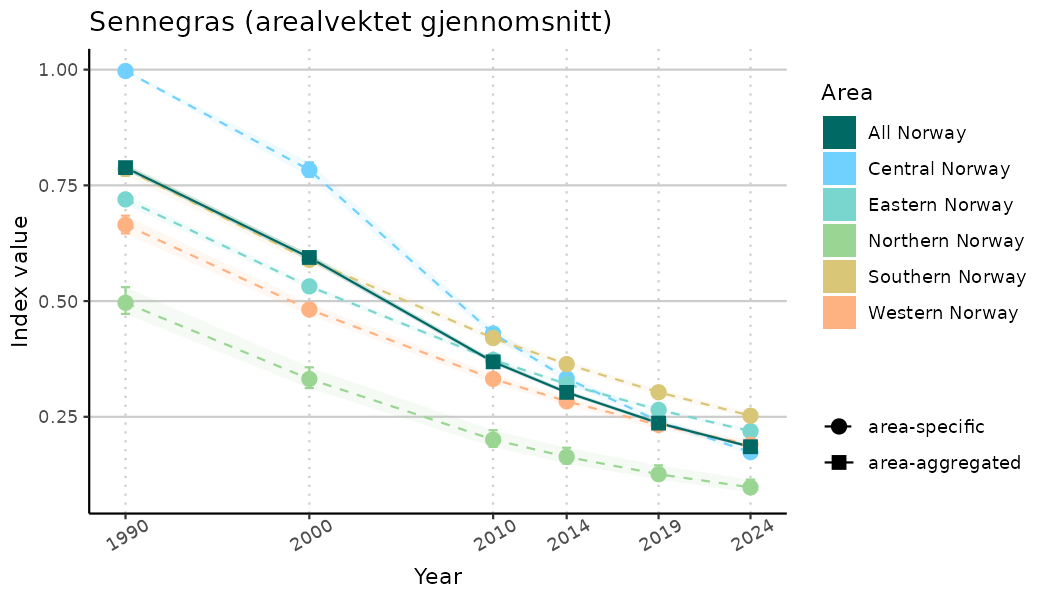
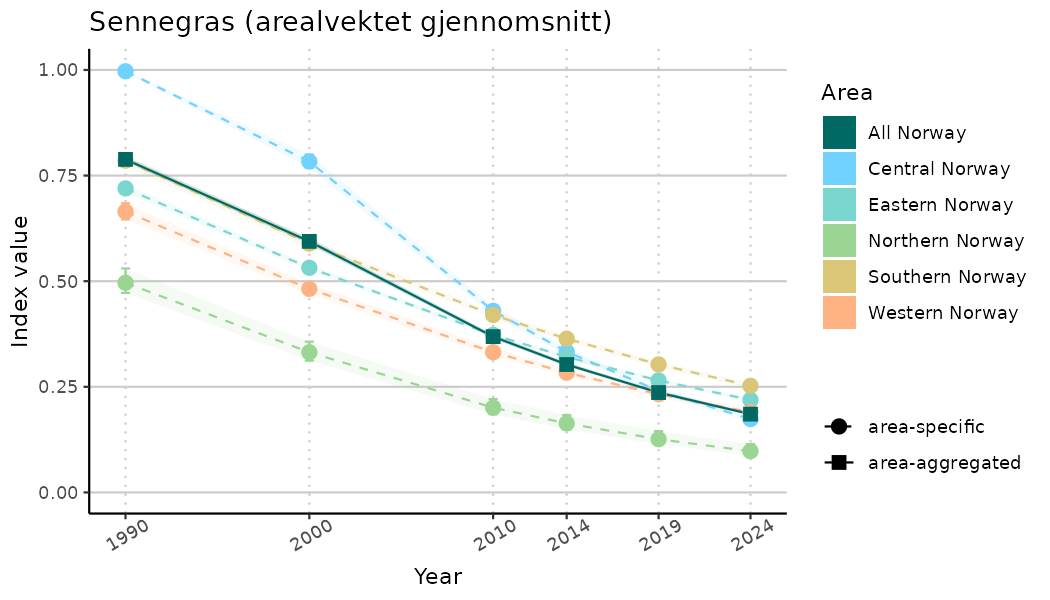










/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)

_%E2%80%93_arealandel/TimeSeries_raw.png)
_%E2%80%93_arealandel/TimeSeries_scaled.png)
_%E2%80%93_arealandel/TimeSeries_indIdx.png)
_%E2%80%93_arealandel/TimeSeries_indIdx_std.png)












/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)
/TimeSeries_raw.png)
/TimeSeries_scaled.png)
/TimeSeries_indIdx.png)
/TimeSeries_indIdx_std.png)